Abstract
Purpose
Skin cutaneous melanoma (SKCM) is a malignant tumor responsible for over 75% of skin cancer deaths, the relationship between fibrosis and cancer has been increasingly appreciated. The aim of this study is to investigate the fibrotic gene signature (FGS) in melanoma and construct a prognostic model based on FGS.
Methods
SKCM-related datasets were obtained from the Gene Expression Omnibus (GEO) database and The Cancer Genome Atlas (TCGA) database. By weighted gene co-expression network analysis (WGCNA) of the TCGA-SKCM cohort and GSE65904 cohort, core modules and central genes highly associated with fibrotic features were identified and intersecting genes were defined as fibrotic gene signature (FGS). The least absolute shrinkage and selection operator (LASSO) regression analysis and the Akaike information criterion (AIC) method were conducted to construct a prognostic model based on the FGS gene set. The fibrotic gene signature enrichment score (FGES) and fibrotic gene signature risk score (FGRS) were used to analyze immune infiltration. For FGRS, the correlation between clinical characteristics and the expression of immune checkpoint genes between different risk groups was also analyzed in depth.
Results
A total of 301 genes were defined as FGS, and a robust eight-gene prediction model was constructed based on FGS, these 8 genes are SV2A, HEYL, OLFML2A, PROX1, ACOX2, PRRX1, PHACTR1 and LHX6. On the basis of the model, a nomogram consisting of FGRS could accurately predict prognosis. In addition, patients in the high-risk group showed immunosuppression, while patients in the low-risk group may benefit more from immunotherapy. However, there was no significant difference between the immune infiltration of different FGES groups.
Conclusion
In this study, taken together, we developed a fibrotic gene signature in melanoma, and construct an eight-gene prognostic model based on the FGS to provide a reference for prognosis estimation and treatment selection for melanoma patients.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
1 Introduction
Although skin cutaneous melanoma (SKCM) accounts for only a small percentage of skin cancers, it is responsible for more than 75% of skin cancer deaths with an increasing incidence [1, 2].
Fibrosis is the result of a tissue repair response and is a common pathological feature of chronic inflammatory diseases. As a normal and important stage of organ tissue repair, fibrosis is characterized by excessive accumulation of the extracellular matrix (ECM). When tissues are injured, local tissue fibroblasts are activated with increased contractility, secreting inflammatory mediators and ECM components to initiate the wound healing response [3].
The relationship between fibrosis and cancer has been demonstrated in a variety of cancers [4,5,6,7,8,9], where it induces and promotes the development of cancer through a "non-healing wound" like mechanism, both activated tumor stroma and fibrosis are characterized by large numbers of activated fibroblasts and increased deposition of extracellular matrix [7]. In addition, chemotherapies and radiotherapies can induce the formation of a fibrotic-like microenvironment [10].
In studies related to melanoma, co-morbidity between cancer and the fibrotic disease scleroderma is very frequent, and the incidence of melanoma is 3.3 times higher in patients diagnosed with scleroderma than in the general population [7]. When using therapies combining BRAF and mitogen-activated protein kinase (MEK) inhibitors for metastatic melanoma, BRAF-mutant melanomas will develop a paradoxical fibro-mechanic reprogramming that produces a therapy-induced fibrotic-like phenotype, giving cancer cells the cell-autonomous ability to resist treatment and escape the challenging tumor microenvironment, resulting in drug resistance [10].
In recent years, there have been many studies on model construction based on immune-related genes [11], metastasis-related genes [12], and chemokine members [13] in the field of melanoma, and Bagaev et al. constructed conserved pan-cancer tumor microenvironment (TME) subtypes containing fibrosis information based on melanoma [14]. However, studies on the fibrotic gene signature of melanoma have not been reported, and prognostic models of melanoma based on fibrotic signature genes are still lacking. Therefore, in this study, we constructed the melanoma fibrotic gene signature for the first time and constructed and validated an 8-gene prognostic model based on the fibrotic signature genes to explore the potential value of fibrotic-related genes in melanoma.
2 Materials and methods
2.1 Data Source and Preprocessing
The RNA-seq counts data and clinical information of patients with SKCM from The Cancer Genome Atlas (TCGA) were downloaded from the TCGA database, then 1) convert Ensemble IDs to gene symbols, 2) take the median expression of genes with multiple gene symbols, 3) remove non-protein-coding genes, and 4) retain the primary and metastatic melanoma samples.
The GSE65904 cohort, which is a microarray that contains gene expression profiles and clinical follow-up information of metastatic melanoma samples, was downloaded from the Gene Expression Omnibus (GEO) database, the downloaded data is in a normalized format. Then 1) convert Probe IDs to gene symbols, 2) take the median expression of genes with multiple gene symbols, 3) remove non-protein-coding genes.
The fibrotic information of the samples was obtained from the TME subtype in the study of Bagaev et al. [14].
2.2 WGCNA and Construction of Fibrotic Gene Signature
After preprocessing, we normalized the gene expression data, calculated the median absolute deviation, and selected the top 5000 genes, and only those samples with both TME subtype information and expression data were retained. Then we used the “goodSamplesGenes” function in the R package “WGCNA” for a further filter, and outlier samples were removed according to the sample clustering results. After the above steps were completed, 462 samples in TCGA-SKCM and 209 samples in the GSE65904 cohort were used in the follow-up study.
Firstly, we used “pickSoftThreshold” function to selected suitable soft thresholds, which was 3 in both the TCGA and GSE65904 cohort, to construct a gene co-expression scale-free network and calculated the adjacency matrix and topological overlap matrix (TOM). Additionally, we performed hierarchical clustering of modules with at least 30 genes per module (minModuleSize = 30). Subsequently, the dynamic tree cut algorithm was used to identify similar modules, and those with similarities greater than 0.75 were merged (abline = 0.25). Finally, we calculated correlations between modules and the fibrotic trait to identify important modules and extract genes within modules. The intersection between extracted genes of TCGA-SKCM and GSE65904 were defined as fibrotic gene signature (FGS).
2.3 Construction of the Prognostic Model
Firstly, we defined 0.01 as the p value to filter genes when employing the univariate Cox proportional hazard regression model to identify prognostic genes in the FGS gene set. Subsequently, the least absolute shrinkage and selection operator (LASSO) regression analysis was used to narrow the range and five-fold cross-validation was used to build the model. Finally, the Akaike information criterion (AIC) method was employed to conduct stepwise regression, by which the multivariate COX model obtains an adequate degree of fit with fewer genes.
2.4 Functional Enrichment Analysis
The gene ontology (GO) functional annotation and Kyoto encyclopedia of genes and genomes (KEGG) pathway analysis were performed using the R package “clusterProfiler” (v4.0.5) to analyze the functional enrichment of the gene set in FGS.
2.5 Analysis of Tumor Microenvironment and Signatures
R package “IOBR” was used in several parts of our study. We use the function “calculate_sig_score” to evaluate the fibrotic gene signature enrichment score (FGES) of samples, TME deconvolution methods to assess the immune infiltration, and function “iobr_cor_plot” to calculate the expression of signatures or genes and draw the plots [15].
2.6 Statistical Analysis
All statistical analyses were performed using the R software version 4.1.0.
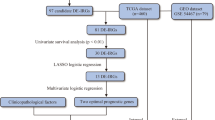
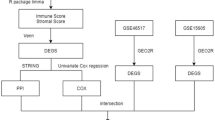
2.7 Technical Route and Flow Chart
The overall analysis process and flow chart of the experiment are shown in Fig. S8.
3 Results
3.1 Construction, Analysis, and Validation of Fibrotic Gene Signature
In the analysis of the TCGA-SKCM data, we selected 1234 genes in the blue module (Fig. S1A), while 2037 genes in the blue and green modules were co-selected in the WGCNA results of GSE65904 (Fig. S1B), additional WGCNA results are shown in Fig. S2, Tables S1 and S2. After selecting the intersection, 301 genes were used to construct the fibrotic gene signature (FGS) (Table S3).
The GO analysis was divided into the biological process (BP), cell component (CC), and molecular function (MF). The results showed that the genes in the BP group showed a significant relationship with extracellular matrix and cell-substrate adhesion. In addition, cell components associated with collagen and organelle lumen were enriched in multiple categories. Moreover, MF group genes were mainly related to extracellular matrix structural constituent and multiple binding functions (Fig. 1A). Notably, the results of KEGG analysis showed that the mainly enriched pathways (Fig. 1B), such as ECM-receptor interaction, focal adhesion, and signaling pathways like PI3K-Akt, PPAR, and Wnt, were tumor-related pathways.
In order to quantify the enrichment degree of the FGS, we calculated the signature enrichment score for each sample to represent the degree to which the genes in the gene set are coordinately regulated. Samples without follow-up information were removed, with 454 samples in TCGA and 210 samples in GSE65904 were selected. The remaining samples were divided into two groups by the cutpoint in the Kaplan–Meier survival analysis, KM survival curves are shown in Figs. 2A and 3A. Besides, the receiver operating characteristic (ROC) curves indicated that the FGES could predict TME subtype with approximately over 85% accuracy (Figs. 2B and 3B, Fig. S3), and the violin plot further verified the significant difference between FGES of TME subtype (Figs. 2C and 3C).
Effectiveness of FGES and immune infiltration analysis of FGES groups in TCGA-SKCM. A KM survival curve. B ROC curve of the classification effectiveness of FGES to TME subtype. C Violin plot showed the difference of FGES between two TME subtypes. D Comparison of ESTIMATE scores between two FGES groups. E Comparison of MCPcounter scores between two FGES groups. F Heatmap for the comparison of TME deconvolution scores between two FGES groups (NS, no significant, *P < 0.05, **P < 0.01, ***P < 0.001, ****P < 0.0001)
Effectiveness of FGES and immune infiltration analysis of FGES groups in GSE65904. A KM survival curve. B ROC curve of the classification effectiveness of FGES to TME subtype. C Violin plot showed the difference of FGES between two TME subtypes. D Comparison of ESTIMATE scores between two FGES groups. E Comparison of MCPcounter scores between two FGES groups. F Heatmap for the comparison of TME deconvolution scores between two FGES groups (NS, no significant, *P < 0.05, **P < 0.01, ***P < 0.001, ****P < 0.0001)
3.2 Differences in Immune Infiltration Between FGES Groups
ESTIMATE [16] and MCPcounter [17] are commonly used deconvolution methods to analyze immune infiltration. The ESTIMATE results showed that the group with higher FGES has a higher stromal score, ESTIMATE score, and lower tumor purity (Fig. 2D), which showed the same trend in GSE65904 (Fig. 3D). However, the MCPcounter displayed in Figs. 2E and 3E only showed significant differences in neutrophils, endothelial cells, and fibroblasts. Figures 2F and 3F combined these results and presented them as a heatmap.
3.3 Construction of a Prognostic Model Based on FGS Genes
The TCGA cohort was used to construct the prognostic model. After univariate Cox proportional hazard regression, 16 genes in the FGS gene set were obtained as the prognostic hub genes. Then we used LASSO regression analysis and five-fold cross-validation, the coefficient trajectory of each gene and the confidence interval under each lambda are shown in Fig. 4A separately. According to the results, we selected 13 genes as candidates with the optimal model at lambda = 0.01595426 (log(lambda) = -4.138) (Fig. 4B). Finally, as a result of multivariate COX analysis, eight genes were selected to construct the model to calculate the fibrotic gene signature risk score (FGRS):
LASSO regression analysis and prediction effect in the TCGA-SKCM cohort. A The changing trajectory of each independent variable, the abscissa represents log value of lambda, and the ordinate represents the coefficient of the independent variable. B The log value lambda, the abscissa represents the confidence interval of each lambda, and the ordinate represents errors in cross validation. C KM survival curve of two risk groups. D ROC curve of the prognostic model. E FGRS, survival status and time, and expression trend of eight genes
According to the model, samples were divided into two risk groups by the median of the FGRS, the KM curve presented a significant difference in survival between the high-risk group (HRG) and low-risk group (LRG) (Fig. 4C), Survival analyses for each gene are shown in Fig. S4. Further, we used the R package “timeROC” to analyze the prognostic efficiency of the FGRS, the prediction performance of our model was represented by the area under the curves (AUCs), which reached 0.695 (1 year), 0.653 (3 years), and 0.686 (5 years) (Fig. 4D). In addition, as the risk score increased, the changing trend of gene expression levels was correlated with their interpretable coefficients, which means, specifically, that genes with positive coefficient had an upward trend while those with negative coefficient showed a downward performance (Fig. 4E).
3.4 External Validation of the Robustness of the Model
To determine the robustness of our model, GSE65904 was introduced as an independent external cohort for validation. Each sample in GSE65904 obtained a FGRS through the same formula operated in the TCGA cohort and categorized into a risk group according to its FGRS. Like is in the TCGA cohort, the high-risk group had a significantly lower survival probability (Fig. S5A), and the AUCs of 1-year, 3-year and 5-year survival were 0.673, 0.649, and 0.611 respectively (Fig. S5B). Besides, a similar trend was observed for gene expression (Fig. S5C).
3.5 Correlation Between FGRS and Clinical Characteristics
To investigate the correlation between FGRS and clinical characteristics, we studied the following clinical features in the TCGA-SKCM cohort: age, gender, stage, and Clark level (Table 1). The results showed that there was no difference between FGRS of male and female (Fig. 5A), and no significant relationship was discovered between FGRS and stage (Fig. 5B). Interestingly, older patients (> 65 years old) had a higher FGRS (Fig. 5C), and patients in Clark level V were more likely to receive a high score (Fig. 5D). However, due to a large number of missing values in the Clark level information, the results may not always be reliable.
Correlation between FGRS and clinical characteristics, and clinical value of the model. A Correlation diagram between FGRS and gender. B Correlation diagram between FGRS and stage. C Correlation diagram between FGRS and age. D Correlation diagram between FGRS and Clark level. E Nomogram, the red element shows the points and the probability of 1-, 3-, or 5-year survival of a sample in the TCGA cohort. D Calibration curves, demonstrate the relationship between nomogram-predicted 1-, 3-, and 5-year OS and actual survival
3.6 Construction and Evaluation of Nomograms Comprising FGRS
The univariate and multivariate Cox regression analysis were used to evaluate the independence of our model in clinical application. The results revealed that age, Clark level, stage, and FGRS were significant and independent risk factors for prognosis, while FGRS in multivariate Cox regression analysis showed a strong significance (P < 0.001, HR = 2.15, Table 2), indicating that our eight-gene prognostic model had a good capability in clinical prediction. To further improve the clinical significance of our study, we combined various risk factors based on the results of multivariate Cox regression analysis to construct a nomogram that is more comprehensive and intuitive in clinical decision-making (Fig. 5E). The consistencies between the nomogram-predicted overall survival and actual survival are shown in Fig. 5F by calibration curves, the results indicated that the nomogram had a good calibrating ability.
3.7 Differences in Immune Infiltration Between FGRS Groups
The differences in overall immune infiltration were assessed by ESTIMATE and MCPcounter. Different from the results represented in the FGES group, almost all of the immune infiltrating cells in the MCP results were scored higher in the low-risk group, both in the TCGA and GSE65904 cohort (Fig. S6A and S7A), which indicated that the degree of inflammatory response and immune response was more active in the low-risk group. Meanwhile, the ESTIMATE results of cohort GSE65904 showed that immune score and ESTIMATE score were lower in the high-risk group, and tumor purity of the high-risk group is higher (Fig. S7B). However, this result was not that significant in the TCGA cohort (Fig. S6B).
The difference in immune infiltration between the two groups indicated that our model might be helpful for predicting the immunotherapeutic responses. Therefore, we used the R package “IOBR” to calculate and demonstrate the differences in expression of some immune checkpoint genes between the two FGRS groups. Not surprisingly, almost all of the immune checkpoint genes we tested were upregulated in the low-risk group (Fig. 6A and B), which predicted that patients in the low-risk group may benefit more from immunotherapy.
4 Discussion
Based on the growing number of studies focused on the relationship between fibrosis and cancer, the frequent co-morbidity between melanoma and fibrotic disease, and the lack of research on fibrotic gene signature in melanoma, we did the study to fill the gap.
Orthogonal partial least squares discrimination analysis(OPLS-DA) is a commonly used method to screen for differential metabolites [18,19,20], and it was used to screen for survival-associated differential Long non-coding RNAs (lncRNAs) in the study by Cao et al.[21]. Based on the fact that there is no current concept of gene sets for fibrosis-related traits, in our study we chose the more appropriate approach, weighted gene co-expression network analysis(WGCNA) [22], to screen gene sets as fibrotic gene signature(FGS) when identifying gene sets that vary highly synergistically with the target trait, i.e., fibrosis. We used the information provided by the previous research on melanoma TME subtype, then the intersect hub 301 genes we screened from the TCGA cohort and GEO cohort were selected and defined as fibrotic gene signature. To further analyze the functional enrichment of genes in FGS, we did functional enrichment analysis, the enrichment of ECM-receptor interaction, cell-substrate adhesion, focal adhesion, and signaling pathway like PI3K-Akt, PPAR, and Wnt suggested that the genes in FGS involved in the process of tumor shedding, adhesion, degradation, movement, and hyperplasia.
The fibrotic gene signature enrichment score was used to quantify the degree of FGS enrichment in the samples, based on the results that FGES can be used as an effective classifier for TME subtype as the ROC curve demonstrated, KM survival analysis suggested that samples with high FGES had a worse prognosis, further proved that the degree of tissue fibrosis correlates with poor patient prognosis as described in the study by Piersma B et al. [23]. In the results of the immune infiltration analysis, the group with higher FGES has a higher stromal score and lower tumor purity, suggesting a higher degree of stromal cell infiltration in this group of samples, which is thought to have an important role in tumor growth, disease progression and drug resistance [24,25,26]. In addition, in several solid tumors, the high expression level of neutrophils has been associated with detrimental outcomes [17], matching the results of MCPcounter and OS curves.
Based on the gene set of FGS, univariate Cox regression identified 16 prognostic genes in the TCGA-SKCM cohort, and then we used the LASSO algorithm for dimensionality reduction to select 13 candidates, followed by stepwise regression using the AIC method to finally construct an 8-gene prognostic model. In selecting the method of gene reduction, Cao et al. [21] used a 1.5-fold change as the threshold for screening target lncRNAs in differential lncRNAs to narrow down the number of lncRNAs for constructing the prediction panel. In our study, considering the possible multicollinearity in the FGS gene set, we used the least absolute shrinkage and selection operator (LASSO) regression analysis to compress the estimated parameters before constructing the model. In the validation of model validity, there was a significant survival difference between risk groups classified by FGRS, and the predictive performance of the model was consistently higher than 0.65 at 1 year, 3 years, and 5 years. In addition, we analyzed the GSE65904 cohort and the results showed that the 8-gene model has convincing validity and stability.
After the construction of the prognostic model, we analyzed the correlation between FGRS and clinical traits. It was suggested that FGRS was associated with the age and Clark level of the patients. The FGRS was higher in patients over 65 years old, indicating that our model can identify risk groups by age to some extent. Clark level is a classification system to describe the level of invasion in human cutaneous melanomas. Clark I-IV indicates that tumors located in the skin layer(Clark I represents carcinoma in situ), and tumors invaded the subcutaneous layer are defined as Clark V [27]. Our study showed that patients in Clark level V were more likely to get a high score, but because of the missing values of Clark level, the universality of the results is still debatable. Subsequently, the univariate and multivariate Cox regression analysis suggested that FGRS was just one of the independent risk factors for prognosis, so we combined all the risk factors to construct a nomogram with well-calibrating ability.
Finally, we assessed the overall immune infiltration of two FGRS groups. The differences in immune infiltration landscape between HRG and LRG showed that there was a suppressed immune microenvironment of HRG. Previous studies demonstrated that immune infiltration of the tumor microenvironment can be used to assess the prognostic of multiple tumors, such as glioblastoma [28], breast cancer [29], and melanoma [30], which may mechanistically explain the poor prognosis of patients in HRG.
Immune checkpoint inhibitors have provided durable responses and improved survival in the treatment of patients with melanoma [31,32,33] and many other solid tumors [34,35,36].
To further analyze the potential of our model for immunotherapy response prediction, we compared the differences in expression of immune checkpoint genes between the two groups. Interestingly, almost all immune checkpoint genes were upregulated in LRG, suggesting that this group may be more likely to benefit from immune checkpoint inhibitor therapy like anti-CTLA-4 and anti-PD-1 or anti-PD-L1 therapies.
The current study does have some limitations. First, we did not score the degree of fibrosis of the tissue to validate the relationship between FGES and the degree of fibrosis, and second, we were unable to validate the model in a larger cohort due to the lack of data with a sufficient sample size with clinical information.
Availability of data and materials
The publicly available datasets analyzed in this study can be found in The Cancer Genome Atlas (https://portal.gdc.cancer.gov/), Gene Expression Omnibus (GSE65904, GSE54467, GSE22153, GSE59455, GSE91061, GSE8401, GSE43081, GSE35640, GSE19293, and GSE19234)( https://www.ncbi.nlm.nih.gov/geo/). The information of protein-coding genes was from GENCODE (https://www.gencodegenes.org/human/). TME subtype information for melanoma samples was obtained from the study of Bagaev et al. (https://doi.org/10.1016/j.ccell.2021.04.014). Survival analysis of 8 genes in melanoma were provided by GEPIA(http://gepia.cancer-pku.cn/).
References
Rebecca VW, Somasundaram R, Herlyn M. Pre-clinical modeling of cutaneous melanoma. Nat Commun. 2020;11(1):2858. https://doi.org/10.1038/s41467-020-15546-9.
Siegel RL, Miller KD, Fuchs HE, Jemal A. Cancer Statistics, 2021. CA Cancer J Clin. 2021;71(1):7–33. https://doi.org/10.3322/caac.21654.
Henderson NC, Rieder F, Wynn TA. Fibrosis: from mechanisms to medicines. Nature. 2020;587(7835):555–66. https://doi.org/10.1038/s41586-020-2938-9.
Ballester B, Milara J, Cortijo J. Idiopathic Pulmonary Fibrosis and Lung Cancer: Mechanisms and Molecular Targets. Int J Mol Sci. 2019;20(3):593. https://doi.org/10.3390/ijms20030593.
Chen IX, Chauhan VP, Posada J, Ng MR, Wu MW, Adstamongkonkul P, et al. Blocking CXCR4 alleviates desmoplasia, increases T-lymphocyte infiltration, and improves immunotherapy in metastatic breast cancer. Proc Natl Acad Sci U S A. 2019;116(10):4558–66. https://doi.org/10.1073/pnas.1815515116.
Costa A, Kieffer Y, Scholer-Dahirel A, Pelon F, Bourachot B, Cardon M, et al. Fibroblast Heterogeneity and Immunosuppressive Environment in Human Breast Cancer. Cancer Cell. 2018;33(3):463-79.e10. https://doi.org/10.1016/j.ccell.2018.01.011.
Hutchenreuther J, Leask A. Why target the tumor stroma in melanoma? J Cell Commun Signal. 2018;12(1):113–8. https://doi.org/10.1007/s12079-017-0419-1.
Park J, Kim DS, Shim TS, Lim CM, Koh Y, Lee SD, et al. Lung cancer in patients with idiopathic pulmonary fibrosis. Eur Respir J. 2001;17(6):1216–9. https://doi.org/10.1183/09031936.01.99055301.
Thomas D, Radhakrishnan P. Tumor-stromal crosstalk in pancreatic cancer and tissue fibrosis. Mol Cancer. 2019;18(1):14. https://doi.org/10.1186/s12943-018-0927-5.
Diazzi S, Tartare-Deckert S, Deckert M. Bad Neighborhood: Fibrotic Stroma as a New Player in Melanoma Resistance to Targeted Therapies. Cancers (Basel). 2020;12(6):1364. https://doi.org/10.3390/cancers12061364.
Huang R, Mao M, Lu Y, Yu Q, Liao L. A novel immune-related genes prognosis biomarker for melanoma: associated with tumor microenvironment. Aging (Albany NY). 2020;12(8):6966–80. https://doi.org/10.18632/aging.103054.
Wan Q, Liu C, Liu C, Liu W, Wang X, Wang Z. Discovery and Validation of a Metastasis-Related Prognostic and Diagnostic Biomarker for Melanoma Based on Single Cell and Gene Expression Datasets. Front Oncol. 2020;10:585980. https://doi.org/10.3389/fonc.2020.585980.
Huang B, Han W, Sheng ZF, Shen GL. Identification of immune-related biomarkers associated with tumorigenesis and prognosis in cutaneous melanoma patients. Cancer Cell Int. 2020;20:195. https://doi.org/10.1186/s12935-020-01271-2.
Bagaev A, Kotlov N, Nomie K, Svekolkin V, Gafurov A, Isaeva O, et al. Conserved pan-cancer microenvironment subtypes predict response to immunotherapy. Cancer Cell. 2021;39(6):845-65.e7. https://doi.org/10.1016/j.ccell.2021.04.014.
Zeng D, Ye Z, Shen R, Yu G, Wu J, Xiong Y, et al. IOBR: Multi-Omics Immuno-Oncology Biological Research to Decode Tumor Microenvironment and Signatures. Front Immunol. 2021;12:687975. https://doi.org/10.3389/fimmu.2021.687975.
Yoshihara K, Shahmoradgoli M, Martínez E, Vegesna R, Kim H, Torres-Garcia W, et al. Inferring tumour purity and stromal and immune cell admixture from expression data. Nat Commun. 2013;4:2612. https://doi.org/10.1038/ncomms3612.
Becht E, Giraldo NA, Lacroix L, Buttard B, Elarouci N, Petitprez F, et al. Estimating the population abundance of tissue-infiltrating immune and stromal cell populations using gene expression. Genome Biol. 2016;17(1):218. https://doi.org/10.1186/s13059-016-1070-5.
Kang C, Zhang Y, Zhang M, Qi J, Zhao W, Gu J, et al. Screening of specific quantitative peptides of beef by LC-MS/MS coupled with OPLS-DA. Food Chem. 2022;387:132932. https://doi.org/10.1016/j.foodchem.2022.132932.
Du Y, Fan P, Zou L, Jiang Y, Gu X, Yu J, et al. Serum Metabolomics Study of Papillary Thyroid Carcinoma Based on HPLC-Q-TOF-MS/MS. Front Cell Dev Biol. 2021;9:593510. https://doi.org/10.3389/fcell.2021.593510.
Zhao S, Liu H, Su Z, Khoo C, Gu L. Identifying Cranberry Juice Consumers with Predictive OPLS-DA Models of Plasma Metabolome and Validation of Cranberry Juice Intake Biomarkers in a Double-Blinded, Randomized, Placebo-Controlled, Cross-Over Study. Mol Nutr Food Res. 2020;64(11):e1901242. https://doi.org/10.1002/mnfr.201901242.
Cao W, Liu JN, Liu Z, Wang X, Han ZG, Ji T, et al. A three-lncRNA signature derived from the Atlas of ncRNA in cancer (TANRIC) database predicts the survival of patients with head and neck squamous cell carcinoma. Oral Oncol. 2017;65:94–101. https://doi.org/10.1016/j.oraloncology.2016.12.017.
Langfelder P, Horvath S. WGCNA: an R package for weighted correlation network analysis. BMC Bioinformatics. 2008;9:559. https://doi.org/10.1186/1471-2105-9-559.
Piersma B, Hayward MK, Weaver VM. Fibrosis and cancer: A strained relationship. Biochim Biophys Acta Rev Cancer. 2020;1873(2):188356. https://doi.org/10.1016/j.bbcan.2020.188356.
Kalluri R, Zeisberg M. Fibroblasts in cancer. Nat Rev Cancer. 2006;6(5):392–401. https://doi.org/10.1038/nrc1877.
Hanahan D, Weinberg RA. Hallmarks of cancer: the next generation. Cell. 2011;144(5):646–74. https://doi.org/10.1016/j.cell.2011.02.013.
Straussman R, Morikawa T, Shee K, Barzily-Rokni M, Qian ZR, Du J, et al. Tumour micro-environment elicits innate resistance to RAF inhibitors through HGF secretion. Nature. 2012;487(7408):500–4. https://doi.org/10.1038/nature11183.
Clark WH Jr, From L, Bernardino EA, Mihm MC. The histogenesis and biologic behavior of primary human malignant melanomas of the skin. Cancer Res. 1969;29(3):705–27.
Fortis SP, Mahaira LG, Anastasopoulou EA, Voutsas IF, Perez SA, Baxevanis CN. Immune profiling of melanoma tumors reflecting aggressiveness in a preclinical model. Cancer Immunol Immunother. 2017;66(12):1631–42. https://doi.org/10.1007/s00262-017-2056-1.
Priedigkeit N, Watters RJ, Lucas PC, Basudan A, Bhargava R, Horne W, et al. Exome-capture RNA sequencing of decade-old breast cancers and matched decalcified bone metastases. JCI Insight. 2017;2(17):e95703. https://doi.org/10.1172/jci.insight.95703.
Jia D, Li S, Li D, Xue H, Yang D, Liu Y. Mining TCGA database for genes of prognostic value in glioblastoma microenvironment. Aging (Albany NY). 2018;10(4):592–605. https://doi.org/10.18632/aging.101415.
Twyman-Saint Victor C, Rech AJ, Maity A, Rengan R, Pauken KE, Stelekati E, et al. Radiation and dual checkpoint blockade activate non-redundant immune mechanisms in cancer. Nature. 2015;520(7547):373–7. https://doi.org/10.1038/nature14292.
Gibney GT, Weiner LM, Atkins MB. Predictive biomarkers for checkpoint inhibitor-based immunotherapy. Lancet Oncol. 2016;17(12):e542–51. https://doi.org/10.1016/s1470-2045(16)30406-5.
Carlino MS, Larkin J, Long GV. Immune checkpoint inhibitors in melanoma. Lancet. 2021;398(10304):1002–14. https://doi.org/10.1016/s0140-6736(21)01206-x.
Chen D, Menon H, Verma V, Guo C, Ramapriyan R, Barsoumian H, et al. Response and outcomes after anti-CTLA4 versus anti-PD1 combined with stereotactic body radiation therapy for metastatic non-small cell lung cancer: retrospective analysis of two single-institution prospective trials. J Immunother Cancer. 2020;8(1):e000492. https://doi.org/10.1136/jitc-2019-000492.
Braun DA, Hou Y, Bakouny Z, Ficial M, Sant’ Angelo M, Forman J, et al. Interplay of somatic alterations and immune infiltration modulates response to PD-1 blockade in advanced clear cell renal cell carcinoma. Nat Med. 2020;26(6):909–18. https://doi.org/10.1038/s41591-020-0839-y.
Rotte A. Combination of CTLA-4 and PD-1 blockers for treatment of cancer. J Exp Clin Cancer Res. 2019;38(1):255. https://doi.org/10.1186/s13046-019-1259-z.
Acknowledgements
The authors thank TCGA and GEO database for sharing research materials and the researchers and patients who contributed data.
Funding
This work was supported by National Natural Science Foundation of China grants (No. 8197102109).
Author information
Authors and Affiliations
Contributions
SC, CS, BL and LL designed the study, SC and CS analyzed the data, SC and CS drafted the manuscript, BL and LL revised the manuscript. All authors contributed to the article and approved the submitted version.
Corresponding authors
Ethics declarations
Ethics approval and consent to participate
The patient information and data sources for this study have been described, and the study did not involve experiments that required additional ethical approval.
Competing interests
The authors declare that they have no conflict of interest.
Additional information
Publisher’s Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Additional file 2:
Table S1. 1234 genes in the blue module of WGCNA in TCGA-SKCM.
Additional file 3:
Table S2. 2037 genes in the blue module and green module of WGCNA in GSE65904.
Additional file 4:
Table S3. Fibrotic Gene Signature.
Additional file 5:
Table S4. Clinical information of TCGA-SKCM cohort.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Chen, S., Shi, C., Li, B. et al. Development of fibrotic gene signature and construction of a prognostic model in melanoma. Holist Integ Oncol 2, 11 (2023). https://doi.org/10.1007/s44178-023-00035-y
Received:
Accepted:
Published:
DOI: https://doi.org/10.1007/s44178-023-00035-y