Abstract
Cyanobacteriochromes (CBCRs) are phytochrome-related photosensory proteins that play an essential role in regulating phototaxis, chromatic acclimation, and cell aggregation in cyanobacteria. Here, we apply solid-state NMR spectroscopy to the red/green GAF2 domain of the CBCR AnPixJ assembled in vitro with a uniformly 13C- and 15N-labeled bilin chromophore, tracking changes in electronic structure, geometry, and structural heterogeneity of the chromophore as well as intimate contacts between the chromophore and protein residues in the photocycle. Our data confirm that the bilin ring D is strongly twisted with respect to the B–C plane in both dark and photoproduct states. We also identify a greater structural heterogeneity of the bilin chromophore in the photoproduct than in the dark state. In addition, the binding pocket is more hydrated in the photoproduct. Observation of interfacial 1H contacts of the photoproduct chromophore, together with quantum mechanics/molecular mechanics (QM/MM)-based structural models for this photoproduct, clearly suggests the presence of a biprotonated (cationic) imidazolium side-chain for a conserved histidine residue (322) at a distance of ~2.7 Å, generalizing the recent theoretical findings that explicitly link the structural heterogeneity of the dark-state chromophore to the protonation of this specific residue. Moreover, we examine pH effects on this in vitro assembled holoprotein, showing a substantially altered electronic structure and protonation of the photoproduct chromophore even with a small pH drop from 7.8 to 7.2. Our studies provide further information regarding the light- and pH-induced changes of the chromophore and the rearrangements of the hydrogen-bonding and electrostatic interaction network around it. Possible correlations between structural heterogeneity of the chromophore, protonation of the histidine residue nearby, and hydration of the pocket in both photostates are discussed.
Graphical abstract

Similar content being viewed by others
Avoid common mistakes on your manuscript.
1 Introduction
Cyanobacteriochromes (CBCRs), only known from cyanobacteria to date, represent a subgroup in the superfamily of phytochrome photosensors [1,2,3]. CBCRs share with the phytochromes the capability to covalently bind a bilin as chromophore in a GAF (cGMP phosphodiesterase/adenylate cyclase/FhlA) domain, and they also follow the common photochemistry, e.g., undergoing a light-induced isomerization around the 15,16-double bond between rings C and D (Z ↔ E) of the bilin (see Fig. 1 and refs. [4,5,6,7,8]). However, in contrast to canonical phytochromes that request a PAS (Period/Arnt/Single-minded)–GAF–PHY (phytochrome-specific) tridomain arrangement in order to maintain the spectral and kinetic properties [9,10,11], CBCRs are capable of utilizing only a single bilin-binding GAF domain for full photoconversion [12,13,14,15,16]. With the covalent attachment of bilins having different lengths of the π-conjugation systems, CBCRs exhibit diverse absorption maxima ranging from the near-ultraviolet to the near-infrared for their dark states and from blue to red for photoproducts [13,14,15,16,17,18,19,20,21]. Their compact size and the ability to sense various wavelengths and intensities have made CBCR–GAF domains targets for in-depth structural investigations employing protein crystallization [4,5,6, 16, 22, 23] as well as extensive theoretical, spectroscopic, and mechanistic studies [24,25,26,27,28,29,30,31,32,33], in particular through the use of both solution [7, 12, 34,35,36] and solid-state magic-angle spinning (MAS) NMR [8, 37,38,39]. These studies provided a wealth of structural information that has stimulated the discussion which parameters regulate the diverse spectral shifts in CBCRs between the dark state and photoproduct. Moreover, this photochromicity can arrive at more than 100 nm in certain species, e.g., red/green-switching CBCRs represented by AnPixJ from Anabaena 7210 [5, 15] as the focus of this study (Fig. 1A). Four color-tuning mechanisms have been proposed so far for CBCR–GAF domains including bilin variation [14,15,16,17,18] and deprotonation [14, 22], adduct formation by a second Cys [4, 5, 40,41,42], and twisting of the bilin ring D [7, 15, 24, 43]. Among these, the ‘D-ring control’ model appears most favorable for the red/green CBCR subgroup proposing that the bilin is more twisted in the photoproduct (green-absorbing, Pg) than in the dark state (red-absorbing, Pr) with substantial loss of D-ring conjugation, therefore causing a blue shift in the absorption [7, 24, 26, 34, 44, 45].
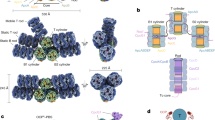
Pr dark-state structure and absorption properties of AnPixJg2. A Ribbon diagram of the AnPixJg2 Pr structure (PDB code 3W2Z, yellow) superimposed on the structures of two other representatives from this CBCR subclass, Slr1393g3 (PDB code 5DFX, green) and NpR6012g4 (PDB code 6BHN, blue) in their respective Pr dark states. The PCB chromophore and its thioether linkage to the conserved Cys residue of the protein are shown as sticks using the same color-coding scheme as for the ribbons. B Close-up view of the chromophore pocket in AnPixJg2. Key hydrogen-bond contacts between the chromophore and its binding pocket are indicated by green dashed lines, water molecules in the cavity as red spheres. Two conserved Phe residues, F268 and F329 proposed to be important for the Pg photoproduct formation are shown in sphere, and the two His residues, H293 and H322 flanking the B–C plane, shown different protonation states in the photoproduct are shaded. C Tetrapyrrole rings A–D are shaded in purple, pink, orange, and green, respectively, and the representative chromophore carbon and nitrogen atoms are labeled for reference. D UV-vis spectra of AnPixJg2 in the Pr (red) and Pg (green) photostates
As members of the red/green CBCR–GAF domains, AnPixJg2 (λmax = 648 nm and 542 nm for Pr and Pg, respectively) together with two other orthologs, Slr1393 GAF3 domain of Synechocystis 6803 (Slr1393g3, λmax = 649 nm and 535 nm; ref. [6]) and NpR6012 GAF4 domain of Nostoc punctiforme (NpR6012g4, λmax = 650 nm and 542 nm; ref. [7]) are the best-studied representatives so far. These sequentially similar photosensors show nearly congruent crystal structures of their respective Pr states, in particularly of the binding pocket (Fig. 1A). However, they show different dark-reversion kinetics from the photoproducts, and more prominently, the in vitro phycocyanobilin (PCB) assembly of holoproteins produce distinct effects on the bilin protonation and absorption maximum of the photoproduct. Specifically, the in vitro PCB-assembled holo-AnPixJg2 exhibits at pH 7.8 a deprotonated photoproduct with a minor λmax shift (548 nm, in vitro vs 542 nm, in vivo; ref. [37]). For Slr1393g3, irrespective of the in vivo and in vitro preparations, the photoproduct bilin remains protonated in the pH range from 6.0 to 9.0. However, an astonishing red-shift of 55 nm was detected for the photoproduct (orange-absorbing) of this CBCR comparing the in vitro assembled protein to its native Pg form (590 nm vs 535 nm; ref. [6]). Taken together, these results demonstrate that a highly precise bilin assembly is essential for the isolated CBCR domains to be properly functional, but how the assembly-related structural perturbations in the bilin-binding pocket respond to a pH change (e.g., within the physiologically relevant pH range) is not yet clear.
CBCRs naturally contain heterogeneous mixtures of populations (for both the chromophore and nearby residues) that are structurally, photochemically, and spectrally distinguishable [7, 8, 25, 27,28,29,30,31, 37, 38, 46]. Inherent links between the structural variations and photochemical/spectral heterogeneity in the red/green CBCR subfamily caught speed by the growing number of high-resolution crystal and solution NMR structures in various photostates [5,6,7]. Indeed, on the basis of solution structures of NpR6012g4 in both Pr and Pg photostates, Ames and coworkers have correlated the minority population of a highly conserved Asp residue (D657) to the photochemically impotent orange-absorbing population observed in its Pr state [7]. Our recent combined experimental and theoretical studies on AnPixJg2 and Slr1393g3 demonstrated that loss of a hydrogen bond between a conserved Tyr residue (e.g., 302 in AnPixJg2, Fig. 1B) and the structurally heterogeneous C-ring propionate side-chain causes the occurrence of an additional far-red-absorbing (Pfr) population in both Pr dark states [46]. Through molecular dynamics (MD) simulations of AnPixJg2 in the Pr dark state, Schapiro and coworkers further proposed that the appearance of the cationic imidazolium form of the conserved H322 (Fig. 1B) induces structural heterogeneity of the chromophore [25], thereby providing clues for understanding the molecular origin of the dark-state heterogeneity.
Here we performed a series of MAS NMR experiments with the in vitro u-[13C,15N]-PCB-assembled holo-AnPixJg2 in both Pr dark state and Pg photoproduct (Fig. 1C, D) to further our mechanistic understanding of red/green CBCR photocycle. In addition to identifying geometrical changes of the bilin associated with photoconversion, the complete and unambiguous 1H, 13C, and 15N assignments of the PCB atoms provide clear evidence for a structurally heterogeneous bilin in both photostates, in particular for the Pg bilin which manifests itself as the mixture of a vast number of conformational states. We also investigated protein–chromophore interactions in the Pg photoproduct aiming to a detailed description of protonation dynamics and charge distribution associated with the residues in the proximity of the chromophore. Both tentative 1H assignment for the surrounding protein and our quantum mechanics/molecular mechanics (QM/MM)-optimized structural model for the photoproduct suggest the presence of a cationic imidazolium side-chain for H322 in this photostate, thus generalizing the recent MD results that dealt solely with the effect of the protonation state of this residue on the structural heterogeneity of the AnPixJg2 PCB in the Pr dark state [25]. We further examined pH effects on this in vitro assembled protein. Analysis of the chemical shift changes associated with a 0.6-unit pH drop (7.8 → 7.2) demonstrated a substantially altered electronic structure of the photoproduct bilin chromophore, and more strikingly, a fully protonated (cationic) bilin π system in both photostates instead of the deprotonated (neutral) photoproduct bilin that is present at pH 7.8 [37].
2 Methods
2.1 Sample preparation
2.1.1 Protein preparation for MAS NMR
Cells transformed with the plasmid pET28a_AnPixJg2 [15] were kindly provided by Dr. R. Narikawa (Tokyo Metropolitan University, Japan). Protein expression was induced by addition of isopropyl-β-d-1-thiogalactopyranoside (IPTG, final concentration 0.8 mM) at OD600 = 0.8 for 16 h at 16℃. The cell pellets were resuspended in ice-cold 50 mM Tris–HCl (pH 7.2), 0.2 M NaCl buffer and disrupted ultrasonically (0℃, pulse on: 1 s, pulse off: 2 s, total duration time of 24 min; Nanjing Safer Biotech, China). The lysate was then clarified by centrifugation at 28,000 rpm for 60 min (fixed angle rotor Ti60 by Beckman-Coulter, Krefeld, Germany) at 4℃, and u-[13C,15N]-PCB (500 nmol, i.e., ~311.7 μg) was added to the supernatant at 20℃ in darkness. The labeled PCB chromophore was prepared as described [8]. Autoassembly was controlled by UV–vis spectroscopy and was complete within 30 min. The samples were then purified via Ni2+-affinity chromatography on chelating Sepharose (GE Healthcare), holoproteins being eluted with 50 mM Tris–HCl (pH 8.0), 0.5 M imidazole. Subsequently, imidazole was removed by dialysis against 50 mM Tris–HCl (pH 7.2), 0.2 M NaCl. The holoprotein solutions were concentrated using Amicon filters (Ultra Centrifugation Filters, MWCO 10 kDa) to 100 μL at 4℃ by centrifugation at 4500 rpm. Prior to the NMR measurement, the protein was loaded into a ~1.4 mm diameter glass capillary using a 100 μL Hamilton syringe and irradiated with either 530 nm or 650 nm light for 2 min from an array of LEDs to prepare the Pr and Pg at near 100% occupancy [8, 37]. Approximately 3.0 mg and 2.5 mg of AnPixJg2 holoproteins as Pr dark state and Pg photoproduct were used in the study. The samples were then transferred into 4-mm ZrO2 MAS rotors and snap-frozen in liquid N2 for subsequent NMR measurements. No additional illumination was applied to the samples during the data acquisition.
2.1.2 Protein UV–vis spectroscopic analyses
Diluted aliquots of all samples were investigated by UV spectrophotometer (Shimadzu 1900i, Duisburg, Germany). Full photoconversions were ensured by irradiation with 530 nm and 650 nm light for 2 min (Fig. 1D). Reversibility of photochemistry was confirmed via 530 → 650 → 530 nm irradiation cycles conducted at 20℃.
2.2 MAS NMR data collection
All MAS NMR data reported in this work were obtained on a Bruker AVANCE-III 400 MHz WB spectrometer (9.4T, Rheinstetten, Germany) equipped with a 4-mm double-resonance MAS probe at −25 ± 0.2℃ (readout temperature). The sample rotor was inserted into the precooled MAS stator and spun at 500–700 Hz upon freezing. The MAS rate of 8 kHz in 13C–13C dipolar correlation (DARR) experiments and 8.176 kHz in conventional 1H–13C and 1H–15N heteronuclear correlation (HETCOR) and 1H–13C medium- and long-distance HETCOR (MELODI–HETCOR) experiments was maintained ±3 Hz with a pneumatic control unit. Optimized 1H, 13C, and 15N π/2 pulse lengths were 2.5, 4.1, and 4.7 µs, respectively. CP was optimized to satisfy n = ±1 Hartmann–Hahn conditions with 1H power ramped 70–100% and a spin-lock field of 74.7 and 32.4 kHz for 13C and 15N, respectively. Swept-frequency two-pulse phase modulation heteronuclear decoupling [47] with 100 kHz radio-frequency (RF) strength was used during data acquisition. 13C and 15N chemical shifts were externally referenced with respect to the C(O)O– signal of solid L-tyrosine·HCl at 172.1 ppm and the NH4+ signal of solid 15NH4NO3 at 23.5 ppm, respectively.
2D DARR experiments were carried out with an optimized mixing time of 50 ms and a CP contact time of 1.8 ms. The n = +2 rotary-resonance condition was achieved with 1H continuous wave irradiation at RF field strength of 16 kHz. The spectra were acquired with 114 t1 increments, accumulating 2016 scans in each indirect slice with a relaxation delay time of 2.09 s. Both 1H–13C/15N HETCOR experiments were carried out with a CP contact time of 1 ms. Homonuclear 1H dipolar decoupling was achieved with the supercycled phase-modulated Lee–Goldburg (PMLG5-S2) scheme [48]. The optimized PMLG5 pulse was 1.33 µs with 88.5 kHz r.f. strength. Optimization was done by observing the J-splitting in adamantane powder by recoding PMLG5-S2-decoupled CP spectra, and further fine-optimized by monitoring the splitting between the CH2 protons of solid α-glycine in the indirect dimension in 1H{PMLG5-S2}–1H{wPMLG5-S2} homonuclear correlation experiment [49]. The scaling factor of 0.32 was experimentally determined [48]. The 2D HETCOR spectra were acquired with 48 t1 increments, accumulating between 3024 and 4480 scans in each indirect slice with a relaxation delay time of 2.5 s for recording 1H–13C and 1H–15N correlations, respectively. A 45° shifted squared sine-bell window function was applied in the indirect dimension, and further zero-filled to 1024 points prior to Fourier transformation. A 90° shifted squared sine-bell window function was applied in the t2 dimension and zero-filled to 4096 data points. For 1H–13C MELODI–HETCOR experiments on the photoproduct, an LG–CP contact time of 2.1 ms was used allowing for selective transfer of the 1H magnetization within the region of ~4.0–4.5 Å, as experimentally determined. The spectrum was recorded with 60 t1 increments, accumulating 4200 scans in each indirect slice with a relaxation delay time of 2.2 s. The t1 data were recorded in an off-resonance manner and linear predicted by 32 points using 40 LP coefficients. A 90° shifted squared sine-bell window function was applied, and zero-filled to 2048 points prior to Fourier transformation. The t2 data were zero-filled to 4096 points and 25 Hz line broadening for exponential multiplication.
2.3 Structural modeling
Owing to the lack of a crystal structure for the AnPixJg2 Pg photoproduct and for the sake of comparability, we took equilibrated protein structures of both Pr and Pg photostates from the closely-related Slr1393g3 [6]. Our starting points for creating the structural models were the last snapshots of the DFTB2+D/AMBER MD simulations reported in the previous study [24]. These structures were optimized via QM/MM by utilizing the ORCA-ChemShell interface [50, 51]. For optimizations, a 66-atom QM region including only the conjugated system of the chromophore (C4–C19) was employed. This region was described with the BLYP functional [52, 53] including Grimme dispersion correction with Becke-Johnson damping [54, 55] combined with the def2-SV(P) basis set [56] and employing the resolution-of-identity approximation with corresponding auxiliary basis [57]. The rest of the protein was described with the AMBER force field [58], whereas TIP3P [59] was used for the water molecules. In all calculations, a cut-off of 12 Å for electrostatic interactions was applied. The final optimized protein models for both photostates are available as PDB files and can be provided by the authors on request.
These structures were then taken to calculate excited states and absolute shieldings. The vertical excited-state calculations employed the simplified time-dependent density functional theory method [60,61,62] based on ground-state calculations with CAM-B3LYP functional [63] and def2-SV(P) basis by utilizing Orca version 4.2.1 [50]. The gauge-including atomic orbital (GIAO) method [64] was chosen to obtain chemical shieldings. The NMR calculations were realized with the Gaussian 16 software suite [65] and employed the B3LYP functional [66] with Jensen’s pcSseg-2 basis [67] retrieved from the Basis Set Exchange database [68]. To obtain reliable results for excited states and absolute shieldings, the QM calculations did not only include the chromophore, but also side-chains of nearby amino acids and water molecules. The calculations were performed with 310 and 331 atoms for Pr and Pg, respectively. Both models included the identical side-chain atoms and seven additional water molecules were included for Pg. The structures of the active sites for both Pr and Pg photostates can also be provided by the authors on request. For the NMR-refined structural model of the AnPixJg2 photoproduct, H293 from AnPixJg2 was positioned similarly to Y500 from Slr1393g3, the subsequent refinement of protonation and side-chain geometry of this His residue was guided by the NMR-determined chromophore interfacial 1H contacts.
3 Results
3.1 NMR spectroscopic analyses
3.1.1 Manifold conformational states of the bilin chromophore in both photostates
Our previous MAS NMR study on AnPixJg2 predicted the coexistence of at least two conformational subpopulations of the PCB chromophore in each photostate deduced from the lineshape analysis of the unusually broadened 13C signals in the 1D spectra [27]. However, atomic resolution of the chromophore subpopulations was not provided due to the low amount of protein available and the poor chromophore incorporation ratio which precluded observation of well-resolved spectral lines in a 2D mode [8, 27]. We here report unambiguous 1H, 13C, and 15N assignments of the AnPixJg2 PCB atoms in both Pr dark and Pg photoproduct states using an improved sample preparation method (see Sect. 2.1.1), thus providing the basis for spectral identification of different chromophore subpopulations. Our assignment strategy relies on a series of 2D homo- and heteronuclear MAS NMR dipolar correlation experiments that enabled complete chromophore assignments for the canonical and Cph2 phytochromes in various photostates [38, 39, 69, 70]. We initially performed 13C–13C DARR experiments to assign the PCB carbons in both photostates with a mixing time of 50 ms (Fig. 2A, B for Pr and Pg, respectively). Both DARR spectra are dominated by medium- and long-range carbon pairs having internuclear C···C distances of ~3.1–3.8 Å; these values concur with the crystal structure of AnPixJg2 in the Pr dark state (PDB code 3W2Z) and in analogy with that of Slr1393g3 in the Pg photoproduct (PDB code 5M82). A similar detection limit of 3.3–3.6 Å at this mixing time for chromophore correlations was defined in the isolated Cph2-type All2699g1 construct from Nostoc [39].
Heterogeneity at bilin carbons in both photostates. A 2D 13C–13C DARR spectrum of AnPixJg2 holoprotein assembled with the u-[13C,15N]-PCB chromophore in the Pr dark state. B DARR spectrum of the corresponding Pg photoproduct as in (A). A mixing time of 50 ms was employed for 13C homonuclear recoupling. Both directly and indirectly bonded carbon pairs of the chromophore (see Fig. 1C for numbering) are indicated by arrows and labeled in blue. Observed signal splittings of a subset of 13C resonances are indicated by superscripts a, b, c, and so on from the high- to low-field side. C 13C chemical shifts of the PCB carbons in both photostates (see also Table S1). Each 13C resonance is represented by a solid stick. Multiple resonances from a given PCB carbon are colored the same and their mean chemical shift values are indicated by solid triangles filled using the same color-coding scheme as in Fig. 1C and labeled
For the DARR analysis, we selected the spectral region of 85–115 ppm for bilin methines (C5, C10, and C15, Fig. 1C for numbering) as the starting point. For example, the Pr correlations appearing at ~90 ppm can be unambiguously assigned to C5 by virtue of its distinct high-field 13C chemical shift among those three carbons seen in the form of free PCB [71] and also after the binding into the protein pockets of other bilin-based photosensors [35, 39, 72]. The slice corresponding to C5 (90.3 ppm) showed two correlations at 39.4 and 38.1 ppm (Fig. 2A) that can be solely assigned to C31 by considering their chemical shift values and the maximal detection of ~3.8 Å. Using the same logic, we attributed the two unassigned correlations in the C5 slice at 149.7 ppm to C6 and at 54.0 ppm to C3, the latter in turn gave an expected correlation with the slice matching C31 at 38.1 ppm, thereby corroborating these assignments. The complete 13C bilin assignments in both photostates were complicated by signal splitting of a subset of carbon resonances, indicative of possible conformational states retained by the bilin. Taking the known C5 slice at 90.3 ppm as an example: the correlations from C31 in the Pr state is doubled with two resonances (termed as C31a and C31b from high- to low-field side) separated by 1.3 ppm, and moreover, C31b (38.1 ppm) further revealed an additional C5 component (C5b) at 88.6 ppm via the correlation network connecting C31b–C3a–C5b–C9 (Fig. 2A). Also, the correlation network in the A-ring region revealed a splitting for C3 (C4 [149.2 ppm]–C3a [54.0 ppm]/C3b [51.1 ppm]) and C32 (C31a [39.4 ppm]–C32a [25.1 ppm]/C32b [23.9 ppm]) as well as a doubling for both C2 and C21 (C2a [37.3 ppm]–C21a [18.5 ppm] and C2b [36.7 ppm]–C21b [16.1 ppm]) and a tripling for C1 (C21a–C1a [184.6 ppm]/C1b [183.5 ppm] and C2a–C1c [182.2 ppm]) with a chemical shift separation of 2.4 ppm. Although signal doubling is observed for the C5-methine carbon with a resonance separation of 1.7 ppm, the splitting of its A- and B-ring neighbors C4 and C6 was not apparent. Similarly, the correlation network in/around the D-ring region showed a single set of chemical shift for C14 (146.8 ppm) and C16 (144.4 ppm), while revealing a splitting for the C15-methine bridge (C15a [97.9 ppm]/C15b [96.7 ppm]). No such splitting was resolved for the central C10-methine carbon (116.0 ppm) bridging rings B and C.
Interestingly, in the Pr dark state, the B-ring propionate side-chain carbons (C81, C82, and C83) showed a correlation network split for C81 (C8 [147.7 ppm]–C81a [21.7 ppm]/C81b [19.8 ppm]–C82 [38.6 ppm]–C83 [182.5 ppm]), while for their C-ring counterpart, the signals from all three carbons (C121, C122, and C123) are evidently doubled (C121a [24.8 ppm]/C121b [23.9 ppm], C122a [40.0 ppm]/C122b [38.9 ppm], and C123a [181.9 ppm]/C123b [180.0 ppm]). The concurrent 13C splitting of the C-ring propionate resonances adds the second independent line of evidence for the heterogeneity and flexibility of this side-chain. Moreover, the most pronounced 13C signal splitting of the Pr chromophore in terms of the number of split components of the 13C resonances and magnitude of their separations occurs at C12, the pyrrolic carbon carrying this side-chain, for which we identified a quadruple set of chemical shifts via the correlations of C11a (131.1 ppm)–C12a (147.0 ppm), C11b (129.4 ppm)–C12b (145.5 ppm), C122b–C12c (143.9 ppm), and C122a–C12d (142.5 ppm) with a separation of 4.5 ppm (Fig. 2A). The former two connecting C11 and C12 also revealed a clear doubling for C11 with a smaller separation of 1.7 ppm. However, the C11b–C12b correlation is much more intense than that of C11a–C12a, having a relative intensity of ~5.5:1, as estimated from the integrated peak areas of the two structural components. Such signal splittings of the pyrrolic carbons in the π system (C4–C19) is also seen for C18 and C19 in the ring D: the correlations of C18 with C181 and C182a (C18a [135.3 ppm]/C182a [13.9 ppm], and C18b [134.8 ppm]/C181 [17.1 ppm]) revealed a small but resolved doubling of the C18 resonances with a separation of 0.5 ppm, while the two C19 resonances (C19a [174.1 ppm]/C182a [17.1 ppm], and C19b [171.2 ppm]/C182b [13.9 ppm]) are separated by 2.9 ppm.
An overall similar strategy was adopted for complete 13C assignment of the photoproduct bilin (Fig. 2B). However, the DARR spectrum showed more complexity in the correlation network split for most carbon atoms which seems to arise from increased conformational flexibility of the bilin chromophore in the photoproduct. Unlike the Pr dark state, in which the 13C signal splittings mainly occur at the carbons in the A-ring region, the C-ring propionate side-chain, and the C5- and C15-methine bridges, nearly all 13C resonances of the photoproduct bilin are at least split up into a doubling except for C8 and C82 locating solely at 149.0 and 39.0 ppm, respectively (Fig. 2C). More features are as follows: (i) 13C doubling is the dominant splitting mode for the π-conjugated C4–C19 system (12 out of 16), with the exceptions being C4, C13 and C19, for which a tripling was resolved with a separation of 3.5, 1.9, and 4.0 ppm, respectively. Among these pyrrolic carbons, the largest separation was seen for the C5 resonances (5.3 ppm), and also its B-ring neighbor C6 showed a large separation of 4.9 ppm. Separations of ≥1.9 ppm are also seen for C9, C15, and C17. (ii) The splitting is most significant in the 13C resonances in/around A-ring region, analogous to those observed in the Pr dark state. A total of 21 split components corresponding to eight carbons in this region were detected. More specifically, like C4, we identified a triple set of chemical shifts for C1, C21, C3, and C31, with a 13C separation of 1.9, 3.4, 1.4, and 3.5 ppm, respectively. Moreover, although 13C splitting patterns at other A-ring components like C2, C32, and C5 in the photoproduct are retained, their resonances are larger separated than those in the dark state (Fig. 2C). (iii) Contrary to nearly all bilin carbons showing additional split components in the photoproduct, the C12 resonances split up into a small doubling instead of the quadrupling identified in the dark state with a 3.7-ppm reduction of the resonance separation. For the propionate side-chain at C12, however, the resonances of C121 and C123 were further split into a tripling upon photoproduct formation, with their separations being at least 0.7 ppm larger than those in the dark state (0.9 vs 3.1 ppm for C121 and 0.9 vs 1.6 ppm for C123). No such complications arise in the B-ring counterpart, for which the splitting patterns of C8, C81, and C82 remained unaffected in the photoproduct, while an additional doubling was resolved for C83 (C83a [182.2 ppm]/C83b [181.1 ppm]). (iv) The largest change in the 13C resonance splitting occurs at the B-ring methyl group, C71 which exhibited a single DARR correlation network in the dark state (9.6 ppm), but split into a quadrupling in the photoproduct at 9.9, 9.3, 8.3, and 7.4 ppm. Similar state-related changes were also seen at the C131 and C171 methyl groups of rings C and D. For C171, although only an additional doubling was resolved in the photoproduct, the separation is comparable to that of C71 (2.6 vs 2.5 ppm, Fig. 2C).
Following the complete set of 13C assignments of the chromophore, we performed 1H–13C correlation experiments to build up intramolecular correlations between the bilin NH protons and carbons (with a cutoff distance of ~3.2 Å by choosing a contact time of 1.7 ms, ref. [39]), permitting straightforward 1H assignment for all four NH protons in the dark and photoproduct states. The spectral region of 8–15 ppm is characteristic for NH protons, in which we identified at least seven 1H slices for each photostate (Fig. 3A). For instance, in the Pr dark state, the NH resonating at 13.0 ppm can be assigned to H22 of ring B via the well-resolved correlations with C9 in the same ring and the neighboring C10-methine bridge. The NH resonance at 10.7 ppm is present in the slices corresponding to C10, C11a&b, and C12a–c which can be solely attributed to the intramolecular magnetization transfer from H23 of ring C, whereas a contribution from H22 is virtually impossible, since the shortest interatomic distance is of 4.4 Å between H22 and C12 (extracted from the 3W2Z structure), well beyond the ~3.2-Å detection range experimentally determined for the intramolecular transfer events. The spectrum also revealed the doubly split 1H resonances of H21 (H21a [11.1 ppm]/H21b [10.8 ppm]) of ring A via the correlations with its neighboring C1a–c and C4 in the same ring, and also the D-ring H24 resonances are evidently tripled (H24a [12.6 ppm]/H24b [12.2 ppm]/H24c [11.0 ppm]) with a large 1H separation of 1.6 ppm (Fig. 3B). The corresponding photoproduct spectrum revealed an increased total number of 1H slices. Among these, H22 retained a single set of resonance with the 1H chemical shift of 13.2 ppm, slightly downfield shifted relative to that of the dark state. Besides H22, the resonances of the remaining three NH protons were clearly split: the H21 and H23 protons each showing one more 1H split component in the photoproduct while the H24 resonances splitting up into a small but resolved doubling (with the 1H separation of 0.1 ppm) instead of the pronounced tripling seen in the dark state (Fig. 3B).
Heterogeneity at bilin NH moieties in both photostates. A 2D 1H–13C supercycled PMLG-decoupled dipolar correlation spectra of the u-[13C,15N]-PCB-AnPixJg2 in the Pr dark state (red) and Pg photoproduct (green). The characteristic spectral region of δH = 8–15 ppm is displayed for tracing intramolecular 1HN21–N24–13CPCB correlations. B 1H chemical shifts of the 1HN21–N24 protons for both photostates (see also Table S2). C 2D 1H–15N supercycled PMLG-decoupled dipolar correlation spectra of the u-[13C,15N]-PCB-AnPixJg2 in the Pr (red) and Pg (green) states for 1H–15N connectivities of the four NH moieties. D 15N chemical shifts of the bilin nitrogens (N21–N24) for both photostates (see also Table S3). For a subset of bilin nitrogens and the protons attached showing multiple sets of chemical shifts, their 15N and 1H signals are indicated by superscripts a, b, and c from the high- to low-field side. Each resonance is represented by a solid stick and labeled with chemical shift. Multiple resonances from a given atom are colored the same and their mean chemical shift values are indicated by solid triangles filled using the same color-coding scheme as in Fig. 1C and labeled
We further extended the experiments by using a short-range 1H–15N HETCOR experiment to correlate bilin nitrogens to the known NH protons via their direct connectivities, thereby permitting the complete 15N assignment of all four nitrogens in both photostates resonating at 130–165 ppm (Fig. 3C). In the dark state, no 15N resonance splittings are apparent for N22 and N23 of the rings B and C which are located at 148.4 and 155.2 ppm, respectively. However, both N21 and N24 of the two outer rings A and D are clearly doubled with a 15N separation of 1.1 and 1.6 ppm, respectively (Fig. 3D). In the photoproduct, a tripling with a large separation of 2.7 ppm was resolved for N21, and moreover, the further split also occurs at N23, with the two resonances separated by 2.8 ppm (Fig. 3D). For the other two bilin nitrogens, although no such additional 1H split components were detected, the 1H separation of N24 doubling is 0.5 ppm larger than that for the dark state, and, strikingly, the single resonance of N22 is downfield shifted from 147.4 to 162.0 ppm, representing the largest 15N chemical shift change upon photoconversion (ΔδNlight) for the bilin nitrogens in AnPixJg2.
3.1.2 Light- and pH-dependent chemical shift changes of the bilin chromophore in AnPixJg2
We have obtained complete 1H, 13C, and 15N assignments for the PCB atoms of AnPixJg2 in both photostates (summarized in Tables S1–S3) at a pH of 7.2. The state-related changes in their chemical shifts upon photoconversion are illustrated in Fig. 4A, with the size of the spheres for a given PCB atom referring to the difference in mean chemical shift between the dark state and photoproduct resonances. We also analyzed the pH effect on the bilin in the in vitro assembled AnPixJg2 by comparing current NMR data acquired from the sample at pH 7.2 to the values previously reported for the same protein at a higher pH of 7.8 [37], although for both photostates there are no detectable changes in the absorption spectra with respect to the peak positions and the band shapes upon this pH shift (Fig. S1). Instead, an unusual temperature effect has been observed for both Slr1393g3 [32] and NpR6012g4 [7], in which cooling of the sample onsets the formation of an orange-absorbing species that is distinct from the major Pr dark state. For the in vitro assembled AnPixJg2 at pH 7.2, however, we found no indication for such changes of the 1D 13C MAS NMR spectra between −45 and −25℃ (Fig. S2).
Light- and pH-induced changes in 13C, 15N, and 1H chemical shifts of the PCB chromophore in AnPixJg2. A Chemical shift changes associated with the photoconversion at pH 7.2 are represented as red and blue circles for down- and upfield shifts as Pg photoproduct, respectively. B Corresponding changes of the Pr chromophore upon tuning down the sample pH from 7.8 to 7.2. C Corresponding changes of the Pg chromophore as in (B). Red and blue circles represent for down- and upfield shifts at pH 7.2 (B and C)
Unlike the formation of Pfr photoproduct in canonical phytochromes that affects most the pyrrolic carbons in/around the ring D [5], nearly all AnPixJg2 resonances of the π-conjugated C4–C19 system (15 out of 16) show only subtle changes of −1.9 ≤ ΔδClight ≤ 2.5 ppm upon Pg formation at pH 7.2 (Fig. 4A). In particular, the pyrrolic quaternary carbons including C6, C9, C11–C13, and C19 appear to be only marginally affected with |ΔδClight|≤ 1.0 ppm. Exception is C18 of ring D, showing a downfield shift of 4.4 ppm. Other prominent ΔδClight changes of the bilin chromophore were found at the ethylidene C31 and C32 positions which are oppositely shifted by 5.4 (to downfield) and 8.9 ppm (to upfield), respectively, and also a 2.8-ppm downfield shift was detected for the A-ring methyl side-chain, C21. Although the photo-conversion produced only marginal 13C changes at C6 and C9 of ring B (0.3 and 0.9 ppm, respectively), a striking 13.6-ppm 15N downfield shift was found for their adjacent pyrrole nitrogen N22. Further prominent 15N changes occur at N21 and N24 of rings A and D that are upfield shifted by 7.5 and 4.4 ppm, respectively, and moreover, ΔδHlight changes of these two NH groups are of 1.3 and 1.9 ppm, both are downfield shifted. Intriguingly, although the strikingly large ΔδNlight change at N22, its attached proton (H22) is downfield shifted by only 0.2 ppm.
AnPixJg2 shows a nearly identical photocycle to that of NpR6012g4, for which complete 1H, 13C, and 15N assignments for the PCB atoms are available in both photostates from solution NMR spectroscopy [34, 35]. Compared to NpR6012g4 the photoconversion affects the AnPixJg2 chromophore in a similar manner in terms of the sign and magnitude of ΔδClight changes (Fig. S3). Conservation of the sign of ΔδClight changes is observed for 11 (out of 16) bilin carbons in the π-conjugated C4–C19 system. Three notable exceptions include C6 of ring B and C12 and C13 of ring C, showing an opposite sign of ΔδClight upon photoconversion with the large ΔδClight deviations for the two proteins of 4.0, 3.6, and 2.3 ppm, respectively. Other prominent deviations are mostly localized in ring A: the signs of ΔδClight changes at C1, C2, and C31 are inverted, with the deviations between the two proteins of 1.4, 5.1, and 5.9 ppm, respectively, and at the ethylidene C32 position, although the sign is conserved, the photoproduct resonance in AnPixJg2 is 5.5 ppm less shifted (upfield) than that of NpR6012g4. By contrast, the two ΔδClight changing patterns in the region including C14 of ring C, C15-methine bridge as well as the entire bilin ring D are surprisingly similar. The sign of the pattern is the same for both CBCR proteins and the largest ΔδClight deviation in this region amounts to only 1.9 ppm, as seen for C14 and C171. Moreover, the ΔδClight deviations of the rest of carbons are of ≤1.2 ppm (Fig. S3). Similar to NpR6012g4, the largest ΔδClight changes in the AnPixJg2-conjugated system occur at C10 and C18, although an overall decrease in magnitude at both positions is detected (2.0 and 0.6 ppm for C10 and C18, respectively). There is also a good agreement between the bilins of the two proteins for the strikingly large ΔδClight changes of the ethylene side-chain, again implying the movement of the A-ring linkage (to the rest of the ring) during photoconversion in both proteins. For the bilin NH moieties, the ΔδN,Hlight changes in the two proteins are quantitatively similar with a largest deviation seen at ring A but amounted to only 2.3 ppm for N21 and 0.5 ppm for H21. Moreover, although the ΔδNlight change is particularly noticeable for N22 of ring B in both cases (>13 ppm, Fig. S3), a small deviation of 1.3 ppm was detected in the two proteins at this position.
The bilin 13C and 15N chemical shift changes associated with the 0.6-unit pH drop (ΔδpH) are illustrated in Fig. 4B, C for the Pr dark state and Pg photoproduct, respectively (see also Tables S1–S3). Overall, 15N chemical shifts of all four bilin nitrogens in the respective Pr dark states are hardly altered by the pH shift (2.2 ≤|ΔδNpH|≤ 2.7 ppm, Fig. 4B), whereas the 13C resonances of 12 (out of 16) bilin carbons across the conjugated system are downfield shifted with large ΔδCpH changes (≥3.1 ppm) occurring at the C5-methine bridge, C7–C9, and C18. For the peripheral bilin side-chain carbon atoms, changes of ≥ 2.4 ppm are seen for C82 and C121 of the two propionates as well as the D-ring methyl C171, with a particularly large ΔδCpH change of 12.1 ppm at the ethylidene C31 position (Fig. 4B). Surprisingly, this small pH drop produced much more pronounced ΔδpH changes for most bilin carbon and nitrogen atoms of the two Pg photoproducts (Fig. 4C), despite their identical absorption properties (λmax = 542 nm, see Fig. S1). In particular, upon the pH shift from 7.8 to 7.2, the B-ring N22 resonance shifts dramatically by 38.9 ppm toward the upfield region which distinctly narrows the δN dispersion of the 15N resonances from 55.6 to 31.5 ppm (calculated as δN22 − δN24). The narrow dispersion of the four bilin nitrogens, together with the 1H and 15N photoproduct data of the bilin NH moieties (Fig. 3C) provides clear evidence that AnPixJg2 has a protonated cationic bilin ring system at pH 7.2 instead of the deprotonated one found at pH 7.8 [37]. Intriguingly, the large ΔδCpH changes associated with the reprotonation at the B-ring nitrogen are not confined at the adjacent carbons like C6 (3.6 ppm) and C9 (2.8 ppm), but spread over the entire bilin molecule. Specifically, almost all bilin carbons (15 out of 16) in the conjugation system show ΔδCpH changes ≥ 2.8 ppm, among which prominent changes ≥ 5.4 ppm were seen for C5, C8, C13, C16, and C17 (Fig. 4C). Moreover, most 13C resonances of these pyrrolic carbons move downfield (as represented by red circles), implying a decrease of electron density at pH 7.2, in particular for ring C. By contrast, most carbons at the periphery of the bilin are oppositely upshifted, e.g., the A-ring ethylidene side-chain (C31 and C32), the C-ring methyl (C131), and the D-ring methyl and ethyl side-chains (C171, C181 and C182), where the largest changes occur at the ethylidene C31 and C32 positions for 8.6 and 6.1 ppm, respectively. No such consistent changing pattern occurs with any of the propionate side-chains, and additionally, we observed a significant downshift of 7.2 ppm for the carboxylate moiety at the C-ring propionate (C123), most likely arising from the strongly altered hydrogen-bonding interactions with the protein environment.
3.1.3 Characterization of protonation states of H322 and H293 in the photoproduct pocket
As for plant and Cph1-like phytochromes [70, 72], we identified the binding site protein–chromophore interactions in the photoproduct state of AnPixJg2 using 1H–13C MELODI–HETCOR experiments which allow the 13C-labeled bilin carbons to probe the nearby residues (and also structural water molecules) located within a sphere up to ~4.5 Å (1Hresi.–13CPCB). The tentative 1H assignments of the protein surrounding were guided by the photoproduct structural data of Slr1393g3 (PDB code 5M82) and NpR6012g4 (PDB code 6BHO) and performed as described [70]. For instance, the highly conserved H322-homologous residue in Slr1393g3 (H529) forms strong hydrogen-bonding interactions with the C-ring carboxylate moiety (C123) via its imidazole ring at Nε2 (two O···N distances of 2.7 and 3.5 Å) in the photoproduct. The 5M82 model also reveals one more imidazole proton, Hε1 at a distance of ~3.9 Å to C123, well within the detection range. Indeed, in the MELODI–HETCOR slices corresponding to C123 (182.9, 182.1, and 181.3 ppm), both interfacial transfer events from the H322 imidazole are present within the 1H spectral region of 7–14 ppm (Fig. 5A). This consistency justifies the use of 5M82 model as the template for 1H assignment of H322 in AnPixJg2, whereas no H322 1H correlations with the B-ring carboxylate moiety (C83) were detected (Fig. 5A). Such a situation is not consistent with the NpR6012g4 6BHO model, in which the H322-homologous residue (H688) is hydrogen-bonded to the B-ring carboxylate [7].
Protonation states of the two His residues flanking the B–C plane in the Pg photoproduct. A The 2D 1H–13C MELODI–HETCOR data of AnPixJg2 in its photoproduct. The spectral region is shown for recognition of H293 and H322 in the binding pocket via their interfacial contacts with the PCB carbons. 13C resonances are indicated by vertical lines and the PCB 1H contacts originating from H293 and H322 are labeled in brown and teal, respectively. B NMR-refined QM/MM-optimized model of AnPixJg2 photoproduct carried out with a cationic H322 protonated at both the ε- and δ-positions and a neutral ε-protonated H293. The associated hydrogen-bonding interactions with the chromophore are indicated by yellow dashed lines
As the buildup rate of a C–H correlation relies primarily on the shortest possible distance for interfacial magnetization transfer, we therefore assigned the stronger 1H signal at 13.0 ppm provisionally to Hε2 as hydrogen bonding to the C123-carboxylate. The same proton also correlates with C6 of ring B (151.5 and 147.2 ppm) and C171 of ring D (11.6 and 9.0 ppm). Considering the maximal detection range of ~4.5 Å, Hε2 is the only candidate in the imidazole ring that allows magnetization transfer from both carbons (the interatomic distances of 4.3 and 4.5 Å for H322-Hε2···C6 and Hε2···C171, respectively), thus corroborating the assignment. The other 1H signal at 8.6 ppm in the slices corresponding to C123b and C123c was assigned to Hε1 of the imidazole. However, the correlation from this proton to C123a is not resolved, implying structural flexibility of the C-ring propionate. This assignment can be confirmed via the correlations with a number of carbons in/around the B-ring region including C5a&b, C7a&b, C8, C81a&b, C82, C9a&b, and C10a&b (Fig. 5A). The distance constraints involving Hε1 are consistent with the proximity of Hε1 to ring B as seen in the Slr1393g3 5M82 model. In addition to Hε1, 1H signal resonating at 11.6 ppm could be assigned to Hδ1 of H322 by virtue of its strong correlations with the B-ring carbons, in particular for C5a, C6a, C7b, and C71b–d. All these correlated C–H pairs are predicted to be <4.0 Å based on the 5M82 model. We also assigned the remaining imidazole 1H signal at 7.7 ppm to Hδ2 which is observed to correlate with C6a&b and C11b, the H322-Hδ2···C11b pair, however, is predicted to be 4.9 Å apart. This could arise from the structural difference between AnPixJg2 and Slr1393g3, e.g., a positional/rotational shift of the H322 imidazole ring relative to the B–C plane. Overall, the data do not support the coexistence of neutral ε- and δ-protonated tautomers of this His residue, but suggest a cationic imidazolium moiety protonated at both nitrogens. Were two tautomeric states to coexist, one would expect at least one more set of 1H imidazole signals from Hδ2 and Hε1 in the region below 10 ppm. Potential hydrogen-bonding interactions involving the cationic H322 and the PCB chromophore in the photoproduct pocket are illustrated in Fig. 5B. This NMR-refined structural model is built on the basis of the QM/MM-optimized structure for the solvated Slr1393g3 photoproduct.
In the 1H spectral region above 10 ppm, the 13C slices matching C8 and C81a also exhibit the correlation to a proton resonating 12.0 ppm (Fig. 5A). With insights from available structural data of red/green CBCRs [5,6,7], this 1H signal can only be associated with the imidazole Hδ1/Hε2 protons of another His residue, H293, on the opposite β-face of the B–C plane (Figs. 1A and 5B). The His residue is conserved in NpR6012g4, but replaced by a Tyr residue in Slr1393g3 (Y500). We therefore used the NpR6012g4 6BHO photoproduct NMR structure to aid in 1H assignment of the H293 imidazole Hδ1/Hε2 protons. The H293-homologous residue in NpR6012g4 (H659) was found to be neutral and place its imidazole side-chain in proximity to the B-ring carbons such as C8, C81, and C9 as well as the C10-methine bridge. For instance, nine out of 10 lowest-energy photoproduct models show the Nδ1 atom sitting within the ~4.5-Å detection range of C10 with an average interatomic distance of 3.3 Å. Similarly, Nδ1 is also close to C9 with an average distance of 3.5 Å. By contrast, the Nε2 atom on the other side of imidazole is predominantly located at >4.5 Å away from C9 (9/10) and C10 (7/10), thus the intermolecular transfer from Hε2 to C9 or C10 is unlikely. The apparent absence of 1H signal at 12.0 ppm in the 13C slices matching these two carbons indicates that the imidazole of H293 in AnPixJg2 is not protonated at its Nδ1-nitrogen, but most likely at the Nε2 position. Moreover, the NpR6012g4 photoproduct structural models are also compatible with the H293-Hε2···C8/C81a contacts, for example in the latter case, seven of the 10 models showing the contact within 4.5 Å. The AnPixJg2 photoproduct model (Fig. 5B) also accounts for the possible magnetization transfers from Hε1 of this imidazole to C71, C8, and C81 (Hε1···C71 representing the longest intramolecular distance of ~3.8 Å), and indeed, the four slices matching C71a&c, C8, and C81b exhibited the expected 1H correlation of Hε1 at 8.1 ppm (Fig. 5A). Intriguingly, the Hε1···C71 contact is inconsistent with the distance restraints extracted from the NpR6012g4 structural models (8/10) which show an interatomic distance >4.6 Å, as such beyond the detection range. It is thus clear that H293 carries a neutral ε2-protonated imidazole, however, the protonation state of its backbone amino group is not yet known. In the C1a slice (38.1 ppm), there is a moderately strong correlation at δH of 9.3 ppm (Fig. 5A), however, the side-chains of nearby residues (within the detection range) showing no such signals in this 1H shift region, but instead indicating the backbone amino protons (HN) of H293 with the corresponding C···H distance of ~4.1 Å. The observed 1H chemical shift is consistent with that of the NH3+ group in a neutral backbone of histidine [73]. Were this 1H resonance to originate from the deprotonated NH2 group in an anionic backbone, the amino 1H chemical shift would be most likely 1–2 ppm upfield shifted.
3.1.4 Identification of structural water molecules around photoproduct chromophore
The 1H chemical shift range of 3.0–6.0 ppm in the correlation spectrum (Fig. 6A) is mainly associated with the Hα of all detected residues (within a sphere up to ~4.5-Å), the Hβ protons of Thr and Ser, the side-chain hydroxyl protons of Thr and Tyr, the ring Hζ protons of Phe, and moreover, the structural water molecules in the photoproduct pocket. Overall, at least six 1H signals located at 4.5–5.2 ppm can be attributed to the magnetization transfer from water protons (Wat1–Wat6). The 1H chemical shifts of these water molecules are in accordance with previously reported data of 4.7 ppm for the neutral pyrrole water in the canonical Cph1 phytochrome [69]. We used a similar strategy for identification of the water molecules based on their respective interactions with the bilin carbons. As an example, each of the MELODI–HETCOR slices matching C14a&b, C15a, and C16a exhibited a 1H signal at 4.5 ppm (Fig. 6A), however, in both Slr1393g3 5M82 and NpR6012g4 6BHO photoproduct models, these three carbons show no common residual 1H partners locating within the 4.5-Å detection range. Specifically, the central aspartate in Slr1393g3 places its Hβ proton at a distance of 4.3 Å to C15, but the proton is too far away to interact with C14 and C16 (at distances of 4.8 and 5.3 Å, respectively). Similarly, the Hζ proton of the ‘β2 Phe’ (F268) locates roughly within the detection range of C16 (4.6 Å), but it is much too distant to be detected by C14 and C15 (at distances of 5.4 and 6.0 Å, respectively). We therefore attributed the 1H signal at 4.5 ppm to a water molecule in the pocket (denoted as ‘Wat6’). Indeed, in the photoproduct structure for Slr1393g3, a number of water molecules, e.g., Wat816, Wat817, Wat827, and Wat863 were found to be in close contact with the C14–C16 segment. Among them, Wat817 is of particular interest because it is roughly equidistant (3.9–4.0 Å) from all three carbons, well within the detection range. As shown in Fig. 6A for AnPixJg2, the integrated intensity ratio of the four contacts involving Wat6 with C14a&b, C15a, and C16a is ~1.1:1:1.2:1, in harmony with the distance restraints extracted from the Slr1393g3 5M82 model, thus implying a similar spatial disposition of Wat6 relative to the bilin C14–C16 segment of the AnPixJg2 photoproduct (Fig. 6B). It is of course possible that other locations are present because of a limited number of unambiguous distance restraints available from the MELODI–HETCOR spectrum. We also determined the possible locations of all other water molecules nearby the chromophore (Wat1–Wat5). Unlike Wat6 that is completely buried in the protein interior and tightly bound to the central aspartate, Wat1, proximal to C71 and C81 at the periphery of the bilin, would not be sealed off from the solvent. The local electronic environment for Wat1 may thus be less shielded relative to Wat6, consistent with the 1H resonance of Wat1 being downfield displaced by 0.7 ppm (Fig. 6A).
Pg formation allowing hydration of the chromophore-binding pocket in AnPixJg2. A The 2D 1H–13C MELODI–HETCOR data of AnPixJg2 in its photoproduct. The 1H spectral region 3–6 ppm is shown for recognition of structural water molecules in the binding pocket via their interfacial contacts with the PCB carbons. 13C resonances are indicated by vertical lines and 1H resonances of multiple water molecules (Wat1–Wat6) are indicated by horizontal lines and labeled in blue. B In the Pr 3W2Z structure, three water molecules are detected in the pocket, however, entry of additional water molecules into the binding pocket of the Pg photoproduct is observed with at least six water molecules around the positions indicated by asterisks in red. The interfacial 1HWat–13CPCB contacts (as resolved in A) are indicated by yellow dashed line
3.2 Spectroscopic properties of the simulated protein structures
The extent of conjugation in the PCB chromophore has been found to be decisive for its lowest-energy absorption band [24]. To assess in how far the chromophore structures of both photostates are representative, we calculated absorption spectra for the QM/MM-optimized models of Slr1393g3 using two QM regions: (i) QM106, including the chromophore and the side-chains of D498, C528, and H529 as used in our previous studies [24, 26], and (ii) QM310/331 for Pr dark and Pg photoproduct states, respectively, including more side-chains of nearby residues and water molecules. As shown in Fig. S4, the simulated absorption of a given photostate for both QM regions does not strongly depend on this choice. In particular, the Q band of the photoproduct is nearly unchanged in intensity and position, whereas this absorption is slightly shifted to higher energies for the dark state when using a larger QM region. Overall, the simulated absorption in this region is in good agreement with experimental data. Therefore, the conjugation of the chromophore in the derived models appears to be representative for both photostates.
To further assess the structural models, we determined the absolute chemical shieldings of the pyrrole nitrogens which are sensitive to the chromophore geometry [74]. To compare our NMR calculations with the experimental data, we employed linear regression of the calculated absolute shieldings versus the experimental chemical shifts [75, 76]. As shown in Fig. S5, we obtained significant linear correlation with squared correlation coefficients of 0.88 and 0.82 for Pr and Pg, respectively (Table S4). Moreover, the slopes of the fits do not deviate much from the ideal value of −1. Using the equation from linear regression, we obtained chemical shifts from the computed absolute shieldings. Besides the value for the A-ring nitrogen atom, all calculated 15N chemical shift values deviate by <5 ppm from the experimental ones. Therefore, the QM/MM-optimized models of both Slr1393g3 photostates also appear in this regard to be a reasonable approximation for both AnPixJg2 photostates underscoring the similarity between the two proteins.
4 Discussion
4.1 The bilin ring D being strongly twisted in both photostates
In AnPixJg2, major ΔδClight changes occur at the A-ring ethylidene linkage to the protein (e.g., C31 and C32, see Fig. 4A), implying a mechanical distortion of the bilin ring A upon photoproduct formation. Indeed, paired Pr/Pg crystal structures of Slr1393g3 show that the ring A is predominantly in coplanar conformation with the B–C plane in the Pr dark state, but with a substantial torsion with respect to the ring B by ~69°Footnote 1 in the Pg photoproduct [6]. Such effects associated with ring A as well as its linkage to the protein would arise due to reframing of surrounding protein matrix as a direct consequence of the D-ring photoflip rather than autonomous movement of the bilin ring A. The strain and/or torsion of the A–B methine bridge is expected not to exert a strong influence on the absorbance properties of red/green CBCRs as the C–D methine bridge, as shown in a recent theoretical study of Slr1393g3 [24]. The ΔδClight changes seen here for the bilin–protein linkage seem to be widespread in the phytochrome family, albeit quantitatively larger than changes at these atoms in the case of canonical and Cph2-like phytochromes (Fig. S3 and ref. [77]). These results suggest that the covalent linkage might play a role in the molecular mechanisms of post light-absorption events that have not been fully elucidated.
The ΔδClight changes associated with the rings C and D seen in this study are clearly distinct from those in canonical phytochromes. For example, C14 and C16 of the C–D methine bridge in both AnPixJg2 and NpR6012g4 show a ΔδC change <2.4 ppm, much smaller than those of Cph1, in which prominent changes of >5.4 ppm for C14 and most D-ring pyrrolic carbons were found (Fig. S3). Although the ΔδClight at the C–D bridge are difficult to reconcile with those characteristic of the D-ring photoflip in Cph1, the appearance of the C13–C171 DARR contact in the Pg photoproduct (Fig. 2B) and the dramatic ΔδHlight change of −1.9 ppm of the D-ring NH proton (H24, Fig. 4A), showed conclusively that the same occurs in AnPixJg2 upon photoconversion. By considering the maximal detection of ~3.8 Å, this Pg-specific carbon pair can be observed as only the bilin adopts a twisted C15-E,anti-geometry. Such a situation is indeed seen in our Pg structural model which predicts a torsional angle of 125° for the C–D bridge (τC–D). We also noted a large torsion of ring D in the corresponding Pr state as revealed by the 3W2Z structure (49°) and QM/MM calculations (44°). In agreement with the AnPixJg2 models, the solution NMR ensembles of NpR6012g4 indicate an averaged torsion of the ring D by 47° and 122° in the 6BHN Pr and 6BHO Pg structures. Such a strongly twisted D-ring geometry in both photostates may interpret the mild ΔδClight changes at C14 and C16 with a C15-Z/E isomerization upon light absorption. Previous ab initio calculations have demonstrated that δC for bilin pyrrolic carbons are largely dependent on the conjugated π-electron system and the overall geometry and protonation of the bilin but less sensitive to surrounding protein matrix [35]. According to the semiempirical Hückel method, the bilin conjugation is roughly in proportion to cosine squared of the angle between the π-electron axes. In the case of AnPixJg2, the τC–D varies from 44° over 90° to 125° in association with the D-ring photoflip which would not largely modify its conjugation with the rest of the π-electron system, thus rationalizing all observed ΔδClight changes of AnPixJg2. For Cph1, the 2VEA structure indicates a shallower τC–D of 30° in its Pr dark state which would permit the ring D to be fully conjugated (i.e., <40°), whereas the structural models for the Pfr photoproduct imply ~125°–135° [70]. Such a twisted D-ring geometry in the photoproduct would only retain weak conjugation across the C–D methine bridge, and the loss of the D-ring conjugation in the photoproduct would thereby localize to some extent the ΔδClight changes to this region and ring D (Fig. S3).
Our AnPixJg2 Pg structural model provides support for the pivotal role of the bilin D-ring twist in tuning the photoproduct absorption of the red/green CBCR subfamily [24]. As predicted, the ring D in AnPixJg2 photoproduct is twisted (τC–D of 125°) to a similar extent to that of NpR6012g4 (122°), consistent with the fact that the absorption properties of both Pg photoproducts are essentially the same (e.g., both λmax ~ 542 nm). Additionally, in Slr1393g3 and NpR6012g4, several residues which are prone to have a direct impact on the D-ring twist have been identified by mutagenesis, e.g., the removal of the bulky phenyl ring of the ‘β2 Phe’ (a Phe → Leu mutation) resulting in an extended D-ring conjugation in the photoproduct relative to the wild-type (WT), in good agreement with the much less blue-shifted photoproduct absorption (Δ ~ 50 nm) seen in these mutants [44, 79]. D531T and N532Y of Slr1393g3 also produced a less blue-shifted photoproduct absorption as compared to the WT (of 18 nm and 11 nm, respectively). However, according to the crystal structures of both photostates [6] these two residues would neither be in close proximity of the ring D (>8.0 Å) nor involved in forming a hydrogen-bonding network with this ring. The possible shortcoming of the ‘D-ring control’ model to predict the effect of these two mutations might underscore the significance of some environmental factors which would also contribute specifically to the fine-tuning of photoproduct absorption. For example, the variation in the polarity of the bilin-binding pocket due to the influx of water molecules after the geometrical changes of the bilin [33], and rearrangement of the hydrogen-bonding network around the bilin [8, 37] might also be involved. The former effect was given greater attention in the CBCR–GAF domains, as the inspection of crystal structures of red/green CBCRs reveals that the bilin-binding cleft is exposed to the solvent, potentially allowing more readily access of water molecules [5,6,7]. Moreover, the bilin chromophore in the CBCR–GAF domains shows a much larger conformational flexibility within the pocket than that of canonical phytochromes, as has been proven by NMR spectroscopy, e.g., the presence of multiple conformations for both the chromophore and the key residues in its proximity [7, 8, 27], giving the impact of the solvent access in CBCRs much greater relevance (see below).
4.2 The structural heterogeneity of Pr bilin and a stretched-to-bent conformational switching of the C-ring propionate in association with the D-ring photoflip
Our NMR analysis also revealed the structural flexibility of the bilin chromophore in both photostates, as manifested in the resolved multiple 1H, 13C, and 15N signals for a large set of bilin atoms (Figs. 2C and 3B,D). It is evident that, in the Pr dark state, signal splittings of the bilin atoms are largely localized at the A-ring region and the C-ring propionate side-chain. It is well known that the extra flexibility of these two bilin regions is not the intrinsic property of AnPixJg2 or other related red/green CBCRs, but widespread in both canonical and Cph2-like phytochromes in their respective Pr dark states [70, 72, 77]. For phytochromes, the N-terminal extension (NTE), together with the tip of the tongue-like hairpin protruding from the PHY domain forms a hydrophobic wall around rings A and B, thus shielded from solvent access. The local dynamics of the ring A seems to be influenced critically by interactions with the protein residues in these two regions [9]. Whereas recent MELODI–HETCOR data for oat phyA3 showed at least 16 1H NTE–A-ring contacts in Pfr, no such interactions detected in Pr [70]. The most plausible interpretation is that the Pr chromophore, in particularly for the ring A is not compact with the NTE region, thus to be only loosely constrained in the binding pocket. However, as the isolated GAF-domain pocket of AnPixJg2 is open to the solvent (Fig. 1A), the loss of electrostatic and hydrogen-bonding interactions with the pocket (from the hydrophobic side of ring A) would further release the dynamic constraints imposed on the bilin, thus leading to a broader distribution of bilin conformational states relative to the canonical phytochromes. Unlike predicted by the AnPixJg2 Pr structure (Fig. 1A), our previous MD simulations on the same construct revealed that the bilin ring A forms no hydrogen-bonding with the protein residues but is stabilized mainly by its interaction with a ‘persistent’ water molecule in the α-face of the bilin which in turn hydrogen-bonds to the central W289 [27]. Moreover, the indole ring of this Trp residue and its homolog in NpR6012g4 (W655, ref. [7]) were found to adopt multiple dramatically different orientations which would nevertheless suffice to further increase the local structural plasticity of protein environment around the ring A, consistent with the large number of sub-conformations specifically detected at this region (Tables S1–S3).
The Pr chromophores in red/green CBCRs and canonical phytochromes share another common feature in dynamics for the propionate side-chains: the B-ring one is pretty rigid, but the C-ring counterpart is more flexible and heterogeneous [7, 27, 46, 72]. The latter side-chain is known to be centrally involved in formation of the Cph1 and oat phyA3 bilin conformers distinguished by hydrogen-bonding networks and charge distribution patterns within the binding pocket [70, 72]. Moreover, the greater flexibility of the C-ring side-chain is seen in the 10 lowest-energy NMR ensembles for the Pr dark state of NpR6012g4 which interconverts radically between a stretched (7/10) and bent (3/10) conformation and participates in different hydrogen-bonds with adjacent protein residues [7]. Similar conformational flexibility can be also expected for AnPixJg2, in which we identified two long-range DARR correlations of the C-ring propionate carboxylate group (C123a) with C11 and C13 (Fig. 2A) at a C···C distance of >4.6 Å (extracted from the 3W2Z structure). The appearance of these two correlations does not imply that the maximal detection range (~3.8 Å) is underestimated, but it may arise due to downward bending of the C-ring propionate pointing toward the pyrrole rings that would cause a reduction of the corresponding interatomic distances, thus becoming detectable in DARR. Our recent spectroscopic and theoretical results for the AnPixJg2-Y302F mutant revealed the key role of this Tyr residue in stabilizing the stretched conformation of the C-ring propionate, and moreover, the removal of the hydrogen-bond from Y302-Oη to C123-O leads to the occurrence of an unusual 687-nm-absorbing population in the Pr dark state which is photochemically impotent [46]. In the photoproduct, the DARR spectrum revealed one more split component for C123 (Fig. 2B), however, the 13C separation of the three resonances is 0.3 ppm narrower than that of the Pr state (Table S1). This result is not consistent with the coexistence of stretched and bent conformations of the C-ring propionate analogous to those seen in the Pr state. Indeed, a uniformly bent conformation is favorable in the Pg photoproduct, noting that a set of DARR correlations implying populations with C123 pointing downward to interact with e.g., C10, C11, and C13, is clearly resolved (Fig. 2B). The Pg population involving C123c most likely corresponds to the bend conformation in the Pr state (C123a), since in both cases their correlations with C11 and C13 are observed. By contrast, the stretched form involving C123a in the Pr state is not maintained upon photoproduct formation but further splits into two subpopulations with most likely bent conformation. This stretched-to-bent transformation of the C-ring propionate was also clear from the paired Pr/Pg crystal structures of Slr1393g3 [6] and the Pr/Pfr structural models of oat phyA3 [70]. Although the need for this process after primary photo-isomerization is not yet clear, the MD-simulated structures of AnPixJg2-Y302F revealed a significantly decreased planarity of ring D in association with the C-ring propionate bending [46]. We thus envision that the downward bent propionate, together with the conserved ‘β2 Phe’ and ‘helix Phe’ (F268 and F329, Fig. 1B) forms a steric blockade to finely tune the D-ring geometry of the 15E photoproduct. Despite of an instrumental role in achieving geometric stabilization of the ring D, bending of the C-ring propionate has little or no impact on its facial disposition. As discussed above, the Pfr bilin of Cph1 exhibits a similar D-ring twist (τC–D of 125°–135°), however, the ring is predicted to be β-facial [70], opposite to AnPixJg2 (as in Fig. 5B) and related red/green CBCRs [6, 7].
4.3 A further structurally heterogeneous bilin chromophore with increased solvent accessibility for the photoproduct
Unlike Pr, nearly all 1H, 13C, and 15N resonances of the Pg bilin chromophore are at least doubly split with the exceptions being only C8, C82, and the NH moiety associated with the ring B (Figs. 2C and 3B, D), implicating that the bilin as a whole in the photoproduct is remarkably flexible. The most prominent increase in conformational flexibility of the bilin associated with photoconversion was detected for the D-ring atoms, as these atoms exhibit an almost uniform increase in the number of split components with larger resonance separations (Tables S1–S3). These results could be interpreted as arising from the radical side-chain movement of the ‘lid’ Trp residue (W289 in AnPixJg2), as implied by the Slr1393g3 [6] and NpR6012g4 [7] structures which further opens the Pg photoproduct pocket from the D-ring side, thus weakening hydrophobic packing contacts for the bilin, in particularly for ring D. The presence/absence of the C17/C171–C13 DARR contacts in the photoproduct is a sensitive parameter of the D-ring geometry (Fig. 2B): for one population at C13 (C13c) both contacts are clearly resolved (with C17b and C171a), however, none such contacts are present involving the other population of these two D-ring carbons (such as with C17a and C171b) which could arise due to large conformational flexibility of this bilin ring in the photoproduct, such as a further D-ring twist or a change of its disposition from α- to β-face of the B–C plane. The latter interpretation is also consistent with our recent MD simulations on the WT and Y302 mutant of AnPixJg2 in their respective Pr dark states [27, 46]. Such state-related flexibility increase associated with the Pg bilin was also evident in the results from the ultrafast spectroscopic studies of other related red/green CBCRs, as implied by inhomogeneous broadening of the transient absorption of the back-reaction from Pg to Pr [80, 81]. NMR studies with an isolated GAF-domain fragment from the cyanobacterium Synechococcus OS-B’ (SyB.Cph2[GAF]) revealed that most bilin resonances showed considerable broadening in the Pfr photoproduct, implicating increased flexibility relative to the Pr dark state [77, 82]. By contrast, a wealth of NMR data on the Pr/Pfr photoconversion in the tripartite photosensory module of canonical phytochromes showed a concerted sharpening of the resonances in Pfr relative to Pr for the entire bilin, the whole binding pocket, and even the regions deeply into the PHY domain [72, 83, 84]. Such a collective flexibility change implies a ‘soft-to-hard’ mesoscopic phase transition that is of functional relevance [83, 84]. Taken together, these results indicate the importance of a compact binding pocket in stabilizing the GAF-bound bilin chromophore, with in particular the PHY domain being relevant for fixation of the bilin conformation [69, 84]. Intriguingly, although there is a large distinction between the red/green CBCRs and canonical phytochromes in conformational heterogeneity of the bilin chromophore in the photoproducts, the numbers of conformers populated in their Pr states are comparable irrespective of intactness of binding pocket.
The 3W2Z structure of AnPixJg2 implies partial solvent access to the Pr chromophore and three strategically-placed water molecules around it (Fig. 6B). The MD simulation on this protein additionally highlighted an intruded water molecule persistent in the center of the Pr pocket for ~1 μs [27]. By contrast, our interfacial 1H contacts of the Pg chromophore revealed at least six water molecules within a 4.5-Å sphere in the pocket (Fig. 6A), consistent with the Pg QM/MM model reported here (Fig. 6B). The data provide good evidence of inclusion of additional water molecules for the Pg photoproduct that might be associated with the radical side-chain rearrangement of the ‘lid’ Trp upon photoconversion, as implied by the paired Pr/Pg structures of Slr1393g3 [6] and NpR6012g4 [7]. In the MELODI–HETCOR spectrum (maximal detection range of ~4.5 Å), no 1H contacts involving the side-chain of this Trp are detected for the PCB carbons in the photoproduct. That is in line with both structures that show the corresponding long-range heteronuclear pairs at least 5.8 Å apart. The hydration model was postulated to explain the photoproduct blue shift of the red/green CBCRs by the increased solvation of the binding pocket, in which a loosely bound and structurally flexible chromophore is embedded [33]. However, the formation of red-shifted Pfr photoproduct in the case of oat phyA3 also allows entry of additional water molecules into the binding pocket [70], therefore ruling out the one-to-one correlation between photoproduct spectral tuning and pocket solvation. Moreover, the results obtained for oat phyA3 also demonstrated an inverse correlation between pocket solvation and rigidity of the photoproduct as seen in red/green CBCRs like AnPixJg2 (see above) and Slr1393g3 [6, 81]. So far, little is known about the actual role of the solvation process and its dynamics in CBCRs. However, recent femtosecond-resolved spectroscopic data and site-directed mutagenesis presented by Zhong and coworkers showed for a unique red/green CBCR from Leptolyngbya JSC-1 that the excited Pg-state isomerization dynamics is tightly coupled to the local protein solvation dynamics and moreover, a loosely ordered binding pocket results in faster solvation relaxations and subsequent isomerization [85].
4.4 The photoproduct chromophore sandwiched between cationic H322 and ε-protonated neutral H293
In this context, it is of great interest to refer to a recent MD investigation of the effect of the imidazole protonation state of H322 on the conformation of AnPixJg2 in the Pr dark state [25]. These authors characterized the cationic form (biprotonation) of this conserved His residue responsible for structural heterogeneity in the Pr dark state and found the neutral form that coincides with the homogeneous chromophore structure. The finding of the role of this His residue in formation of the Pr heterogeneity is in accord with earlier experimental and theoretical studies for phytochromes. Taking Cph1 as an example [72], interfacial 1Hresi.–13CPCB correlation spectroscopy revealed at least two coexisting Pr populations which differ in their hydrogen-bonding networks around the chromophore, in particular with respect to the imidazole protonation state and tautomeric structure of H260 (homologous to H322 in AnPixJg2). This His residue has also been found to control the conformational equilibrium of different Pr substates by switching its side-chain protonation state, e.g., between neutral and cationic forms [86]. However, it remains speculative whether the correlation between the histidine protonation and the chromophore heterogeneity also holds for the photoproduct. Our 1H interfacial correlation data (Fig. 5A) provide unequivocal evidence for the biprotonated imidazolium of H322 in the photoproduct pocket of AnPixJg2 (Fig. 5B) with respect to distinct conformations in the Pg chromophore (Figs. 2C and 3B, D). This observation, together with the fact that a neutral imidazole of H260 is present in the Pfr photoproduct of Cph1 with a structurally homogenous chromophore [72], suggests that this correlation may be generalized to various photoproducts in both CBCRs and phytochromes.
This histidine residue is perfectly conserved in biliproteins and adopts very similar imidazole side-chain positions relative to the chromophore (as in Fig. 5B). Specifically, as seen from available crystal structures [4,5,6,7, 9,10,11], the side-chain is placed in close contact with the inner rings B and C (usually <3.5 Å), thus providing steric packing constraints on the B–C plane from its α-face. Despite this close proximity, the NMR-refined QM/MM structural models for AnPixJg2 reported in this study suggest that there will be no hydrogen-bonding interaction involving the His side-chain and the NH moieties of rings B and C in both photostates (Fig. 5B). It could arise from the absence of pyrrole water which, in phytochromes, serves as a hydrogen-bond bridge between the His residue and both of the inner ring nitrogens. Instead, the side-chain of H322 in the Pr state, irrespective of the protonation state, is bridged via a water molecule (Wat301 as in Fig. 1B) to the D-ring nitrogen [25]. By contrast, C123 interacts in the Pg photoproduct directly with the biprotonated imidazolium of H322 at Hε2 (Fig. 5A), and accordingly, our NMR-refined QM/MM model based on the Pg state of Slr1393g3 predicts a C123-O···H322-Nε2 distance of ~2.7 Å, consistent with a hydrogen bond (Fig. 5B). The formation of such a hydrogen bond in Pg is facilitated by the positional shift of the inner rings and the downward bending of the C-ring propionate upon photoconversion. Similarly, the protonated Nδ1 swaps its hydrogen-bonding partner from the B-ring carboxylate in Pr to a water molecule in Pg, as implied by the structural model. Instead of H322, H293 on the opposite bilin β-face plays the role to stabilize the B-ring carboxylate in the photoproduct, where only the interfacial correlations involving Hε2 were detected (Fig. 5A). The protonated Nε2 atom of this neutral τ tautomer would thus serve as the hydrogen-bond donor to a water molecule (equivalent to ‘Wat2’ as in Fig. 6B) which is further connected to the B-ring carboxylate and thence to the side-chain of R301, as seen in the equivalent Pg structure of Slr1393g3 [6]. The formation of a strong hydrogen-bond (salt bridge) with the guanidinium group which is invariably positively charged under all physiological conditions presumably provides a rigidifying effect on the anionic B-ring propionate. This could thus very well explain, at least in part our finding that the conformational flexibility of the B-ring propionate remains nearly unaffected upon photoproduct formation even though the rest of the bilin becomes less constrained. The increased flexibility of the C-ring propionate, for example, would be associated with the state-related rupture of the bidentate salt-bridge interaction with R301 [6]. Although the C-ring propionate in the photoproduct is neutralized by the cationic imidazolium of H322 at a distance <3.5 Å (Fig. 5B), the resulting hydrogen bonding is essentially weakened which could be attributed to the delocalized positive charge creating a less significant ionic or Coulombic component [87].
The bilin ring B is laterally clamped between the aromatic side-chains of H322 from its α-face and H293 on the opposite β-face with the center-to-center distance of 3.5 and 2.9 Å, respectively (Fig. 5B). Given their different protonation and charged states, it is reasonable to infer that these two histidines experience different microenvironments immediately around the ring B, and their pKa values would adjust accordingly. For H322, the pKa value of the side-chain would be at least 1.5 units higher relative to the buffer pH value of 7.2, e.g., >8.7 so that it can exist at pH 7.2 predominately in its biprotonated form (with a proportion of >97%). Unlike H322, H293 mostly carries a neutral ε-protonated imidazole at pH 7.2 which is favored in case assuming that H293 possesses a pKa lowered by 1.5 units to ~5.7 (Fig. 5B), in total 3.0 units below that of H322. As a result, the β-facial microenvironment would enhance the solvent interactions with the ring B. However, the imidazole side-chain of H293 is placed at only 2.9 Å apart from the B-ring plane which may provide a tight steric packing from the β-face. Further B-ring stabilization from the same face is achieved through hydrogen-bonding interactions with the acidic side-chain of highly conserved D291 (see Fig. 1B). By contrast, the microenvironment of the opposite α-face is more hydrophobic which would result in a poor solvent accessibility of the ring B, however according to the Pg structural model, a water molecule (‘Wat3’ in Fig. 6B) is found in proximity (2.8 Å). This could arise due to the side-chain arrangement of H322 which is placed 0.6 Å more distant from the B-ring plane relative to that of the β-facial H293, thereby loosening the steric packing of the α-face. Indeed, the close juxtaposition of two positively charged constituents (H322 and the bilin rings B and C) seems unfavorable. Moreover, the B-ring nitrogen atom shows no direct or water-mediated hydrogen-bonding interactions with nearby polar residues on the same facial side which presumably leads to a weaker confinement of the ring B. Such differences in hydrophobicity, hydrogen-bonding interactions and packing constraints on the two facial sides of the ring B would cause a small-amplitude inter-planar tilt between the two inner rings arising due to the local movement around the C10-methine bridge that naturally explains the apparent flexibility of the bilin B-ring pyrrolic carbons in the photoproduct (Fig. 2C).
4.5 A 0.6-unit pH drop substantially altering protonation and electronic structure of the in vitro assembled chromophore in the photoproduct
Another salient result from this study is the finding that the Pg chromophore of the in vitro assembled AnPixJg2 becomes protonated at pH 7.2 (Fig. 3C) instead of a deprotonated form at pH 7.8 [37]. More strikingly, their absorption spectra are hardly altered by the change in bilin protonation with respect to the band shapes and λmax of ~542 nm (see also Fig. S1). These results fully support the notion that bilin protonation is not the fundamental parameter controlling the shift of electronic transition of the bilin chromophore in the red/green CBCR subfamily [37]. Unlike the Pg photoproduct, the Pr dark state harbors a fully protonated (cationic) bilin chromophore at both pH values. Despite the Pr bilin retains its protonation upon the decrease of the pH from 7.8 to 7.2, the electronic structure of the π-conjugated system, the overall geometry, and its interactions with the binding pocket are largely modified, as evidenced by the prominent ΔδCpH changes of the bilin carbons, particularly at C31, C5, C7–C9, and C121 (Fig. 4B). The observed change at the ethylidene C31 position (ΔδCpH = 12.1 ppm) is not consistent with a cleavage of the thioether linkage with the Cys, during which much more drastic δC changes at the ethylidene side-chain would be expected [71]. The occurrence of a stereochemical conversion of C31 carbon (R ↔ S) during the pH drop can also be ruled out since it would require the cleavage and reformation of the thioether bond [38]. Such a ΔδCpH change at C31 (shift upfield at pH 7.2) could be interpreted as arising from stretching of the S–C bond from its covalent length which has been known to be susceptible to its environment. Indeed, structures of the Pr state of the in vivo and the in vitro assembled Slr1393g3 (PDB codes 5DFX and 5DFY, respectively) are nearly identical (0.13 Å rmsd over 158 Cα atoms), but showing a S–C bond stretching by ~0.2 Å with the bilin assembled in vitro [6]. In our case, the bond stretching might be due to a movement of the bilin ring A relative to the rest of the molecule, or due to the partial positive charge held by the S atom that results from exposure to a more acidic medium upon the pH drop. The former scenario seems unlikely because no ΔδCpH changes >2.0 ppm are seen for other A-ring carbons (Table S1). Regarding the latter aspect, the solution at pH 7.2 with a higher concentration of H+ ions (four-fold) would facilitate the attraction to the lone-pair electrons on the S atom which would consequently gain a partial positive charge. The increased electronegativity of this S atom would cause an electron withdrawal from the bonding C atom of the thioester linkage. At pH 7.2, we also identified collective 13C downfield shifts of the pyrrolic quaternary carbons of bilin rings B–C (with the exception being C14) as well as the C5- and C15-methine bridge carbons (Fig. 4B). Such a global effect induced by pH on the π-system electronic structure is unlikely to arise through the local modification of the bilin interaction with specific protein residues but attributed to packing effects between the bilin and its binding pocket. More specifically, the downfield shift (stronger deshielding) seen for this region can result from a denser packing of the bilin due to proximity of larger amount of H+ ions in a more acidic environment. This interpretation is supported by the large ΔδCpH changes of ≥3.3 ppm at C5 and C7–C9 of ring B that are all solvent-exposed (Fig. 1A). By contrast, the ring C is less accessible to the solvent and all pyrrolic carbons of the ring showed a downfield shift of ≤2.5 ppm (Fig. 4B). Moving further into the pocket, this effect vanishes for the D-ring carbons in which only C18 moves downfield. Moreover, the dramatic downfield shift at C121 (ΔδCpH = 5.3 ppm) might be accounted for by a stretched/bent conformational transformation of the C-ring propionate analogous to that seen in association with the Pg formation, in which C121 shows the most prominent change among the side-chain carbons (Fig. 4A). This result indicates that the conformation (and flexibility) of the C-ring propionate is not only state-related, but also pH-regulated.
Like Pr (Fig. 4B), the pH drop leads to the prevalence of downfield shifts (red circles) for most atoms in the bilin B–C region of the Pg photoproduct (Fig. 4C), indicating an overall decrease in electron density of this region. Despite the similarity, however, a global increase in ΔδCpH magnitude was seen in the Pg state, in particular for C12–C14 of ring C which are at least 3.0 ppm more downfield shifted relative to those in Pr (Table S1). These ‘extra’ pH-induced shifts in the Pg state cannot be explained solely by a loosely ordered and solvated photoproduct pocket as we observed, but the effects of B-ring protonation on δC of certain pyrrolic carbons must be taken into account. In Pr, both C6 and C9 atoms adjacent to the B-ring nitrogen undergo a downfield shift, however, the sign of ΔδCpH changes is reversed in the photoproduct (Fig. 4C) to be consistent with the modification induced by the B-ring protonation at N22 occurring during the PCB assembly into the Cph1 pocket [71]. Besides C6, C9, and C12–C14, the ΔδCpH change at C16 of ring D can be also directly related to the protonation of N22 which is shifted downfield by 8.8 ppm upon the 0.6-unit pH drop, comparable to that resulting from the assembly of PCB into Cph1 (11.4 ppm, ref. [71]). This overall agreement implies that the pH-induced ΔδCpH changes for these pyrrolic carbons in the photoproduct are largely determined by the B-ring protonation rather than by packing of the bilin with nearby protein residues which is predominantly responsible for the bilin ΔδCpH changes in Pr. Moreover, the ΔδCpH changes of the D-ring carbons (e.g., C16) associated with the B-ring protonation support the existence of at least weak D-ring conjugation of the Pg bilin in red/green CBCRs [24].
Interpretation of the 13.8-ppm ΔδNpH change at N24 of ring D is not straightforward. The isotropic chemical shift of N24 is not ‘sensitive’ to the protonation of N22 which for example upon assembly of PCB in Cph1 (as Pr) moves downfield only by 0.6 ppm [71]. In the absence of the 15N chemical shift anisotropy (CSA) principal values of this bilin nitrogen, we cannot rule out the changes on its local electronic structure and intermolecular interactions [73, 88]. In general, CSA principal values provide more complete information than isotropic shifts on these aspects. Taking the pyrrole as an example, the two principal components tangential and radial to the ring (δT and δR) are sensitive for probing intermolecular interactions such as intermolecular interactions associated with hydrogen bonding and to changes in the hybridization of the nitrogen atom, while the remaining component perpendicular to the ring (δP) is dominated by lone-pair transition of the nitrogen [88]. Although we cannot conclude with certainty which of the factors is the cause of the large ΔδNpH change at N24 upon the pH drop to 7.2 (Fig. 4C), it is reasonable to assume that the proximity of H+ ions to the D-ring nitrogen in the photoproduct would lead to a prominent nitrogen lone-pair hybridization change (represented by δP), thus dominating the observed isotropic ΔδNpH change. This working hypothesis is based on an increased solvent accessibility of the D-ring pocket in the Pg state due to the drastic change of the protein surface upon photoconversion of Pr to Pg, as implied by both the paired Pr/Pg structures of Slr1393g3 [6] and our AnPixJg2 structural models.
The pH-induced protonation process of the in vitro Pg photoproduct is probably due to the reversal of misfolded protein structure which may alter the hydrogen-bonding networks around the chromophore and/or the rotational/conformational arrangement of certain nearby protein residue(s) as compared to the native form. These changes may exert a stabilizing effect on the chromophore as reported here. We also noted that the pH drop to 6.0 causes a substantial destabilization of the in vitro photoproduct, however, the native form of the protein retains stability at pH 5.0 (Fig. S6). It has been known that multiple factors are capable of modulating the protein stability such as hydrophobic and electrostatic interactions, and moreover, the signature of the latter is its pH-dependence [89]. Although a number of mutational and structural analyses are available for AnPixJg2 [5, 15, 25, 27, 33] and other related red/green CBCRs [6, 7, 44, 79], we cannot pinpoint the regions or specific residues as the cause for this in vitro photoproduct destabilization. Moreover, it has been proposed that, e.g., at pH 7.8 the central D291, whose side-chain is within the hydrogen-bonding distance to the B-ring nitrogen, acts as the acceptor of the released proton from the bilin [37]. At pH 7.2, however, no 1H contacts of the bilin in the MELODI–HETCOR spectrum could be attributed to the proton of its side-chain carboxyl moiety (Fig. 5A). Were this proton to exist, one would expect a 1H slice in the downfield region of 7–9 ppm due to the interactions with nearby pyrrolic carbons (i.e., within 4.5 Å). This observation implicates a pKa of ~ 7.2–7.8 for the side-chain of D291. Moreover, in absence of the MELODI–HETCOR data of the in vitro photoproduct at pH 7.8, we cannot rule out the presence of a hydronium ion (H3O+) due to charge rearrangement between D291 and the structural water molecules nearby. It has also to be noted that the current data do not allow us to assess the conformational heterogeneity of D291 which was discussed for the homologous residue in the NpR6012g4 Pr dark state [7].
5 Conclusion
To conclude, in in vitro assembled AnPixJg2, our results are consistent with the ring D being highly twisted relative to the B–C plane in both Pr and Pg photostates in particular for the Pg bilin which support the essential role for the D-ring twist in photoproduct tuning of red/green CBCRs [24]. Our results also demonstrate a downward bending of the C-ring propionate upon Pg formation which presents additional steric hindrance to promote a further twisted D-ring geometry. We observe a heterogeneous bilin in both photostates of AnPixJg2, however, unlike the Pr heterogeneity localized selectively in the A-ring region and C-ring propionate, the Pg bilin as a whole is remarkably heterogeneous. We ascribe this prominent increase in structural flexibility of the Pg bilin to a loosely ordered and more solvated photoproduct pocket, as evident from both MELODI–HETCOR spectrum and QM/MM structural models (Fig. 6). Observation of interfacial 1H contacts of the Pg bilin also reveals that the two His residues, clamping the ring B from the opposite facial sides, have different protonation states (H322, biprotonated imidazolium; and H293, ε-protonated imidazole; Fig. 5). Differences in the pKa values (~3.0 units) and hydrophobicity between the two bilin sides imply a special local environment of the ring B. Moreover, the presence of a cationic imidazolium for H322 in the Pg photoproduct generalizes the recent results correlating the structural heterogeneity of the Pr dark-state bilin explicitly to the protonation of this His residue [25]. The pH effects on this in vitro assembled protein (7.8 → 7.2) reveal surprisingly that the Pg bilin becomes fully protonated upon the 0.6-unit pH drop with only subtle changes of the absorption spectrum (Fig. S1). These results support our previous notion that the bilin protonation is not the largest determinant to control the electronic transitions of the chromophore in the red/green CBCRs [37].
Notes
The A–B methine bridge torsional angle (τA–B) represents the sum of the deviations of the dihedral angles (ψ) from the ideal planar geometry, that is, τA–B = ψ(N21–C4=C5–C6) + ψ(C4=C5–C6–N22), and 0° for Z and 180° for E, respectively (as in ref. 78).
References
Anders, K., & Essen, L.-O. (2015). The family of phytochrome-like photoreceptors: Diverse, complex and multi-colored, but very useful. Current Opinion in Structural Biology, 35, 7–16. https://doi.org/10.1016/j.sbi.2015.07.005
Fushimi, K., & Narikawa, R. (2019). Cyanobacteriochromes: Photoreceptors covering the entire UV-to-visible spectrum. Current Opinion in Structural Biology, 57C, 39–46. https://doi.org/10.1016/j.sbi.2019.01.018
Rockwell, N. C., & Lagarias, J. C. (2020). Phytochrome evolution in 3D: Deletion, duplication, and diversification. New Phytologist, 225, 2283–2300. https://doi.org/10.1111/nph.16240
Burgie, E. S., Walker, J. M., Phillips, G. N., Jr., & Vierstra, R. D. (2013). A photo-labile thioether linkage to phycoviolobilin provides the foundation for the blue/green photocycles in DXCF-cyanobacteriochromes. Structure, 21, 88–97. https://doi.org/10.1016/j.str.2012.11.001
Narikawa, R., Ishizuka, T., Muraki, N., Shiba, T., Kurisu, G., & Ikeuchi, M. (2013). Structures of cyanobacteriochromes from phototaxis regulators AnPixJ and TePixJ reveal general and specific photoconversion mechanism. Proceedings of the National Academy of Sciences of the United States of America, 110, 918–923. https://doi.org/10.1073/pnas.1212098110
Xu, X., Höppner, A., Wiebeler, C., Zhao, K.-H., Schapiro, I., & Gärtner, W. (2020). Structural elements regulating the photochromicity in a cyanobacteriochrome. Proceedings of the National Academy of Sciences of the United States of America, 117, 2432–2440. https://doi.org/10.1073/pnas.1910208117
Lim, S., Yu, Q., Gottlieb, S. M., Chang, C.-W., Rockwell, N. C., Martin, S. S., Madsen, D., Lagarias, J. C., Larsen, D. S., & Ames, J. B. (2018). Correlating structural and photochemical heterogeneity in cyanobacteriochrome NpR6012g4. Proceedings of the National Academy of Sciences of the United States of America, 115, 4387–4392. https://doi.org/10.1073/pnas.1720682115
Song, C., Narikawa, R., Ikeuchi, M., Gärtner, W., & Matysik, J. (2015). Color tuning in red/green cyanobacteriochrome AnPixJ: Photoisomerization at C15 causes an excited-state destabilization. The Journal of Physical Chemistry B, 119, 9688–9695. https://doi.org/10.1021/acs.jpcb.5b04655
Essen, L.-O., Mailliet, J., & Hughes, J. (2008). The structure of a complete phytochrome sensory module in the Pr ground state. Proceedings of the National Academy of Sciences of the United States of America, 105, 14709–14714. https://doi.org/10.1073/pnas.0806477105
Yang, X., Ren, Z., Kuk, J., & Moffat, K. (2011). Temperature-scan cryocrystallography reveals reaction intermediates in bacteriophytochrome. Nature, 479, 428–432. https://doi.org/10.1038/nature10506
Takala, H., Björling, A., Berntsson, O., Lehtivuori, H., Niebling, S., Hoernke, M., Kosheleva, I., Henning, R., Menzel, A., Ihalainen, J. A., & Westenhoff, S. (2014). Signal amplification and transduction in phytochrome photosensors. Nature, 509, 245–248. https://doi.org/10.1038/nature13310
Lim, S., Rockwell, N. C., Martin, S. S., Dallas, J. L., Lagarias, J. C., & Ames, J. B. (2014). Photoconversion changes bilin chromophore conjugation and protein secondary structure in the violet/orange cyanobacteriochrome NpF2163g3. Photochemical & Photobiological Sciences, 13, 951–962. https://doi.org/10.1039/c3pp50442e
Rockwell, N. C., Martin, S. S., Gulevich, A. G., & Lagarias, J. C. (2012). Phycoviolobilin formation and spectral tuning in the DXCF cyanobacteriochrome subfamily. Biochemistry, 51, 1449–1463. https://doi.org/10.1021/bi201783j
Hirose, Y., Rockwell, N. C., Nishiyama, K., Narikawa, R., Ukaji, Y., Inomata, K., Lagarias, J. C., & Ikeuchi, M. (2013). Green/red cyanobacteriochromes regulate complementary chromatic acclimation via a protochromic photocycle. Proceedings of the National Academy of Sciences of the United States of America, 110, 4974–4979. https://doi.org/10.1073/pnas.1302909110
Narikawa, R., Fukushima, Y., Ishizuka, T., Itoh, S., & Ikeuchi, M. (2008). A novel photoactive GAF domain of cyanobacteriochrome AnPixJ that shows reversible green/red photoconversion. Journal of Molecular Biology, 380, 844–855. https://doi.org/10.1016/j.jmb.2008.05.035
Bandara, S., Rockwell, N. C., Zeng, X., Ren, Z., Wang, C., Shin, H., Martin, S. S., Moreno, M. V., Lagarias, J. C., & Yang, X. (2021). Crystal structure of a far-red–sensing cyanobacteriochrome reveals an atypical bilin conformation and spectral tuning mechanism. Proceedings of the National Academy of Sciences of the United States of America, 118, e2025094118. https://doi.org/10.1073/pnas.2025094118
Rockwell, N. C., Martin, S. S., & Lagarias, J. C. (2012). Red/green cyanobacteriochromes: Sensors of color and power. Biochemistry, 51, 9667–9677. https://doi.org/10.1021/bi3013565
Narikawa, R., Nakajima, T., Aono, Y., Fushimi, K., Enomoto, G., Ni-Ni-Win, Itoh, S., Sato, M., & Ikeuchi, M. (2015). A biliverdin-binding cyanobacteriochrome from the chlorophyll d-bearing cyanobacterium Acaryochloris marina. Scientific Reports, 5, 7950. https://doi.org/10.1038/srep07950
Rockwell, N. C., Martin, S. S., & Lagarias, J. C. (2017). There and back again: Loss and reacquisition of two-Cys photocycles in cyanobacteriochromes. Photochemistry and Photobiology, 93, 741–754. https://doi.org/10.1111/php.12708
Fushimi, K., Nakajima, T., Aono, Y., Yamamoto, T., Ni-Ni-Win, Ikeuchi, M., Sato, M., & Narikawa, R. (2016). Photoconversion and fluorescence properties of a red/green-type cyanobacteriochrome AM1_C0023g2 that binds not only phycocyanobilin but also biliverdin. Frontiers in Microbiology, 7, 588. https://doi.org/10.3389/fmicb.2016.00588
Moreno, M. V., Rockwell, N. C., Mora, M., Fisher, A. J., & Lagarias, J. C. (2020). A far-red cyanobacteriochrome lineage specific for verdins. Proceedings of the National Academy of Sciences of the United States of America, 117, 27962–27970. https://doi.org/10.1073/pnas.2016047117
Nagae, T., Unno, M., Koizumi, T., Miyanoiri, Y., Fujisawa, T., Masui, K., Kamo, T., Wada, K., Eki, T., Ito, Y., Hirose, Y., & Mishima, M. (2021). Structural basis of the protochromic green/red photocycle of the chromatic acclimation sensor RcaE. Proceedings of the National Academy of Sciences of the United States of America, 118, e2024583118. https://doi.org/10.1073/pnas.2024583118
Fushimi, K., Miyazaki, T., Kuwasaki, Y., Nakajima, T., Yamamoto, T., Suzuki, K., Ueda, Y., Miyake, K., Takeda, Y., Choi, J.-H., Kawagishi, H., Park, E. Y., Ikeuchi, M., Sato, M., & Narikawa, R. (2019). Rational conversion of chromophore selectivity of cyanobacteriochromes to accept mammalian intrinsic biliverdin. Proceedings of the National Academy of Sciences of the United States of America, 116, 8301–8309. https://doi.org/10.1073/pnas.1818836116
Wiebeler, C., Rao, A. G., Gärtner, W., & Schapiro, I. (2019). The effective conjugation length is responsible for the red/green spectral tuning in the cyanobacteriochrome Slr1393g3. Angewandte Chemie International Edition, 58, 1934–1938. https://doi.org/10.1002/anie.201810266
Rao, A. G., Wiebeler, C., Sen, S., Cerutti, D. S., & Schapiro, I. (2021). Histidine protonation controls structural heterogeneity in the cyanobacteriochrome AnPixJg2. Physical Chemistry Chemical Physics, 23, 7359–7369. https://doi.org/10.1039/d0cp05314g
Wiebeler, C., & Schapiro, I. (2019). QM/MM benchmarking of cyanobacteriochrome Slr1393g3 absorption spectra. Molecules, 24, 1720. https://doi.org/10.3390/molecules24091720
Scarbath-Evers, L. K., Jähnigen, S., Elgabarty, H., Song, C., Narikawa, R., Matysik, J., & Sebastiani, D. (2017). Structural heterogeneity in a parent ground-state structure of AnPixJg2 revealed by theory and spectroscopy. Physical Chemistry Chemical Physics, 19, 13882–13894. https://doi.org/10.1039/c7cp01218g
Gottlieb, S. M., Kim, P. W., Chang, C.-W., Hanke, S. J., Hayer, R. J., Rockwell, N. C., Martin, S. S., Lagarias, J. C., & Larsen, D. S. (2015). Conservation and diversity in the primary forward photodynamics of red/green cyanobacteriochromes. Biochemistry, 54, 1028–1042. https://doi.org/10.1021/bi5012755
Kim, P. W., Freer, L. H., Rockwell, N. C., Martin, S. S., Lagarias, J. C., & Larsen, D. S. (2012). Second-chance initiation dynamics of the cyanobacterial photocycle in the NpR6012 GAF4 domain of Nostoc punctiforme. Journal of the American Chemical Society, 134, 130–133. https://doi.org/10.1021/ja209533x
Slavov, C., Fischer, T., Barnoy, A., Shin, H., Rao, A. G., Wiebeler, C., Zeng, X., Sun, Y., Xu, Q., Gutt, A., Zhao, K.-H., Gärtner, W., Yang, X., Schapiro, I., & Wachtveitl, J. (2020). The interplay between chromophore and protein determines the extended excited state dynamics in a single-domain phytochrome. Proceedings of the National Academy of Sciences of the United States of America, 117, 16356–16362. https://doi.org/10.1073/pnas.1921706117
Wang, D., Li, X., Wang, L., Yang, X., & Zhong, D. (2020). Elucidating ultrafast multiphasic dynamics in the photoisomerization of cyanobacteriochrome. The Journal of Physical Chemistry Letters, 11, 8819–8824. https://doi.org/10.1021/acs.jpclett.0c02467
Buhrke, D., Battocchio, G., Wilkening, S., Blain-Hartung, M., Baumann, T., Schmitt, F.-J., Friedrich, T., Mroginski, M.-A., & Hildebrandt, P. (2020). Red, orange, green: Light- and temperature-dependent color tuning in a cyanobacteriochrome. Biochemistry, 59, 509–519. https://doi.org/10.1021/acs.biochem.9b00931
Velazquez Escobar, F., Utesch, T., Narikawa, R., Ikeuchi, M., Mroginski, M. A., Gärtner, W., & Hildebrandt, P. (2013). Photoconversion mechanism of the second GAF domain of cyanobacteriochrome AnPixJ and the cofactor structure of its green-absorbing state. Biochemistry, 52, 4871–4880. https://doi.org/10.1021/bi400506a
Rockwell, N. C., Martin, S. S., Lim, S., Lagarias, J. C., & Ames, J. B. (2015). Characterization of red/green cyanobacteriochrome NpR6012g4 by solution nuclear magnetic resonance spectroscopy: A hydrophobic pocket for the C15-E, anti chromophore in the photoproduct. Biochemistry, 54, 3772–3783. https://doi.org/10.1021/acs.biochem.5b00438
Rockwell, N. C., Martin, S. S., Lim, S., Lagarias, J. C., & Ames, J. B. (2015). Characterization of red/green cyanobacteriochrome NpR6012g4 by solution nuclear magnetic resonance spectroscopy: A protonated bilin ring system in both photostates. Biochemistry, 54, 2581–2600. https://doi.org/10.1021/bi501548t
Lim, S., Rockwell, N. C., Martin, S. S., Lagarias, J. C., & Ames, J. B. (2014). 1H, 15N, and 13C chemical shift assignments of cyanobacteriochrome NpF2164g3 in the photoproduct state. Biomolecular NMR Assignments, 8, 259–262. https://doi.org/10.1007/s12104-013-9496-0
Song, C., Velazquez Escobar, F., Xu, X.-L., Narikawa, R., Ikeuchi, M., Siebert, F., Gärtner, W., Matysik, J., & Hildebrandt, P. (2015). A red/green cyanobacteriochrome sustains its color despite a change in the bilin chromophore’s protonation state. Biochemistry, 54, 5839–5848. https://doi.org/10.1021/acs.biochem.5b00735
Xu, Q.-Z., Bielytskyi, P., Otis, J., Lang, C., Hughes, J., Zhao, K.-H., Losi, A., Gärtner, W., & Song, C. (2019). MAS NMR on a red/far-red photochromic cyanobacteriochrome All2699 from Nostoc. International Journal of Molecular Sciences, 20, 3656. https://doi.org/10.3390/ijms20153656
Kim, Y., Xu, Q.-Z., Zhao, K.-H., Gärtner, W., Matysik, J., & Song, C. (2020). Lyophilization reveals a multitude of structural conformations in the chromophore of a Cph2-like phytochrome. The Journal of Physical Chemistry B, 124, 7115–7712. https://doi.org/10.1021/acs.jpcb.0c03431
Rockwell, N. C., Martin, S. S., Feoktistova, K., & Lagarias, J. C. (2011). Diverse two-cysteine photocycles in phytochromes and cyanobacteriochromes. Proceedings of the National Academy of Sciences of the United States of America, 108, 11854–11859. https://doi.org/10.1073/pnas.1107844108
Narikawa, R., Enomoto, G., Ni-Ni-WinFushimi, K., & Ikeuchi, M. (2014). A new type of dual-Cys cyanobacteriochrome GAF domain found in cyanobacterium Acaryochloris marina, which has an unusual red/blue reversible photoconversion cycle. Biochemistry, 53, 5051–5059. https://doi.org/10.1021/bi500376b
Rockwell, N. C., Njuguna, S. L., Roberts, L., Castillo, E., Parson, V. L., Dwojak, S., Lagarias, J. C., & Spiller, S. C. (2008). A second conserved GAF domain cysteine is required for the blue/green photoreversibility of cyanobacteriochrome Tlr0924 from Thermosynechococcus elongatus. Biochemistry, 47, 7304–7316. https://doi.org/10.1021/bi800088t
Ishizuka, T., Kamiya, A., Suzuki, H., Narikawa, R., Noguchi, T., Kohchi, T., Inomata, K., & Ikeuchi, M. (2011). The cyanobacteriochrome, TePixJ, isomerizes its own chromophore by converting phycocyanobilin to phycoviolobilin. Biochemistry, 50, 953–961. https://doi.org/10.1021/bi101626t
Rockwell, N. C., Martin, S. S., Gulevich, A. G., & Lagarias, J. C. (2014). Conserved phenylalanine residues are required for blue-shifting of cyanobacteriochrome photoproducts. Biochemistry, 53, 3118–3130. https://doi.org/10.1021/bi500037a
Rockwell, N. C., Martin, S. S., Gan, F., Bryant, D. A., & Lagarias, J. C. (2015). NpR3784 is the prototype for a distinctive group of red/green cyanobacteriochromes using alternative Phe residues for photoproduct tuning. Photochemical & Photobiological Sciences, 14, 258–269. https://doi.org/10.1039/c4pp00336e
Altmayer, S., Jähnigen, S., Köhler, L., Wiebeler, C., Song, C., Sebastiani, D., & Matysik, J. (2021). The hydrogen bond between a tyrosine residue and the C-ring propionate has a direct influence on conformation and absorption of the bilin cofactor in red/green cyanobacteriochromes. The Journal of Physical Chemistry B, 125, 1331–1342. https://doi.org/10.1021/acs.jpcb.0c08518
Thakur, R. S., Kurur, N. D., & Madhu, P. K. (2006). Swept-frequency two-pulse phase modulation for heteronuclear dipolar decoupling in solid-state NMR. Chemical Physics Letters, 426, 459–463. https://doi.org/10.1016/j.cplett.2006.06.007
Vinogradov, E., Madhu, P. K., & Vega, S. (2001). Phase modulated Lee-Goldburg magic angle spinning proton nuclear magnetic resonance experiments in solid state: A bimodal Floquet theoretical treatment. The Journal of Chemical Physics, 115, 8983–9000. https://doi.org/10.1063/1.1408287
Coelho, C., Rocha, J., Madhu, P. K., & Marfa, L. (2007). Practical aspects of Lee–Goldburg based CRAMPS techniques for high-resolution 1H NMR spectroscopy in solids: Implementation and applications. Journal of Magnetic Resonance, 194, 264–282. https://doi.org/10.1016/j.jmr.2008.07.019
Neese, F. (2018). Software update: The ORCA program system, version 4.0. WIREs Computational Molecular Science, 8, e1327. https://doi.org/10.1002/wcms.1327
Metz, S., Kästner, J., Sokol, A. A., Keal, T. W., & Sherwood, P. (2014). ChemShell—a modular software package for QM/MM simulations. WIREs Computational Molecular Science, 4, 101–110. https://doi.org/10.1002/wcms.1163
Becke, A. D. (1988). Density-functional exchange-energy approximation with correct asymptotic behavior. Physical Review A, 38, 3098–3100. https://doi.org/10.1103/PhysRevA.38.3098
Lee, C., Yang, W., & Parr, R. G. (1988). Development of the Colle-Salvetti correlation-energy formula into a functional of the electron density. Physical Review B, 37, 785–789. https://doi.org/10.1103/PhysRevB.37.785
Grimme, S., Antony, J., Ehrlich, S., & Krieg, H. (2010). A consistent and accurate ab initio parametrization of density functional dispersion correction (DFT-D) for the 94 elements H-Pu. The Journal of Chemical Physics, 132, 154104. https://doi.org/10.1063/1.3382344
Grimme, S., Ehrlich, S., & Goerigk, L. (2011). Effect of the damping function in dispersion corrected density functional theory. Journal of Computational Chemistry, 32, 1456–1465. https://doi.org/10.1002/jcc.21759
Weigend, F., & Ahlrichs, R. (2005). Balanced basis sets of split valence, triple zeta valence and quadruple zeta valence quality for H to Rn: Design and assessment of accuracy. Physical Chemistry Chemical Physics, 7, 3297–3305. https://doi.org/10.1039/B508541A
Weigend, F. (2006). Accurate Coulomb-fitting basis sets for H to Rn. Physical Chemistry Chemical Physics, 8, 1057–1465. https://doi.org/10.1039/B515623H
Maier, J. A., Martinez, C., Kasavajhala, K., Wickstrom, L., Hauser, K. E., & Simmerling, C. (2015). ff14SB: Improving the accuracy of protein side chain and backbone parameters from ff99SB. Journal of Chemical Theory and Computation, 11, 3696–3713. https://doi.org/10.1021/acs.jctc.5b00255
Jorgensen, W. L. (1981). Transferable intermolecular potential functions for water, alcohols, and ethers. Application to liquid water. Journal of the American Chemical Society, 103, 335–340. https://doi.org/10.1021/ja00392a016
Grimme, S. (2013). A simplified Tamm-Dancoff density functional approach for the electronic excitation spectra of very large molecules. The Journal of Chemical Physics, 138, 244104. https://doi.org/10.1063/1.4811331
Bannwarth, C., & Grimme, S. (2014). A simplified time-dependent density functional theory approach for electronic ultraviolet and circular dichroism spectra of very large molecules. Computational and Theoretical Chemistry, 1040–1041, 45–53. https://doi.org/10.1016/j.comptc.2014.02.023
Risthaus, T., Hansen, A., & Grimme, S. (2014). Excited states using the simplified Tamm–Dancoff-Approach for range-separated hybrid density functionals: Development and application. Physical Chemistry Chemical Physics, 16, 14408–14419. https://doi.org/10.1039/C3CP54517B
Yanai, T., Tew, D. P., & Handy, N. C. (2004). A new hybrid exchange–correlation functional using the Coulomb-attenuating method (CAM-B3LYP). Chemical Physics Letters, 393, 51–57. https://doi.org/10.1016/j.cplett.2004.06.011
Cheeseman, J. R., Trucks, G. W., Keith, T. A., & Frisch, M. J. (1996). A comparison of models for calculating nuclear magnetic resonance shielding tensors. The Journal of Chemical Physics, 104, 5497–5509. https://doi.org/10.1063/1.471789
Frisch, M. J., Trucks, G. W., Schlegel, H. B., Scuseria, G. E., Robb, M. A., Cheeseman, J. R., Scalmani, G., Barone, V., Petersson, G. A., Nakatsuji, H., Li, X., Caricato, M., Marenich, A. V., Bloino, J., Janesko, B. G., et al. (2016). Gaussian 16 Revision B.01.
Becke, A. D. (1993). Density-functional thermochemistry. III. The role of exact exchange. The Journal of Chemical Physics, 98, 5648–5652. https://doi.org/10.1063/1.464913
Jensen, F. (2015). Segmented contracted basis sets optimized for nuclear magnetic shielding. Journal of Chemical Theory and Computation, 11, 132–138. https://doi.org/10.1021/ct5009526
Pritchard, B. P., Altarawy, D., Didier, B., Gibson, T. D., & Windus, T. L. (2019). New basis set exchange: An open, up-to-date resource for the molecular sciences community. Journal of Chemical Information and Modeling, 59, 4814–4820. https://doi.org/10.1021/acs.jcim.9b00725
Stöppler, D., Song, C., van Rossum, B.-J., Geiger, M.-A., Lang, C., Mroginski, M.-A., Jagtap, A. P., Sigurdsson, S. T., Matysik, J., Hughes, J., & Oschkinat, H. (2016). Dynamic nuclear polarization provides new insights into chromophore structure in phytochrome photoreceptors. Angewandte Chemie International Edition, 55, 16017–16020. https://doi.org/10.1002/anie.201608119
Song, C., Mroginski, M. A., Lang, C., Kopycki, J., Gärtner, W., Matysik, J., & Hughes, J. (2018). 3D structures of plant phytochrome A as Pr and Pfr from solid-state NMR: Implications for molecular function. Frontiers in Plant Sciences, 9, 498. https://doi.org/10.3389/fpls.2018.00498
Rohmer, T., Lang, C., Gärtner, W., Hughes, J., & Matysik, J. (2010). Role of the protein cavity in phytochrome chromoprotein assembly and double-bond isomerization: A comparison with model compounds. Photochemistry and Photobiology, 86, 856–861. https://doi.org/10.1111/j.1751-1097.2010.00740.x
Song, C., Rohmer, T., Tiersch, M., Zaanen, J., Hughes, J., & Matysik, J. (2013). Solid-state NMR spectroscopy to probe photoactivation in canonical phytochromes. Photochemistry and Photobiology, 89, 259–273. https://doi.org/10.1111/php.12029
Li, S., & Hong, M. (2011). Protonation, tautomerization, and rotameric structure of histidine: A comprehensive study by magic-angle-spinning solid-state NMR. Journal of the American Chemical Society, 133, 1534–1544. https://doi.org/10.1021/ja108943n
Watermann, T., Elgabarty, H., & Sebastiani, D. (2014). Phycocyanobilin in solution – a solvent triggered molecular switch. Physical Chemistry Chemical Physics, 16, 6146–6152. https://doi.org/10.1039/C3CP54307B
Forsyth, D. A., & Sebag, A. B. (1997). Computed 13C NMR chemical shifts via empirically scaled GIAO shieldings and molecular mechanics geometries. Conformation and configuration from 13C shifts. Journal of the American Chemical Society, 119, 9483–9494. https://doi.org/10.1021/ja970112z
Mulder, F. A. A., & Filatov, M. (2010). NMR chemical shift data and ab initio shielding calculations: Emerging tools for protein structure determination. Chemical Society Reviews, 39, 578–590. https://doi.org/10.1039/B811366C
Song, C., Psakis, G., Kopycki, J., Lang, C., Matysik, J., & Hughes, J. (2014). The D-ring, not the A-ring, rotates in Synechococcus OS-B’ phytochrome. Journal of Biological Chemistry, 289, 2252–2262. https://doi.org/10.1074/jbc.M113.520031
Mroginski, M. A., Kaminski, S., von Stetten, D., Ringsdorf, S., Gärtner, W., Essen, L.-O., & Hildebrandt, P. (2011). Structure of the chromophore binding pocket in the Pr state of plant phytochrome phyA. The Journal of Physical Chemistry B, 115, 1220–1231. https://doi.org/10.1021/jp108265h
Xu, X.-L., Gutt, A., Mechelke, J., Raffelberg, S., Tang, K., Miao, D., Valle, L., Borsarelli, C. D., Zhao, K.-H., & Gärtner, W. (2014). Combined mutagenesis and kinetics characterization of the bilin-binding GAF domain of the protein Slr1393 from the cyanobacterium Synechocystis PCC6803. ChemBioChem, 15, 1190–1199. https://doi.org/10.1002/cbic.201400053
Kim, P. W., Freer, L. H., Rockwell, N. C., Martin, S. S., Lagarias, J. C., & Larsen, D. S. (2012). Femtosecond photodynamics of the red/green cyanobacteriochrome NpR6012g4 from Nostoc punctiforme. 2. Reverse dynamics. Biochemistry, 51, 619–630. https://doi.org/10.1021/bi2017365
Slavov, C., Xu, X., Zhao, K.-H., Gärtner, W., & Wachtveitl, J. (2015). Detailed insight into the ultrafast photoconversion of the cyanobacteriochrome Slr1393 from Synechocystis sp. Biochimica et Biophysica Acta, 1847, 1335–1344. https://doi.org/10.1016/j.bbabio.2015.07.013
Ulijasz, A. T., Cornilescu, G., Cornilescu, C. C., Zhang, J., Rivera, M., Markley, J. L., & Vierstra, R. D. (2010). Structural basis for the photoconversion of a phytochrome to the activated Pfr form. Nature, 463, 250–254. https://doi.org/10.1038/nature08671
Song, C., Psakis, G., Lang, C., Mailliet, J., Zaanen, J., Gärtner, W., Hughes, J., & Matysik, J. (2011). On the collective nature of phytochrome photoactivation. Biochemistry, 50, 10987–10989. https://doi.org/10.1021/bi201504a
Gustavsson, E., Isaksson, L., Persson, C., Mayzel, M., Brath, U., Vrhovac, L., Ihalainen, J. A., Karlsson, B. G., Orekhov, V., & Westenhoff, S. (2020). Modulation of structural heterogeneity controls phytochrome photoswitching. Biophysical Journal, 118, 415–421. https://doi.org/10.1016/j.bpj.2019.11.025
Wang, D., Li, X., Zhang, S., Wang, L., Yang, X., & Zhong, D. (2020). Revealing the origin of multiphasic dynamic behaviors in cyanobacteriochrome. Proceedings of the National Academy of Sciences of the United States of America, 117, 19731–19736. https://doi.org/10.1073/pnas.2001114117
Velazquez Escobar, F., Lang, C., Takiden, A., Schneider, C., Balke, J., Hughes, J., Alexiev, U., Hildebrandt, P., & Mroginski, M. A. (2017). Protonation-dependent structural heterogeneity in the chromophore binding site of cyanobacterial phytochrome Cph1. The Journal of Physical Chemistry B, 121, 47–57. https://doi.org/10.1021/acs.jpcb.6b09600
Jacobs, D. J., Rader, A. J., Kuhn, L. A., & Thorpe, M. F. (2001). Protein flexibility predications using graph theory. Proteins, 44, 150–165. https://doi.org/10.1002/prot.1081
Solum, M. S., Altmann, K. L., Strohmeier, M., Berges, D. A., Zhang, Y. L., Facelli, J. C., Pugmire, R. J., & Grant, D. M. (1997). 15N chemical shift principal values in nitrogen heterocycles. Journal of the American Chemical Society, 119, 9804–9809. https://doi.org/10.1021/ja964135+
Matthews, B. W. (1993). Structural and genetic analysis of protein stability. Annual Review of Biochemistry, 62, 139–160. https://doi.org/10.1146/annurev.bi.62.070193.001035
Acknowledgements
The authors thank Dr. J. Hughes and C. Lang (Justus-Liebig-Universität Gießen) for providing the labeled PCB compound and for fruitful insights and stimulating discussions. We also thank the Deutsche Forschungsgemeinschaft (DFG) for funding the project. The quantum mechanical and QM/MM calculations were performed on resources provided by the Leipzig University Computing Center and by the Paderborn Center for Parallel Computing, respectively. C.W. thanks E. Paltanin and A. Belgara (Universität Leipzig) for preliminary NMR calculations on related tetrapyrrole chromophores.
Funding
Open Access funding enabled and organized by Projekt DEAL. This work was supported by the DFG (project ID 282144690 to J.M. and project ID 417685888 to C.S.).
Author information
Authors and Affiliations
Contributions
W.G., J.M., and C.S. conceived the experiments; S.A., L.K., P.B., C.W., and C.S. performed the experiments; S.A., W.G., C.W., and C.S. analyzed the data; S.A., C.W., and C.S. drafted the paper; L.K., P.B., W.G., J.M., C.W., and C.S. revised and edited the paper; W.G., J.M., and C.S. supervised the experiments and gave the final approval of the manuscript supervision; J.M., and C.S. acquired the funding. All authors read and approved the manuscript as submitted.
Corresponding authors
Ethics declarations
Conflict of interest
The authors have no conflicts of interest or competing interests to declare.
Additional information
This publication is dedicated to Prof. Silvia E. Braslavsky, a pioneer in photobiology and photobiophysics, on the occasion of her 80th birthday.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Altmayer, S., Köhler, L., Bielytskyi, P. et al. Light- and pH-dependent structural changes in cyanobacteriochrome AnPixJg2. Photochem Photobiol Sci 21, 447–469 (2022). https://doi.org/10.1007/s43630-022-00204-4
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s43630-022-00204-4