Abstract
Phosphoinositides are important regulatory membrane lipids, with a role in plant development and cellular function. Emerging evidence indicates that phosphoinositides play crucial roles in plant defence and are also utilized by pathogens for infection. In this review, we highlight the role of phosphoinositides in plant-pathogen interaction and the implication of this remarkable convergence in the battle against plant diseases.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Lipids derived from Phosphatidylinositol (PtdIns) i.e., phosphoinositides are an important class of cellular phospholipids which play an important role in cell-signalling pathways, actin-cytoskeletal remodelling and membrane dynamics. Further, lipid-protein interaction also forms the basis of phospholipid-based signalling in plant immunity. Here, we discuss phosphoinositides biosynthesis and mode of action, and their key functions in plant defence. This review highlights the significant findings in the field of phosphoinositides in plant-pathogen interactions and offers a fresh perspective on the importance of these macromolecules in the battle against plant diseases.
Biosynthesis of phosphoinositides in plants
In a eukaryotic cell, phosphoinositides are synthesized via the cytidine diphosphate – diacylglycerol (CDP-DAG) pathway. The nucleotide intermediate CDP-DAG reacts with free inositol to form PtdIns in the endoplasmic reticulum (ER) (Fig. 1). Phosphorylated derivatives of PtdIns are then produced by a set of phosphatidylinositol kinases (PIKs) and phosphatidylinositol phosphatases. PIKs phosphorylate the -hydroxyl (OH) group of inositol ring at positions 3′, 4′, and 5′ to generate seven phosphatidylinositol phosphates (PtdInsPs or PIPs) – three monophosphates [PtdIn(3) P, PtdIn(4) P and PtdIn(5)P], three bisphosphates [PtdIn(3,4)P2, PtdIn(4,5)P2 and PtdIn(3,5)P2] and one triphosphate [PtdIn(3,4,5)P3]. However, in plants, only five of the seven PIPs have been detected. The presence of PtdIn(3,4)P2 and PtdIn(3,4,5)P3 has not been confirmed in plants. PtdIn(4) P is the most abundant in the plant cell (Munnik and Vermeer 2010).
Pathway for the biosynthesis of phosphoinositides. Phosphoinositides are synthesized via the cytidine diphosphate–Diacylglycerol (CDP-DAG) pathway. Gro-3-P, glycerol-3-phosphate; Lyso-PA, lysophosphatidic acid; PA, phosphatidic acid; CDP-DAG, cytidine diphosphate diacylglycerol; PtdIns, phosphatidylinositol; PI4K, Phosphatidylinositol-4 kinase; PI4P or PtdIn(4)P, phosphatidylinositol-4-phosphate; PI4P5K, phosphatidylinositol 4-phosphate 5-kinase; PI(4,5)P2 or PtdIn(4,5)P2, phosphatidylinositol-4,5-bisphosphate; PS, phosphatidylserine; PE, phosphatidylethanolamine; PC, phosphatidylcholine
In plants, PtdIn(3) P and PtdIn(4) P are formed from PtdIns by the action of specific kinases: the PI3-kinase (VPS34) and 2 of PI4-kinases, namely PI4Kα, PI4Kβ (Mueller-Roeber and Pical 2002). Twelve isoforms of PI4K have been identified in Arabidopsis, which can be divided into two subfamilies, type II (PI4Kγ 1–8) and type III (PI4K α1, α2, β1 and β2) (Mueller-Roeber and Pical, 2002). The type III PI4Ks members contain a PI3K/PI4K catalytic domain and contain PtdIns-binding domains such as Pleckstrin Homology (PH) domain (in PI4Kα) or PI4K charged (PPC) domain (in PI4Kβ) (Mueller-Roeber and Pical, 2002; Stevenson-Paulik et al., 2003, Fig. 2). Further, type II PI4Ks contain PI3/4 kinase domain but lack a PtdIns-binding domain and have not been shown to synthesize PtdIn(4) P via a PtdIns-catalyzed pathway (Galvão et al., 2008). However, type II PI4Ks have been shown to phosphorylate various other proteins and only a few reports of the physiological function exist (Tang et al., 2016; Galvão et al., 2008; Liu et al., 2013; Alves-Ferreira et al., 2007). For detail on PI4Ks, we refer the readers to Szumlanski and Nielsen (2009) and Mueller-Roeber and Pical (2002). While PI3K and PI4Ks have been identified in plants, an enzyme catalyzing the generation of PtdIn(5) P directly from PtdIns has not been identified. Further, PtdIns can be dephosphorylated by phosphatases and phospholipases. Although the A. thaliana genome encodes several PtdIn phosphatases and phospholipases, only a few of these have been assigned particular catalytic or physiological functions. Important families of PtdIn phosphatases include Suppressor of Actin (SAC) phosphatases which hydrolyse phosphate group of phosphoinositides at multiple positions (Zhong and Ye, 2003), Phosphatase and Tensin homolog deleted on chromosome 10 (PTEN) related enzyme that dephosphorylates PtdIn(3) P, PtdIn(3,4)P2, and PtdIn(3,5)P2 (Pribat et al., 2012) and PtdIn-5 phosphatase (5PTase) family which hydrolyses PtdIn(4,5)P2 (Gunesekera et al., 2007). Further, PtdIn can also be cleaved by PtdIn-specific phospholipase C (PI-PLC) (Chen et al., 2011). The A. thaliana genome encodes nine isoforms of PI-PLCs (Mueller-Roeber and Pical, 2002) which are activated by Ca2+ (Pokotylo et al., 2013).
Domain architectures and multiple sequence alignment of Arabidopsis thaliana phosphatidylinositol 4- kinases. PI4Ks are characterized by a predicted catalytic domain, and are divided into three groups: alpha (α), beta (β), and gamma (γ), based on additional domains. The α and β groups, with the exception of AtPI4Kα2, have a lipid kinase unique (LKU) domain or the helical domain. The β group members have a novel homology (NH) domain, and also contain an N-terminal LKU domain. Both α and β groups contain a PtdIns-binding domain, Pleckstrin Homology (PH) and Pi4K charged (PPC) domain, respectively
Dynamics of phosphoinositides biosynthesis and distribution
PIKs and PtdIn phosphatases are responsible for the rapid and dynamic turnover and distribution of phosphoinositides (Reviewed Heilmann 2016a). The PtdIn pathway enzymes exhibit different patterns of expression, with some being expressed ubiquitously, whereas others are restricted to certain developmental stages or cell types (Heilmann, 2016b). Spatio-temporal dynamics of PtdIn change rapidly in response to various developmental and environmental cues. For instance, changes in PtdIn level have been observed in response to osmotic stress (DeWald et al., 2001; Einspahr et al., 1988a, b; Heilmann et al., 1999, 2001; König et al., 2007, 2008), heat (Mishkind et al., 2009), nutritional deficiency, developmental cues such as gravitropic curvature (Perera et al., 1999, 2001), ER stress response (Kanehara et al., 2015), and pollen tube growth (Bloch et al., 2016) (Table 1). Further, biotic stress including haustoria biogenesis (Qin et al., 2020), bacterial as well as a defence response to the fungal pathogen (Antignani et al., 2015; Qin et al., 2020) also results in changes in the PtdIn levels (Table 2). The changes in the PtdIn levels are generally associated with membrane deformation, endomembrane organization, cytoskeletal rearrangement and changes in membrane trafficking Gerth et al. 2017b; Heilmann 2016a, b; Ischebeck et al. 2008, 2011, 2013; Noack and Jaillais, 2017; Sousa et al., 2008; Tejos et al., 2014; Thole and Nielsen, 2008 König et al., 2008; Zhao et al., 2010; Mei et al., 2012, Stenzel et al., 2008, Munnik and Nielsen, 2011; Heilmann and Heilmann, 2013; Heilmann and Ischebeck, 2016).
The mechanism underlying the dynamic turnover of PtdIn under various environmental cues remains unclear. It is interesting to note that no significant change in the transcription of genes encoding enzymes for PtdIn metabolism was observed upon challenge by a wide range of stress (Zimmermann et al., 2004; Heilmann, 2016b). However, transcription of some genes such as PLC1 & PIP5K1 is induced by osmotic stress and exogenous auxin application respectively in A. thaliana (Hirayama et al., 1995, Tejos et al., 2014). Some studies indicate post-translational modifications of PtdIn-catalysing enzymes in response to stress (Westergren et al., 2001). For example, phosphorylation of PIP5K6 by mitogen-activated protein kinase (MAPK) MPK6 upon PAMP (Pathogen-associated molecular patterns) perception reduces the activity of PIP5K6 kinases (Menzel et al., 2019). Further, PIP5K6 expression on PIP5K6 protein abundance largely remained unchanged upon PAMP perception.
Modes of phosphoinositides action
In the cell, PtdIn function either as intact or hydrolyzed phosphoinositides. Intact PtdIn is found predominantly at the cytosolic face of the cellular membrane and acts as ligand for partner proteins and can influence the biophysical properties of the plasma membrane (PM, Falkenburger et al., 2010). The characteristic head groups of PtdIn with distinct phosphorylation patterns protrude into the cytosol and can bind to partner proteins containing specific phosphoinositides-recognition domains (Takenawa, 2010) such as Pleckstrin homology (PH) domains, Fab1 YOTB Vac1 EEA1 (FYVE) domains and phagocytic oxidase (PX) domains (Lemmon, 2003). The binding of a PtdIn ligand to a protein could affect the activity or function of the target protein and can result in the recruitment of a particular protein to membrane areas enriched in particular (spatial distribution) phosphoinositides. However, only some proteins having PtdIn recognition domains display a high degree of specificity for particular phosphoinositides. For example, during endocytosis, epsin NH2-terminal homology (ENTH) domain-containing proteins bind preferentially to PtdIn(4,5)P2 (Itoh & Takenawa, 2004), other regulatory proteins prefer PtdIn(4) P, while most PH domains do not exhibit such preferences and can bind to any phosphoinositides. A recent study has identified PtdIn(4)P as the main PtdIn required to establish the surface charge on the plasma membrane (Simon et al., 2016). Further, phosphatidylserine (PS) in combination with PtdIn(4)P organizes the intracellular electrostatic gradient along the endocytic pathway (Platre et al., 2018). This change in the biophysical property of the membrane controls the localization and function of many proteins. In the case of plant development, PtdIn can act as a recruiter to facilitate auxin transport. For example, D6 protein kinase (D6PK), a serine/threonine kinase which helps in PIN-mediated auxin transport, binds to both phosphoinositide and phosphatidic acid through its domain K-rich motif. Further, this binding helps to recruit D6PK towards the plasma membrane to maintain directionality in a polar manner (Barbosa et al., 2016). Another report highlighted the role of phospholipids in endomembrane reticulum (ER) - PM contact and cell-to-cell signalling. Here, MCTP (Multiple C2 domain and Transmembrane region Protein), a plasmodesmata protein binds to both ER and PM by a transmembrane region and anionic phospholipid. This enables MCTPs to respond in cell-to-cell signalling by changing to their different conformations (Brault et al., 2019). In another example, the vesicle tethering complex exocysts subunit EXO70A1 interacts with PtdIn(4)P and phosphatidic acid and recruits the exocyst to the PM, thereby driving plant cell polarity and morphogenesis (Synek et al., 2021). Further, plant-specific membrane protein REMORIN (REMs) anchors itself to the PM by its specific interaction with PtdIn(4)P (Gronnier et al., 2017).
Further, hydrolyzed PtdIns could serve as a precursor for further downstream signalling. The hydrolysis of PtdIn(4,5)P2 by PLC yields DAG and Inositol 1,4,5-triphosphate (IP3). Unlike the mammalian system, a signalling function for DAG in plants has not been reported. In plants, DAG is rapidly phosphorylated to phosphatidic acid (PA), which has been shown to have several regulating functions. Similarly, the role of soluble IP3 is only beginning to be uncovered in plants.
Role of phosphoinositides in regulating plant nuclear function
PtdIns are important regulators of cellular and physiological processes in eukaryotes. In addition to PtdIn and PtdIn-pathway enzymes in PM, their presence in the nucleus was also reported in animals, plants and yeast. The fate of the nuclear PtdIn pathway is generally governed by nuclear kinases and phosphatases rather than PM-bound PtdIn-enzymes, illustrating the novel intra-nuclear signalling pathway composed of nuclear-generated PtdIn. It is well established that the nuclear PtdIns regulate many processes including nuclear size, nuclear membrane reassembly, chromatin structure, DNA replication, localization of transcription factors, transcription, RNA splicing, mRNA export, and cell cycle progression in animal and yeast cells.
In plants, the role of PtdIns in membrane trafficking events and endomembrane organization and cytoskeletal rearrangement is well documented, however, their critical signalling role in nuclear functions is only being discovered. A few PtdIns pathway enzymes or enzyme activities and their products also have been found in the nucleus of plants. The A. thaliana PIP5K isoforms, 4, 5, 6, 9, and 11 contain possible nuclear localization sequence (NLS) sequences (by WoLF PSORT, Horton et al., 2007). GFP-tagged AtPIP5K isoform 9 has been shown to localize to both the nucleus and the PM (Lou et al., 2007). Other PtdIn pathway enzymes reported in plant nuclei include myo-inositol phosphate synthase (MIPS), PI3K, PI4K, PI4P5K, PI-PLC, Diacylglycerolkinase (DAGK) and Inositol 5-phosphatase (Dieck et al., 2012a, b). Further, the presence of phospholipids including phosphoinositides, phosphatidylserine (PS), phosphatidylethanolamine (PE) and phosphatidylcholine (PC) in plant nuclei has been confirmed (Philipp et al., 1976). Lipid profiles of Nicotiana tabacum cells indicate that PIs account for a higher molecular percentage of total lipids compared to the protoplast PM lipids (Dieck et al., 2012a, b). In another study Gerth et al. (2017a), characterize the four putative NLSs present in Arabidopsis PIP5K2. Mutation in the basic residue of NLS precludes AtPIP5K2 from importing into the nucleus. AtPIP5K2 also interacts with selected alpha-importin isoforms, an important component of nuclear-import machinery. Further, the immunofluorescence experiment confirmed the presence of AtPIP5K2 substrate, PtdIn(4)P and the reaction product PtdIn(4,5)P2 in the plant nucleus (Gerth et al., 2017a). These experiments indicate that PtdIns can be synthesised in the plant nuclei using nuclear-import of enzymes of the PtdIns pathway.
Changes in PtdIn levels during the cell cycle have been documented in plants. Induction of somatic embryogenesis in coffee plants causes a transient change in levels of PtdIn(3)P and PtdIn(4)P (Ek-Ramos et al., 2003). In a recent study, Von der Mark et al. (2022) show that the exclusion of PIP5K1 from the nucleus disrupts xylem differentiation (Von der Mark et al., 2022). Further, the expression of the human phosphatidylinositol (4)-phosphate 5-kinase (HsPIP5K) 1α in N. tabacum cells resulted in an increase in PtdIn(4,5)P2 levels in their isolated nuclei, which led to altered nuclear lipids and nuclear functions (Dieck et al., 2012a, b). The increase in PtdIn(4,5)P2 levels resulted in a significant reduction in DNA replication, histone 3 lysine 9 acetylation (H3K9ac) and phosphorylation of the retinoblastoma protein (pRb) (Dieck et al., 2012a, b). An increase in PtdIn(4,5)P2 in the nucleus of the transformed N. tabacum cells could have inhibited the DNA polymerase activity and/or Topoisomerase activity, thus slowing down DNA polymerase progression. A direct binding of human Topoisomerase IIα to PtdIn(4,5)P2 and the consequently decreased topoisomerase activity have already been reported (Lewis et al., 2011). Additionally, studies have shown PtdIns to decrease α, δ, and ε DNA polymerase activity in animals (Shoji-Kawaguchi et al., 1995). In animals, chromatin remodelling proteins are sensitive to the PtdIns pathway lipid. In one such example in plants, exogenously added PtdIn(5)P causes relocalization of histone trimethytransferase ATX1 from the nucleus to the PM and subcellular vesicles resulting in a decrease in ATX1-dependent H3K4 trimethylation levels (Alvarez-Venegas et al., 2003). Cross-talk between H3K4 trimethylation is known to enhance H3K9 acetylation in plants, hence, it is plausible that perturbation of the PtdIn(4,5)P2 levels in the cell could also alter H3K9 acetylation. Further, a role for PtdIns in regulating transcription and RNA processing has been described in the animal system. For example, PtdIn(4,5)P2 decreased the inhibition of RNA polymerase activity by H1 (Yu et al., 1998) and mutation in the nuclear PIP5K skittles in D. melanogaster lead to a decrease in transcriptionally active chromatin (Cheng & Shearn, 2004). Although the exogenous addition of PtdIn(4)P and PtdIn(5)P causes differential gene expression in A. thaliana, the role of plant nuclear phosphoinositides pathway in transcription and RNA processing remains to be described.
Role of phosphoinositides in plant-pathogen interaction
Many lipids and lipid-related compounds are known to have a role in plant defence signalling and responses to biotrophic and necrotrophic diseases. However, our knowledge of the possible roles of phosphoinositide in plant defence is limited. The role of PtdIns in biotic stress is rapidly being uncovered in bacterial, fungal and viral infections. Here, we discuss the current understanding of the direct and indirect role of PtdIns in plant immunity and various stages of pathogen infection.
Plant defence signalling
The transient over-expression of tomato PLCs in N. benthamiana enhances the biotrophic fungal pathogen Cladosporium fulvum effector protein, Avr4-triggered hypersensitive response (Abd-El-Haliem et al., 2016). Expression of Oryza sativa specific PI-PLC1(OsPI-PLC1) was induced by various chemical and biological inducers of the plant defence pathway including salicylic acid (SA), benzo-(1,2,3)-thiadiazole-7-carbothioic (BTH), an SA analogue, jasmonic acid (JA), Pseudomonas syringae pv syringae and wounding (Song & Goodman, 2002). Perception of the pathogen-associated molecular pattern (PAMP), flg22, by the pattern-recognition receptor, FLAGELLIN SENSITIVE 2 (FLS2) at the surface of A. thaliana cells, led to a decrease in the levels and reduced plasma membrane association of PtdIn(4,5)P2 (Menzel et al., 2019). In mesophyll protoplasts, flg22 triggers the phosphorylation of phosphatidylinositol 4-phosphate 5-kinase (PI4P5K) by MPK6 resulting in reduced PtdIn(4,5)P2 production and reduced catalytic activity of PI4P5K. Mutation of the MPK6-targeted residues or protoplasts from mpk6 mutants does not affect the catalytic activity of PI4P5K (Menzel et al., 2019). Further, this PI4P5K expression and protein abundance were very slightly affected by the flg22 treatment, indicating that PI4P5K activity is controlled post-translationally (Menzel et al., 2019). In another example, the recognition of two P. syringae Avr proteins, AvrRpm1 and AvrRpt2, causes biphasic accumulation of PA before cell death in Arabidopsis (Andersson et al., 2006). This increase in PA is mediated by PI-PLC and PLD. PI-PLC hydrolyses PtdIn(4)P and PtdIn(4,5)P2 to produce monophosphatidylinositol, inositol di-phosphate(IP2) and IP3 and subsequent influx of extracellular Ca2+ and in concert with DAGK produces PA. Interestingly, inhibition of phospholipase reduced PA accumulation and the severity of the hypersensitive response (HR, Andersson et al., 2006).
The application of fungal elicitor xylanase in tomato causes accumulation of PtdIn(4)P in the extracellular medium in a rapid, dose- and time-dependent manner and the exogenous application of PtdIn(4)P mimics xylanase effects, suggesting its putative role as an intercellular signalling molecule (Gonorazky et al., 2008). Transgenic Brassica napus overexpressing phospholipase C2 (BnPLC2) showed enhanced tolerance to the fungus Sclerotinia sclerotiorum (Nokhrina et al., 2014, Fig. 3). Solanum lycopersicum PLC, SlPLC4 and SlPLC6 were differentially regulated in response to infection with the pathogenic fungus Cladosporium fulvum in C. fulvum-resistant Cf-4 tomato (Vossen et al., 2010). Silencing of SlPLC4 impaired the C. fulvum effector Avr4/Cf-4 (R gene) induced HR and resulted in increased colonization of Cf-4 plants by C. fulvum. However, silencing of SlPLC6 did not affect HR but caused increased colonization by the fungus (Vossen et al., 2010, Fig. 3). Interestingly, resistance to another fungus Verticillium dahlia and bacteria P. syringae was mediated only by SlPLC6, indicating the differential requirement of PLC isoforms for the plant immune response (Vossen et al., 2010). PtdIns have also been shown to play a role in SA-dependent defence responses in A. thaliana. PUB13, a Plant U-Box (PUB) family of E3 ligase proteins and a negative regulator of SA-dependent defence responses, interact with Rab effector proteins (RabA4B) through N-terminal domains and with PtdIn(4)P through a C-terminal armadillo domain. RabA4B then recruits the closely related PI4K, PI4Kβ1 and PI4Kβ2 to repress SA-dependent gene expression and defence response (Fig. 3). Mutations in PUB13 and PI4Kβ1/PI4Kβ2 result inactivation of the SA-dependent defence gene and enhanced resistance against P. syringae pathovar tomato (Pst) DC3000 (Antignani et al., 2015).
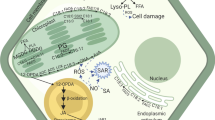
Role of phosphoinositides in fungal, oomycetes and bacterial infection in plants. a Brassica napus phospholipase C2 (BnPLC2) and Solanum lycopersicum PLC4 (SlPLC4) negatively regulate fungal infection. Solanum lycopersicum PLC6 (SlPLC6) positively regulates effector Avr4/Cf-4 (R gene) induced HR. b Phosphatidylinositol-3-phosphate (PI(3)P) mediates the entry of oomycete effector Avr3a into the host cell. c In bacterial infection, PUB13, a Plant U-Box (PUB) family of E3 ligase proteins and a negative regulator of SAdependent defense responses, interacts with Rab effector proteins (RabA4B) and PI(4)P and then recruits phosphatidylinositol 4- kinase (PI4K) to repress SA-dependent gene expression and defense response. BnPLC2, Brassica napus phospholipase C2; SlPLC4, Solanum lycopersicum PLC4; SlPLC6, Solanum lycopersicum; PI(3)P, phosphatidylinositol-3-phosphate; PUB, Plant U-Box; PI(4)P, phosphatidylinositol-4-phosphate; PI4K, phosphatidylinositol 4- kinase; Rab, Ras-related protein; HR, hypersensitive response; SA; salicylic acid; PM; plasma membrane; ER, endoplasmic reticulum; TGN, trans-golgi network; TF, transcription factor
Pathogen/effector entry
Besides mediating defence signalling, PtdIns play a role in pathogen effector entry into the host cell. Oomycete RXLR-dEER host-targeting signals, and similar signals in fungal effectors, bind to host cell surface PtdIn(3)P and mediate the entry of the effector into the host cell (Kale et al., 2010). PtdIn(3)P binding proteins and exogenous inositol phosphates inhibited the entry of the effector into the cell (Kale et al., 2010, Fig. 3). Further, the Pseudomonas sojae effectors Avr1b, Avh5, and Avh331 were reported to bind PtdIn(3)P, and binding required an intact RxLR motif (Kale et al., 2010). The effector domain of Phytophthora capsici effector protein AVR3a4 contains a conserved, positively charged patch and this region, rather than the RXLR domain, is required for binding to PtdIns in vitro (Yaeno et al., 2011). Mutations in the positively charged patch of AVR3a4 do not prevent AVR3a from being recognized by the resistance protein R3a, but they do diminish R3a’s capacity to inhibit INF1-induced cell death in plants (Yaeno et al., 2011). Since the level of PtdIn(3) is low in plants, Phytophthora species produce PtdIn(3)P to promote the entry of RxLR effectors into host cells (Lu et al., 2013). The accumulation of pathogen-derived PtdIn(3)P was detected using PtdIn(3)P-specific GFP biosensors, which could bind to P. parasitica and P. sojae hyphae during infection of N. benthamiana leaves transiently secreting the biosensors (Lu et al., 2013). The pathogen’s virulence on soybean was decreased by either silencing the PI3K genes, the treatment with LY294002, or the production of PtdIn(3)P-binding proteins, demonstrating that pathogen-synthesized PtdIn(3)P was necessary for maximal virulence. N. benthamiana leaves secreting PtdIn(3)P-binding proteins or a PI3P5K considerably enhanced resistance to infection by P. parasitica and P. capsici (Lu et al., 2013).
Colonization and hyphal growth
Lipid reporters have been used to study the role of PtdIns in polar tip growth in plants. For example, fluorescent reporters indicate that PtdIn(4,5)P2 accumulates in the apical region of the pollen tubes and root hair (Dowd et al., 2006; Tejos et al., 2014; Ischebeck et al., 2013; Helling et al., 2006; Kusano et al., 2008; Sousa et al., 2008; Stenzel et al., 2008; Ischebeck et al., 2011). A similar distribution of PtdIns has been reported during the development of intracellular branched hyphae or arbuscule of the arbuscular mycorrhizal (AM) fungi within root cortical cells of Medicago truncatula roots during symbiosis with Rhizophagus irregularis, AM fungi. PtdIn(4)P reporters accumulated all along on the periarbuscular membrane (PAM), while PtdIn(4,5)P2 reporters show accumulation at small, discrete patches on the PAM on arbuscule trunk (Ivanov and Harrison, 2019), reflecting changes in endomembrane secretion and activity.
In another example of rapid polarized membrane expansion, PtdIn(4,5)P2 accumulation was enriched in the extra-invasive hyphal membrane (EIHM) of the hemibiotrophic fungus Colletotrichum higginsianum (Ch) in Arabidopsis. Further, PIP5K, enzymes which catalyze PtdIn(4,5)P2 production also enriched at the EIHM (Shimada et al., 2019), Interestingly extra haustorial membrane (EHM) of biotrophic powdery mildew fungus did not accumulate PtdIn(4,5)P2. In contrast EHM of oomycete Hyaloperonospora arabidopsis, the causal pathogen of downy mildew was enriched with PtdIn(4)P indicating that PtdIn content of plant-pathogen interfacial membrane is specific for a pathosystem (Shimada et al., 2019). In contrast, Raushe et al. (2021), have identified a Solanum tuberosum phosphoinositide 5-phosphatase (StIPP) which specifically dephosphorylates PtdIn(4,5)P2. Upon Phytophthora infestans infection, StIPP relocalizes from PM to EHM and may antagonize PtdIn(4,5) P2-mediated plant immunity at the infection site (Raushe et al., 2021).
Intracellular movement and transmission
Lipids and proteins in the plasma membrane dynamically assemble into membrane domains giving rise to fluid molecular patchwork (Gronnier et al., 2017, Jaillais and Ott, 2020). Emerging evidence suggests that these “nanodomains” provide a dedicated biochemical and biophysical environment for cell-signalling (Gronnier et al., 2019; Jaillais and Ott, 2020). In plants, members of REMORIN (REM) protein families have been characterized as membrane markers for these “nanodomains” (Raffaele et al., 2007, 2009; Jarsch et al., 2014). The physiological functions of REMs have largely been reported in plant-virus interaction and hormone crosstalk. For example, in Solanaceae, REM organizes into a cytosolic domain of ~ 70 nm in PM and is also present in the plasmodesmata. Further, REM directly interacts with Potato virus X (PVX) movement protein TRIPLE GENE BLOCK PROTEIN 1 (TGBP1) and limits PVX cell-to-cell movement without affecting virus replication (Raffaele et al., 2007 and 2009, Fig. 4). In another example, the loss of function mutant of Arabidopsis REMORIN group 4 (AtREM4) shows reduced susceptibility to geminivirus Beet Curly Top Virus (BCTV) and Beet Severe Curly Top Virus (BSCTV) (Son et al., 2014). REMs have been shown to promote susceptibility to P. infestans (Bozkurt TO et al., 2014) and also play a positive role during the nodulation process in M truncatula (Lefebvre et al., 2010). In a recent study, Gronnier et al., (2022) showed that perception of an endogenous peptide hormone RAPID ALKALANIZATION FACTOR 23 (RALF23) by PM receptor kinase FERONIA (FER) and leucine rich repeat extension protein (LRXs) actively modulates PM nanoscale organization of other ligand-binding receptor kinase and inhibits immune signalling (Gronnier et al., 2022). To fulfil these functions, REM needs to localize to the PM and they achieve PM anchoring by interacting with PtdIn(4)P through the C-terminal domain of REM called REM-CA (REM C-terminal anchor) (Gronnier et al., 2017). Interestingly, REM-CA mutant lose their ability to restrict PVX viral intercellular movement and reduce PD permeability (Gronnier et al., 2017).
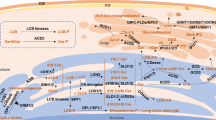
Role of phosphoinositides in virus infection in plants. a REMORINs need to localize to the PM and they achieve PM anchoring by interacting with PI(4)P, where they affect plasmodesmata (PD) size exclusion limit and limit viral cell-to cell movement. b RNA viruses such TBSV form VRCs for replication. TBSV replication protein p33 helps the recruitment of PE rich Rab5 GTPase-positive endosomes in the TBSV VRC. TBSV p33 replication protein re-localizes the yeast SNX-BAR Vps5p into VRC as a permanent component of the viral replicase complex and the binding Vps5p to PI(3)P is required for this relocalization. PI(3)P, ssDNA, single-stranded DNA; PE, Phosphatidylethanolamine; RdRP, RNA-dependent RNA polymerase, p33,Tomato bushy stunt virus (TBSV) replication protein; VRC, viral replicase complexes; SNX-BAR, Sorting nexins-Bin/Amphiphysin/Rvs; PI(3)P phosphatidylinositol-3-phosphate; PI(4)P, phosphatidylinositol-4-phosphate; PI4K, phosphatidylinositol 4- kinase; Rab, Ras-related protein; REM, REMORIN; PM; plasma membrane; ER, endoplasmic reticulum; TGN, trans-golgi network
In another example, high-affinity binding between Rice black streaked dwarf virus (RBSDV) main capsid protein, P10 and PtdIn(3,5)P2 lipid layer was observed using biolayer interferometry (BLI) and subcellular co-localization of PtdIn(3,5)P2 and P10 was observed on membranous structures in vector insect cells (Liu et al., 2022). Interestingly, P10 binds and elevates the levels of PtdIn(3,5)P2 in the insect vector Laodelphax striatellus cell. The virus induced PtdIn(3,5)P2 inhibits insect autophagy by preventing the fusion of autophagosomes and lysosomes through activation and helps the virus evade autophagic degradation in the vector (Wang et al., 2022).
Replication
The crucial role of membrane lipids in the life cycle of human, animal and plant positive-strand RNA is well established. Plant RNA viruses such as Tobacco mosaic virus (TMV), Tomato bushy stunt virus (TBSV) and Brome mosaic virus (BMV) typically harness cellular membranes for replication and viral RNA synthesis (Miller and Krijnse-Locker, 2008; den Boon et al., 2010). These viruses assemble numerous membrane-bound viral replicase complexes (VRCs) with the help of viral replication proteins and host proteins within large viral replication compartments in the cytosol of infected cells. VRCs of many animal and human viruses appear to be enriched in PtdIn and PC (Banerjee et al., 2018; Berger et al., 2011; Hsu et al., 2010; Reiss et al., 2011), whereas many plant and insect vectored animal viruses utilize PE and PC enriched membrane for the same purpose (Xu and Nagy, 2015; Zhang et al., 2016; Zhang et al., 2018). However, the role of PtdIns in plant virus replication is relatively a recent finding. A recent paper (Sasvari et al., 2020) demonstrates the enrichment of PtdIn(4)P within the replication compartment of TBSV as PtdIn(4)P partially co-localized with the TBSV replication protein p33 in yeast cells; reduction in the PtdIn(4)P level due to chemical inhibition in plant protoplasts; or sequestration of free PtdIn(4)P using PtdIn(4)P -binding protein in yeast inhibited TBSV replication (Sasvari et al., 2020). Further, the depletion of two crucial PI4P kinases, Stt4p and Pik1p, also inhibited TBSV replication in yeast cells (Sasvari et al., 2020). In the same study, (Sasvari et al., 2020) dissected the proviral function of Sac1 (suppressor of actin mutations 1-like protein). Sac1 is an ER-localized PtdIn(4)P lipid phosphatase, hence converts PtdIn(4)P to PtdIn and plays a crucial role in virus-induced membrane contact sites (MCSs) formation/function between the ER and peroxisomes, which are required for sterol/PtdIns enrichment within the replication compartment. Sac1 interacts with TBSV replication protein p33 and is present in the MCSs and TBSV replication compartments. Depletion of Sac1 inhibited the recruitment of PE-rich Rab5 GTPase-positive endosomes and the subsequent enrichment of PE in the TBSV VRC. Depletion of Sac1 also negatively affected the recruitment of syntaxin18-like Ufe1p SNARE, which play an important role in TBSV VRC formation (Sasvari et al., 2020, Fig. 4). In another study, using artificial giant unilamellar vesicles (GUVs)-based in-vitro approach to reconstitute TBSV VRCs (Kovalev et al., 2020), showed that PtdIn(3)P, a minor signalling lipid facilitates TBSV replication and incorporation of PtdIn(3)P into the GUV enhanced TBSV replication 2–3 folds (Kovalev et al., 2020). On the same lines, the deletion of Vps34, the PI3K in yeast also reduced TBSV replication (Kovalev et al., 2020). Through interaction with the p33 replication protein, Vps34 is directly recruited into the TBSV VRC and the vesicle transport activity of Vps34 is required for TBSV replication (Feng et al., 2019). In the absence of Vps34, TBSV is unable to successfully recruit PE-rich Rab5-positive early endosomes, which supply PE-rich membranes for TBSV replication compartment membrane synthesis (Feng et al., 2019). In a eukaryotic cell, Sorting nexins (SNXs) are regulators of endosomal sorting. For the SNX-BAR subgroup, a Bin/Amphiphysin/Rvs (BAR) domain is vital for the formation/stabilization of tubular subdomains that mediate cargo recycling (Kovtun et al., 2018). SNXs are recruited to specific subdomains of endosomes by binding to PtdIn(3)P via their Phox-homology (PX) domain and sensing positive membrane curvature through their banana-shaped BAR domains (Chandra et al., 2019). TBSV recruits the SNX-BAR proteins for VRC formation. In yeast, the p33 replication protein of TBSV re-localizes the yeast SNX-BAR Vps5p into VRC as a permanent component of the viral replicase complex and the binding of Vps5p to PtdIn(3)P is required for this relocalization (Feng et al., 2020). The local enrichment of PtdIn(3)P at the VRCs results in positive membrane curvature. SNX-BAR proteins sense, bind and reshape membranes into positive curvature (tube-like forms), which likely stabilizes the narrow neck structure of the VRCs (Feng et al., 2020, Fig. 4). SNX-BAR protein depletion makes the viral double-stranded (ds) RNA replication intermediate ribonuclease-sensitive, confirming the role of SNX-BAR protein and PtdIn(3)P in the stabilization of VRCs (Feng et al., 2020).
The retromer complex plays a role in various cellular processes, including autophagy and lysosome maturation. Tubular transport carriers produced from endosomes recycle cargo goes to the Golgi and ER or to the PM with the aid of the help retromer complex (Cui et al., 2019; Kovtun et al., 2018, Johannes and Wunder, 2016). The core retromer complex consists of three conserved proteins, Vps26, Vps29, and Vps35, which are involved in cargo sorting (Cui et al., 2019; Kovtun et al., 2018, Johannes and Wunder, 2016). TBSV p33 replication protein interacts with the retromer complex, including Vps26, Vps29, and Vps35 (Feng et al., 2021). The TBSV p33-driven retargeting of the retromer into VRCs results in the delivery of important cellular enzymes such as Psd2 (phosphatidylserine decarboxylase), Vps34 (yeast PI3K) and PI4Kα-like to the VRCs enabling the de novo production and enrichment of PE, PtdIn(3)P and PtdIn(4)P within the VRCs (Feng et al., 2021).
Summary
Emerging findings have greatly expanded our knowledge of the central role PtdIns play in plant-pathogen interaction. The general theme that emerges from these studies suggests the role of PtdIns in pathogen entry, recognition, replication, movement and transmission in the host. However, the data available is preliminary and fragmented, leaving important questions unanswered. The examples discussed in this review emphasize the importance of PtdIns for the pathogen life cycle, however, our current understanding of the precise molecular targets of PtdIn actions is still limited. For instance, numerous proteins with PtdIns-binding domains have been identified, making them prime candidates for regulation. Functional validation of these proteins would help link relevant cellular processes to PtdIns regulation.
Entry of pathogen effector, hyphal growth and defence mechanism such as HR would involve multiple steps of exocytosis and endocytosis. Several studies in plants confirmed the role of PtdIn in vesicle trafficking. For instance, in endocytosis, PI(4,5)P2 affects the formation of the clathrin face and in the case of exocytosis, PI(4,5)P2 interacts with exocyst subunit SEC3 facilitating the fusion of secretory vesicles (Ischebeck et al., 2013; Bloch et al., 2016).
Further, PI(4,5)P2 binding proteins such as Munc-13-1/2 and Sty1 interact with SNARE proteins for assembly at the exocytic site (Martin, 2015). PtdIns have been implicated in the entry of pathogen effector and hyphal growth, but the detailed mechanism involved in PtdIns mediated regulation of endo/exocytosis in response to pathogen attack remains to be studied.
Alteration in levels of PtdIns or PtdIn- pathway enzymes result in a reduction in viral replication (Feng et al., 2019, 2020 Sasvari et al., 2020). It is essential to recall that viruses are obligate cellular parasites and depend on the host machinery for their replication and the discussion in this review on the role of PtdIns in the regulation of nuclear function was added to put into context the emerging pieces of evidence that indicate a direct/indirect interaction of PtdIns with the components DNA replication, transcription and translational machinery (Shoji-Kawaguchi et al., 1995; Ishimaru et al., 2010; Lewis et al., 2011). It would be of special interest to study whether the nuclear function of the PtdIns directly/indirectly affects viral replication. The replication of geminiviruses, a group of plant DNA viruses, is epigenetically regulated by DNA histone methylation. It will be interesting to elucidate the cross-talk between PtdIns and epigenetic regulation of viral replication. Further, future studies are needed to determine the role of PtdIns in plant virus movement and transmission, particularly in the context of REM mediated regulation of PD permeability. Some insight into the function of the PtdIns pathway in vesicle-mediated viral transmission, cytoskeletal re-arrangement, virus movement, and transmission can also be gleaned from animal viruses.
Studies discussed in the above section clearly emphasized the role of PIs in plant-pathogen interactions, however, information on the direct role of PtdIns in regulating defence-responsive genes is limiting, although sufficient evidence exists for such regulation in mammalian systems (Zhang et al., 2019). Defence responses in plants are regulated by plant hormones and PtdIns have also been shown to play a role in SA-dependent defence responses in A. thaliana (Antignani et al., 2015). The biological relevance of PtdIn-mediated regulation of plant hormones in plant defence remains to be determined.
Finally, progress in these areas will enable the identification of important candidates and new strategies for crop protection and improvement. Rapidly involving repertoire of cutting-edge microscopy, lipidomics, proteomics and the genetic tool will help shed light on the detailed mechanism coordinating PtdIn-mediated defence response.
Availability of data and materials
Not applicable.
References
Abd-El-Haliem AM, Vossen JH, van Zeijl A, Dezhsetan S, Testerink C, Seidl MF, Beck M, Strutt J, Robatzek S, Joosten MHAJ (2016) Biochemical characterization of the tomato phosphatidylinositol-specific phospholipase C (PI-PLC) family and its role in plant immunity. Biochim Biophys Acta 1861(9):1365–1378. https://doi.org/10.1016/j.bbalip.2016.01.017
Akhter S, Uddin MN, Jeong IS, Kim DW, Liu XM, Bahk JD (2016) Role of Arabidopsis at PI 4Kγ3, a type II phosphoinositide 4-kinase, in abiotic stress responses and floral transition. Plant Biotechnol J 14(1):215–230. https://doi.org/10.1111/pbi.12376
Alvarez-Venegas R, Pien S, Sadder M, Witmer X, Grossniklaus U, Avramova Z (2003) ATX-1, an Arabidopsis homolog of Trithorax, activates flower homeotic genes. Curr Biol 13(8):627–637. https://doi.org/10.1016/S0960-9822(03)00243-4
Alves-Ferreira M, Wellmer F, Kumar V, Riechmann JL, Meyerowitz EM (2007) Global expression profiling appliedto the analysis of Arabidopsis stamen development. Plant Physiol 145(3):747–762. https://doi.org/10.1104/pp.107.104422
Andersson MX, Kourtchenko O, Dangl JL, Mackey D, Ellerström M (2006) Phospholipase-dependent signalling during the AvrRpm1- and AvrRpt2-induced disease resistance responses in Arabidopsis thaliana. Plant J 47(6):947–959. https://doi.org/10.1111/j.1365-313X.2006.02844.x
Antignani V, Klocko AL, Bak G, Chandrasekaran SD, Dunivin T, Nielsen E (2015) Recruitment of PLANT U-BOX13 and the PI4Kβ1/β2 Phosphatidylinositol-4 kinases by the small GTPase RabA4B plays important roles during salicylic acid-mediated plant defense signaling in Arabidopsis. Plant Cell 27(1):243–261. https://doi.org/10.1105/tpc.114.134262
Banerjee S, Aponte-Diaz D, Yeager C, Sharma SD, Ning G, Oh HS, Han Q, Umeda M, Hara Y, Wang RYL, Cameron CE (2018) Hijacking of multiple phospholipid biosynthetic pathways and induction of membrane biogenesis by a picornaviral 3CD protein. PLoS Pathog 14(5):e1007086. https://doi.org/10.1371/journal.ppat.1007086
Barbosa IC, Shikata H, Zourelidou M, Heilmann M, Heilmann I, Schwechheimer C (2016) Phospholipid composition and a polybasic motif determine D6 PROTEIN KINASE polar association with the plasma membrane and tropic responses. Development 143(24):4687–4700. https://doi.org/10.1242/dev.137117
Berger KL, Kelly SM, Jordan TX, Tartell MA, Randall G (2011) Hepatitis C virus stimulates the phosphatidylinositol 4-kinase III alpha-dependent phosphatidylinositol 4-phosphate production that is essential for its replication. J Virol 85(17):8870–8883. https://doi.org/10.1128/JVI.00059-11
Bloch D, Pleskot R, Pejchar P, Potocký M, Trpkošová P, Cwiklik L, Vukašinović N, Sternberg H, Yalovsky S, Žárský V (2016) Exocyst SEC3 and phosphoinositides define sites of exocytosis in pollen tube initiation and growth. Plant Physiol:00690.2016. https://doi.org/10.1104/pp.16.00690
Bozkurt TO, Richardson A, Dagdas YF, Mongrand S, Kamoun S, Raffaele S (2014) The plant membrane-associated remorin1.3 accumulates in discrete perihaustorial domains and enhances susceptibility tophytophthora infestans. Plant Physiol 165:1005–1018. https://doi.org/10.1104/pp.114.235804
Brault ML, Petit JD, Immel F, Nicolas WJ, Glavier M, Brocard L et al (2019) Multiple C2 domains and transmembrane region proteins (MCTP s) tether membranes at plasmodesmata. EMBO Rep 20(8):e47182. https://doi.org/10.15252/embr.201847182
Chandra M, Chin YK-Y, Mas C, Feathers JR, Paul B, Datta S, Chen K-E, Jia X, Yang Z, Norwood SJ, Mohanty B, Bugarcic A, Teasdale RD, Henne WM, Mobli M, Collins BM (2019) Classification of the human phox homology (PX) domains based on their phosphoinositide binding specificities. Nat Commun 10(1):1528. https://doi.org/10.1038/s41467-019-09355-y
Chen G, Snyder CL, Greer MS, Weselake RJ (2011) Biology and biochemistry of plant phospholipases. Crit Rev Plant Sci 30(3):239–258. https://doi.org/10.1080/07352689.2011.572033
Cheng MK, Shearn A (2004) The direct interaction between ASH2, a Drosophila Trithorax group protein, and SKTL, a nuclear phosphatidylinositol 4-phosphate 5-kinase, implies a role for phosphatidylinositol 4,5-bisphosphate in maintaining transcriptionally active chromatin. Genetics 167(3):1213–1223. https://doi.org/10.1534/genetics.103.018721
Contento AL, Kim SJ, Bassham DC (2004) Transcriptome profiling of the response of Arabidopsis suspension culture cells to Suc starvation. Plant Physiol 135(4):2330–2347. https://doi.org/10.1104/pp.104.044362
Cote GG, Crain RC (1993) Biochemistry of phosphoinositides. Annu Rev Plant Biol 44(1):333–356.https://doi.org/10.1146/annurev.pp.44.060193.002001
Cui Y, Carosi JM, Yang Z, Ariotti N, Kerr MC, Parton RG, Sargeant TJ, Teasdale RD (2019) Retromer has a selective function in cargo sorting via endosome transport carriers. J Cell Biol 218(2):615–631. https://doi.org/10.1083/jcb.201806153
Darwish E, Testerink C, Khalil M, El-Shihy O, Munnik T (2009) Phospholipid signaling responses in salt-stressed rice leaves. Plant Cell Physiol 50(5):986–997. https://doi.org/10.1093/pcp/pcp051
den Boon JA, Diaz A, Ahlquist P (2010) Cytoplasmic viral replication complexes. Cell Host Microbe 8(1):77–85. https://doi.org/10.1016/j.chom.2010.06.010
DeWald DB, Torabinejad J, Jones CA, Shope JC, Cangelosi AR, Thompson JE, Prestwich GD, Hama H (2001) Rapid accumulation of phosphatidylinositol 4,5-bisphosphate and inositol 1,4,5-trisphosphate correlates with calcium mobilization in salt-stressed Arabidopsis. Plant Physiol 126(2):759–769. https://doi.org/10.1104/pp.126.2.759
Dieck CB, Boss WF, Perera IY (2012a) A role for Phosphoinositides in regulating plant nuclear functions. Front Plant Sci 3. https://doi.org/10.3389/fpls.2012.00050
Dieck CB, Wood A, Brglez I, Rojas-Pierce M, Boss WF (2012b) Increasing phosphatidylinositol (4,5) bisphosphate biosynthesis affects plant nuclear lipids and nuclear functions. Plant Physiol Biochem 57:32–44. https://doi.org/10.1016/j.plaphy.2012.05.011
Dove SK, Cooke FT, Douglas MR, Sayers LG, Parker PJ, Michell RH (1997) Osmotic stress activates phosphatidylinositol-3,5-bisphosphate synthesis. Nature 390(6656):187-192. https://doi.org/10.1038/36613
Dowd PE, Coursol S, Skirpan AL, Kao T-h, Gilroy S (2006) Petunia phospholipase C1 is involved in pollen tube growth. Plant Cell 18(6):1438–1453. https://doi.org/10.1105/tpc.106.041582
Einspahr KJ, Maeda M, Thompson GA (1988a) Concurrent changes in Dunaliella salina ultrastructure and membrane phospholipid metabolism after hyperosmotic shock. J Cell Biol 107(2):529–538. https://doi.org/10.1083/jcb.107.2.529
Einspahr KJ, Peeler TC, Thompson GA (1988b) Rapid changes in polyphosphoinositide metabolism associated with the response of Dunaliella salina to hypoosmotic shock. J Biol Chem 263(12):5775–5779
Ek-Ramos MJ, Racagni-Di Palma G, Hernández-Sotomayor SMT (2003) Changes in phosphatidylinositol and phosphatidylinositol monophosphate kinase activities during the induction of somatic embryogenesis in Coffea arabica. Physiol Plant 119(2):270–277. https://doi.org/10.1034/j.1399-3054.2003.00171.x
Falkenburger BH, Jensen JB, Dickson EJ, Suh BC, Hille B (2010) Phosphoinositides: lipid regulators of membrane proteins. J Physiol 588(Pt 17):3179–3185. https://doi.org/10.1113/jphysiol.2010.192153
Feng Z, Inaba JI, Nagy PD (2021) The retromer is co-opted to deliver lipid enzymes for the biogenesis of lipid-enriched tombusviral replication organelles. Proc Natl Acad Sci U S A 118(1):e2016066118. https://doi.org/10.1073/pnas.2016066118
Feng Z, Kovalev N, Nagy PD (2020) Key interplay between the co-opted sorting nexin-BAR proteins and PI3P phosphoinositide in the formation of the tombusvirus replicase. PLoS Pathog 16(12):e1009120. https://doi.org/10.1371/journal.ppat.1009120
Feng Z, Xu K, Kovalev N, Nagy PD (2019) Recruitment of Vps34 PI3K and enrichment of PI3P phosphoinositide in the viral replication compartment is crucial for replication of a positive-strand RNA virus. PLoS Pathog 15(1):e1007530. https://doi.org/10.1371/journal.ppat.1007530
Frank W, Munnik T, Kerkmann K, Salamini F, Bartels D (2000) Water deficit triggers phospholipase D activity in the resurrection plant Craterostigma plantagineum. Plant Cell 12(1):111–123. https://doi.org/10.1105/tpc.12.1.111
Galvão RM, Kota U, Soderblom EJ, Goshe MB, Boss WF (2008) Characterization of a new family of protein kinases from Arabidopsis containing phosphoinositide 3/4-kinase and ubiquitin-like domains. Biochem J 409(1):117–127. https://doi.org/10.1042/BJ20070959
Gerth K, Lin F, Daamen F, Menzel W, Heinrich F, Heilmann M (2017a) Arabidopsis phosphatidylinositol 4-phosphate 5-kinase 2 contains a functional nuclear localization sequence and interacts with alpha-importins. Plant J Cell Mole Biol 92(5):862–878. https://doi.org/10.1111/tpj.13724
Gerth K, Lin F, Menzel W, Krishnamoorthy P, Stenzel I, Heilmann M, Heilmann I (2017b) Guilt by association: a phenotypebased view of the plant phosphoinositide network. Annu Rev Plant Biol 68:349–374. https://doi.org/10.1146/annurev-arplant-042916-041022
Gonorazky G, Laxalt AM, Testerink C, Munnik T, La Canal D, l. (2008) Phosphatidylinositol 4-phosphate accumulates extracellularly upon xylanase treatment in tomato cell suspensions. Plant Cell Environ 31(8):1051–1062. https://doi.org/10.1111/j.1365-3040.2008.01818.x
Gronnier J, Crowet JM, Habenstein B, Nasir MN, Bayle V, Hosy E et al (2017) Structural basis for plant plasma membrane protein dynamics and organization into functional nanodomains. Elife 6:e26404. https://doi.org/10.7554/elife.26404
Gronnier J, Franck CM, Stegmann M, DeFalco TA, Abarca A, von Arx M, Dünser K, Lin W, Yang Z, Kleine-Vehn J, Ringli C, Zipfel C (2022) Regulation of immune receptor kinase plasma membrane nanoscale organization by a plant peptide hormone and its receptors. eLife 11:e74162. https://doi.org/10.7554/eLife.74162
Gronnier J, Legrand A, Loquet A, Habenstein B, Germain V, Mongrand S (2019) Mechanisms governing sub compartmentalization of biological membranes. Curr Opin Plant Biol 52:114–123. https://doi.org/10.1016/j.pbi.2019.08.003
Gunesekera B, Torabinejad J, Robinson J, Gillaspy GE (2007) Inositol polyphosphate 5-phosphatases 1 and 2 are required for regulating seedling growth. Plant Physiol 143(3):1408–1417. https://doi.org/10.1104/pp.106.089474
Ha KS, Thompson GA Jr (1991) Diacylglycerol metabolism in the green alga Dunaliella salina under osmotic stress. Possible role of diacylglycerols in phospholipase C-mediated signal transduction. Plant Physiol 97(3):921–927. https://doi.org/10.1104/pp.97.3.921
Heilmann I (2016a) Plant phosphoinositide signaling - dynamics on demand. Biochim Biophys Acta 1861(9 Pt B):1345–1351. https://doi.org/10.1016/j.bbalip.2016.02.013
Heilmann I (2016b) Phosphoinositide signaling in plant development. Development 143(12):2044–2055. https://doi.org/10.1242/dev.136432
Heilmann M, Heilmann I (2013) Arranged marriage in lipid signalling? The limited choices of PtdIns(4,5)P2 in finding the right partner. Plant Biol (Stuttg) 15(5):789–797. https://doi.org/10.1111/plb.12025
Heilmann I, Ischebeck T (2016) Male functions and malfunctions: the impact of phosphoinositides on pollen development and pollen tube growth. Plant Reprod 29(1-2):3–20. https://doi.org/10.1007/s00497-015-0270-6
Heilmann I, Perera IY, Gross W, Boss WF (1999) Changes in phosphoinositide metabolism with days in culture affect signal transduction pathways in Galdieria sulphuraria. Plant Physiol 119(4):1331–1340. https://doi.org/10.1104/pp.119.4.1331
Heilmann I, Perera IY, Gross W, Boss WF (2001) Plasma membrane phosphatidylinositol 4,5-bisphosphate levels decrease with time in culture. Plant Physiol 126(4):1507–1518. https://doi.org/10.1104/pp.126.4.1507
Helling D, Possart A, Cottier S, Klahre U, Kost B (2006) Pollen tube tip growth depends on plasma membrane polarization mediated by tobacco PLC3 activity and endocytic membrane recycling. Plant Cell 18(12):3519–3534. https://doi.org/10.1105/tpc.106.047373
Hirayama T, Ohto C, Mizoguchi T, Shinozaki K (1995) A gene encodinga phosphatidylinositol-specific phospholipase C is induced by dehydration andsalt stress in Arabidopsis thaliana. Proc Natl Acad Sci U S A 92(9):3903–3907. https://doi.org/10.1073/pnas.92.9.3903
Horton P, Park K-J, Obayashi T, Fujita N, Harada H, Adams-Collier CJ, Nakai K (2007) WoLF PSORT: protein localization predictor. Nucleic Acids Res 35(Web Server):W585–W587. https://doi.org/10.1093/nar/gkm259
Hou Q, Ufer G, Bartels D (2016) Lipid signalling in plant responses to abiotic stress. Plant Cell Environ 39(5):1029–1048. https://doi.org/10.1111/pce.12666
Hsu N-Y, Ilnytska O, Belov G, Santiana M, Chen Y-H, Takvorian PM, Pau C, van der Schaar H, Kaushik-Basu N, Balla T, Cameron CE, Ehrenfeld E, van Kuppeveld FJM, Altan-Bonnet N (2010) Viral reorganization of the secretory pathway generates distinct organelles for RNA replication. Cell 141(5):799–811. https://doi.org/10.1016/j.cell.2010.03.050
Ischebeck T, Stenzel I, Heilmann I (2008) Type B phosphatidylinositol-4-phosphate 5-kinases mediate Arabidopsis and Nicotiana tabacum pollen tube growth by regulating apical pectin secretion. Plant Cell 20(12):3312–3330. https://doi.org/10.1105/tpc.108.059568
Ischebeck T, Stenzel I, Hempel F, Jin X, Mosblech A, Heilmann I (2011) Phosphatidylinositol-4, 5-bisphosphate influences Nt-Rac5-mediated cell expansion in pollen tubes of Nicotiana tabacum. Plant J 65(3):453–468. https://doi.org/10.1111/j.1365-313x.2010.04435.x
Ischebeck T, Werner S, Krishnamoorthy P, Lerche J, Meijón M, Stenzel I, Löfke C, Wiessner T, Im YJ, Perera IY (2013) Phosphatidylinositol 4,5- bisphosphate influences PIN polarization by controlling clathrin-mediated membrane trafficking in Arabidopsis. Plant Cell 25(12):4894–4911. https://doi.org/10.1105/tpc.113.116582
Ishimaru C, Takeuchi T, Sugawara F, Yoshida H, Mizushina Y (2010) Inhibitory effects of diacylglyceride phospholipids on DNA polymerase and topoisomerase activities, and human cancer cell growth. Med Chem (Shariqah (United Arab Emirates)) 6(3):114–122. https://doi.org/10.2174/1573406411006030114
Itoh T, Takenawa T (2004) Regulation of endocytosis by phosphatidylinositol 4,5-bisphosphate and ENTH proteins (pp. 31–47). https://doi.org/10.1007/978-3-642-18805-3_2
Ivanov S, Harrison MJ (2019) Accumulation of phosphoinositides in distinct regions of the periarbuscular membrane. New Phytol 221(4):2213–2227. https://doi.org/10.1111/nph.15553
Jaillais Y, Ott T (2020) The nanoscale Organization of the Plasma Membrane and its Importance in signaling: A proteolipid perspective. Plant Physiol 182:1682–1696. https://doi.org/10.1104/pp.19.01349
Jarsch IK, Konrad SS, Stratil TF, Urbanus SL, Szymanski W, Braun P, Braun KH, Ott T (2014) Plasma membranes are subcompartmentalized into a plethora of coexisting and diverse microdomains in Arabidopsis and Nicotiana benthamiana. Plant Cell 26:1698–1711. https://doi.org/10.1105/tpc.114.124446
Johannes L, Wunder C (2016) Retromer sets a trap for endosomal cargo sorting. Cell 167(6):1452–1454. https://doi.org/10.1016/j.cell.2016.11.026
Kale SD, Gu B, Capelluto DGS, Dou D, Feldman E, Rumore A, Arredondo FD, Hanlon R, Fudal I, Rouxel T, Lawrence CB, Shan W, Tyler BM (2010) External lipid PI3P mediates entry of eukaryotic pathogen effectors into plant and animal host cells. Cell 142(2):284–295. https://doi.org/10.1016/j.cell.2010.06.008
Kanehara K, Yu C-Y, Cho Y, Cheong W-F, Torta F, Shui G, Wenk MR, Nakamura Y (2015) Arabidopsis AtPLC2 is a primary phosphoinositide specific phospholipase C in phosphoinositide metabolism and the endoplasmic reticulum stress response. PLoS Genet 11:e1005511. https://doi.org/10.1371/journal.pgen.1005511
Katagiri T, Takahashi S, Shinozaki K (2001) Involvement of a novel Arabidopsis phospholipase D, AtPLDδ, in dehydration-inducible accumulation of phosphatidic acid in stress signalling. Plant J 26(6):595–605. https://doi.org/10.1046/j.1365-313x.2001.01060.x
König S, Ischebeck T, Lerche J, Stenzel I, Heilmann I (2008) Salt-stress-induced association of phosphatidylinositol 4,5-bisphosphate with clathrin-coated vesicles in plants. Biochem J 415(3):387–399. https://doi.org/10.1042/BJ20081306
König S, Mosblech A, Heilmann I (2007) Stress-inducible and constitutive phosphoinositide pools have distinctive fatty acid patterns in Arabidopsis thaliana. FASEB J 21(9):1958–1967. https://doi.org/10.1096/fj.06-7887com
Kovalev N, Pogany J, Nagy PD (2020) Reconstitution of an RNA virus replicase inartificial giant unilamellar vesicles supports fullreplication and provides protection for thedouble-stranded RNA replication intermediate. J Virol 94:e00267–e00220. https://doi.org/10.1128/JVI.00267-20
Kovtun O, Leneva N, Bykov YS, Ariotti N, Teasdale RD, Schaffer M, Engel BD, Owen DJ, Briggs JAG, Collins BM (2018) Structure of the membrane-assembled retromer coat determined by cryo-electron tomography. Nature 561(7724):561–564. https://doi.org/10.1038/s41586-018-0526-z
Kusano H, Testerink C, Vermeer JE, Tsuge T, Shimada H, Oka A, Munnik T, Aoyama T (2008) The Arabidopsis phosphatidylinositol phosphate 5-kinase PIP5K3 is a key regulator of root hair tip growth. Plant Cell 20(2):367–380. https://doi.org/10.1105/tpc.107.056119
Laxalt AM, Ter Riet B, Verdonk JC, Parigi L, Tameling WI, Vossen J et al (2001) Characterization of five tomato phospholipase D cDNAs: rapid and specific expression of LePLDβ1 on elicitation with xylanase. Plant J 26(3):237–247. https://doi.org/10.1046/j.1365-313x.2001.01023.x
Lefebvre B, Timmers T, Mbengue M, Moreau S, Herve’ C, To´ t K, Bittencourt-Silvestre J, Klaus D, Deslandes L, Godiard L, Murray JD, Udvardi MK, Raffaele S, Mongrand S, Cullimore J, Gamas P, Niebel A, Ott T (2010) A remorin protein interacts with symbiotic receptors and regulates bacterial infection. PNAS 107:2343–2348. https://doi.org/10.1073/pnas.0913320107
Lemmon MA (2003) Phosphoinositide recognition domains. Traffic 4(4):201–213. https://doi.org/10.1034/j.1600-0854.2004.00071.x
Lewis AE, Sommer L, Arntzen MØ, Strahm Y, Morrice NA, Divecha N, D’Santos CS (2011) Identification of nuclear phosphatidylinositol 4,5-bisphosphate-interacting proteins by neomycin extraction. Mol Cell Proteomics 10(2):S1–S15. https://doi.org/10.1074/mcp.M110.003376
Li L, Wang F, Yan P, Jing W, Zhang C, Kudla J, Zhang W (2017) A phosphoinositide-specific phospholipase C pathway elicits stress-induced Ca2+ signals and confers salt tolerance to rice. New Phytol 214(3):1172–1187. https://doi.org/10.1111/nph.14426
Liu H, Peck XY, Choong YK, Ng WS, Engl W, Raghuvamsi PV, Zhao ZW, Anand GS, Zhou Y, Sivaraman J, Xu Q, Wong S-M (2022) Identification of putative binding interface of PI(3,5)P2 lipid on rice black-streaked dwarf virus (RBSDV) P10 protein. Virology 570:81–95. https://doi.org/10.1016/j.virol.2022.03.010
Liu P, Xu ZS, Pan-Pan L, Hu D, Chen M, Li LC, Ma YZ (2013) A wheat PI4K gene whose product possesses threonine autophophorylation activity confers tolerance to drought and salt in Arabidopsis. J Exp Bot 64(10):2915–2927. https://doi.org/10.1093/jxb/ert133
Lou Y, Gou J-Y, Xue H-W (2007) PIP5K9, an Arabidopsis phosphatidylinositol monophosphate kinase, interacts with a cytosolic Invertase to negatively regulate sugar-mediated root growth. Plant Cell 19(1):163–181. https://doi.org/10.1105/tpc.106.045658
Lu S, Chen L, Tao K, Sun N, Wu Y, Lu X, Wang Y, Dou D (2013) Intracellular and extracellular phosphatidylinositol 3-phosphate produced by Phytophthora species is important for infection. Mol Plant 6(5):1592–1604. https://doi.org/10.1093/mp/sst047
Maarouf HE, Zuily-Fodil Y, Gareil M, d'Arcy-Lameta A, Thu Pham-Thi A (1999) Enzymatic activity and gene expression under water stress of phospholipase D in two cultivars of Vigna unguiculata L. Walp differing in drought tolerance. Plant Mole Biol 39(6):1257–1265. https://doi.org/10.1023/a:1006165919928
Mansi, Kushwaha NK, Singh AK, Karim MJ, Chakraborty S (2019) Nicotiana benthamiana phosphatidylinositol 4-kinase type II regulates chilli leaf curl virus pathogenesis. Mol Plant Pathol 20(10):1408–1424. https://doi.org/10.1111/mpp.12846
Martin TFJ (2015) PI(4,5)P2-binding effector proteins for vesicle exocytosis. Biochimica et Biophysica Acta (BBA) - molecular and cell biology of. Lipids 1851(6):785–793. https://doi.org/10.1016/j.bbalip.2014.09.017
Mei Y, Jia WJ, Chu YJ, Xue HW (2012) Arabidopsis phosphatidylinositol monophosphate 5-kinase 2 is involved in root gravitropism through regulation of polar auxin transport by affecting the cycling of PIN proteins. Cell Res 22(3):581–597. https://doi.org/10.1038/cr.2011.150
Meijer HJ, Berrie CP, Iurisci C, Divecha N, Musgrave A, Munnik T (2001) Identification of a new polyphosphoinositide in plants, phosphatidylinositol 5-monophosphate (PtdIns5P), and its accumulation upon osmotic stress. Biochem J 360(2):491–498. https://doi.org/10.1042/0264-6021:3600491
Meijer HJ, Divecha N, van den Ende H, Musgrave A, Munnik T (1999) Hyperosmotic stress induces rapid synthesis of phosphatidyl-D-inositol 3, 5-bisphosphate in plant cells. Planta 208(2):294–298. https://doi.org/10.1007/s004250050561
Menzel W, Stenzel I, Helbig L, Krishnamoorthy P, Neumann S, Eschen-Lippold L, Heilmann M, Lee J, Heilmann I (2019) A PAMP−triggered MAPK cascade inhibits phosphatidylinositol 4,5-bisphosphate production by PIP5K6 in Arabidopsis thaliana. New Phytol 224(2):833–847. https://doi.org/10.1111/nph.16069
Miller S, Krijnse-Locker J (2008) Modification of intracellular membrane structures for virus replication. Nat Rev Microbiol 6(5):363–374. https://doi.org/10.1038/nrmicro1890
Mishkind M, Vermeer JEM, Darwish E, Munnik T (2009) Heat stress activates phospholipase D and triggers PIP 2 accumulation at the plasma membrane and nucleus. Plant J 60(1):10–21. https://doi.org/10.1111/j.1365-313X.2009.03933.x
Morales JA, Gonzalez-Kantun WA, Rodriguez-Zapata LC, Ramón-Ugalde J, Castano E (2019) The effect of plant stress on phosphoinositides. Cell Biochem Funct 37(7):553–559. https://doi.org/10.1002/cbf.3432
Mueller-Roeber B, Pical C (2002) Inositol phospholipid metabolism in Arabidopsis. Characterized and putative isoforms of inositol phospholipid kinase and phosphoinositide-specific phospholipase C. Plant Physiol 130(1):22–46. https://doi.org/10.1104/pp.004770
Munnik T (2014) PI-PLC: phosphoinositide-phospholipase C in plant signaling. In: Phospholipases in plant signaling. Springer, Berlin, pp 27–54
Munnik T, Nielsen E (2011) Green light for polyphosphoinositide signals in plants. Curr Opin Plant Biol 14(5):489–497. https://doi.org/10.1016/j.pbi.2011.06.007
Munnik T, Vermeer JE (2010) Osmotic stress-induced phosphoinositide and inositol phosphate signalling in plants. Plant Cell Environ 33(4):655–669. https://doi.org/10.1111/j.1365-3040.2009.02097.x
Ndamukong I, Jones DR, Lapko H, Divecha N, Avramova Z (2010) Phosphatidylinositol 5-phosphate links dehydration stress to the activity of ARABIDOPSIS TRITHORAX-LIKE factor ATX1. PLoS One 5(10):e13396. https://doi.org/10.1371/journal.pone.0013396
Noack LC, Jaillais Y (2017) Precision targeting by phosphoinositides: how PIs direct endomembrane trafficking in plants. Curr Opin Plant Biol 40:22–33. https://doi.org/10.1016/j.pbi.2017.06.017
Nokhrina K, Ray H, Bock C, Georges F (2014) Metabolomic shifts in Brassica napus lines with enhanced BnPLC2 expression impact their response to low temperature stress and plant pathogens. GM Crops Food 5(2):120–131. https://doi.org/10.4161/gmcr.28942
Perera IY, Heilmann I, Boss WF (1999) Transient and sustained increases in inositol 1,4,5-trisphosphate precede the differential growth response in gravistimulated maize pulvini. Proc Natl Acad Sci 96(10):5838–5843. https://doi.org/10.1073/pnas.96.10.5838
Perera IY, Heilmann I, Chang SC, Boss WF, Kaufman PB (2001) A role for inositol 1,4,5-trisphosphate in Gravitropic signaling and the retention of cold-perceived gravistimulation of oat shoot Pulvini. Plant Physiol 125(3):1499–1507. https://doi.org/10.1104/pp.125.3.1499
Perera IY, Hung CY, Moore CD, Stevenson-Paulik J, Boss WF (2008) Transgenic Arabidopsis plants expressing the type 1 inositol 5-phosphatase exhibit increased drought tolerance and altered abscisic acid signaling. Plant Cell 20(10):2876–2893. https://doi.org/10.1105/tpc.108.061374
Philipp EI, Franke WW, Keenan TW, Stadler J, Jarasch ED (1976) Characterization of nuclear membranes and endoplasmic reticulum isolated from plant tissue. J Cell Biol 68, 11–29. https://doi.org/10.1083/jcb.68.1.11
Pical C, Westergren T, Dove SK, Larsson C, Sommarin M (1999) Salinity and hyperosmotic stress induce rapid increases in phosphatidylinositol 4, 5-bisphosphate, diacylglycerol pyrophosphate, and phosphatidylcholine in Arabidopsis thaliana cells. J Biol Chem 274(53):38232–38240. https://doi.org/10.1074/jbc.274.53.38232
Platre MP, Noack LC, Doumane M, Bayle V, Simon MLA, Maneta-Peyret L, Fouillen L, Stanislas T, Armengot L, Pejchar P, Caillaud MC, Potocký M, Čopič A, Moreau P, Jaillais Y (2018) A combinatorial lipid code shapes the electrostatic landscape of plant endomembranes. Dev Cell 45(4):465–480. https://doi.org/10.1016/j.devcel.2018.04.011
Pokotylo I, Pejchar P, Potocký M, Kocourková D, Krčková Z, Ruelland E, Kravets V, Martinec J (2013) The plant non-specific phospholipase C gene family. Novel competitors in lipid signalling. Prog Lipid Res 52(1):62–79. https://doi.org/10.1016/j.plipres.2012.09.001
Pribat A, Sormani R, Rousseau-Gueutin M, Julkowska MM, Testerink C, Joubès J, Castroviejo M, Laguerre M, Meyer C, Germain V, Rothan C (2012) A novel class of PTEN protein in Arabidopsis displays unusual phosphoinositide phosphatase activity and efficiently binds phosphatidic acid. Biochem J 441(1):161–171. https://doi.org/10.1042/BJ20110776
Qin L, Zhou Z, Li Q, Zhai C, Liu L, Quilichini TD, Gao P, Kessler SA, Jaillais Y, Datla R, Peng G, Xiang D, Wei Y (2020) Specific recruitment of phosphoinositide species to the plant-pathogen interfacial membrane underlies Arabidopsis susceptibility to fungal infection. Plant Cell 32(5):1665–1688. https://doi.org/10.1105/tpc.19.00970
Raffaele S, Bayer E, Lafarge D, Cluzet S, German Retana S, Boubekeur T, Leborgne-Castel N, Carde JP, Lherminier J, Noirot E, Satiat-Jeunemaıˆtre B, Laroche-Traineau J, Moreau P, Ott T, Maule AJ, Reymond P, Simon-Plas F, Farmer EE, Bessoule JJ, Mongrand S (2009) Remorin, a solanaceae protein resident in membrane rafts and plasmodesmata, impairs potato virus X movement. Plant Cell 21:1541–1555. https://doi.org/10.1105/tpc.108.064279
Raffaele S, Mongrand S, Gamas P, Niebel A, Ott T (2007) Genome-wide annotation of remorins, a plant-specific protein family: evolutionary and functional perspectives. Plant Physiol 145:593–600. https://doi.org/10.1104/pp.107.108639
Rausche J, Stenzel I, Stauder R, Fratini M, Trujillo M, Heilmann I, Rosahl S (2021) A phosphoinositide 5‐phosphatase from Solanum tuberosum is activated by PAMP‐treatment and may antagonize phosphatidylinositol 4,5‐bisphosphate at Phytophthora infestans infection sites. New Phytol 229(1):469-487. https://doi.org/10.1111/nph.16853
Reiss S, Rebhan I, Backes P, Romero-Brey I, Erfle H, Matula P, Kaderali L, Poenisch M, Blankenburg H, Hiet M-S, Longerich T, Diehl S, Ramirez F, Balla T, Rohr K, Kaul A, Bühler S, Pepperkok R, Lengauer T et al (2011) Recruitment and activation of a lipid kinase by hepatitis C virus NS5A is essential for integrity of the membranous replication compartment. Cell Host Microbe 9(1):32–45. https://doi.org/10.1016/j.chom.2010.12.002
Sang Y, Cui D, Wang X (2001) Phospholipase D and phosphatidic acid-mediated generation of superoxide in Arabidopsis. Plant Physiol 126(4):1449–1458. https://doi.org/10.1104/pp.126.4.1449
Sasvari Z, Lin W, Inaba J-I, Xu K, Kovalev N, Nagy PD (2020) Co-opted cellular Sac1 lipid phosphatase and PI(4)P phosphoinositide are key host factors during the biogenesis of the Tombusvirus replication compartment. J Virol 94(12). https://doi.org/10.1128/JVI.01979-19
Shigaki T, Bhattacharyya MK (2000) Decreased inositol 1, 4, 5-trisphosphate content in pathogen-challenged soybean cells. Mol Plant-Microbe Interact 13(5):563–567. https://doi.org/10.1094/mpmi.2000.13.5.563
Shimada TL, Betsuyaku S, Inada N, Ebine K, Fujimoto M, Uemura T et al (2019) Enrichment of phosphatidylinositol 4, 5-bisphosphate in the extra-invasive hyphal membrane promotes colletotrichum infection of Arabidopsis thaliana. Plant Cell Physiol 60(7):1514–1524. https://doi.org/10.1093/pcp/pcz058
Shoji-Kawaguchi M, Izuta S, Tamiya-Koizumi K, Suzuki M, Yoshida S (1995) Selective inhibition of DNA polymerase ε by phosphatidylinositol. J Biochem 117(5):1095–1099. https://doi.org/10.1093/oxfordjournals.jbchem.a124812
Simon ML, Platre MP, Marques-Bueno MM, Armengot L, Stanislas T, Bayle V, Caillaud MC, Jaillais Y (2016) A PtdIns(4)P-driven electrostatic field controls cell membrane identity and signalling in plants. Nat Plants 2:16089. https://doi.org/10.1038/nplants.2016.89
Son S, Oh CJ, An CS (2014) Arabidopsis thaliana remorins interact with SnRK1 and play a role in susceptibility to beet curly top virus and beet severe curly top virus. Plant Pathol J 30:269–278. https://doi.org/10.5423/PPJ.OA.06.2014.0061 PMID: 25289013
Song F, Goodman RM (2002) Molecular cloning and characterization of a rice phosphoinositide-specific phospholipase C gene, OsPI-PLC1, that is activated in systemic acquired resistance. Physiol Mol Plant Pathol 61(1):31–40. https://doi.org/10.1006/pmpp.2002.0414
Song MF, Han YZ (2008) Molecular cloning and characterization of a phosphoinositide-specific phospholipase C from Torenia fournieri. Russ J Plant Physiol 55(3):385–389. https://doi.org/10.1134/S1021443708030151
Sousa E, Kost B, Malh OR (2008) Arabidopsis phosphatidylinositol-4-monophosphate 5-kinase 4 regulates pollen tube growth and polarity by modulating membrane recycling. Plant Cell 20(11):3050–3064. https://doi.org/10.1105/tpc.108.058826
Stenzel I, Ischebeck T, K€onig S, Hołubowska A, Sporysz M, Hause B, Heilmann I (2008) The type B phosphatidylinositol-4-phosphate 5-kinase 3 is essential for root hair formation in Arabidopsis thaliana. Plant Cell 20(1):124–141. https://doi.org/10.1105/tpc.107.052852
Stevenson-Paulik J, Love J, Boss WF (2003) Differential regulation of two Arabidopsis type III phosphatidylinositol 4-kinase isoforms. A regulatory role for the Pleckstrin homology domain. Plant Physiol 132(2):1053–1064. https://doi.org/10.1104/pp.103.021758
Synek L, Pleskot R, Sekereš J, Serrano N, Vukašinović N, Ortmannová J, Klejchová M, Pejchar P, Batystová K, Gutkowska M, Janková-Drdová E, Marković V, Pečenková T, Šantrůček J, Žárský V, Potocký M (2021) Plasma membrane phospholipid signature recruits the plant exocyst complex via the EXO70A1 subunit. Proc Natl Acad Sci U S A 118(36):e2105287118. https://doi.org/10.1073/pnas.2105287118
Szumlanski A, Nielsen E (2009) Phosphatidylinositol 4-phosphate is required for tip growth in Arabidopsis thaliana. Plant Cell Monographs. https://doi.org/10.1007/978-3-642-03873-0_4
Takenawa T (2010) Phosphoinositide-binding interface proteins involved in shaping cell membranes. Proc. Jpn Acad Ser B Phys Biol Sci 86(5):509–523. https://doi.org/10.2183/pjab.86.509
Tang Y, Zhao C-Y, Tan S-T, Xue H-W (2016) Arabidopsis type II phosphatidylinositol 4-kinase PI4Kγ5 regulates auxin biosynthesis and leaf margin development through interacting with membrane-bound transcription factor ANAC078. PLoS Genetic 12(8):e1006252. https://doi.org/10.1371/journal.pgen.1006252
Tejos R, Sauer M, Vanneste S, Palacios-Gomez M, Li HJ, Heilmann M, vanWijk R, Vermeer JEM, Heilmann I, Munnik T et al (2014) Bipolar plasma membrane distribution of phosphoinositides and their requirement for auxin mediated cell polarity and patterning in Arabidopsis. Plant Cell 26:2114–2128. https://doi.org/10.1105/tpc.114.126185
Thole JM, Nielsen E (2008) Phosphoinositides in plants: novel functions in membrane trafficking. Curr Opin Plant Biol 11(6):620–631. https://doi.org/10.1016/j.pbi.2008.10.010
Toyoda, K., Shiraishi, T., Yamada, T., Ichinose, Y., & Oku, H. (1993) Rapid changes in polyphosphoinositide metabolism in pea [Pisum sativum] in response to fungal signals. Plant Cell Physiol (Japan) ISSN : 0032-0781
Van Leeuwen W, Vermeer JE, Gadella TW Jr, Munnik T (2007) Visualization of phosphatidylinositol 4, 5-bisphosphate in the plasma membrane of suspension-cultured tobacco BY-2 cells and whole Arabidopsis seedlings. Plant J 52(6):1014–1026. https://doi.org/10.1111/j.1365-313x.2007.03292.x
Von der Mark C, Cruz TMD, Blanco-Touriñan N, Rodriguez-Villalon A (2022) Bipartite phosphoinositide-dependent modulation of auxin signaling during xylem differentiation in Arabidopsis thaliana roots. The New Phytologist 236(5):1734–1747. https://doi.org/10.1111/nph.18448
Vossen JH, Abd-El-Haliem A, Fradin EF, van den Berg GCM, Ekengren SK, Meijer HJG, Seifi A, Bai Y, ten Have A, Munnik T, Thomma BPHJ, Joosten MHAJ (2010) Identification of tomato phosphatidylinositol-specific phospholipase-C (PI-PLC) family members and the role of PLC4 and PLC6 in HR and disease resistance. Plant J 62(2):224–239. https://doi.org/10.1111/j.1365-313X.2010.04136.x
Wang H, Zhang J, Liu H, Wang M, Dong Y, Zhou Y, Wong SM, Xu K, Xu Q (2022) A plant virus hijacks phosphatidylinositol-3,5-bisphosphate to escape autophagic degradation in its insect vector. Autophagy:1–16. Advance online publication. https://doi.org/10.1080/15548627.2022.2116676
Westergren T, Dove SK, Sommarin M, Pical C (2001) AtPIP5K1, an Arabidopsis thaliana phosphatidylinositol phosphate kinase, synthesizes PtdIns(3,4)P2 and PtdIns(4,5)P2 in vitro and is inhibited by phosphorylation. Biochem J 359(3):583. https://doi.org/10.1042/0264-6021:3590583
Xu K, Nagy PD (2015) RNA virus replication depends on enrichment of phosphatidylethanolamine at replication sites in subcellular membranes. Proc Natl Acad Sci 112(14). https://doi.org/10.1073/pnas.1418971112
Yaeno T, Li H, Chaparro-Garcia A, Schornack S, Koshiba S, Watanabe S, Kigawa T, Kamoun S, Shirasu K (2011) Phosphatidylinositol monophosphate-binding interface in the oomycete RXLR effector AVR3a is required for its stability in host cells to modulate plant immunity. Proc Natl Acad Sci 108(35):14682–14687. https://doi.org/10.1073/pnas.1106002108
Young SA, Wang X, Leach JE (1996) Changes in the plasma membrane distribution of rice phospholipase D during resistant interactions with Xanthomonas oryzae pv oryzae. Plant Cell 8(6):1079–1090. https://doi.org/10.1105/tpc.8.6.1079
Yu H, Fukami K, Watanabe Y, Ozaki C, Takenawa T (1998) Phosphatidylinositol 4,5-bisphosphate reverses the inhibition of RNA transcription caused by histone H1. Eur J Biochem 251(1–2):281–287. https://doi.org/10.1046/j.1432-1327.1998.2510281.x
Zhai S, Gao Q, Liu X, Sui Z, Zhang J (2013) Overexpression of a Zea mays phospholipase C1 gene enhances drought tolerance in tobacco in part by maintaining stability in the membrane lipid composition. Plant Cell, Tissue Organ Culture (PCTOC) 115(2):253–262. https://doi.org/10.1007/s11240-013-0358-3
Zhang Z, He G, Filipowicz NA, Randall G, Belov GA, Kopek BG, Wang X (2019) Host lipids in positive Strand RNA virus GenomeReplication. Front Microbiol 10:286. https://doi.org/10.3389/fmicb.2019.00286
Zhang Z, He G, Han G-S, Zhang J, Catanzaro N, Diaz A, Wu Z, Carman GM, Xie L, Wang X (2018) Host Pah1p phosphatidate phosphatase limits viral replication by regulating phospholipid synthesis. PLoS Pathog 14(4):e1006988. https://doi.org/10.1371/journal.ppat.1006988
Zhang K, Jin C, Wu L, Hou M, Dou S, Pan Y (2014) Expression analysis of a stress-related phosphoinositide-specific phospholipase C gene in wheat (Triticum aestivum L.). PLoS One 9(8):e105061. https://doi.org/10.1371/journal.pone.0105061
Zhang J, Zhang Z, Chukkapalli V, Nchoutmboube JA, Li J, Randall G, Belov GA, Wang X (2016) Positive-strand RNA viruses stimulate host phosphatidylcholine synthesis at viral replication sites. Proc Natl Acad Sci 113(8). https://doi.org/10.1073/pnas.1519730113
Zhao Y, Yan A, Feijó JA, Furutani M, Takenawa T, Hwang I, Fu Y, Yang Z (2010) Phosphoinositides Regulate Clathrin-Dependent Endocytosis at the Tip of Pollen Tubes in Arabidopsis and Tobacco. Plant Cell 22(12): 4031-4044. https://doi.org/10.1105/tpc.110.076760
Zhong R, Ye Z-H (2003) The SAC domain-containing protein gene family in Arabidopsis. Plant Physiol 132(2):544–555. https://doi.org/10.1104/pp.103.021444
Zimmermann P, Hirsch-Hoffmann M, Hennig L, Gruissem W (2004) GENEVESTIGATOR. Arabidopsis microarray database and analysis toolbox. Plant Physiol 136(1):2621–2632. https://doi.org/10.1104/pp.104.046367
Acknowledgements
Financial support from DST FIST II and DBT BUILDER to SC (SC/DBT-BUILDER/2022) is gratefully acknowledged. FZ and KK acknowledge SERB and CSIR for providing National Post-doctoral fellowship and Ph.D. scholarship, respectively.
Author information
Authors and Affiliations
Contributions
FZ, KK and SC conceptualized the study. Data acquisition was done by FZ and KK. Data analysis was performed by FZ, KK and SC. FZ prepared the original draft of the manuscript. SC, KK and FZ edited the manuscript. SC arranged the funds. The author(s) read and approved the final manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors have no competing financial interests.
Additional information
Handling editor: Rosa Lozano-duran.
Publisher’s Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Zarreen, F., Kumar, K. & Chakraborty, S. Phosphoinositides in plant-pathogen interaction: trends and perspectives. Stress Biology 3, 4 (2023). https://doi.org/10.1007/s44154-023-00082-5
Received:
Accepted:
Published:
DOI: https://doi.org/10.1007/s44154-023-00082-5