Abstract
Biological structures, spanning the spectrum from cells to multicellular organisms, organs, and the human body, exhibit exquisite hierarchical precision in three dimensions. Modern biomedicine necessitates precise detection and manipulation on the surfaces of organisms with complex morphology. The assembly of medical devices into three-dimensional structures requires a complex design and preparation process. Origami and Kirigami, rooted in historical craftsmanship, provide a viable solution to the challenges posed by the intricate world of biomedical engineering. Through simple geometric design, these traditional arts can impart novel mechanical properties, including but not limited to precise 3D deformation, hyper-stretching capabilities, and the creation of multistable structures. Origami and Kirigami designs present innovative opportunities for the high-volume assembly of 3D bio-integrated systems. This review aims to summarize and highlight the advantages offered by Origami and Kirigami design in biomedical engineering. Specifically, we will delve into their applications in diverse areas such as biosensing, scaffolds, medical instruments, and bioassembly. By elucidating these applications, we intend to underscore the versatility and transformative potential these traditional arts bring to the forefront of cutting-edge biomedical technologies. Lastly, we will discuss the challenges and future perspectives of origami and Kirigami design.

Copyright 2018 Wiley. Microfluidics. Microfluidics. Reprinted with permission from Ref. [16]. Copyright 2014 ACS Publication. Cell Culture Stents. Reprinted with permission from Ref. [17]. Copyright 2023 Springer Nature. Bone stents. Reprinted with permission from Ref. [18]. Copyright 2020 Elsevier. Manipulators. Reprinted with permission from Ref. [19]. Copyright 2020 Springer Nature. Robots. Reprinted with permission from Ref. [20]. Copyright 2022 Springer Nature. DNA Origami. Reprinted with permission from Ref. [21]. Copyright 2006 Springer Nature. Molecular Origami. Reprinted with permission from Ref. [22]. Copyright 2018 AAAS

Copyright 2018 Wiley. b A 3D clamp platform that gently wraps and mechanically constrains organoids in ways that facilitate mechanical evaluations without damage. Reprinted with permission from Ref. [38]. Copyright 2021 Wiley. c FET probe that can be inserted into a single cell. Reprinted with permission from Ref. [43]. Copyright 2010 AAAS d Strain-insensitive graphene-based stretchable electrode with a Kirigami structure. Reprinted with permission from Ref. [44]. Copyright 2020 Elsevier

Copyright 2011 ACS Publication. b Capillary force-assembly microfluidic device. Reprinted with permission from Ref. [49]. Copyright 2017 Wiley

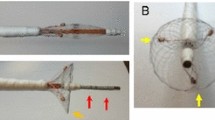
Copyright 2023 Springer Nature. b Bilayer hydrogel system for the simultaneous cultivation of epithelial and fibroblast cells. Reprinted with permission from Ref. [56]. Copyright 2023 Wiley. c Origami stent. Reprinted with permission from Ref. [57]. Copyright 2006 Elsevier. d The Bioadhesive patch folded into a triangular sleeve and then adhered the patch to the inside of the trachea and esophagus in conjunction with a balloon catheter. Reprinted with permission from Ref. [58]. Copyright 2021 Wiley e Bone scaffolds. Reprinted with permission from Ref. [60]. Copyright 2018 RSC publishing. f Multistable bone scaffold using a Kirigami structure. Reprinted with permission from Ref. [18]. Copyright 2020 Elsevier

Copyright 2015 ACS Publication. b A folding nanoinjector for injection into mouse zygotes. Reprinted with permission from Ref.[19]. Copyright 2020 Springer Nature. c magnetically driven amphibious origami micro-robot based on origami design. Reprinted with permission from Ref. [20]. Copyright 2022 Springer Nature. d soft manipulators. Reprinted with permission from Ref. [68]. Copyright 2023 Springer Nature

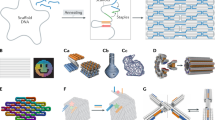
Copyright 2006 Springer Nature. c A DNA box. Reprinted with permission from Ref.[72]. Copyright 2009 Springer Nature. d A three-dimensional DNA spherical shell. Reprinted with permission from Ref. [73]. Copyright 2011 AAAS. e A clamp using DNA origami to test molecular force. Reprinted with permission from Ref. [75]. Copyright 2016 AAAS. f The motor consists of two rigid DNA origami arms. Reprinted with permission from Ref. [76]. Copyright 2023 Springer Nature. g Synthesizing DNA nanoshells on cell membranes. Reprinted with permission from Ref. [78]. Copyright 2023 ACS Publication. h A glycan hairpin. Reprinted with permission from Ref. [80]. Copyright 2023 Springer Nature
Similar content being viewed by others
Data Availability
Manuscript has no associated data.
References
D. Rus, M.T. Tolley, Design, fabrication and control of origami robots. Nat. Rev. Mater. 3(6), 101–112 (2018). https://doi.org/10.1038/s41578-018-0009-8
A.K. Brooks, S. Chakravarty, M. Ali, V.K. Yadavalli, Kirigami-inspired biodesign for applications in healthcare. Adv. Mater. 34(18), 2109550 (2022). https://doi.org/10.1002/adma.202109550
J.Y. Tao, H. Khosravi, V. Deshpande, S.Y. Li, Engineering by cuts: how kirigami principle enables unique mechanical properties and functionalities. Adv. Sci. 10(1), 2204733 (2023). https://doi.org/10.1002/advs.202204733
C.A. Fairchild, Origami design secrets: mathematical methods for an ancient art. Libr. J. 129(3), 124 (2004)
E.D. Demaine, M.L. Demaine, J.S.B. Mitchell, Folding flat silhouettes and wrapping polyhedral packages: new results in computational origami. Comp. Geom. Theor. Appl. 16(1), 3–21 (2000). https://doi.org/10.1016/S0925-7721(99)00056-5
L.A. Bowen, C.L. Grames, S.P. Magleby, L.L. Howell, R.J. Lang, A classification of action origami as systems of spherical mechanisms. J. Mech. Des. 135(11), 111008 (2013). https://doi.org/10.1115/1.4025379
S.Y. Li, H.B. Fang, S. Sadeghi, P. Bhovad, K.W. Wang, Architected origami materials: how folding creates sophisticated mechanical properties. Adv. Mater. 31(5), 1805282 (2019). https://doi.org/10.1002/adma.201805282
T. Chen, O.R. Bilal, R. Lang, C. Daraio, K. Shea, Autonomous deployment of a solar panel using elastic origami and distributed shape-memory-polymer actuators. Phys. Rev. Appl. 11(6), 064069 (2019). https://doi.org/10.1103/PhysRevApplied.11.064069
M. Meloni, J.G. Cai, Q. Zhang, D.S.H. Lee, M. Li, R.J. Ma et al., Engineering origami: a comprehensive review of recent applications, design methods, and tools. Adv. Sci. 8(13), 2000636 (2021). https://doi.org/10.1002/advs.202000636
M. Runciman, A. Darzi, G.P. Mylonas, Soft robotics in minimally invasive surgery. Soft Robot. 6(4), 423–443 (2019). https://doi.org/10.1089/soro.2018.0136
C. Py, P. Reverdy, L. Doppler, J. Bico, B. Roman, C. Baroud, Capillary origami. Phys. Fluids (2007). https://doi.org/10.1063/1.2775288
J.H. Cho, D.H. Gracias, Self-assembly of lithographically patterned nanoparticles. Nano Lett. 9(12), 4049–4052 (2009). https://doi.org/10.1021/nl9022176
A.J. Cui, Z. Liu, J.F. Li, T.H.H. Shen, X.X. Xia, Z.Y. Li et al., Directly patterned substrate-free plasmonic "nanograter’’ structures with unusual Fano resonances. Light-Sci. Appl. 4(7), e308 (2015). https://doi.org/10.1038/lsa.2015.81
J.Z. Cui, T.Y. Huang, Z.C. Luo, P. Testa, H.R. Gu, X.Z. Chen et al., Nanomagnetic encoding of shape-morphing micromachines (vol 575, pg 164, 2019). Nature (2020). https://doi.org/10.1038/s41586-019-1888-6
J. Cools, Q.R. Jin, E. Yoon, D.A. Burbano, Z.X. Luo, D. Cuypers et al., A micropatterned multielectrode shell for 3D spatiotemporal recording from live cells. Adv. Sci. 5(4), 1700731 (2018). https://doi.org/10.1002/advs.201700731
C. Renault, M.J. Anderson, R.M. Crooks, Electrochemistry in hollow-channel paper analytical devices. J. Am. Chem. Soc. 136(12), 4616–4623 (2014). https://doi.org/10.1021/ja4118544
G.M. Rodriguez, D. Trueb, J. Köser, J. Schoelkopf, M. Gullo, An origami like 3D patterned cellulose-based scaffold for bioengineering cardiovascular applications. Cellulose 30(16), 10401–10412 (2023). https://doi.org/10.1007/s10570-023-05492-2
F.S.L. Bobbert, S. Janbaz, T. van Manen, Y. Li, A.A. Zadpoor, Russian doll deployable meta-implants: fusion of kirigami, origami, and multi-stability. Mater. Des. 191, 108624 (2020). https://doi.org/10.1016/j.matdes.2020.108624
H. Suzuki, R.J. Wood, Origami-inspired miniature manipulator for teleoperated microsurgery. Nat. Mach. Intell. 2(8), 437 (2020). https://doi.org/10.1038/s42256-020-0203-4
Q.J. Ze, S. Wu, J.Z. Dai, S. Leanza, G. Ikeda, P.C. Yang et al., Spinning-enabled wireless amphibious origami millirobot. Nat. Commun. 13(1), 3118 (2022). https://doi.org/10.1038/s41467-022-30802-w
P.W.K. Rothemund, Folding DNA to create nanoscale shapes and patterns. Nature 440(7082), 297–302 (2006). https://doi.org/10.1038/nature04586
D.D. Prabhu, K. Aratsu, Y. Kitamoto, H. Ouchi, T. Ohba, M.J. Hollamby et al., Self-folding of supramolecular polymers into bioinspired topology. Sci. Adv. 4(9), eaat8466 (2018). https://doi.org/10.1126/sciadv.aat8466
M. Mittelbrunn, F. Sánchez-Madrid, Intercellular communication: diverse structures for exchange of genetic information. Nat. Rev. Mol. Cell Bio. 13(5), 328–335 (2012). https://doi.org/10.1038/nrm3335
A.M. Pappa, O. Parlak, G. Scheiblin, P. Mailley, A. Salleo, R.M. Owens, Organic electronics for point-of-care metabolite monitoring. Trends Biotechnol. 36(1), 45–59 (2018). https://doi.org/10.1016/j.tibtech.2017.10.022
T.F. Xiao, F. Wu, J. Hao, M.N. Zhang, P. Yu, L.Q. Mao, In vivo analysis with electrochemical sensors and biosensors. Anal. Chem. 89(1), 300–313 (2017). https://doi.org/10.1021/acs.analchem.6b04308
S.D. Wang, Y.D. Liu, A.W. Zhu, Y. Tian, In vivo electrochemical biosensors: recent advances in molecular design. Anal. Chem. 95(1), 388–406 (2023). https://doi.org/10.1021/acs.analchem.2c04541
L. Heinrich, D. Bennett, D. Ackerman, W. Park, J. Bogovic, N. Eckstein et al., Whole-cell organelle segmentation in volume electron microscopy. Nature 599(7883), 141 (2021). https://doi.org/10.1038/s41586-021-03977-3
C.S. Xu, S. Pang, G. Shtengel, A. Muller, A.T. Ritter, H.K. Hoffman et al., An open-access volume electron microscopy atlas of whole cells and tissues. Nature 599(7885), E5 (2021). https://doi.org/10.1038/s41586-021-04132-8
X.J. Duan, C.M. Lieber, Nanoelectronics meets biology: from new nanoscale devices for live-cell recording to 3D innervated tissues. Chem. Asian J. 8(10), 2304–2314 (2013). https://doi.org/10.1002/asia.201300630
S. Li, H. Li, Y. Lu, M. Zhou, S. Jiang, X. Du et al., Advanced textile-based wearable biosensors for healthcare monitoring. Biosensors 13(10), 909 (2023). https://doi.org/10.3390/bios13100909
Z.C. Zhang, J.F. Ou, W. Li, A. Amirfazli, Folding characteristics of membranes in capillary origami. J. Colloid Interface Sci. 630, 111–120 (2023). https://doi.org/10.1016/j.jcis.2022.10.046
X.H. Zhao, Y. Kim, Soft microbots programmed by nanomagnets. Nature 575(7781), 58 (2019). https://doi.org/10.1038/d41586-019-03363-0
S. Lim, H.W. Luan, S.W. Zhao, Y.J. Lee, Y.H. Zhang, Y.G. Huang et al., Assembly of foldable 3D microstructures using graphene hinges. Adv. Mater. 32(28), 2001303 (2020). https://doi.org/10.1002/adma.202001303
Q.R. Jin, M. Li, B. Polat, S.K. Paidi, A. Dai, A. Zhang et al., Mechanical trap surface-enhanced raman spectroscopy for three-dimensional surface molecular imaging of single live cells. Angew. Chem. Int. Edit. 56(14), 3822–3826 (2017). https://doi.org/10.1002/anie.201700695
Y. Park, C.K. Franz, H. Ryu, H.W. Luan, K.Y. Cotton, J.U. Kim et al., Three-dimensional, multifunctional neural interfaces for cortical spheroids and engineered assembloids. Sci. Adv. 7(12), eabf9153 (2021). https://doi.org/10.1126/sciadv.abf9153
S. Xu, Z. Yan, K.I. Jang, W. Huang, H.R. Fu, J. Kim et al., Assembly of micro/nanomaterials into complex, three-dimensional architectures by compressive buckling. Science 347(6218), 154–159 (2015). https://doi.org/10.1126/science.1260960
Z. Yan, M.D. Han, Y. Shi, A. Badea, Y.Y. Yang, A. Kulkarni et al., Three-dimensional mesostructures as high-temperature growth templates, electronic cellular scaffolds, and self-propelled microrobots. Proc. Natl. Acad. Sci. USA 114(45), E9455–E9464 (2017). https://doi.org/10.1073/pnas.1713805114
H. Ryu, Y. Park, H.W. Luan, G. Dalgin, K. Jeffris, H.J. Yoon et al., Transparent, compliant 3D mesostructures for precise evaluation of mechanical characteristics of organoids. Adv. Mater. 33(25), 2100026 (2021). https://doi.org/10.1002/adma.202100026
D. Jäckel, D.J. Bakkum, T.L. Russell, J. Müller, M. Radivojevic, U. Frey et al., Combination of high-density microelectrode array and patch clamp recordings to enable studies of multisynaptic integration. Sci. Rep. UK 7(1), 978 (2017). https://doi.org/10.1038/s41598-017-00981-4
Y. Cao, K. Zhang, Z. Huang, S. Li, A unique and robust physically unclonable function based on bionic tunable ion gel-gated synaptic transistors. IEEE Electr. Device Lett. (2023). https://doi.org/10.1109/LED.2023.3322165
J. Gao, C.Y. Liao, S.J. Liu, T. Xia, G.B. Jiang, Nanotechnology: new opportunities for the development of patch-clamps. J. Nanobiotechnol. 19(1), 97 (2021). https://doi.org/10.1186/s12951-021-00841-4
J. Abbott, T.Y. Ye, D. Ham, H. Park, Optimizing nanoelectrode arrays for scalable intracellular electrophysiology. Acc. Chem. Res. 51(3), 600–608 (2018). https://doi.org/10.1021/acs.accounts.7b00519
B.Z. Tian, T. Cohen-Karni, Q. Qing, X.J. Duan, P. Xie, C.M. Lieber, Three-dimensional, flexible nanoscale field-effect transistors as localized bioprobes. Science 329(5993), 830–834 (2010). https://doi.org/10.1126/science.1192033
K. Yong, S. De, E.Y. Hsieh, J. Leem, N.R. Aluru, S. Nam, Kirigami-inspired strain-insensitive sensors based on atomically-thin materials. Mater. Today 34, 58–65 (2020). https://doi.org/10.1016/j.mattod.2019.08.013
E.K. Sackmann, A.L. Fulton, D.J. Beebe, The present and future role of microfluidics in biomedical research. Nature 507(7491), 181–189 (2014). https://doi.org/10.1038/nature13118
S. Battat, D.A. Weitz, G.M. Whitesides, An outlook on microfluidics: the promise and the challenge. Lab Chip 22(3), 530–536 (2022). https://doi.org/10.1039/d1lc00731a
F. Lan, B. Demaree, N. Ahmed, A.R. Abate, Single-cell genome sequencing at ultra-high-throughput with microfluidic droplet barcoding. Nat. Biotechnol. 35(7), 640 (2017). https://doi.org/10.1038/nbt.3880
H. Liu, R.M. Crooks, Three-dimensional paper microfluidic devices assembled using the principles of origami. J. Am. Chem. Soc. 133(44), 17564–17566 (2011). https://doi.org/10.1021/ja2071779
Z.X. Lao, Y.L. Hu, D. Pan, R.Y. Wang, C.C. Zhang, J.C. Ni et al., Self-sealed bionic long microchannels with thin walls and designable nanoholes prepared by line-contact capillary-force assembly. Small 13(23), 1603957 (2017). https://doi.org/10.1002/smll.201603957
A.R. Ahmed, O.C. Gauntlett, G. Camci-Unal, Origami-inspired approaches for biomedical applications. ACS Omega 6(1), 46–54 (2021). https://doi.org/10.1021/acsomega.0c05275
A. Abbott, Cell culture: biology’s new dimension. Nature 424(6951), 870–872 (2003). https://doi.org/10.1038/424870a
L. Moroni, J.A. Burdick, C. Highley, S.J. Lee, Y. Morimoto, S. Takeuchi et al., Biofabrication strategies for 3D in vitro models and regenerative medicine (vol 3, pg 21, 2018). Nat. Rev. Mater. (2018). https://doi.org/10.1038/s41578-018-0020-01
E. Cukierman, R. Pankov, D.R. Stevens, K.M. Yamada, Taking cell-matrix adhesions to the third dimension. Science 294(5547), 1708–1712 (2001). https://doi.org/10.1126/science.1064829
M. Montgomery, S. Ahadian, L.D. Huyer, M. Lo Rito, R.A. Civitarese, R.D. Vanderlaan et al., Flexible shape-memory scaffold for minimally invasive delivery of functional tissues. Nat. Mater. 16(10), 1038 (2017). https://doi.org/10.1038/Nmat4956
M.R. Visetsouk, E.J. Falat, R.J. Garde, J.L. Wendlick, J.H. Gutzman, Basal epithelial tissue folding is mediated by differential regulation of microtubules. Development 145(22), dev167031 (2018). https://doi.org/10.1242/dev.167031
A. Roy, Z.H. Zhang, M.K. Eiken, A.L. Shi, A. Pena-Francesch, C. Loebel, Programmable tissue folding patterns in structured hydrogels. Adv. Mater. (2023). https://doi.org/10.1002/adma.202300017
K. Kuribayashi, K. Tsuchiya, Z. You, D. Tomus, M. Umemoto, T. Ito et al., Self-deployable stent grafts as a biomedical application of Ni-rich TiNi shape memory alloy foil. Mater. Sci. Eng. A-Struct. 419(1–2), 131–137 (2006). https://doi.org/10.1016/j.msea.2005.12.016
S.J. Wu, H. Yuk, J.J. Wu, C.S. Nabzdyk, X.H. Zhao, A multifunctional origami patch for minimally invasive tissue sealing. Adv. Mater. 33(11), 2007667 (2021). https://doi.org/10.1002/adma.202007667
X. Mei, D.S. Zhu, J.L. Li, K. Huang, S.Q. Hu, Z.H. Li et al., A fluid-powered refillable origami heart pouch for minimally invasive delivery of cell therapies in rats and pigs. Med-Cambridge 2(11), 1253 (2021). https://doi.org/10.1016/j.medj.2021.10.001
F.S.L. Bobbert, S. Janbaz, A.A. Zadpoor, Towards deployable meta-implants. J. Mater. Chem. B 6(21), 3449–3455 (2018). https://doi.org/10.1039/c8tb00576a
G. Camci-Unal, A. Laromaine, E. Hong, R. Derda, G.M. Whitesides, Biomineralization guided by paper templates. Sci. Rep.-UK (2016). https://doi.org/10.1038/srep27693
Y.M. Huang, V. Fitzpatrick, N. Zheng, R. Cheng, H.Y. Huang, C. Ghezzi et al., Self-folding 3D silk biomaterial rolls to facilitate axon and bone regeneration. Adv. Healthc. Mater. (2020). https://doi.org/10.1002/adhm.202000530
J.C. Breger, C. Yoon, R. Xiao, H.R. Kwag, M.O. Wang, J.P. Fisher et al., Self-folding thermo-magnetically responsive soft microgrippers. Acs Appl. Mater. Inter. 7(5), 3398–3405 (2015). https://doi.org/10.1021/am508621s
Q.T. Aten, B.D. Jensen, S.H. Burnett, L.L. Howell, A self-reconfiguring metamorphic nanoinjector for injection into mouse zygotes. Rev. Sci. Instrum. (2014). https://doi.org/10.1063/1.4872077
M. Sitti, H. Ceylan, W.Q. Hu, J. Giltinan, M. Turan, S. Yim et al., Biomedical applications of untethered mobile milli/microrobots. Proc. IEEE 103(2), 205–224 (2015). https://doi.org/10.1109/Jproc.2014.2385105
Q.J. Ze, S. Wu, J. Nishikawa, J.Z. Dai, Y. Sun, S. Leanza et al., Soft robotic origami crawler. Sci. Adv. (2022). https://doi.org/10.1126/sciadv.abm7834
L.S. Novelino, Q.J. Ze, S. Wu, G.H. Paulino, R.K. Zhao, Untethered control of functional origami microrobots with distributed actuation. Proc. Natl. Acad. Sci. U.S.A. 117(39), 24096–24101 (2020). https://doi.org/10.1073/pnas.2013292117
H.R. Gu, M. Möckli, C. Ehmke, M. Kim, M. Wieland, S. Moser et al., Self-folding soft-robotic chains with reconfigurable shapes and functionalities. Nat. Commun. 14(1), 1263 (2023). https://doi.org/10.1038/s41467-023-36819-z
T. Li, X.M. Lu, M.R. Zhang, K. Hu, Z. Li, Peptide-based nanomaterials: self-assembly, properties and applications. Bioact. Mater. 11, 268–282 (2022). https://doi.org/10.1016/j.bioactmat.2021.09.029
M. Abbas, Q.L. Zou, S.K. Li, X.H. Yan, Self-assembled peptide- and protein-based nanomaterials for antitumor photodynamic and photothermal therapy. Adv. Mater. (2017). https://doi.org/10.1002/adma.201605021
Y.T. Zhang, R.M. Malamakal, D.M. Chenoweth, A single stereodynamic center modulates the rate of self-assembly in a biomolecular system. Angew. Chem. Int. Edit. 54(37), 10826–10832 (2015). https://doi.org/10.1002/anie.201504459
E.S. Andersen, M. Dong, M.M. Nielsen, K. Jahn, R. Subramani, W. Mamdouh et al., Self-assembly of a nanoscale DNA box with a controllable lid. Nature 459(7243), 73-U5 (2009). https://doi.org/10.1038/nature07971
D.R. Han, S. Pal, J. Nangreave, Z.T. Deng, Y. Liu, H. Yan, DNA origami with complex curvatures in three-dimensional space. Science 332(6027), 342–6 (2011). https://doi.org/10.1126/science.1202998
S.M. Douglas, H. Dietz, T. Liedl, B. Högberg, F. Graf, W.M. Shih, Self-assembly of DNA into nanoscale three-dimensional shapes. Nature 459(7245), 414–8 (2009). https://doi.org/10.1038/nature08016
P.C. Nickels, B. Wünsch, P. Holzmeister, W. Bae, L.M. Kneer, D. Grohmann et al., Molecular force spectroscopy with a DNA origami-based nanoscopic force clamp. Science 354(6310), 305–7 (2016). https://doi.org/10.1126/science.aah5974
M. Centola, E. Poppleton, S. Ray, M. Centola, R. Welty, J. Valero et al., A rhythmically pulsing leaf-spring DNA-origami nanoengine that drives a passive follower. Nat. Nanotechnol. (2023). https://doi.org/10.1038/s41565-023-01516-x
W. Siti, H.L. Too, T. Anderson, X.R. Liu, I.Y. Loh, Z.S. Wang, Autonomous DNA molecular motor tailor-designed to navigate DNA origami surface for fast complex motion and advanced nanorobotics. Sci. Adv. 9(38), eadi8444 (2023). https://doi.org/10.1126/sciadv.adi8444
W.T. Wang, P.R. Hayes, X. Ren, R.E. Taylor, Synthetic cell armor made of DNA origami. Nano Lett. 23(15), 7076–85 (2023). https://doi.org/10.1021/acs.nanolett.3c01878
I. Seitz, S. Saarinen, E.P. Kumpula, D. McNeale, E. Anaya-Plaza, V. Lampinen et al., DNA-origami-directed virus capsid polymorphism. Nat. Nanotechnol. 18(10), 1205 (2023). https://doi.org/10.1038/s41565-023-01443-x
G. Fittolani, T. Tyrikos-Ergas, A. Poveda, Y. Yu, N. Yadav, P.H. Seeberger et al., Synthesis of a glycan hairpin. Nat. Chem. 15(10), 1461 (2023). https://doi.org/10.1038/s41557-023-01255-5
Acknowledgements
This work was supported by the National Key Research and Development Program of China under Grant No. 2021YFA1401103; The National Science Fund for Distinguished Young Scholars under Grant No. 61825403; The National Natural Science Foundation of China under Grants No. 61921005 and 61674078.
Author information
Authors and Affiliations
Corresponding authors
Ethics declarations
Conflict of interest
The authors declare no conflict of interest.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Wu, J., Guo, X., Pan, X. et al. Origami-Kirigami Structures and Its Applications in Biomedical Devices. Biomedical Materials & Devices (2024). https://doi.org/10.1007/s44174-024-00168-2
Received:
Accepted:
Published:
DOI: https://doi.org/10.1007/s44174-024-00168-2