Abstract
The ability of some knotless phytochromes to photoconvert without the PHY domain allows evaluation of the distinct effect of the PHY domain on their photodynamics. Here, we compare the ms dynamics of the single GAF domain (g1) and the GAF-PHY (g1g2) construct of the knotless phytochrome All2699 from cyanobacterium Nostoc punctiforme. While the spectral signatures and occurrence of the intermediates are mostly unchanged by the domain composition, the presence of the PHY domain slows down the early forward and reverse dynamics involving chromophore and protein binding pocket relaxation. We assign this effect to a more restricted binding pocket imprinted by the PHY domain. The photoproduct formation is also slowed down by the presence of the PHY domain but to a lesser extent than the early dynamics. This indicates a rate limiting step within the GAF and not the PHY domain. We further identify a pH dependence of the biphasic photoproduct formation hinting towards a pKa dependent tuning mechanism. Our findings add to the understanding of the role of the individual domains in the photocycle dynamics and provide a basis for engineering of phytochromes towards biotechnological applications.
Graphical abstract

Similar content being viewed by others
Avoid common mistakes on your manuscript.
1 Introduction
The photoreceptor superfamily of phytochromes is involved in the regulation of a variety of biologically relevant processes like photomorphogenesis, photoprotection and phototaxis in a multitude of different organisms [1, 2]. Their regulatory function is based on the photoconversion between a dark-adapted and a light-adapted state. Specifically, irradiation induces Z ↔ E isomerization of the embedded, covalently bound bilin chromophore that in turn triggers formation of further intermediates characterized by reorganization of the protein scaffold.
Phytochromes can be divided into three groups based on the domain structure and composition of their photosensory core module (PCM). The PCM of Group I phytochromes, comprising canonical plant (e.g. OatPhyA), bacterial (e.g. Agp1, DrBphP), cyanobacterial (e.g. CphA, SynCph1) and fungal phytochromes (e.g. FphA, NcFph1), consists of a PAS-GAF-PHY array [3,4,5]. Group II phytochromes, known as knotless phytochromes, lack the PAS domain with its domain specific figure-of-eight knot formation with the GAF domain [1, 2, 6, 7], while Group III phytochromes, called cyanobacteriochromes (CBCRs), consist only of GAF domains [8]. So far, the latter two groups are found only in cyanobacteria [1]. In contrast to CBCRs, which cover the whole near-IR to UV spectral range, Group I and Group II phytochromes mainly switch between Pr (red-absorbing) and Pfr (far-red-absorbing) states [9,10,11,12,13,14,15,16,17].
Some knotless phytochromes retain their photoconversion abilities even in their isolated GAF domain (after removal of the PHY domain), and thus can be utilized as model systems to investigate the influence of the PHY domain on the photoconversion process [18, 19]. This is of particular interest since the PHY domain interacts directly with the chromophore embedded in the GAF domain via a structural element referred to as “tongue” (Fig. 1A). The latter undergoes secondary structure reorganization during photoconversion [6, 7]. This reorganization not only affects the slower, protein driven dynamics of the later intermediates in the photocycle, but also plays a key role in the signal transduction towards the output domain [20, 21].
A Schematic drawing of the structural homology model of the Pr state of All2699g1g2. The homology model is based on the crystal structure of the All2699g1 domain (PDB ID 6OZA) [19] and the crystal structure of the structurally similar knotless phytochrome SynCph2 (PDB ID 4BWI) [22]. The PHY-like GAF2 domain (yellow) interacts with the chromophore (orange) embedded in the GAF1 domain (green) via the tongue-like protrusion displayed in pink. Taken from [23]. B Stationary absorption spectra of the Pr and Pfr state in All2699g1 (dotted lines) and All2699g1g2 (solid lines). The method to obtain the pure spectrum of the Pfr state was described elsewhere in detail [18]
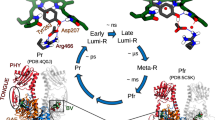
The forward (Pr → Pfr) reaction in the photocycle of canonical phytochromes typically involves three intermediates termed Lumi-R, Meta-Ra and Meta-Rc. The Lumi-R intermediate is formed directly after the decay of the excited state on the timescale of tens to hundreds of ps [24]. It contains the isomerized chromophore and a reorganized hydrogen-bonding network. Some studies distinguish Lumi-RI and Lumi-RII intermediates [24,25,26,27,28,29]. The structural changes in Lumi-RI occur on the ps timescale and are limited to the ring D vicinity, while the following hydrogen bonding reorganization and structural changes in the wider vicinity of the chromophore are attributed to Lumi-RII [24, 28, 29]. On the µs timescale, Lumi-R converts into the Meta-Ra intermediate via relaxation of the strained chromophore [21, 26, 27, 30,31,32]. Within a few ms, in plant and some cyanobacterial and bacterial phytochromes, the chromophore undergoes deprotonation to form the Meta-Rc intermediate [30, 31, 33, 34], which then forms the final state, Pfr, upon re-protonation of the chromophore on the tens to hundreds of ms timescale [21, 26, 27, 30,31,32]. However, the de-/re-protonation steps of the chromophore are not observed in all phytochromes, and thus the role of this mechanism in the dynamics of Meta-Rc is still under debate [24]. This formation of Pfr involves large-scale protein changes required for the final accommodation of the isomerized chromophore, reorganization of the hydrogen-bonding network and refolding of the tongue [20, 21, 24, 32]. In SynCph1 and OatPhyA, this transition has been observed to be biphasic [27, 30].
Generally, the reverse photoconversion reaction (Pfr → Pr) proceeds via two intermediates – Lumi-F and Meta-F. The Lumi-F intermediate contains an isomerized chromophore and is directly formed by the ultrafast photoreaction on the ps timescale [35,36,37,38,39]. Following ground state chromophore relaxation and minor side chain readjustments, a spectrally Pr-like intermediate termed Meta-F is formed on the ps-to-ns timescale [36, 40, 41]. The Meta-F intermediate undergoes rearrangement of the hydrogen-bonding network and the binding pocket, as well as refolding of the tongue to form the Pr state [41]. These steps occur on the µs-ms timescale and do not involve major spectral changes in the visible spectral range [40, 42, 43].
In this work, we expand our ultrafast studies on knotless phytochromes [18, 19] to the ms timescale. We compare the ms dynamics of the knotless phytochrome All2699g1g2 and its isolated GAF domain All2699g1 to reveal the effect of the PHY-like g2 domain on the forward (Pr → Pfr) and reverse (Pfr → Pr) photoconversion of knotless phytochromes. We further compare our results to the previously investigated knotless phytochrome SynCph2 and discuss the similarities and differences observed for these structurally related knotless phytochromes.
2 Materials and methods
2.1 Protein preparation
Preparation of All2699g1 and All2699g1g2 was described elsewhere in detail [19, 44]. The final buffer conditions for the experiments were set to 50 mM TRIS, 200 mM NaCl, 5% Glycerol, pH 8.0. For the pH dependence measurements of All2699g1, the following buffer conditions were used: 20 mM MES, 150 mM NaCl, 5% Glycerol, pH 6.1; 20 mM KPB, 150 mM NaCl, 5% Glycerol, pH 7.2; 20 mM TRIS, 150 mM NaCl, 5% Glycerol, pH 8.4.
Preparation of SynCph2 was described elsewhere [9]. The final buffer conditions for the experiments were set to 50 mM TRIS, 300 mM NaCl, 5 mM EDTA, pH 8.0.
Samples were measured in a 2 × 10 mm quartz cuvette with an optical density between 0.4 and 0.8 on 10 mm.
2.2 Stationary characterization
Stationary absorption spectra of the investigated samples were obtained using a Specord S600 absorption spectrometer (Analytik Jena). Conversion between the Pr and the Pfr state was achieved by illumination with appropriate LEDs (730 nm and 590 nm respectively, 1 W, Thorlabs).
2.3 Transient flash photolysis experiments
The pump pulses were provided by an optical parametric oscillator (OPO) (preciScan, GWU-Lasertechnik) pumped by a Nd:YAG laser (SpitLight 600, Innolas Laser). The OPO was set to 640 nm for Pr and 702 nm for Pfr measurements with an average pulse energy of 2.4–2.6 mJ/cm2. The probe light was generated by a spectrally broad continuous wave Xenon lamp (Hamamatsu LC-8) and two matched monochromators before and after the sample. The absorbance changes were then detected with a photomultiplier tube (PMT) and the resulting signal was converted into an electrical signal and recorded using an oscilloscope (DPO5204B-10RL, Tektronix). Time points were measured every 100 ns, except for the g1g2 Pfr measurements where a point was measured every 200 ns. For all samples absorbance changes were recorded between 500 and 750 nm with at least 30 averaged scans per wavelength. Between each individual scan the sample was illuminated for 4 s using an appropriate LED (Pr → Pfr, 730 nm; Pfr → Pr, 590 nm) to avoid accumulation of photoproducts. For analysis, the single transients were reduced by averaging data points in a combined linear and logarithmic timescale to obtain feasible data sizes.
For the pH dependence measurements of the ms dynamics of All2699g1, transients at six key wavelengths (510, 540, 600, 630, 680, and 710 nm) were measured and analyzed like the complete datasets. To validate the reduction to the 6-wavelength datasets, a comparison of the analysis to a complete dataset is shown exemplarily in the SI for pH 7.2 (SI Fig. S1).
2.4 Data analysis
Data analysis was performed using OPTIMUS (http://www.optimusfit.org/) [45]. We performed global target analysis (GTA) [45, 46] on all datasets using a sequential model. This model is fitted directly to the experimental data and results in the lifetimes, the decay-associated spectra (DAS) and the so-called evolution-associated difference spectra (EADS) for the respective kinetic components. The DAS contain both spectral and kinetic information, i.e. the positive amplitudes in the DAS account for decay of photoproduct absorption (PA) while negative amplitudes account for rise of absorption or recovery of the ground state bleach. Unlike DAS, EADS contain pure spectral information.
3 Results and discussion
3.1 Forward (Pr → Pfr) ms dynamics
3.1.1 Comparison between All2699g1 and All2699g1g2
The flash photolysis data of the Pr state of All2699g1 and All2699g1g2 (Fig. 2) appear very similar. Both show a dominant negative difference signal visible throughout the entire detected time range, which can be straightforwardly assigned to the ground state bleach (GSB) of the Pr state after excitation. Additionally, several distinct positive signals, at 660–690 nm, 660–700 nm, 500–560 nm and 660–750 nm, are present at different delay times in the data. These absorption changes are associated with the different intermediates observed during the photocycle. We performed global target analysis using a sequential model to reveal the kinetic details of the Pr → Pfr conversion.
Pr → Pfr ms dynamics of All2699g1g2 (A–C) and All2699g1 (D–F) at pH 8. A, D Flash photolysis data of the Pr → Pfr ms dynamics after excitation at 640 nm. B, E Corresponding decay-associated spectra (DAS) and C, F evolution-associated spectra (EADS) obtained after global target analysis of (A,D). G Schematic representation of the proposed photocycle for the Pr → Pfr ms dynamics of All2699g1g2 and All2699g1 at pH 8. The dashed line indicates the thermal reversion from Pfr to Pr, while the dotted line represents the initial photoreaction
Five components were required to fit the Pr data of All2699g1g2. Based on the resemblance to the primary photoproduct formed on the ultrafast timescale [18], we assign the S1 evolution-associated difference spectrum (EADS, Fig. 2C), peaking at 675 nm, to the Lumi-R intermediate. The positive (575–660 nm) and negative amplitudes (660–730 nm) of the 150 µs decay-associated spectrum (DAS, Fig. 2B), indicate the Lumi-R-to-Meta-Ra transition, which occurs with a red spectral shift (S1 vs. S2 EADS) induced by the bilin chromophore relaxation. In a next step, the chromophore undergoes a prominent spectral blue shift with a 1.5 ms lifetime to form the Meta-Rc intermediate (positive amplitude DAS at 575–750 nm and negative amplitude DAS at 500–575 nm in Fig. 2B, S2 → S3 EADS transition in Fig. 2C). We attribute this blue spectral shift to a chromophore deprotonation step following the assignment made previously for other phytochromes [31, 33, 34]. The following re-protonation of the chromophore is biphasic (S3 and S4 EADS). The corresponding DAS show similar spectral features representing the decay of the Meta-Rc absorption (positive amplitude at about 550 nm in τ3 (26 ms) and in τ4 (210 ms)) and the rise of the absorption of the final photoproduct Pfr (negative amplitude 650–750 nm). In contrast to τ3, the DAS amplitude of τ4 is negative in the range of 610–650 nm, potentially indicating some recovery of the spectrally coinciding Pr ground state via a shunt pathway as reported in some bacteriophytochromes [47, 48].
The major difference between the Pr flash photolysis data of All2699g1g2 (Fig. 2A) and All2699g1 (Fig. 2D) is that for g1 the kinetics is accelerated and can be described with only four components (Fig. 2E-F). Following the assignment above, those four components correspond to Meta-Ra (660–700 nm), Meta-Rc (500–560 nm), Pfr (660–740 nm) and the Pr GSB. The Lumi-R-to-Meta-Ra transition occurs on a timescale shorter than what is detected in our experiment (i.e. < 20 µs). Thus, the 150 µs component represents instead the Meta-Ra-to-Meta-Rc transition as indicated by the negative amplitude between 500 and 550 nm in the corresponding DAS (Fig. 2E). The Meta-Rc decay in All2699g1, albeit accelerated, is also biphasic (11 ms and 91 ms lifetimes, see below for more detailed discussion).
3.1.2 Effect of the PHY domain on the forward dynamics
The Lumi-R-to-Meta-Ra and Meta-Ra-to-Meta-Rc transitions are significantly slower (~ 10-fold) in All2699g1g2 as compared to All2699g1, which indicates that the presence of the PHY domain imposes a higher energetic barrier for the associated chromophore relaxation and proximal protein structural rearrangements. This conclusion is in line with previous NMR [23, 49] and RR [28] studies, where it was proposed that the interaction with the PHY tongue makes the chromophore-binding pocket more rigid. In contrast, the final chromophore transition (Meta-Rc-to-Pfr), typically associated with major structural changes (tongue refolding [20, 21, 24, 32]) in the protein environment distal to the chromophore, is affected to a smaller extent (twofold) by the lacking PHY domain in All2699g1. Furthermore, the preserved biphasic character of the Meta-Rc-to-Pfr transition in All2699g1 suggests that the chromophore transitions associated with this step are predominantly determined by factors within the binding pocket of the GAF domain. Previously, a similar conclusion was derived based on time-resolved X-ray and UV/Vis studies on DrBphP [21].
3.1.3 Origin of the biphasic meta-Rc to Pfr transition
To further investigate the origin of the biphasic Meta-Rc-to-Pfr kinetics in All2699g1 and All2699g1g2, we performed flash photolysis measurements at selected detection wavelengths (510, 540, 600, 630, 680, and 710 nm) and at three different pH values (6.1, 7.2, and 8.4). A direct comparison of the transients at 710 nm reveals that the Pfr formation is significantly accelerated at high pH values (Fig. 3A). This clear pH dependence is further corroborated by the DAS of the three datasets (Fig. 3B).
pH dependence of the Pr → Pfr ms dynamics of All2699g1 at pH 6.1, 7.2 and 8.4. A Transients of the UV/Vis flash photolysis measurements of All2699g1 at 710 nm at different pH values. B DAS obtained by global lifetime analysis of 6 transients at key wavelengths (510, 540, 600, 630, 680 and 710 nm) of All2699g1 after excitation at 640 nm at different pH values
While the lifetime components do not vary significantly, their corresponding amplitudes and especially the amplitude ratio between the shorter and the longer component (correspondingly marked blue and green in Fig. 3B) change with the pH. At pH 6.1, the amplitude of the longer component is higher than that of the shorter one resulting in the slowest overall Meta-Rc-to-Pfr transition (Fig. 3A). At pH 8.4 the ratio is reversed, the shorter component clearly dominates, thus making the overall Meta-Rc decay fastest. At pH 7.2 the two components have nearly identical amplitude. Consequently, the process occurring with the shorter lifetime component is favored at high pH, while the process described by the longer lifetime component is favored at low pH. Furthermore, the monoexponential formation of the Meta-Rc and the lack of spectral differences in the two DAS describing its decay imply that there is a single chromophore population in Meta-Rc and that the following re-protonation reaction is dependent on the pKa value of the proton donor residue and the state of the hydrogen-bonding network linking it to the solvent. Thereby, the faster component represents a chromophore re-protonation reaction occurring from a protonated residue, while for the slower component the residue is deprotonated, and thus the re-protonation requires proton uptake from the environment. Such a proton uptake step has been reported for Cph1 [30]. In effect, the role of the slower component is prominent at low pH, while at higher (H+ deficient) pH, its contribution is diminished.
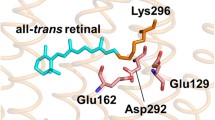
While we cannot directly identify the relevant residue, based on the nearly identical amplitudes of the DAS of the two Meta-Rc decay components, we can deduce that its pKa value is close to 7.2. Mutational studies on other phytochromes (SynCph1, DrBphP, Agp1) identified histidine residues that are part of the extended hydrogen-bonding network of the chromophore and affect its protonation state [33, 50,51,52,53]. The respective residues in All2699g1, His139 and His169, are therefore potential candidates for the origin of the observed pH dependence. His139 is in direct contact with the pyrrole nitrogens of the bilin chromophore and was reported to be directly involved in the proton uptake leading to the formation of the Pfr state as some mutants of this residue do not progress past Meta-Rc in their photocycle [33, 52]. In addition, the His139-homologous residue H260 exhibits a pKa value of 7.4 in the Pr state of SynCph1, strikingly similar to the value we derive here for the biphasic transition to Pfr [51]. His169 is part of the extended hydrogen-bonding network, and thus it is in an indirect contact with His139. It is also in contact with the chromophore in both Pr and Pfr interacting with the D-ring carbonyl or the C-ring propionate, respectively [54]. As such, its protonation state could influence the hydrogen-bonding network potentially contributing to the biphasic kinetics. However, we consider His139 a more likely candidate since mutants of His169 exhibit only minor impact on the chromophore and/or hydrogen-bonding network pKa [50]. The highly conserved Asp87 may also be considered as a candidate involved in the protonation steps. This residue interacts with the chromophore through the pyrrole water and similar to His139 is involved in tuning the chromophore pKa and its hydrogen-bonding network [33, 52]. However, it is reported that this Asp87 retains its protonation state during the photocycle rendering a direct involvement in the reprotonation unlikely [24]. Indirectly, an increase in the protein mobility, such as that observed in SynCph1 [34], may also be contributing to the overall acceleration of Pfr formation at higher pH.
3.1.4 Comparison of the Pr dynamics to other phytochromes
The lifetimes of the intermediates observed in the knotless phytochrome All2699g1g2 match well with previously reported lifetimes for SynCph1 [30], SynCph2 [32], and Agp1 [31] indicating high similarities between the dynamics of knotless and knotted phytochromes on the ms timescale. In the only investigated knotless phytochrome SynCph2, the lifetimes for the Lumi-R-to-Meta-Ra and Meta-Ra-to-Meta-Rc transition (170 µs and 1.6 ms, SI Fig. S2) are nearly identical to those found here for All2699g1g2 (150 µs and 1.5 ms, Fig. 2B). However, the Meta-Rc-to-Pfr transition is significantly faster in SynCph2 and proceeds monoexponentially with a lifetime of 17 ms. In contrast, the Meta-Rc-to-Pfr transition in SynCph1 is similarly biphasic with lifetimes close to those we find in All2699g1g2. A comparison between SynCph2 and All2699g1 shows a different involvement of water molecules and propionic side chains in the extended hydrogen-bonding network, which can result in different pKa values and explain the occurrence or lack of biphasic Meta-Rc decay in different phytochromes.
3.2 Reverse (Pfr → Pr) ms dynamics
3.2.1 Comparison of All2699g1 and All2699g1g2
To provide a complete picture of the influence of the PHY domain on the dynamics of knotless phytochromes, we investigated the ms reverse reaction (Pfr → Pr) in All2699g1 and All2699g1g2. The GSB (negative absorption difference) signal in the Pfr flash photolysis data of both samples is located above 650 nm (Fig. 4). However, reflecting the difference in the Pfr absorption spectra (Fig. 1), the GSB of All2699g1g2 is red shifted. In addition to the GSB signal, we observe a spectral evolution of the positive absorption difference signal associated with the transitions of the intermediate states towards the final photoproduct Pr (i.e. from ~ 550–650 nm through ~ 575–660 nm to ~ 500–630 nm, Fig. 1). This evolution is better discerned for All2699g1g2.
Pfr → Pr ms dynamics of All2699g1g2 (A–C) and All2699g1 (D–F) at pH 8. A, D Flash photolysis data of the Pfr → Pr ms dynamics after excitation at 702 nm. (B,E) Corresponding decay-associated spectra (DAS) and C, F evolution-associated spectra (EADS) obtained after global target analysis of A, D. G Schematic representation of the proposed photocycle for the Pfr → Pr ms dynamics of All2699g1g2 and All2699g1 at pH 8. The vertical black dashed line indicates the thermal reversion from Pfr to Pr, while the dotted line represents the initial photoreaction. The short dashed purple line represents the shunt pathway present in All2699g1
The global target analysis with a sequential scheme yields four lifetime components for both samples. In All2699g1g2, the S1 EADS (Fig. 4C) resembles the spectrum of the species present at the end of the ultrafast Pfr dynamics in SynCph2 [37] or SynCph1 [36], and thus can be assigned to the Meta-F intermediate. Meta-F decays with a lifetime of 590 µs to form a blue-shifted intermediate (S2 EADS) as described by the positive–negative amplitudes of the corresponding DAS (Fig. 4B). A similar blue-shifted species has also been observed in SynCph2 and termed F2 [32]. The F2 intermediate then decays to form the final Pr state with two lifetimes where the amplitude of the 860 ms component is significantly higher than that of the 58 ms component (Fig. 4B).
The Pfr flash photolysis data of All2699g1 exhibit similar signals (Fig. 4D). However, the S1 EADS is broadened on the blue spectral side, which indicates that the F2 intermediate is already present at the beginning of the measurement in equilibrium with Meta-F (Fig. 4F). Thus, in analogy to Meta-Ra from the forward dynamics of All2699g1, in the reverse direction we cannot clearly detect the rise of F2 in the single domain sample. Instead, the first lifetime component (2 ms) in All2699g1 indicates a shunt pathway to the Pfr ground state from a leftover Meta-F population. Such a pathway is not present in the reverse dynamics of All2699g1g2 and is not observed in SynCph2, but has been observed in bacteriophytochromes [47, 48]. The following transition from F2 to the Pr state is described again by two lifetimes. However, in contrast to All2699g1g2, the difference in the amplitude of the two components is much smaller. In addition, the second lifetime component is significantly shorter in All2699g1 (i.e. 220 ms). As a result, the overall Pr formation is accelerated in All2699g1. Interestingly, in both samples, there appears to be an equilibrium between the Meta-F and F2 intermediates. This equilibrium is shifted towards Meta-F in All2699g1, while in All2699g1g2 the F2 and Meta-F intermediates have similar amplitudes. The S3 EADS of All2699g1g2 (Fig. 4C) suggests that at longer timescales the equilibrium shifts towards high F2 populations.
3.2.2 Effect of the PHY domain on the reverse dynamics
In both the forward and the reverse dynamics, the photoreaction is accelerated in absence of the PHY domain, i.e. the Meta-Ra in the forward and F2 in the reverse dynamics are formed in All2699g1 earlier than in All2699g1g2. We attribute this to fewer restrictions in the binding pocket in absence of the PHY domain as revealed by previous NMR and cryo-trapping experiments [23, 28, 49]. The photoreaction then follows similar steps where an intermediate spectrally close to the final photoproduct (Meta-Ra and Meta-F) converts into a blue-shifted one (Meta-Rc and F2). The final photoproduct (Pfr or Pr) is formed with a biphasic kinetics, as indicated by a corresponding red shift on the ms timescale. Similar to the spectral shift of Meta-Rc in the forward direction, we observe a spectral shift of F2 in the reverse direction. This resemblance suggests that the formation and the decay of F2 may also occur via deprotonation and re-protonation steps. For consistency, we propose that F2 is termed Meta-Fc by analogy to Meta-Rc.
3.2.3 Comparison of the Pfr dynamics to other phytochromes
It is particularly remarkable that the Meta-F-to-Meta-Fc transition in All2699g1g2 is on a timescale (hundreds of µs) similar to that found in SynCph2 and PhyA while the Pr formation step is two orders of magnitude slower [32, 40]. Even in comparison to other phytochromes like CphA [40] or PstBphP1 and PaBphP1 [43] that exhibit slightly different reverse kinetics, the formation of the photoproduct is still exceptionally slow in both All2699g1g2 and All2699g1. The unknown origin of the striking temporal difference for the final photoproduct formation step between the two structurally very similar knotless phytochromes (All2699g1g2 and SynCph2) should be addressed in future studies.
4 Conclusion
Our work provides a dynamic picture of the effect of the PHY domain on the ms dynamics of knotless phytochromes. We show that the occurrence and spectral signature of the involved intermediates are not significantly affected by the presence of the PHY domain. However, the µs dynamics involving relaxation of the chromophore are slowed down for both the forward and the reverse direction. We attribute this effect to a more confined chromophore binding pocket in presence of the PHY domain. Quite remarkably, the lifetimes describing the transition to the final photoproduct are much less affected by the presence/absence of the PHY domain. This supports previous findings that the rate limiting step of the photoproduct formation observed by UV/vis spectroscopy is imposed by internal factors of the GAF (PAS-GAF) domain and are not directly related to tongue refolding.
Interestingly, we find that the Meta-Rc-to-Pfr transition is biphasic in both All2699g1 and All2699g1g2, but not in SynCph2. Based on the pH dependence of the transition in All2699g1, we associate this behavior to the pKa value of the proton donor residue in the Meta-Rc intermediate. The lack of biphasic behavior in some phytochromes suggests that the rate of photoproduct formation can be tweaked through fine tuning of the pKa value of the hydrogen-bonding network.
References
Anders, K., & Essen, L. O. (2015). The family of phytochrome-like photoreceptors: Diverse, complex and multi-colored, but very useful. Current Opinion in Structural Biology, 35, 7–16. https://doi.org/10.1016/j.sbi.2015.07.005
Rockwell, N. C., & Lagarias, J. C. (2017). Phytochrome diversification in cyanobacteria and eukaryotic algae. Current Opinion in Plant Biology, 37, 87–93. https://doi.org/10.1016/j.pbi.2017.04.003
Rockwell, N. C., & Lagarias, J. C. (2006). The structure of phytochrome: A picture is worth a thousand spectra. The Plant Cell, 18(1), 4–14. https://doi.org/10.1105/tpc.105.038513
Rockwell, N. C., Su, Y.-S., & Lagarias, J. C. (2006). Phytochrome structure and signaling mechanisms. Annual Review of Plant Biology, 57(1), 837–858. https://doi.org/10.1146/annurev.arplant.56.032604.144208
Möglich, A., Yang, X., Ayers, R. A., & Moffat, K. (2010). Structure and function of plant photoreceptors. Annual Review of Plant Biology, 61, 21–47. https://doi.org/10.1146/annurev-arplant-042809-112259
Yang, X., Kuk, J., & Moffat, K. (2008). Crystal structure of Pseudomonas aeruginosa bacteriophytochrome: Photoconversion and signal transduction. Proceedings of the National Academy of Sciences of the United States of America, 105(38), 14715–14720. https://doi.org/10.1073/pnas.0806718105
Essen, L. O., Mailliet, J., & Hughes, J. (2008). The structure of a complete phytochrome sensory module in the Pr ground state. Proceedings of the National Academy of Sciences of the United States of America, 105(38), 14709–14714. https://doi.org/10.1073/pnas.0806477105
Ikeuchi, M., & Ishizuka, T. (2008). Cyanobacteriochromes: A new superfamily of tetrapyrrole-binding photoreceptors in cyanobacteria. Photochemical and Photobiological Sciences, 7(10), 1159. https://doi.org/10.1039/b802660m
Anders, K., Von Stetten, D., Mailliet, J., Kiontke, S., Sineshchekov, V. A., Hildebrandt, P., Hughes, J., & Essen, L. O. (2011). Spectroscopic and photochemical characterization of the red-light sensitive photosensory module of Cph2 from synechocystis PCC 6803. Photochemistry and Photobiology, 87(1), 160–173. https://doi.org/10.1111/j.1751-1097.2010.00845.x
Lamparter, T., Mittmann, F., Gartner, W., Borner, T., Hartmann, E., & Hughes, J. (1997). Characterization of recombinant phytochrome from the cyanobacterium Synechocystis. Proceedings of the National Academy of Sciences, 94(22), 11792–11797. https://doi.org/10.1073/pnas.94.22.11792
Lamparter, T., Michael, N., Mittmann, F., & Esteban, B. (2002). Phytochrome from Agrobacterium tumefaciens has unusual spectral properties and reveals an N-terminal chromophore attachment site. Proceedings of the National Academy of Sciences, 99(18), 11628–11633. https://doi.org/10.1073/pnas.152263999
Chai, Y. G., Singh, B. R., Song, P. S., Lee, J., & Robinson, G. W. (1987). Purification and spectroscopic properties of 124-kDa oat phytochrome. Analytical Biochemistry, 163(2), 322–330. https://doi.org/10.1016/0003-2697(87)90231-4
Hirose, Y., Shimada, T., Narikawa, R., Katayama, M., & Ikeuchi, M. (2008). Cyanobacteriochrome CcaS is the green light receptor that induces the expression of phycobilisome linker protein. Proceedings of the National Academy of Sciences, 105(28), 9528–9533. https://doi.org/10.1073/pnas.0801826105
Rockwell, N. C., Martin, S. S., Gulevich, A. G., & Lagarias, J. C. (2012). Phycoviolobilin formation and spectral tuning in the DXCF cyanobacteriochrome subfamily. Biochemistry, 51(7), 1449–1463. https://doi.org/10.1021/bi201783j
Rockwell, N. C., Martin, S. S., Feoktistova, K., & Lagarias, J. C. (2011). Diverse two-cysteine photocycles in phytochromes and cyanobacteriochromes. Proceedings of the National Academy of Sciences of the United States of America, 108(29), 11854–11859. https://doi.org/10.1073/pnas.1107844108
Rockwell, N. C., Martin, S. S., & Lagarias, J. C. (2016). Identification of Cyanobacteriochromes detecting far-red light. Biochemistry, 55(28), 3907–3919. https://doi.org/10.1021/acs.biochem.6b00299
Fushimi, K., Ikeuchi, M., & Narikawa, R. (2017). The expanded red/green Cyanobacteriochrome lineage: An evolutionary hot spot. Photochemistry and Photobiology, 93(3), 903–906. https://doi.org/10.1111/php.12764
Fischer, T., Xu, Q.-Z., Zhao, K.-H., Gärtner, W., Slavov, C., & Wachtveitl, J. (2020). Effect of the PHY domain on the photoisomerization step of the forward Pr→Pfr conversion of a knotless phytochrome. Chemistry A European Journal. https://doi.org/10.1002/chem.202003138
Slavov, C., Fischer, T., Barnoy, A., Shin, H., Rao, A. G., Wiebeler, C., Zeng, X., Sun, Y., Xu, Q., Gutt, A., Zhao, K., Gärtner, W., Yang, X., Schapiro, I., & Wachtveitl, J. (2020). The interplay between chromophore and protein determines the extended excited state dynamics in a single-domain phytochrome. Proceedings of the National Academy of Sciences of the United States of America, 117(28), 16356–16362. https://doi.org/10.1073/pnas.1921706117
Takala, H., Björling, A., Berntsson, O., Lehtivuori, H., Niebling, S., Hoernke, M., Kosheleva, I., Henning, R., Menzel, A., Ihalainen, J. A., & Westenhoff, S. (2014). Signal amplification and transduction in phytochrome photosensors. Nature, 509(7499), 245–248. https://doi.org/10.1038/nature13310
Björling, A., Berntsson, O., Lehtivuori, H., Takala, H., Hughes, A. J., Panman, M., Hoernke, M., Niebling, S., Henry, L., Henning, R., Kosheleva, I., Chukharev, V., Tkachenko, N. V., Menzel, A., Newby, G., Khakhulin, D., Wulff, M., Ihalainen, J. A., & Westenhoff, S. (2016). Structural photoactivation of a full-length bacterial phytochrome. Science Advances, 2(8), e1600920. https://doi.org/10.1126/sciadv.1600920
Anders, K., Daminelli-Widany, G., Mroginski, M. A., Von Stetten, D., & Essen, L. O. (2013). Structure of the cyanobacterial phytochrome 2 photosensor implies a tryptophan switch for phytochrome signaling. Journal of Biological Chemistry, 288(50), 35714–35725. https://doi.org/10.1074/jbc.M113.510461
Xu, Q.-Z., Bielytskyi, P., Otis, J., Lang, C., Hughes, J., Zhao, K.-H., Losi, A., Gärtner, W., & Song, C. (2019). MAS NMR on a red/far-red photochromic Cyanobacteriochrome All2699 from Nostoc. International Journal of Molecular Sciences, 20(15), 3656. https://doi.org/10.3390/ijms20153656
Ihalainen, J. A., Gustavsson, E., Schroeder, L., Donnini, S., Lehtivuori, H., Isaksson, L., Thöing, C., Modi, V., Berntsson, O., Stucki-Buchli, B., Liukkonen, A., Häkkänen, H., Kalenius, E., Westenhoff, S., & Kottke, T. (2018). Chromophore-protein interplay during the phytochrome photocycle revealed by step-scan FTIR spectroscopy. Journal of the American Chemical Society, 140(39), 12396–12404. https://doi.org/10.1021/jacs.8b04659
Braslavsky, S. E., Gärtner, W., & Schaffner, K. (1997). Phytochrome photoconversion. Plant, Cell and Environment, 20(6), 700–706. https://doi.org/10.1046/j.1365-3040.1997.d01-101.x
Schmidt, P., Westphal, U. H., Worm, K., Braslavsky, S. E., Gärtner, W., & Schaffner, K. (1996). Chromophore-protein interaction controls the complexity of the phytochrome photocycle. Journal of Photochemistry and Photobiology B, 34(1), 73–77. https://doi.org/10.1016/1011-1344(95)07269-1
Zhang, C. F., Farrens, D. L., Björling, S. C., Song, P. S., & Kliger, D. S. (1992). Time-resolved absorption studies of native etiolated oat phytochrome. Journal of the American Chemical Society, 114(12), 4569–4580. https://doi.org/10.1021/ja00038a019
Velazquez Escobar, F., Kneip, C., Michael, N., Hildebrandt, T., Tavraz, N., Gärtner, W., Hughes, J., Friedrich, T., Scheerer, P., Mroginski, M. A., & Hildebrandt, P. (2020). The lumi-R intermediates of prototypical phytochromes. Journal of Physical Chemistry B, 124(20), 4044–4055. https://doi.org/10.1021/acs.jpcb.0c01059
Kübel, J., Chenchiliyan, M., Ooi, S. A., Gustavsson, E., Isaksson, L., Kuznetsova, V., Ihalainen, J. A., Westenhoff, S., & Maj, M. (2020). Transient IR spectroscopy identifies key interactions and unravels new intermediates in the photocycle of a bacterial phytochrome. Physical Chemistry Chemical Physics, 22(17), 9195–9203. https://doi.org/10.1039/c9cp06995j
Van Thor, J. J., Borucki, B., Crielaard, W., Otto, H., Lamparter, T., Hughes, J., Hellingwerf, K. J., & Heyn, M. P. (2001). Light-induced proton release and proton uptake reactions in the cyanobacterial phytochrome Cph1. Biochemistry, 40(38), 11460–11471. https://doi.org/10.1021/bi002651d
Borucki, B., von Stetten, D., Seibeck, S., Lamparter, T., Michael, N., Mroginski, M. A., Otto, H., Murgida, D. H., Heyn, M. P., & Hildebrandt, P. (2005). Light-induced proton release of phytochrome is coupled to the transient deprotonation of the tetrapyrrole chromophore. Journal of Biological Chemistry, 280(40), 34358–34364. https://doi.org/10.1074/jbc.M505493200
Anders, K., Gutt, A., Gärtner, W., & Essen, L. O. (2014). Phototransformation of the red light sensor cyanobacterial phytochrome 2 from synechocystis species depends on its tongue motifs. Journal of Biological Chemistry, 289(37), 25590–25600. https://doi.org/10.1074/jbc.M114.562082
Von Stetten, D., Seibeck, S., Michael, N., Scheerer, P., Mroginski, M. A., Murgida, D. H., Krauss, N., Heyn, M. P., Hildebrandt, P., Borucki, B., & Lamparter, T. (2007). Highly conserved residues Asp-197 and His-250 in Agp1 phytochrome control the proton affinity of the chromophore and Pfr formation. Journal of Biological Chemistry, 282(3), 2116–2123. https://doi.org/10.1074/jbc.M608878200
Sadeghi, M., Balke, J., Schneider, C., Nagano, S., Stellmacher, J., Lochnit, G., Lang, C., Weise, C., Hughes, J., & Alexiev, U. (2020). Transient deprotonation of the chromophore affects protein dynamics proximal and distal to the linear tetrapyrrole chromophore in phytochrome Cph1. Biochemistry, 59(9), 1051–1062. https://doi.org/10.1021/acs.biochem.9b00967
Heyne, K., Herbst, J., Stehlik, D., Esteban, B., Lamparter, T., Hughes, J., & Diller, R. (2002). Ultrafast dynamics of phytochrome from the cyanobacterium Synechocystis, reconstituted with phycocyanobilin and phycoerythrobilin. Biophysical Journal, 82(2), 1004–1016. https://doi.org/10.1016/S0006-3495(02)75460-X
Kim, P. W., Rockwell, N. C., Martin, S. S., Lagarias, J. C., & Larsen, D. S. (2014). Heterogeneous photodynamics of the Pfr state in the cyanobacterial phytochrome Cph1. Biochemistry, 53(28), 4601–4611. https://doi.org/10.1021/bi5005359
Fischer, T., van Wilderen, L. J. G. W., Gnau, P., Bredenbeck, J., Essen, L.-O., Wachtveitl, J., & Slavov, C. (2021). Ultrafast photoconversion dynamics of the knotless phytochrome SynCph2. International Journal of Molecular Sciences, 22(19), 10690. https://doi.org/10.3390/ijms221910690
Rohmer, T., Lang, C., Bongards, C., Gupta, K. B. S. S., Neugebauer, J., Hughes, J., Gärtner, W., & Matysik, J. (2010). Phytochrome as molecular machine: Revealing chromophore action during the Pfr → Pr photoconversion by magic-angle spinning NMR spectroscopy. Journal of the American Chemical Society, 132(12), 4431–4437. https://doi.org/10.1021/ja9108616
Yang, X., Ren, Z., Kuk, J., & Moffat, K. (2011). Temperature-scan cryocrystallography reveals reaction intermediates in bacteriophytochrome. Nature, 479(7373), 428–431. https://doi.org/10.1038/nature10506
Chizhov, I., Zorn, B., Manstein, D. J., & Gärtner, W. (2013). Kinetic and thermodynamic analysis of the light-induced processes in plant and cyanobacterial phytochromes. Biophysical Journal, 105(9), 2210–2220. https://doi.org/10.1016/j.bpj.2013.09.033
Kraskov, A., Nguyen, A. D., Goerling, J., Buhrke, D., Velazquez Escobar, F., Fernandez Lopez, M., Michael, N., Sauthof, L., Schmidt, A., Piwowarski, P., Yang, Y., Stensitzki, T., Adam, S., Bartl, F., Schapiro, I., Heyne, K., Siebert, F., Scheerer, P., Mroginski, M. A., & Hildebrandt, P. (2020). Intramolecular proton transfer controls protein structural changes in phytochrome. Biochemistry, 59(9), 1023–1037. https://doi.org/10.1021/acs.biochem.0c00053
Chen, E., Lapko, V. N., Lewis, J. W., Song, P. S., & Kliger, D. S. (1996). Mechanism of native oat phytochrome photoreversion: A time-resolved absorption investigation. Biochemistry, 35(3), 843–850. https://doi.org/10.1021/bi952115z
Consiglieri, E., Gutt, A., Gärtner, W., Schubert, L., Viappiani, C., Abbruzzetti, S., & Losi, A. (2019). Dynamics and efficiency of photoswitching in biliverdin-binding phytochromes. Photochemical and Photobiological Sciences, 18(10), 2484–2496. https://doi.org/10.1039/c9pp00264b
Xu, Q.-Z., Goett-Zink, L., Gärtner, W., Zhao, K.-H., & Kottke, T. (2020). Tongue refolding in the knotless Cyanobacterial phytochrome All2699. Biochemistry, 59(22), 2047–2054. https://doi.org/10.1021/acs.biochem.0c00209
Slavov, C., Hartmann, H., & Wachtveitl, J. (2015). Implementation and evaluation of data analysis strategies for time-resolved optical spectroscopy. Analytical Chemistry, 87(4), 2328–2336. https://doi.org/10.1021/ac504348h
Van Stokkum, I. H. M. M., Larsen, D. S., & van Grondelle, R. (2004). Global and target analysis of time-resolved spectra. Biochimica et Biophysica Acta, 1657(2–3), 82–104. https://doi.org/10.1016/j.bbabio.2004.04.011
Merga, G., Lopez, M. F., Fischer, P., Piwowarski, P., Nogacz, Ż, Kraskov, A., Buhrke, D., Escobar, F. V., Michael, N., Siebert, F., Scheerer, P., Bartl, F., & Hildebrandt, P. (2021). Light-and temperature-dependent dynamics of chromophore and protein structural changes in bathy phytochrome Agp2. Physical Chemistry Chemical Physics, 23(33), 18197–18205. https://doi.org/10.1039/d1cp02494a
Buhrke, D., Kuhlmann, U., Michael, N., & Hildebrandt, P. (2018). The photoconversion of phytochrome includes an unproductive shunt reaction pathway. ChemPhysChem, 19(5), 566–570. https://doi.org/10.1002/cphc.201701311
Kim, Y., Xu, Q.-Z., Zhao, K.-H., Gärtner, W., Matysik, J., & Song, C. (2020). Lyophilization reveals a multitude of structural conformations in the chromophore of a Cph2-like phytochrome. Journal of Physical Chemistry B, 124(33), 7115–7127. https://doi.org/10.1021/acs.jpcb.0c03431
Rumfeldt, J. A., Takala, H., Liukkonen, A., & Ihalainen, J. A. (2019). UV-Vis spectroscopy reveals a correlation between Y263 and BV protonation states in Bacteriophytochromes. Photochemistry and Photobiology, 95(4), 969–979. https://doi.org/10.1111/php.13095
Velazquez Escobar, F., Lang, C., Takiden, A., Schneider, C., Balke, J., Hughes, J., Alexiev, U., Hildebrandt, P., & Mroginski, M. A. (2017). Protonation-dependent structural heterogeneity in the chromophore binding site of Cyanobacterial phytochrome Cph1. The Journal of Physical Chemistry B, 121(1), 47–57. https://doi.org/10.1021/acs.jpcb.6b09600
Wagner, J. R., Zhang, J., Von Stetten, D., Günther, M., Murgida, D. H., Mroginski, M. A., Walker, J. M., Forest, K. T., Hildebrandt, P., & Vierstra, R. D. (2008). Mutational analysis of Deinococcus radiodurans bacteriophytochrome reveals key amino acids necessary for the photochromicity and proton exchange cycle of phytochromes. Journal of Biological Chemistry, 283(18), 12212–12226. https://doi.org/10.1074/jbc.M709355200
Hahn, J., Strauss, H. M., Landgraf, F. T., Gimenez, H. F., Lochnit, G., Schmieder, P., & Hughes, J. (2006). Probing protein-chromophore interactions in Cph1 phytochrome by mutagenesis. FEBS Journal, 273(7), 1415–1429. https://doi.org/10.1111/j.1742-4658.2006.05164.x
Burgie, E. S., Bussell, A. N., Walker, J. M., Dubiel, K., & Vierstra, R. D. (2014). Crystal structure of the photosensing module from a red/far-red light-absorbing plant phytochrome. Proceedings of the National Academy of Sciences of the United States of America, 111(28), 10179–10184. https://doi.org/10.1073/pnas.1403096111
Acknowledgements
Lisa Köhler and Chen Song acknowledge the Deutsche Forschungsgemeinschaft (417685888). Chavdar Slavov and Josef Wachtveitl acknowledge the Deutsche Forschungsgemeinschaft (WA 1850/4-3). The authors thank Wolfgang Gärtner for his insightful comments and his support for the realization of the project. The authors also thank Petra Gnau and Lars-Oliver Essen for providing the SynCph2 samples for benchmark experiments.
Funding
Open Access funding enabled and organized by Projekt DEAL.
Author information
Authors and Affiliations
Corresponding authors
Ethics declarations
Conflict of interest
The datasets generated during and/or analyzed during the current study are available from the corresponding author upon reasonable request. The authors have no competing interests to declare that are relevant to the content of this article.
Additional information
This publication is dedicated to Prof. Silvia E. Braslavsky, a pioneer in photobiology and photobiophysics, on the occasion of her 80th birthday.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Fischer, T., Köhler, L., Ott, T. et al. Influence of the PHY domain on the ms-photoconversion dynamics of a knotless phytochrome. Photochem Photobiol Sci 21, 1627–1636 (2022). https://doi.org/10.1007/s43630-022-00245-9
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s43630-022-00245-9