Abstract
Organophosphate compounds are widely used in agricultural activities to optimize food production. Contamination of field soil by these compounds may result in detrimental effects on soil biota. The aim of the present study was to isolate microorganisms from field soils and evaluate the strains on ability to degrade organophosphates as single isolate and as a consortium. Isolated strains were identified using both biochemical and molecular techniques. Results revealed that, out of the 46 isolated strains, three isolates herein referred to as S6, S36 and S37 showed an average diazinon degradation rate of 76.4%, 76.7% and 76.8% respectively, of the initial dose (50 ppm) within 11 days of incubation in mineral medium. Notably, isolates S36 and S37 were more effective than S6 in degrading diazinon by 40% in soil aliquot after 11 days and therefore were evaluated on biochemical reactions and molecular identification. The isolates showed variable biochemical characteristics. However, both isolates possessed catalase enzyme, but lacked oxidase enzyme. Molecular characterization showed that, the closest species for S36 and S37 were Priestia megaterium and P. arybattia, respectively, based on 16S rRNA gene similarity (> 99%). Combination of the strains increased diazinon degradation ability by 45% compared to single strain treatment. Chlorpyrifos was the most highly degraded organophosphate, compared to phorate and cadusafos. Therefore it is expected that the pesticide-degrading bacteria could be a solution to soil health improvement and contribution to the production of safe agricultural products.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
As a result of modern farming practices, the levels of organic compounds in soil have increased tremendously over the past decades. Among the most highly utilized and detected organic compounds in field soil are the pesticides as integral component of agriculture (Chowdhury et al. 2014). Agrochemicals have being envisaged as one of the key tools for increasing agricultural intensification in producing food and feed globally (Carvalho 2017). However, the strategy has often raised human health, environment and soil degradation concerns ensuing the destruction of the natural self-regulating ecosystem (Ahmed et al. 2021).
One of the highly utilized agrochemicals in farming are the organophosphates (OPs) pesticides, which occupies a significant proportion of the chemical market share (Market Data Forecast 2020). This is attributed to their profound efficacies in managing crop pests among farmers (Arjmandi et al. 2010). Yet, accumulation of these OPs have been apparent in crop fields at unnaturally high concentrations predisposing crop and crop products to residue accumulation (Kang et al. 2015). In addition, the chemical class has been reported as the most toxic substance and habitually associated with endocrine disruption (Aguilar-Garduño et al. 2013), cancer (Catherine et al. 2016) and hypospadias (Michalis et al. 2014), nervous disorders (Lee et al. 2021) following exposure. Indeed, diazinon is among the most detected organophosphate in crop products despite its ban in some countries (Sapbamrer and Hongsibsong 2014). Diazinon residue in crops above allowable limit (0.01 mg kg−1 body weight) have been reported in a number of developed countries (Rahimi et al. 2015). Moreover, extensive application of diazinon in field crops can result in accumulation of the chemical in soil and could be absorbed by growing crop during growing period (Wang et al. 2019, b).
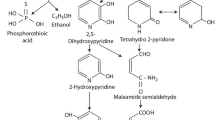
The fate of diazinon in the environment is apparent and includes oxidative decomposition to diazoxon, which rapidly hydrolyze to oxypyrimidine; a more mobile and persistent molecule in the environment than the parent material. The chemical may also be volatilized, absorbed by plants or undergo microbial breakdown (Harper et al. 2009). The hazardous aspects of diazinon and diazoxon to human include; endocrine disruption (ElMazoudy and Attia 2012), nervous disorders (Lee et al. 2021), and liver and kidney damage (Cakici and Akat 2013) and potential to impact on ecosystem balance.
Currently, there are limited approaches in existence for soil revitalization and restoration of soil biological functions. Among the possible mechanisms, is natural degradation processes that include; de-chlorination, cleavage, oxidation and reduction reactions by promiscuous enzymes (Arora and Bae 2014). Weather conditions including light, moisture, heat and humidity, also contribute to pesticides and other xenobiotics degradation or even breaking the chemicals to metabolites that may be less toxic or more harmful to biodiversity and human (Sibanda et al. 2011). Analogous to this, microbes involved in pesticides biodegradation under natural ecosystem reduce substantially, the possibilities of agrochemicals persistence in a contaminated soil (Morales et al. 2021). Under this phenomenon, chemicals and their intermediates are either detoxified or used as a new source of nutritional by the microorganisms (Hayatsu et al. 2000). However, the efficacy of the degradation process, is influenced by physio-chemical properties and residue levels of the chemicals, and duration in which the microbes exposed (Kumar et al. 2017). Thus, optimizing microbes’ bioremediation activities under in situ conditions to provide adequate exposure time for mineralizing of the agrochemicals could be a sustainable approach in revitalization of degraded soil.
Previous studies have revealed successes in bioremediation using microbiomes that form syntrophic associations. The potential of the microorganisms to break down the chemicals has been hypothesized as simple to use, low cost and eco-friendliness (Ibrahim and Essa 2010). In this regard, microorganisms that have metabolic pathway adapted to a wide range of ecological conditions and ability to degrade a number of organophosphates degradation has attracted scientific interests. For instance, Tanmaya et al. (2019) showed that, Ochrabactrum species degraded chlorpyrifos effectively within 28 days. Equally, Amani et al. (2019) and Morales et al. (2021) showed that Pseudomonas and Acinetobacter species degraded up to 80% of the original concentration of both chlorpyrifos and diazinon separately in agricultural soil. The ability of fungi including mycorrhizae to biodegrade pollutants by has also been investigated (Pathak et al. 2017). However, bacteria has gained more study interests over fungi on bioremediation ability due to their higher multiplication rate attribute than fungi (El-Ghany and Ahmed 2016). Unfortunately, bacteria may lose their biodegradation capacity over time as a result of confounding effects, including environmental conditions such as pH, temperature and chemical concentration gradient (Yadav et al. 2016) unlike fungi.
The need to explore for more efficacious microbes remain eminent to enhance tolerance to increased agrochemical residue dosages and climate change impacts. This remain a major challenge in developing microorganism assemblages for bioremediation (Classen et al. 2015). In this phenomenon, plausible explanation could be attributed to the ability of chemicals to exert evolutionary pressure allowing only the species of microorganism to survive under the lethal doses and extreme environmental conditions. Therefore, exploring more promising microorganisms for consortium build up to develop resilient microbial activities in agro-farming system is vital to optimize ‘clean up’. In this study, we therefore sought to isolate and characterize organophosphate degrading bacterial strains to optimize on soil health.
Results
Effects of diazinon on bacterial population dynamics
Results on population dynamics reveals that, increase in diazinon concentration repressed bacteria colony count across days after incubation. The colony count differed significantly among the experimental diazinon concentrations. In the first day after incubation, diazinon at the concentration of 50 and 100 ppm decreased the number of culturable bacterial cells by 50% and 73%, while in the second day, the colony count was reduced by 47% and 70% respectively. Similar results were observed 10 days after incubation. The findings of the present study further show that, in all the treatments, the greatest decline in bacterial colony was observed in the control followed by 50 ppm treatment. On average, over 2 × 108 CFU mL−1 bacterial colonies were observed on the third day, followed by a sharp decline in bacterial count by 10th day after incubation. Slight decline in population count from 9.33 × 106 CFU mL− 1 to 8.78 × 106 CFU mL−1 in soil treated with diazinon at 100 ppm concentration 10 days after incubation was evident (Table 1).
Isolation of diazinon degrading microorganisms
Following successes in diazinon degradation (Fig. 1) in the enrichment culture, a total of 46 strains were isolated. It was observed that, diazinon residue reduction was higher within the first 24 hours of exposure to soil microorganisms than the succeeding period. From the current study, only three isolates herein referred as S6, S36 and S37 amongst the 46 strains isolated from the agricultural field, were found effective and promising in degrading diazinon. Therefore, strains were used for further studies.
Diazinon degradation using isolated bacterial strains in mineral medium and soil aliquot
On average, the strains S6, S36 and S37 showed significant diazinon degradation of 49.5, 49.7 and 49.9% respectively from 1st to 11th day after incubation in the mineral medium. Degradation of diazinon was not significantly different among the isolates across the days after incubation. However, the strains depicted a varying degradation aptitude. Isolate S6 showed the highest diazinon loss (51.8%) after 24 hours but followed by decelerated rate thereafter. On the other hand, isolate S36 and S37 showed constant degradation rate of 39.8 and 39.9% respectively in the mineral medium (Fig. 2). To test efficacy of diazinon degradation in soil environment using the effective strains, an experiment was conducted in soil aliquot. Observed diazinon degradation results, followed similar trajectory as in the mineral medium (Fig. 3). On average, about 75.0% of diazinon residue was degraded from soil aliquot by the isolates discretely. The findings depicts a constant diazinon degradation rate of 45.2% by the isolates from day 0 to 7th after incubation, followed by a declined rate of 20.3% after incubating for 10 days in soil aliquot.
Degradation kinetics of diazinon by isolated bacterial strains
Degradation of diazinon and bacterial strain growth was apparent in all treatments. After 24 hours of incubation, degradation kinetics attributed to the three isolates were not significantly different (p ≤ 0.05) at lower diazinon concentration (50 ppm). After 48 hours, isolates S36 and S37 were more effective in degrading diazinon than isolate S6. Similarly, the two isolates were not significantly different and achieved the highest degradation percentage of 74% after 60 hours. However, the highest diazinon loss was observed within 48 hours (Table 2). Parallel batch experiment showed an increase in the average rate of degradation per day following increase in diazinon concentration from 10 ppm to 100 ppm. Above 50 ppm concentration, isolates S36 and S37 showed a higher average rate of degradation per day than S6 (Fig. 4). The Highest growth rate was exhibited by isolates S36 and S37 exposed in 100 ppm diazinon concentration treatment. Furthermore, the strain S6 reached stationery growth phase 24 hours earlier than the other isolates (Fig. 5), while, a more vigorous growth following increase diazinon concentrations was observed in isolate S36 (Fig. 6) and S37 (Fig. 7). At high optical density of 0.4, 0.8 and 0.84, higher degradation rate of diazinon was evident.
Morphology and biochemical characterization of isolates S36 and S37
Following the higher efficacious ability and resource at hand, isolates S36 and S37 were selected for characterization using morphometric analysis and biochemical techniques. Morphological results following microscopy suggests that, isolate S36 was creamy white and Gram variable, while S37 was creamy yellow and Gram positive. Both isolates were rod shaped, had raised colony and formed paired chain (Fig. 8). Both isolates were able to liquefy gelatin and undergo a spectrum of reactions following 20 NE API tests, revealed the results illustrated in Table 3; while carbohydrate pattern utilization test using API 50 CH are presented in Table 4. The three isolates had distinct difference in carbohydrates fermentation and reactions. Results from API 20 NE and 50 CH tests showed that, isolate S36 and S37 were similar in most reactions by 86% and 80% respectively. Isolate S36 could not utilize D- mannose and D-adipic acid. Isolate S36 was showed urease positive, while S37 responded negatively on the test. Additionally, all the strains exhibited oxidase negative and catalase positive reactions.
Molecular characterization of the diazinon degrading bacteria
Conferring to 16S rRNA gene-based phylogenetic tree, S36 and S37 were similar in most instances and belonged to Bacceliaceae, genus Priestia previously named Bacillus. The isolate S36 and S37 showed the highest sequence similarity of 99.6% and 99.2% with type of strain of species P. megaterium and P. arybhattia respectively. Subsequently, phylogenetic tree constructed with neighbour-joining group showed that isolate S36 and S37 were similar to Priestia species (Fig. 9). The closeness results are in tandem with the findings of biochemical characteristics.
Physiological studies
Isolates S36 and S37 were screened on temperature and pH regimes to establish favourable growth conditions as well as evaluate tolerance to extreme conditions. Our results suggests that, temperature and pH had significant effect on the growth of the bacterial strains. Based on growth curtailing profile, extreme temperatures were 5°C, 8°C and 50°C while optimal temperatures were observed in the range of 25°C to 40°C in both isolates. Isolate S37 showed higher growth rate under similar temperature condition than S36 and therefore seemed more stable within 48 hours. Congruently, the highest growth for isolate S36 (optical density of 0.7) was observed at 28°C while isolate S37 had a higher optical density of 1.2 at 40°C (Fig. 10). Concurrently, influence of pH on isolates growth was evident 2 hours after incubation. Maximum growth was observed in pH 7, in which the isolates S36 and S37 had optical density of 0.7 and 0.6 respectively. Favourable pH condition for growth was in the range of 8 to 10 in both isolates (neutral to alkali conditions). At pH below 4, both isolates’ growth was suppressed. At pH 5 and 6, there was no growth of new cells 10 hours after incubation Fig. 11.
Effects of isolates combination on diazinon degradation in broth medium and soil
From broth medium experiments, overall results showed that combination of the two isolates provided a stronger degradation in the first 10 days after incubation. Remarkably, the isolates S36 and S37 showed a degradation of 47.9 and 58.4% respectively 13 days after incubations, whereas, the isolates combination degraded diazinon by 69.2%. A higher degradation gradient was observed in combined isolate treatment than individual strains. However, the combination of S36 and S37 did not differ significantly from degradation showed by S37 after 13 days of incubation (Fig. 12). In soil medium combination of the two isolates, efficiently degraded diazinon by 69.7% as compared to medium inoculated with single strain, S36 and S37 that yielded 66.3 and 64.0% respectively, 20 days after incubation (Table 5).
Degradation of other organophosphates
Organophosphate degradation dynamics by the isolates are shown in Fig. 13. Results obtained suggest that all isolates significantly degraded the organophosphates. Phorate was the most highly degraded (94%) followed by cadusafos (89%) then chlorpyrifos (75%) after 17 days of incubation by the two isolates. There was no significant differences in organophosphates degradation potential from day 1 to 17th after incubation among the bacterial strains. Isolate S36 degraded chlorpyrifos, cadusafos and phorate by 72%, 88% and 95% respectively while isolate S37 reduced the chemicals by 64%, 88% and 95 respectively.
Discussion
Globally, organophosphates group is among the most widely used pesticides and often associated with irreparable consequences to human and environment. Consequently, the chemicals have attracted a considerable interests in the development of bioremediation technologies (Kaushal et al. 2021). As such, a number of pesticides degrading microorganisms have been isolated and evaluated globally to revitalize contaminated soil under different terrestrial ecosystems to restore the natural self-purifying and self-repair. Previous studies have shown that, the efficacy of the isolate varies with environmental conditions and concentration of the chemical which may limit the number of stable microorganisms capable of degrading the pesticides (Zhang et al. 2020). However, only a fraction of isolated bacterial species have been explored on and proofed effective in degrading pesticides in soil environments (Martı et al. 2004). This study therefore, endeavored to explore for bacterial species with higher degrading ability and capable of constructing a microbial consortium.
In the present study, effects of organophosphate toxicities on microorganisms’ population was evident. Increase in diazinon concentration repressed bacteria colony count across days after incubation. The decline in viable count could be attributed to the effects of diazinon toxicity on the microorganisms. Even though, there is limited literature on effects of diazinon in suppressing microbial populace, Ingram et al. (2005) and Fenlon et al. (2011) reported that, the pesticide influence urease enzyme activities of the soil dwelling organisms and mineralization process respectively. Similarly, Storck et al. (2018) found that, chlorpyrifos which has similar structure as diazinon, restrained growth of bacterial community in soil. The sharp decline in bacterial count observed from third day to 10th day after incubation could be attributed to the increased competition among the bacteria isolates over the declining resources including waning nutrients in the soil. Also, the slight decline in population count in soil amended with diazinon at 100 ppm concentration after 10 days of incubation, suggests that, diazinon was source of nutrition for the microorganisms. Our findings results of this study on effects of diazinon on microorganisms are in agreement with Wang et al. (2019a, 2019b) and (Muturi et al. 2017) report. The reduction in population could be suggesting that, certain microorganisms preferred the pesticide as source of nutrition which decreased with an increase with population. This is in support with previous studies which also hypothesized that, microbes are discerning in the source of carbon (Schrader and Blevins 2001).
Diazinon concentration was reduced by 72% by the end of 7th day in the enrichment culture in which the 46 bacterial strains were isolated. The decline in diazinon toxicity levels in the enrichment culture, further provided an insight on possibility of the presence of microbial strains utilizing the pesticide as source of food. In support of this, research findings have showed that, nutritionally, nitrogen and carbon are among the most important minerals extracted by microbes, essential for their growth and energy (Nisha et al. 2018). Correspondingly, diazinon structure provided sufficient and readily available carbons to microbes upon mineralization. The successes of the microbes to solubilize diazinon, is based on the possession of diverse mechanisms to mineralize the xenobiotic to plummet the residue levels in soil (Thabit and EL-Naggar 2013). Certainly, Drufovka et al. (2008) and Nasrollahi et al. (2020) observed that, bacteria could utilize carbon from pesticide source often reducing the chemical concentration. Our findings supported the hypothesized view that, microbial species differ in their remediation bioactivity on toxins.
Degradation kinetics showed the relationship between exponential growth phase of the bacterial cells, the period in which higher amount of energy and diazinon concentration. This relationship was also reported by Martin et al. (2003). Even though, previous studies suggested a degradation efficiency of up to 90% provided by Pseudomonas (Essa et al. 2016) and Priestia bacteria (Pailan et al. 2015) after 21 days, 74% degradation achieved within 60 hours, connotes possibility of prominent degradation in the long run. Higher growth rate exhibited by isolates S36 and S37 in 100 ppm diazinon concentration treatment suggested that, the strains tolerated the insecticide more than isolate S6. Higher growth rate exhibited by isolates S36 and S37 in 100 ppm diazinon concentration treatment suggested that, the strains tolerated the insecticide more than isolate S6. Its worthy noting that higher inoculum (higher multiplication rate) has potential for compensating for the loss of bacterial cells that reach stationery phase early (He et al. 2018). Significance of diazinon as source of carbon was demonstrated by a sharp decline in isolates growth in medium amended with diazinon at 10 ppm after 48 hours. This could be attributed death of bacterial cells following depletion of carbon source (Wang et al. 2019a, 2019b). Hence isolate S36 and S37 were selected for further study.
Following the evaluation of diazinon degradation ability attributed to the 46 bacterial strains isolated, a varying degradability aptitude in both soil aliquot and mineral media was portrayed. One of the key attribute of promising strain in bioremediation is the ability to have high stable growth rate under varying toxin concentration (Kebede et al. 2021) and thereby providing the best tool for optimization in bioremediation. Physiological studies showed delayed growth at 15°C within the first 20 hours. This could imply presence of adaptability attributes after exposure to shock temperatures. The trajectory could be due to the need for conducive temperature for the bacteria and their enzymes to growth and survive (Sartoros et al. 2005). Hugo et al. (2014) and Arbeli and Fuentes (2010) in their study concluded that the success of biodegradation depends on adaptability of the microbes to the contiguous temperature regimes. in addition, previous study suggest that, pH influence the rate of bacterial growth and pesticides degradation (Ali et al. 2014). Besides, thrilling pH and temperatures affect availability of nutrients which may jeopardize microbial growth (Gupta et al. 2015).
Our isolates showed responses in utilization of carbon sources and enzymatic activities and oftenly differing in merely fewer occasions. Even though, Elarabi et al. (2020) isolated closely related strain to S37, that could degrade glyphosate, our isolate (S27) showed negative oxidase reactions, but similar in catalase metabolism. Similarly, reactions on sorbital, ducitol, rhamnose, adonitol and D- xylose differed, signifying that, the two strains were diverse. On sequence analysis of 16S rRNA, the two isolates were found to belong to Bacceliaceae family, genus Priestia which was previously named Bacillus. Explicitly, isolates S36 and S37 showed the highest sequence similarity with P. megaterium and P. arybhattia respectively. In correspondence, a number of studies have showed that, bacterial strains in the genus Pristia are versatile in degrading pesticides including atrazine (Abd Rani et al. 2022), chlorpyrifos (Varghese et al. 2022) and benzoate (Esikova et al. 2021) using diverse metabolic pathways in mineralizing toxins and pesticides (Li et al. 2022). Moreover, Pailan et al. (2015) reported that B. arybhattia SanPS1 could tolerate chlorpyrifos and parathion at concentrations of 500 μg mL−1. Similarly, Amani et al. (2019) suggested that Acinetobacter bacteria degraded 88.25% of diazinon within 20 days in mineral medium, whilst, our strains could degrade the pesticide up to 75% within 14 days.
Combination of bacterial isolates provided a stronger degradation in the first 10 days after incubation in mineral media and soil aliquot than individual strains. The results were similar to findings (Liao and Xie 2008). It was found that, unexpectedly, combination of 2 isolates did not provide a stronger comparative rate of degradation as observed by Wójcik et al. (2009). In our views, this could be attributed to possibly of confounding factors including materials released by this bacterial isolates that may be antagonistic to each other thereby influencing their synergistic behaviour (Tyc et al. 2017). However, it’s inevitable that consortium presented a higher tendency to increase diazinon degradation than when single strain was applied.
Phorate was the most highly degraded (94%) followed by cadusafos (89%) then chlorpyrifos (75%) after 17 days of incubation. The results are in agreement with Gevao et al. (2000) and Esikova et al. (2021) preposition that, certain bacterial strains may have bioactivity against specific substance. These organophosphate loss phases could be matched with log phase and transition to stationery phase as previously described by Karpouzas and Walker (2000). This trend suggests that the isolates lag phase is short and proliferation of the strains is within 24 hours.
Conclusions
Biological degradation of pesticides using microorganisms is an efficient and low cost tool useful in removing pesticides residues and toxins from soil. We therefore aimed at developing a more stable microorganism for degrading diazinon and other organophosphate. From the present study, we conclude that, a number of bacterial strains were isolated and therefore are key in bioremediation activities. Diazinon tolerant bacterial isolates were isolated from agricultural field which belonged to genus Priestia. The strains degraded diazinon in both broth medium and soil. Degradation kinetics results showed that pesticide degradation by microbes was evident and their potential effects differ with bacterial strain. Combining of the two bacterial isolates improved degradation of diazinon by 67%. Moreover, results suggests that, still some pesticides remain in soil at the end of experiment. This means an extended period for microbial exposure to pesticides is required.
Materials and methods
Soil samples and chemicals
Soil samples were randomly collected from crop field which had potato crop previously. The soil was taken at a depth of 0–20 cm and stored at 4°C in laboratory. From composite sample, 800 g was ground, mixed thoroughly and passed through 2 mm sieve. Analytical organophosphates (diazinon, cadusafos, phorate and chlorpyrifos) grade (95% purity) were utilized in this study. Other chemical reagents with high commercial purity were sourced from local market of Kenya.
Population dynamics of soil microbes
To approximate effects of diazinon toxicity on bacterial count, an experiment was conducted using sieved soil sample (unsterilized). From the sample, 10 g were weighed and moisture adjusted to 15% of the soil moisture using deionized water. The sample was treated with diazinon at a concentration of 50 and 100 ppm and placed in 50 mL bioreactor tubes in three replicates. Control included diazinon-untreated soil. The set up was incubated at 28°C for 7 days. Gentle shaking was conducted daily manually by hands (inverting 4 times) after every 24 hours. Then, 2 g was sampled and suspended in 6 mL of mineral medium and serial diluted using saline solution (0.85% NaCl) to 10−6 and 10−7 concentration. Dropwise, 80 μL was placed at the centre of 90 mm petri dish containing R2A agar media (BD Difco, USA) and spread uniformly on the media. The plates were incubated at 28°C and data on colony count collected after 1, 2 and 10 days.
Isolation of diazinon degrading microorganisms
Microorganisms were isolated from enrichment culture prepared by mixing 5 g of with 20 mL of mineral medium. The composition of the medium was Na2HPO4.12H2O (3.58 g), anhydrous KH2P04 (1.361 g), (NH4)2SO4 (0.3 g), MgSO4 (0.05 g), CaCl2 (0.0058 g), trace minerals (10 mL), vitamin solution (1 mL) and deionized water (1 L) at pH 7 amended with diazinon at concentration of 50 ppm. The suspension was placed in shaker incubator at 28°C at 150 rpm in dark to prevent photodegradation of diazinon. Samples for high performance liquid chromatography (HPLC) analysis were collected after 4 hours and then after 1, 2 and 5 days of incubation. When more than 50% diazinon degradation was observed, 10% of the enrichment culture was serial diluted to 10−6 concentration. Soil aliquot droplet (80 μL) from the culture was spread on 90 mm diameter petri dishes containing mineral medium agar amended with diazinon (50 ppm). Colonies that differed morphologically were subsequently sub-cultured to obtain pure colonies for pesticide degradation.
Diazinon degradation in broth medium and soil aliquot using isolated strains
Successfully isolated candidate strains here in referred as S6, S36 and S37 (standardized to optical density of 1.0 at wavelength of 600 nm using spectrophotometer (Jenway 7315 spectrophotometer, Biosan)) were suspended in sterile 0.85% NaCl solution. Mineral medium (30 mL) was amended with diazinon (50 ppm) and was inoculated separately with each bacterial suspension. Additionally, soil aliquots (10 g of autoclaved soil amended with 20 mL mineral medium) were also inoculated separately with bacterial strains to test the diazinon degradation under soil based environment. Control included diazinon-uninoculated media. The set up was incubated at 28°C and residues analyzed after 4 hours and then 1, 3, 7, 10 and 11 days after incubation for broth medium while in soil aliquot, analysis was extended to 21 and 28 days after incubation.
Diazinon analysis using high performance liquid chromatograph (HPLC)
Diazinon standard was purchased from Kemidas Company limited (dissolved in acetone) whose wavelength was predetermined (192 nm). Absorbance peak and retention time were preliminarily determined before conducting residue analysis. HPLC model (Shimadzu SCL- 10 Avp HPLC System) equipped with a Shimadzu SPD- 10 Avp photodiode array (PDA) detector (Shimadzu) and C18 column (Zobrax Eclipse Xdb-18, 80 Å, 4.5 × 150 mm, 5 μm, Agilent) was used. Analysis conditions were as follows: Ratio of acetonitrile:water (70:30), oven temperature (40°C), injection volume (20 μL), flow rate (0.7 mL min−1) and UV detector (190–300 nm).
Degradation kinetics
To determine degradation kinetics, mineral medium broth was amended with diazinon at concentrations of 10, 50 and 100 ppm separately and isolates inoculated at concentration of 1 × 106 CFU mL−1 separately into the mixture. Control included media without isolate. The treatments were replicated three times and incubated at 28°C in shaking incubator (150 rpm). Analysis for diazinon residue using High performance Liquid Chromatograph analysis (HPLC) (Shimadzu) and optical density at wavelength of 600 nm (OD600) using Spectrophotometer was taken after four, 24, 48 and 60 hours after incubation. Rate of degradation of diazinon was calculated using the formula below as described by Essa et al. (2016).
Where C0 and Ct are the concentration of diazinon at t = 0 and time = t respectively while r is the rate of degradation.
Morphological and biochemical characterization
Morphological identification of the isolates S36 and S37 was conducted by visual description and Gram staining technique as described by Becerra et al. (2016). Biochemical assays were conducted using Analytical Profile Index (API 50 CH and API 20 NE kits; bioMerieux) (Ashraf et al. 2019) following manufacturer’s protocol. To determine if the strains have oxidase and catalase enzyme, presence of oxidase enzyme was tested (colour change) using filter paper soaked in presence of oxidizing reagent and isolate suspension droplet, while catalase test (air bubbles) was conducted by placing isolate drop on a glass slide and adding hydrogen peroxide dropwise.
Molecular and phylogenetic analysis
Molecular characterization was conducted following modified (Kumar et al. 2017) procedure. Deoxyribonucleic acid (DNA) was extracted using NucleoSpin® Microbial DNA protocol. Polymerase chain reaction (PCR) was performed using template of amplified 16S ribosomal RNA gene. The PCR primers included; 27f (5’- AGAGTTTGATCMTGGCTCAG-3’) and 1492r (5’- GGYTACCTTGTTACGACTT-3’). Subsequently, PCR mixture consisted of forward primer (1 μL) reverse primer (1 μL), 5X buffer (5 μL), Taq polymerase (0.25 μL), dNTP (1 μL), BSA (2.5 μL) and sterilized distilled water (13.25 μL). Precisely, 1.0 μL of template was added to make a total volume of 25 μL. PCR conditions were set as follows; denaturation (95°C for 5 minutes and further 1 minute) followed by 30 cycles of annealing (55°C), extension (72°C for 1 minute and 30 seconds) and elongation (72°C for 10 minutes) and then held at 8°C. Cleanup of PCR product was conducted following NucleoSpin® protocol. The PCR product bands were visualized using gel electrophoresis to confirm the amplification via Gel-Doc (Vilber Quarmat, Bioneer) where 1 kb ladder (NC™, NanoCell) was utilized. Sequencing was performed using SolGent co. Ltd. The raw sequences were aligned with reference sequences for phylogenetic dendrogram, which was constructed using neighbor-joining algorithm with bootstrap value of 1000 replicates in MEGA 8.0 software. The working reference sequences were downloaded using EzBioCloud database.
Physiological studies
Physiological studies were performed following Ahn et al. (2018) procedure with slight modification. Isolate concentrations were standardized (OD600= 1) and exposed to 5, 8, 15, 25, 30, 35, 40 and 50°C temperature variations for 48 hours in five replications. Optical density was measured after every 4 hours. Tolerance to pH was conducted using R2A broth adjusted to 2, 3, 4, 5, 6, 7, 8, 9, 10 and 11 pH values and plated in bioscreen microplate reader (SpectraMax 340). Isolates S36 and S37 were then inoculated (1 × 106 CFU mL−1).
Diazinon degradation by isolates S36 and S37, and their consortium in broth medium and soil
The isolates and their consortium was evaluated against diazinon degradation in broth mineral medium and soil (autoclaved at 121°C for 60 minutes) separately. Efficiency of autoclaving was tested by serial diluting 5 g of the autoclaved soil and inoculating R2A media in plated petri dish. The strains’ cells were suspended in saline solution (0.85% NaCl) and standardized to 1 × 108 CFU mL−1 and separately added as a single strain or combined into glass test tubes containing 20 mL of mineral medium amended with diazinon at 50 ppm concentration in triplicates. The set up was incubated at 28°C and samples taken after 1, 3, 7, 10, 14, 17 days of incubation for diazinon analysis. Control included non-inoculated mineral medium. Diazinon degradation in soil was performed using 500 g pre-treated with 50 ppm diazinon in acetone solution under sterile conditions, mixed and aired dried for 3 hours to evaporate acetone under flame hood. Then, 20 g was transferred to Tubespin® Bioreactor and moisture level adjusted to about 16% using deionized water. The isolates suspended in saline solution were separately inoculated into the soil to give a final concentration of 1 × 108 cells g−1 in triplicate and mixed thoroughly using a sterilized spatula. Control tubes included soil without the isolates. Moisture was adjusted and maintained to about 16 ± 2% of water holding capacity of the soil throughout the experiment and incubated in the dark at 28°C. Diazinon residue analysis was conducted by soil sampling 2 g aseptically for residue analysis after 4 hours, and then after 3, 5, 7, 10, 14, 21, and 28 days of incubation.
Degradation of other organophosphates by isolate S3 and S37
Degradation of organophosphates namely; chlorpyrifos, cadusafos and phorate was similarly conducted in mineral medium. The analytical pure grades were obtained from Sigma Aldrich Company while dissolved in acetonitrile and introduced to mineral medium broth as sole source of carbon to give a final concentration of 100 ppm. Isolates suspensions were standardized to 1 × 106 CFU mL−1 and separately inoculated to the broth medium. The treatments were replicated three times. Control included medium without isolates. The set up was incubated at 28°C and residues analyzed after 4 hours and day one, three, seven, 10, 14 and 17 after incubation and then degradation rate determined.
Statistical analysis
Data on colony count from population dynamics experiment was first transformed using log10 before conducting analysis of variance using SAS version 8.2. Similarly, data on diazinon concentration from combination test was conducted. Mean values and standard errors were calculated to separate means between and among treatments.
Availability of data and materials
The correspondence agrees to supply data whenever required by editor.
Abbreviations
- OP:
-
Organophosphate
- HPLC:
-
High Performance Liquid Chromatography
- Na2HPO4. 12H2O:
-
Disodium hydrogen phosphate dodecahydrate
- KH2PO4:
-
Potassium dihydrogen phosphate
- (NH4)2SO4 :
-
Ammonium sulphate
- MgSO4:
-
Magnesium sulphate
- CaCl2 :
-
Calcium chloride
- L:
-
Litre
- mL:
-
milliliter
- NaCl:
-
Sodium chloride
- UV:
-
ultraviolet
- Min:
-
Minute
- CFU:
-
Cell Forming Unit
- OD:
-
Optical density
- API:
-
Analytical Profile Index
- PCR:
-
Polymerase Chain Reaction
- ppm:
-
parts per million
- HSD:
-
Honest Significant Difference
References
Abd Rani NF, Ahmad Kamil K, Aris F, Mohamed Yunus N, Zakaria NA (2022) Atrazine-degrading bacteria for bioremediation strategy: a review. Biocatal Biotransfor 40:233–247. https://doi.org/10.1080/10242422.2021.2000967
Aguilar-Garduño C, Lacasaña M, Blanco-Muñoz J, Rodríguez-Barranco M, Hernández AF, Bassol S, González-Alzaga B, Cebrián ME (2013) Changes in male hormone profile after occupational organophosphate exposure A longitudinal study. GEF Bull Biosci 1:1–6. https://doi.org/10.1016/j.tox.2012.11.001
Ahmed A, Sara TA, Sundas RQ, Man-Qung W (2021) Heavy metals and pesticides toxicity in agricultural soil and plants: ecological risks and human health implications. Toxics 9(42):1–3. https://doi.org/10.3390/toxics9030042
Ahn JH, Lee SA, Kim SJ, You J, Han BH, Weon HY, Lee SW (2018) Biodegradation of organophosphorus insecticides with P–S bonds by two Sphingobium sp. strains. Int Biodeterioration and Biodegradation 132(May):59–65. https://doi.org/10.1016/j.ibiod.2018.05.006
Ali M, Kazmi AA, Ahmed N (2014) Study on effects of temperature, moisture and pH in degradation and degradation kinetics of aldrin, endosulfan, lindane pesticides during full-scale continuous rotary drum composting. Chemosphere 102:68–75. https://doi.org/10.1016/j.chemosphere.2013.12.022
Amani F, Student P, Akbar A, Sinegani S, Ebrahimi F, Nazarian S (2019) Biodegradation of chlorpyrifos and diazinon organophosphates by two bacteria isolated from contaminated agricultural soils. Biol J Microorg 7(28):27–39 (http://bjm.ui.ac.ir/article_22826_6a3890dee1b08c8e1efa308998666dab.pdf)
Arbeli Z, Fuentes CL (2010) Microbial degradation of pesticides in tropical soils. Soil Biol Agric Trop 21:251–274. https://doi.org/10.1007/978-3-642-05076-3_12
Arjmandi R, Tavakol M, Shayeghi M (2010) Determination of organophosphorus insecticide residues in the rice paddies. Int J Environ Sci Technol 7:175–182. https://doi.org/10.1007/BF03326129
Arora PK, Bae H (2014) Role of dehalogenases in aerobic bacterial degradation of chlorinated aromatic compounds. J Chem 2014:1–11. https://doi.org/10.1155/2014/157974
Ashraf MA, Arshad MI, Rahman S, Khan A (2019) Characterization of moderately thermostable α- amylase-producing Bacillus licheniformis from decaying potatoes and sweet potatoes. BioResources 13(3):4931–4945. https://doi.org/10.15376/biores.13.3.4931-4945
Becerra SC, Roy DC, Sanchez CJ, Christy RJ, Burmeister DM (2016) An optimized staining technique for the detection of gram positive and gram negative bacteria within tissue. BMC Res Notes 9(1):1–10. https://doi.org/10.1186/s13104-016-1902-0
Cakici O, Akat E (2013) Effects of oral exposure to diazinon on mice liver and kidney tissues biometric analyses of histopathologic changes. Anal Quant Cytol Histol 35(1):7–16
Carvalho FP (2017) Pesticides, environment, and food safety. Food Energy Secur 6(2):48–60. https://doi.org/10.1002/fes3.108
Catherine CL, Stella K, Gabriella A, Melissa CF, Michael C, Alavanja AB, Jane AH, Dale P, Jay H, Lubin XM, Yawei Z, Laura EB (2016) Spouses of pesticide applicators in the agricultural health study. Occup Environ Med 72(10):736–744. https://doi.org/10.1136/oemed-2014-102798.Organophosphate
Chowdhury MAZ, Jahan I, Karim N, Alam MK, Rahman MA, Moniruzzaman M, Gan SH, Fakhruddin ANM (2014) Determination of carbamate and organophosphorus pesticides in vegetable samples and the efficiency of gamma radiation in their removal. Biomed Res Int 2014:1–9. https://doi.org/10.1155/2014/145159
Classen AT, Sundqvist MK, Henning JA, Newman GS, Moore JAM, Cregger MA, Moorhead LC, Patterson CM (2015) Direct and indirect effects of climate change on soil microbial and soil microbial-plant interactions: what lies ahead? Ecosphere 6(8):1–21. https://doi.org/10.1890/ES15-00217.1
Drufovka K, Danevčič T, Trebše P, Stopar D (2008) Microorganisms trigger chemical degradation of diazinon. Int Biodeterior Biodegrad 62(3):293–296. https://doi.org/10.1016/j.ibiod.2008.02.003
Elarabi NI, Abdelhadi AA, Ahmed RH, Saleh I, Arif IA, Osman G, Ahmed DS (2020) Bacillus aryabhattai FACU : a promising bacterial strain capable of manipulate the glyphosate herbicide residues. Saudi J Biol Sci 27(9):2207–2214. https://doi.org/10.1016/j.sjbs.2020.06.050
El-Ghany A, Ahmed M (2016) Fungal biodegradation of organophosphorus insecticides and their impact on soil microbial population. J Plant Pathol Microbiol 7(5):1–7. https://doi.org/10.4172/2157-7471.1000349
ElMazoudy RH, Attia AA (2012) Endocrine-disrupting and cytotoxic potential of anticholinesterase insecticide, diazinon in reproductive toxicity of male mice. J Hazard Mater 209–210(2012):111–120. https://doi.org/10.1016/j.jhazmat.2011.12.073
Esikova TZ, Anokhina TO, Abashina TN, Suzina NE, Solyanikova IP (2021) Characterization of soil bacteria with potential to degrade benzoate and antagonistic to fungal and bacterial phytopathogens. Microorganisms 9(4):1–16. https://doi.org/10.3390/microorganisms9040755
Essa AM, Reyad A, Radwan T, Ibrahim WM (2016) Biodegradation of the organophosphorus insecticide Diazinon by Pseudomonas aeruginosa isolated from agricultural drainage ditches. Egypt J Bot 56:353–370. https://doi.org/10.21608/ejbo.2016.393
Fenlon KA, Andreou K, Jones KC, Semple KT (2011) The formation of bound residues of diazinon in four UK soils: implications for risk assessment. Environ Pollut 159(3):776–781. https://doi.org/10.1016/j.envpol.2010.11.039
Gevao B, Semple KT, Jones KC (2000) Bound pesticide residues in soils: a review. Environ Pollut 108:3–14. https://doi.org/10.1016/S0269-7491(99)00197-9
Gupta S, Pathak B, Fulekar MH (2015) Molecular approaches for biodegradation of polycyclic aromatic hydrocarbon compounds: a review. Rev Environ Sci Biotechnol 14:241–269. https://doi.org/10.1007/s11157-014-9353-3
Harper B, Luukinen B, Gervais J, Buhl K, Stone D (2009) Diazinon technical fact sheet. National Pesticide Information Center. National Pesticide Information Center, pp 1–14 (http://npic.orst.edu/factsheets/archive/diazinontech.html)
Hayatsu M, Hirano M, Tokuda S (2000) Involvement of two plasmids in fenitrothion degradation by Burkholderia sp. strain NF100. Appl Environ Microbiol 66(4):1737–1740. https://doi.org/10.1128/AEM.66.4.1737-1740.2000
He F, Zhang M, Zhang L, Hu Q (2018) Response surface methodology for the optimization of Chlorpyrifos-degrading conditions by Pseudomonas stutzeri ZH-1. Open Access Libr J 5:1–14. https://doi.org/10.4236/oalib.1104405
Hugo HJ, Mouton C, Malan AP (2014) Accelerated microbial degradation of nematicides in vineyard and orchard soils. South African J Enol Vitic 35:157–167. http://dx.doi.org/10.21548/35-1-998
Ibrahim MW, Essa MA (2010) Tolerance and utilization of organophosphorus insecticide by nitrogen fixing Cyanobacteria. Egypt J Bot 27:225–240. https://doi.org/10.1155/2014/392682
Ingram CW, Coyne MS, Williams DW (2005) Effects of commercial diazinon and imidacloprid on microbial urease activity in soil. J Environ Qual 34(5):1573–1580. https://doi.org/10.2134/jeq2004.0433
Kang N, Kim S, Kang Y, Kim D, Jang J, Won S, Hyun J, Kim D, Jeong IY, Rhee G, Shin Y, Joung DY, Kim SY, Park J, Kwon K, Ji Y (2015) Monitoring and exposure assessment of pesticide residues in domestic agricultural products. Korean J Pestic Sci 19(1):32–40. https://doi.org/10.7585/kjps.2015.19.1.32
Karpouzas DG, Walker A (2000) Factors influencing the ability of Pseudomonas putida strains epI and II to degrade the organophosphate ethoprophos. J Appl Microbiol 89:40–48. https://doi.org/10.1046/j.1365-2672.2000.01080.x
Kaushal J, Khatri M, Arya SKA (2021) Treatise on organophosphate pesticide pollution: current strategies and advancements in their environmental degradation and elimination. Ecotoxicol Environ Saf 207:111–148. https://doi.org/10.1016/j.ecoenv.2020.111483
Kebede G, Tafese T, Abda EM, Kamaraj M, Assefa F (2021) Factors influencing the bacterial bioremediation of hydrocarbon contaminants in the soil: mechanisms and impacts. J Chem 2021:1–17. https://doi.org/10.1155/2021/9823362
Kumar U, Berliner J, Adak T, Rath PC, Dey A, Pokhare SS, Jambhulkar NN, Panneerselvam P, Kumar A, Mohapatra SD (2017) Non-target effect of continuous application of chlorpyrifos on soil microbes, nematodes and its persistence under sub-humid tropical rice-rice cropping system. Ecotoxicol Environ Saf 135(2016):225–235. https://doi.org/10.1016/j.ecoenv.2016.10.003
Lee B, Park SM, Jeong S, Kim K, Jeung EB (2021) Combined exposure to diazinon and nicotine exerts a synergistic adverse effect in vitro and disrupts brain development and behaviors in vivo. Int J Mol Sci 22(14). https://doi.org/10.3390/ijms22147742
Li W, Wilkes RA, Aristilde L (2022) Effects of phosphonate herbicides on the secretions of plant-beneficial compounds by two plant growth-promoting soil bacteria: a metabolomics investigation. ACS Environ Au 2:136–149. https://doi.org/10.1021/acsenvironau.1c00030
Liao M, Xie X (2008) Effects of combination of plant and microorganism on degradation of simazine in soil. J Environ Sci 20(2):195–198. https://doi.org/10.1016/S1001-0742(08)60031-5
Market Data Forecast (2020) Organophosphate Pesticides Market Size & Growth (2021–2026). pp 1–4 (https://www.marketdataforecast.com/market-reports/organophosphate-pesticides-market)
Martı J, Sa ME, Aller A, Mora A (2004) Influence of the application of sewage sludge on the degradation of pesticides in the soil. Chemosphere 57:673–679. https://doi.org/10.1016/j.chemosphere.2004.07.023
Martin JD, Werner BG, Hotchkiss JH (2003) Effects of carbon dioxide on bacterial growth parameters in milk as measured by conductivity. J Dairy Sci 86:1932–1940. https://doi.org/10.3168/jds.s0022-0302(03)73780-1
Michalis M, Manolis NT, Leda KA, Andreas K, Tsakalofd IH (2014) Hypospadias in offspring is associated with chronic exposure of parents to organophosphate and organochlorine pesticides. Toxicol Lett 310:99–100. https://doi.org/10.1016/j.toxlet.2019.04.010
Morales M, Sentchilo V, Hadadi N, Van der Meer JR (2021) Genome-wide gene expression changes of Pseudomonas veronii 1YdBTEX2 during bioaugmentation in polluted soils. Environ Microbiomes 16(1):1–22. https://doi.org/10.1186/s40793-021-00378-x
Muturi EJ, Donthu RK, Fields CJ, Moise IK, Kim CH (2017) Effect of pesticides on microbial communities in container aquatic habitats. Sci Rep 7(2016):1–10. https://doi.org/10.1038/srep44565
Nasrollahi M, Pourbabaei AA, Etesami H, Talebi K (2020) Diazinon degradation by bacterial endophytes in rice plant (Oryzia sativa L.): a possible reason for reducing the efficiency of diazinon in the control of the rice stem–borer. Chemosphere 246:125759. https://doi.org/10.1016/j.chemosphere.2019.125759
Nisha R, Kiran B, Kaushik A, Kaushik CP (2018) Bioremediation of salt affected soils using cyanobacteria in terms of physical structure, nutrient status and microbial activity. Int J Environ Sci Technol 15:571–580. https://doi.org/10.1007/s13762-017-1419-7
Pailan S, Gupta D, Apte S, Krishnamurthi S, Saha P (2015) Degradation of organophosphate insecticide by a novel Bacillus aryabhattai strain SanPS1, isolated from soil of agricultural field in Burdwan, West Bengal. India Int Biodeterior Biodegrad 103:191–195. https://doi.org/10.1016/J.IBIOD.2015.05.006
Pathak S, Kumar P, Mishra PK, Kumar M (2017) Mycorrhiza assisted approach for bioremediation with special reference to biosorption. Pollut Res 36(2):329–332
Rahimi S, Talebi K, Torabi E, Naveh VH (2015) The dissipation kinetics of malathion in aqueous extracts of different fruits and vegetables. Environ Monit Assess 187:693. https://doi.org/10.1007/s10661-015-4865-z)
Sapbamrer R, Hongsibsong S (2014) Organophosphorus pesticide residues in vegetables from farms, markets, and a supermarket around Kwan Phayao Lake of northern Thailand. Arch Environ Contam Toxicol 67(1):60–67. https://doi.org/10.1007/s00244-014-0014-x
Sartoros C, Yerushalmi L, Béron P, Guiot SR (2005) Effects of surfactant and temperature on biotransformation kinetics of anthracene and pyrene. Chemosphere 61:1042–1050. https://doi.org/10.1016/j.chemosphere.2005.02.061
Schrader KK, Blevins WT (2001) Effects of carbon source, phosphorus concentration, and several micronutrients on biomass and geosmin production by Streptomyces halstedii. J Ind Microbiol Biotechnol 26(4):241–247. https://doi.org/10.1038/sj.jim.7000121
Sibanda MM, Focke WW, Labuschagne FJW, Moyo L, Nhlapo NS, Maity A, Muiambo H, Massinga P, Crowther NAS, Coetzee M, Brindley GWA (2011) Degradation of insecticides used for indoor spraying in malaria control and possible solutions. Malar J 10(1):307. https://doi.org/10.1186/1475-2875-10-307
Storck V, Nikolaki S, Perruchon C, Chabanis C, Sacchi A, Pertile G, Baguelin C, Karas PA, Spor A, Devers-Lamrani M, Papadopoulou ES, Sibourg O, Malandain C, Trevisan M, Ferrari F, Karpouzas DG, Tsiamis G, Martin-Laurent F (2018) Lab to field assessment of the ecotoxicological impact of chlorpyrifos, isoproturon, or tebuconazole on the diversity and composition of the soil bacterial community. Front Microbiol 9(June):1–13. https://doi.org/10.3389/fmicb.2018.01412
Tanmaya N, Panda AN, Tapan K, Adhya B, Das VR (2019) Biodegradation of Chlorpyrifos and 3,5,6-trichloro-2-pyridinol (TCP) by Ochrobactrum sp. CPD-03: insights from genome analysis on organophosphorus pesticides degradation, chemotaxis and PGPR activity. bioRxiv. https://doi.org/10.1101/2019.12.12.866210
Thabit T, El-Naggar M (2013) Malathion degradation by soil isolated bacteria and detection of degradation products by GC-MS. Int J Environ Sci 3(5):1467–1476. https://doi.org/10.6088/ijes.2013030500017
Tyc O, Song C, Dickschat JS, Vos M, Garbeva P (2017) The ecological role of volatile and soluble secondary metabolites produced by soil bacteria. Trends Microbiol 25(4):280–292. https://doi.org/10.1016/j.tim.2016.12.002
Varghese EM, Aswani P, Jisha MS (2022) Strategies in microbial degradation enhancement of chlorpyrifos–a review based on the primary approaches in soil bioremediation. Biocatal Biotransfor 40:83–94. https://doi.org/10.1080/10242422.2021.1939693
Wang P, Li H, Hassan MM, Guo Z, Zhang ZZ, Chen Q (2019) Fabricating an acetylcholinesterase modulated UCNPs-cu fluorescence biosensor for ultrasensitive detection of organophosphorus pesticides-diazinon in food. J Agric Food Chem 67(14):4071–4079. https://doi.org/10.1021/acs.jafc.8b07201
Wang X, Xia K, Yang X, Tang C (2019) Growth strategy of microbes on mixed carbon sources. Nat Commun 10(1):1–7. https://doi.org/10.1038/s41467-019-09261-3
Wójcik M, Piotrowska-seget Z, Cycon M (2009) Biodegradation of the organophosphorus insecticide diazinon by Serratia sp . and Pseudomonas sp . and their use in bioremediation of contaminated soil. Chemosphere 76:494–501. https://doi.org/10.1016/j.chemosphere.2009.03.023
Yadav M, Shukla AK, Srivastva N, Upadhyay SN, Dubey SK (2016) Utilization of microbial community potential for removal of chlorpyrifos: a review. Crit Rev Biotechnol 36(4):727–742. https://doi.org/10.3109/07388551.2015.1015958
Zhang H, Yuan X, Xiong T, Wang H, Jiang L (2020) Bioremediation of co-contaminated soil with heavy metals and pesticides: influence factors, mechanisms and evaluation methods. Chem Eng J 398:125–146. https://doi.org/10.1016/j.cej.2020.125657
Acknowledgements
I acknowledge the support from my colleagues in KALRO Tigoni staff, KCSAP and support from KAFACI projects to actualize this study.
Funding
The project funding dependent on personal savings from other projects.
Author information
Authors and Affiliations
Contributions
Dr. Susan Otieno contributed to the writing of this manuscript including reviewing.
Corresponding author
Ethics declarations
Ethics and approval consent to participate
This study did not involve participation of human being.
Consent for publication
Not applicable.
Competing interests
The author declares that there are no competing interests in this study.
Additional information
Handling Editor: Dr. Lei Xu.
Publisher’s Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Kilonzi, J.M., Otieno, S. Degradation kinetics and physiological studies of organophosphates degrading microorganisms for soil bioremediation. Stress Biology 4, 11 (2024). https://doi.org/10.1007/s44154-023-00138-6
Received:
Accepted:
Published:
DOI: https://doi.org/10.1007/s44154-023-00138-6