Abstract
Microphytobenthos and sea ice algae comprise globally significant photosynthetic biofilms. While their microalgal and bacterial constituents are well characterized, there is very little information on their viral communities or on the virus–bacteria and virus–algae interactions within them. While high levels of interaction might be expected because of the high density of cells, infection rates, particularly of microalgae, have been found to be low. It remains unclear whether this is a result of environment characteristics, developed resistance or because of the small number of studies.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Viruses are the most abundant ‘life form’ on earth, with an estimated total abundance in the oceans of ~ 1030 particles (Suttle 2007). They are responsible for 10–30% of bacterial mortality, but in some circumstances, this can reach up to 100% (Danovaro et al. 2008). In fact, it has been estimated that about 1023 phytoplankton and bacterial cells per second are infected by viruses in the world’s oceans (Knowles et al. 2016). Viral lysis of bacteria and microalgae increases the pool of dissolved organic matter and thus turbo charges the microbial loop. These changes in the organic matter flow induced by viral lysis have been termed the ‘viral shunt’ (Wilhelm and Suttle 1998). Although virus particles are abundant in sea water, with up to 106–109/ml, their critical role in marine microbial communities was not established until the late 1980s (Bergh et al. 1989; Proctor and Fuhrman 1990). Initially, most attention was given to the infection of prokaryote hosts, but phytoplankton infection by viruses was identified by Suttle et al. (1990), who estimated that viral infection could reduce primary production by up to 78%. Subsequently, viruses have been found to be widespread in microalgae and found to infect all major phytoplankton groups, including chlorophytes (van Etten 1981), prymnesiophytes (Schroeder et al. 2002), rhaphidophytes (Nagasaki and Yamaguchi 1997), diatoms (Nagasaki et al. 2004; Nagasaki 2008) and dinoflagellates (Nagasaki 2008). Most identified phytoplankton viruses have been classified as dsDNA viruses, although ssDNA and ssRNA viruses have been found to infect a few diatom and dinoflagellate taxa (Nagasaki 2008).
Virus infection rates in marine ecosystems have been thought to be largely host density mediated, i.e. the more abundant a species becomes the more likely it is that it will interact with a virus, become infected and then suffer cell lysis. Thus, viruses were seen to have the ability to control species succession and maintain maximum biodiversity; this became known as the ‘kill the winner’ scenario (Thingstad 2000). However, rather than always causing cell lysis, some viruses transfer their DNA into the host’s genome without killing it, a process known as lysogeny. This mechanism, which enables the virus to coexist with a rare host over many generations, is thought to be increasingly important in oligotrophic environments (Thingstad and Bratbak 2016). Furthermore, it has also been shown that as host cell densities increase, some viruses integrate themselves into their host. The viruses replicate more slowly, but also avoid competing with other viruses and their own host’s immune system; this includes a switch between a lytic and lysogenous mode (also referred to as ‘temperate’), a process termed ‘Piggyback the Winner’ (Knowles et al. 2016; Silveira and Rohwer 2016). However, the assumption that viruses are always able to control biodiversity through selective infection has more recently been questioned (Chi and Goldenfeld 2017; Winter et al. 2010). Instead, it has been proposed that coevolution, or ‘an antagonistic arms race’ mostly maintains biomass and biodiversity (Chi and Goldenfeld 2017). This concept is supported by the ‘Red Queen hypothesis’ (Bidle et al. 2007; Frada et al. 2008; Zhao et al. 2013), who suggested that high recombination rates in some bacterial populations allow for rapid adaptation to novel phage phenotypes.
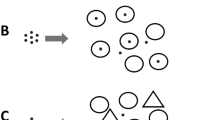
Although phytoplankton are the dominant primary producers in the marine environment, there are two substrate-bound marine ecosystems, namely, microphytobenthos (MPB) and sea ice, that also play important global roles (Fig. 1). Even though both of these ecosystems make a significant contribution to global primary production, the role of viruses in neither has received much attention. Because algal cells, bacteria and viruses are in much closer proximity in these substrate-bound biofilms than in open sea water, it is intuitively hypothesized that virus–bacteria and virus–microalgal interactions would be even more important.
Sea ice and sediment photosynthetic biofilms. The figure shows the relationship between ice, sediments, seawater, algal cells, bacteria, viruses and extracellular polymeric substances (EPS). The EPS surrounds the cells and fills the spaces between cells and substrate. Note the similarity in structure with the ice ecosystem essentially being an inverted benthos
Microphytobenthos
Benthic photosynthetic biofilms (i.e. MPB) are widespread in both marine and non-marine environments. In marine environments they occur in subtidal and intertidal areas within the euphotic zone, i.e. in depths up to ~ 30 m. These communities are often highly productive and in shallow, sunlit waters often contribute more of the total primary production than the phytoplankton (Cahoon 1999). The earth has an estimated total coastline length of ~ 356,000 km (CIA WorldFact book) and so MPB habitats potentially cover ~ 100,000 to 300,000 km2. Even though this compares with a total ocean surface area of ~ 510,000,000 km2, because coastal areas are usually shallow and typically have high nutrient concentrations, they provide a disproportionately large share of global primary and secondary production.
MPB communities occur in the top 1–5 mm of marine sediments, with the depth limited by either light penetration into the sediment or the ability of the microalgal cells, usually diatoms, to migrate. These environments are typically highly vertically structured with steep gradients in oxygen, pH, sulphide and nutrients (Jørgensen 2001). Unlike planktonic environments, the photosynthetic component of sediments is mostly dominated by diatoms with smaller contributions of Cyanobacteria and chlorophytes. MPB biomass is often very high, regularly exceeding 500 mg/m2 chlorophylla (Beardall and Light 1994; Dayton et al. 1986; Underwood 2010).
Most studies of marine benthic viruses have so far focused on those from the deep sea (Danovaro et al. 2008; Dell’anno et al. 2015; Jørgensen and Marshall 2015; Manea et al. 2019; Middleboe et al. 2011) rather than those associated with MPB. Even those that have examined MPB have mostly only examined virus–bacteria interactions (Helton et al. 2009, 2011). Virus abundances in marine sediments are mostly between 107 and 1011/g, roughly 100 times that of the overlying sea water (Carreira et al. 2015; Helton et al. 2011). While most viruses in sediments are usually bacteriophages, in a genomic study of estuarine Chesapeake Bay sediments, up to 11% of sequences were found to belong to Phycodnaviridae, which is associated with algae (Helton and Wommack 2009). Also, Montainie et al. (2015) found more than half the viruses found in a French Atlantic estuary had a capsid size greater than 65 nm, a size range usually associated with eukaryote viruses.
There have been several studies on the seasonal dynamics of viruses in sediments. In a study of a Wadden Sea photosynthetic microbial mat, virus abundance was found, not surprisingly, to track bacterial and microalgal abundances in both sediment depth and season (Carreira et al. 2015). The correlation was weaker when photoautotrophs were absent.
Although viruses can be up to 1000 times more abundant in benthic sediments than in the water column (Danvaro and Serresi 2000), infection rates of prokaryotes, particularly in freshwater systems have been found to be unusually low (Danovaro et al. 2008; Filippini et al. 2006). For example, in a freshwater lake Filipini et al. (2006) found only four out of ~ 15,000 bacteria cells in the sediments were infected compared with 300 of ~ 5000 in the overlying water. Montaine et al. (2015) provided a number of possible explanations including an allochthonous origin for many eukaryote viruses, larger burst size of algal viruses and possible sorption of viral particles onto mineral matter or embedding in the extracellular polymeric substance (EPS) matrix.
Hewson et al. (2001) were the first to demonstrate the effect of elevated virus-like particles (VLP) on benthic microalgae (MPB). They showed that the number of both bacteria and microalgae in eutrophic and oligotrophic conditions declined when exposed to elevated VLP numbers. The response varied taxonomically, however, with euglenophyte cell numbers decreasing but dinoflagellates increasing.
While diatoms tend to dominate marine MPB communities, brackish and freshwater environments are more typically dominated by cyanobacteria (Bolhuis et al. 2013). These environments share most of the physical attributes of their marine counterparts and also have strong prokaryote–virus interactions (Voories et al. 2015).
Sea ice
Each year, sea ice expands to cover 20 million km2 of the Southern Hemisphere and 12 million km2 of the Northern Hemisphere. As the sea ice forms, in late autumn, it traps phytoplankton cells which then grow to become vibrant, high biomass communities. These communities are concentrated at the ice water interface and in trapped brine pockets, often with high concentrations of EPS (Fig. 2). Cells in the brine pockets in particular experience extremes in temperature, salinity, gas and nutrient concentrations (Thomas and Dieckmann 2004). For most of the year, these communities are dominated by diatoms (Ryan et al. 2006), although in summer these are replaced by a phytoflagellate community (McMinn et al. 2017; Stoecker et al. 1992). Biomass can often reach very high levels, greater than 300 mg/m2 chlorophyll a, which can be greater than the integrated concentration in the underlying water column (Meiners et al. 2012, 2018).
Bacteriophages were reported from the sea ice as early as 1994 (Maranger et al. 1994a) and from both the Arctic (Maranger et al. 1994a) and the Antarctic (Maranger et al. 1994b; Patterson and Laybourne-Parry 2008). The first virus to be isolated from polar sea ice belonged to the typical sea ice bacteria genera, Paraglaciecola and Octadecabacter (Luhtanen et al. 2018), although phage–host systems had earlier been isolated and characterized from Baltic Sea ice (Luhtanen et al. 2014). Large viruses, i.e. those thought likely to infect eukaryotes, were first recorded from the sea ice by Gowing et al. (2002, 2003). These viruses, which comprised up to 18% of the virus community, occurred at abundances of 106–108/ml and were strongly correlated with chlorophylla concentration. However, in spite of the large number of eukaryote cells examined (i.e. > 10,000), no diatoms were found to be infected, although other microeukaryotes, such as Pyramimonas and Cryothecomonas, were (Gowings 2003). Virus concentrations of up to 1.7 × 109/ml with exceptionally high enrichment factors of up to 2800 have been reported from Arctic sea ice (Collins and Deming 2011).
In a study of the seasonal dynamics of viruses in Antarctic sea ice, Patterson and Laybourne-Parry (2008) found no seasonal pattern in the abundance of viruses or bacteria, although there was a relationship between season and virus to bacteria ratios (0.2–20.8), which were significantly lower in winter. Much higher ratios, 10–72, have been reported from Arctic sea ice (Maranger et al. 1994a, b). Gowing et al. (2004), likewise found no correlation between virus and bacterial numbers but did identify a correlation with chlorophylla concentration. Interestingly, Patterson and Laybourne-Parry (2008) found that between 40 and 50% of bacteria cells were infected with viruses. Maranger et al. (1994a) found that there were 10–100× more viruses in the underlying water than in the ice, although Patterson and Laybourne-Parry (2008) found the reverse.
Methodological considerations
Progress in understanding virus–bacteria and virus–algae interactions in photosynthetic biofilms has run parallel with the development of new protocols, methods and datasets; in particular, the establishment of large datasets of viral and host parameters, such as viral and microbial abundances, ratios of virus and host cell abundance, gross viral production, viral decay rates, viral diversity and viral genomes (Dell'Anno et al. 2015). The accurate use of these parameters relies on the development of new protocols for highly efficient extraction of viruses from the sediments and sea ice, the development of dilution-based gross viral production procedures, improvement in the extraction efficiency of viral DNA and RNA, development of the viral metagenomic analyses without amplification of nucleic acid and the establishment of specific viral reference genomes datasets for photosynthetic biofilms (Davila-Ramos et al. 2019). Recently, several new protocols for the resuspension of viruses (Trubl et al. 2016), the quantification of gross viral production (Rastelli et al. 2016) and optimized viral DNA metagenomes in soil and sediments (Trubl et al. 2019) have provided possible methods to study the virus–bacteria and virus–algae interactions in benthic photosynthetic biofilms. These would allow abundance estimations, gross production, community structure and elucidate the potential functions of viruses in the environments.
They would also allow the linkage between virus and host cells to be determined. However, whether these protocols are appropriate for sea ice ecosystems still needs to be tested and verified. In addition, the molecular mechanisms of the virus–bacteria and virus–algae interactions requires the isolation and genomic analysis of more virus–host interaction systems in these photosynthetic biofilms (Luhtanen et al. 2018).
In addition to molecular approaches, some advanced chemical methods have been applied to virus–bacteria and virus–algae interactions in the marine environments. These include ultraviolet–visible absorbance, excitation emission matrix fluorescence, ultrahigh-resolution mass spectrometry and nuclear magnetic resonance spectroscopy (Zhao et al. 2017, 2019). The applications of these methods to photosynthetic biofilms will also significantly improve our understanding of the interactions between virus, bacteria and algae in the future.
Conclusions
Despite the importance of shallow benthic and sea ice habitats to global marine primary production and their influence on biogeochemical cycles, there are very few studies that include the role of viruses in these microbial ecosystems. Studies of both ecosystems report high virus numbers but very few reports on infection rates or on the activity of the viral shunt. Likewise, there are currently no reports of lysogeny in either MPB or sea ice algae hosts. Although, because of the close proximity of cells to each other, such virus–host interactions would be expected to be higher than in pelagic systems, the sparse observations so far have in fact indicated the opposite, with mostly low levels of infection, particularly in microalgae.
It is clear that research into the role of viruses in photosynthetic biofilms is still at a preliminary stage. Future work will need to initially focus on identifying virus infection systems and then move on to determine the susceptibility to viral infection and then quantify the role of viruses in biogeochemical cycles and biodiversity regulation. Although these initial observations discussed here are characteristic of all photosynthetic biofilms is of particular interest. If further studies confirm these early results, then understanding what makes cells resistant to infection could have broad implications for viral infections generally. If the results are not confirmed, then virus–microbe infections in these biofilms will necessitate a major rethink of the importance of viruses in controlling microbial biomass, speciation and biogeochemical cycling. Thus, there will need to be many more studies of the role of viruses in marine photosynthetic biofilms before a confident appraisal of their role can be realized.
References
Beardall J, Light B (1994) Biomass, productivity and nutrient requirements of microphytobenthos. CSIRO Tech Rep CSIRO Inst Nat Resour Environ Port Phillip Bay Environ Study 16:1–12
Bergh Ø, Børsheim KY, Bratbak G, Heldal M (1989) High abundance of viruses found in aquatic environments. Nature 340:467–468
Bolhuis H, Fillinger L, Stal LJ (2013) Coastal microbial mat diversity along a salinity gradient. PLoS ONE 8:e63166
Bidle KD, Haramaty L, Barcelos e Ramos J, Falkowski P (2007) Viral activation and recruitment of metacaspases in the unicellular coccolithophore, Emiliania huxleyi. Proc Natl Acad Sci USA 104:6049–6054
Cahoon LB (1999) The role of benthic microalgae in neritic ecosystems. Oceanogr Mar Biol Ann Rev 37:47–86
Carreira C, Piel T, Staal M, Stuut J, Middleboe M, Brussaard CPD (2015) Microscale spatial distributions of microbes and viruses in intertidal photosynthetic microbial mats. SpringerPlus 4:239
Chi X, Goldenfeld N (2017) Coevolution maintains diversity in the stochastic “Kill the Winner” model. Phys Rev Lett 119:268101
Collins RE, Deming JW (2011) Abundant dissolved genetic material in Arctic sea ice. Part II: Viral dynamics during autumn freeze-up. Polar Biol 34:1831–1841
Danovaro R, Corinaldes C, Filippini M, Fischer UR, Gessner MO, Jacquet S, Magagnini M, Velimirov B (2008) Viriobenthos in freshwater and marine sediments: a review. Freshwater Biol 53:1186–1213
Danovaro R, Serresi M (2000) Viral density and virus-to-bacterium ratio in deep-sea sediments of the Eastern Mediterranean. Appl Environ Microbiol 66:1857–1861
Dayton PK, Watson D, Palmisano A, Barry JP, Oliver JS, Rivera D (1986) Distribution patterns of benthic microalgal standing stock at McMurdo Sound, Antarctica. Polar Biol 6:207–213
Dávila-Ramos S, Castelán-Sánchez HG, Martínez-Ávila L, Sánchez-Carbente MDR, Peralta R, Hernández-Mendoza A, Dobson ADW, Gonzalez RA, Pastor N, Batista-García RA (2019) A review on viral metagenomics in extreme environments. Front Microbiol 10:2403
Dell'Anno A, Corinaldesi C, Danovaro R (2015) Virus decomposition provides an important contribution to benthic deep-sea ecosystem functioning. Proc Natl Acad Sci USA 112:E2014–E2019
Filippini M, Buesing N, Battarel Y, Sime-Ngando T, Gessner MO (2006) Infection paradox: high abundance but low impact of freshwater benthic virus. Appl Environ Microbiol 72:4893–4898
Frada M, Probert I, Allen MJ et al. (2008) The “Cheshire Cat” escape strategy of the coccolithophore Emiliania huxleyi in response to viral infection. Proc Natl Acad Sci USA 105:15944–15949
Gowing MM (2003) Large viruses and infected microeukaryotes in Ross Sea summer pack ice habitats. Mar Biol 142:1029–1040
Gowing MM, Garrison DL, Gibson AH, Krupp JM, Jeffries MO, Fritsen CH (2004) Bacterial and viral abundance in Ross Sea summer pack ice communities. Mar Ecol Progr Ser 279:3–12
Gowing MM, Riggs BE, Garrison DL, Gibson AH, Jeffries MO (2002) Large viruses in Ross Sea late autumn pack ice habitats. Mar Ecol Prog Ser 241:1–11
Helton RR, Wang K, Kan J, Powell DH, Wommack KE (2011) Interannual dynamics of viriobenthos abundance and morphological diversity in Chesapeake Bay sediments. FEMS Microbiol Ecol 79:474–486
Helton RR, Wommack KE (2009) Seasonal dynamics and metagenomic characterization of estuarine viriobenthos assemblages by randomly amplified polymorphic DNA PCR. Appl Environ Microbiol 75:2259–2265
Hewson I, O’Neil JO, Heil CA, Bratbak G, Dennison WC (2001) Effects of concentrated viral communities on photosynthesis and community composition of co-occurring benthic microalgae and phytoplankton. Aquat Microb Ecol 25:1–10
Jørgensen BB (2001) Life in the benthic boundary layer. In: Boudreau BP, Jørgensen BB (eds) The boundary layer, transport processes and biogeochemistry. Oxford University Press, Oxford, pp 348–373
Jørgensen BB, Marshall IPG (2016) Slow microbial life in the seabed. Ann Rev Mar Sci 8:311–332
Knowles B, Silveira CB, Bailey BA, Barott K, Cantu VA, Cobián-Güemes AG, Coutinho FW, Dinsdale EA, Felts B, Furby KA, George EE, Green KT, Gregoracci GB, Haas AF, Haggerty JM, Hester ER, Hisakawa N, Kelly LW, Lim YW, Little M et al. (2016) Lytic to temperate switching of viral communities. Nature 531:466–470
Luhtanen AM, Eronen-Rasimus E, Kaartokallo H, Rintala JM, Autio R, Roine E (2014) Isolation and characterization of phage–host systems from the Baltic Sea ice. Extremophiles 18:121–130
Luhtanen A, Eronen-Rasimus E, Oksanen HM, Tilson JL, Delille B, Dieckmann GS, Rintala JM, Bamford DH (2018) The first known virus isolates from Antarctic sea ice have complex infection patterns. FEMS Microbiol Ecol 94:fiy028
Manea E, Dell’Anno A, Rastelli E, Tangherlini M, Nunoura T, Nomaki H, Danovaro R, Corinaldesi C (2019) Viral infections boost prokaryotic biomass production and organic cycling in hadal trench sediments. Front Microbiol 10:1952
Maranger R, Bird DF, Juniper SK (1994a) Viral and bacterial dynamics in Arctic sea ice during the spring algal bloom near Resolute, N.W.T., Canada. Mar Ecol Prog Ser 111:121–127
Maranger R, Bird DF, Karl DM (1994b) Palmer LTER: Spatial distribution of viruses in the Palmer LTER region. Antarctic J US 29:209–211
McMinn A, Muller M, Martin A, Ryan KG (2017) Effects of changing pH and CO2 concentration on a late summer surface sea ice community. Mar Biol 164:86–96
Meiners KM, Vancoppenolle M, Thanassekos S, Dieckmann GS, Thomas DN, Tison J-L, Arrigo KR, Garrison DL, McMinn A, Lannuzel D, van der Merwe P, Swadling KM, Smith WO Jr, Melnikov I, Raymond B (2012) Chlorophyll a in Antarctic sea ice from historical ice core data. Geophys Res Lett 39:L21602
Meiners KM, Vancoppenolle M, Carnat G, Castellani G, Delille B, Delille D, Dieckmann GS, Flores H, Fripiat F, Grotti M, Lange BA, Lannuzel D, Martin A, McMinn A, Nomura D, Peeken I, Rivaro P, Ryan KG, Stefels J, Swadling K et al. (2018) Chlorophyll-a in Antarctic landfast sea ice: a synthesis of historical ice core data. J Geophys Res Oceans 123:8444–8459
Middleboe M, Glud RN, Filippini M (2011) Viral abundance and activity in the deep sub-seafloor biosphere. Aquat Microb Ecol 63:1–8
Montanié H, de Crignis MG, Lavaud J (2015) Viral impact on prokaryotic and microalgal activities in the microphytobenthic biofilm of an intertidal mudflat (French Atlantic coast). Front Microbiol 6:1214
Nagasaki K (2008) Review: dinoflagellates diatoms and their viruses. J Microbiol 46:235–243
Nagasaki K, Tomaru Y, Katanozaka N, Shirai Y, Nishida K, Itakura S, Yamaguchi M (2004) Isolation and characterization of a novel single-stranded RNA virus infecting the bloom-forming diatom Rhizosolenia setigera. Appl Environ Microbiol 70:704–711
Nagasaki K, Yamaguchi M (1997) Isolation of a virus infectious to the harmful bloom causing microalga Heterosigma akashiwo (Raphidophyceae). Aquat Microb Ecol 13:135–140
Patterson H, Laybourn-Parry J (2008) Antarctic sea ice viral dynamics over an annual cycle. Polar Biol 35:491–497
Proctor LM, Fuhrman JA (1990) Viral mortality of marine bacteria and cyanobacteria. Nature 343:60–62
Rastelli E, Dell'Anno A, Corinaldesi C, Middelboe M, Noble RT, Danovaro R (2016) Quantification of viral and prokaryotic production rates in benthic ecosystems: a methods comparison. Front Microbiol 7:1501
Ryan KG, Hegseth EN, Martin A, Davy SK, O’Toole R, Ralph PJ, McMinn A, Thorn CJ (2006) Comparison of the microalgal community within fast ice at two sites along the Ross Sea coast, Antarctica. Antarct Sci 18:583–594
Schroeder DC, Oke J, Malin G, Wilson WH (2002) Coccolithovirus (Phycodnaviridae): characterisation of a new large dsDNA algal virus that infects Emiliania huxleyi. Arch Virol 147:1685–1698
Silveira CB, Rohwer FL (2016) Piggyback-the-Winner in host-associated microbial communities. NPJ Biofilms Microbiomes 2:16010
Stoecker DK, Buck KR, Putt M (1992) Changes in the sea-ice brine community during the spring–summer transition, McMurdo Sound, Antarctica. I. Photosynthetic protists. Mar Ecol Prog Ser 84:265–278
Suttle CA (2007) Marine viruses—major players in the global ecosystem. Nat Rev Microbiol 5:801–812
Suttle CA, Chan AM, Cottrell MT (1990) Infection of phytoplankton by viruses and reduction of primary productivity. Nature 347:467–469
Thingstad TF (2000) Elements of a theory for the mechanisms controlling abundance, diversity, and biogeochemical role of lytic bacterial viruses in aquatic systems. Limnol Oceanogr 45:1320–1328
Thingstad TF, Bratbak G (2016) Viral strategies at sea. Nature 531:454–455
Thomas DN, Dieckmann GS (2004) Sea Ice, an introduction to its physics, chemistry, biology and geology. Blackwell, Oxford
Trubl G, Solonenko N, Chittick L, Solonenko SA, Rich VI, Sullivan MB (2016) Optimization of viral resuspension methods for carbon-rich soils along a permafrost thaw gradient. PeerJ 4:e1999
Trubl G, Roux S, Solonenko N, Li YF, Bolduc B, Rodríguez-Ramos, Eloe-Fadrosh EA, Rich VI, Sullivan MB (2019) Towards optimized viral metagenomes for double-stranded and single-stranded DNA viruses from challenging soils. PeerJ 7:e7265
Underwood GJC (2010) Microphytobenthos and phytoplankton in the Severn estuary, UK: present situation and possible consequences of a tidal energy barrage. Mar Pollut Bull 61:83–91
Van Etten JL, Meints RH, Burbank DE, Kuczmarski D, Cuppels DA, Lane LC (1981) Isolation and characterization of a virus from the intracellular green alga symbiotic with Hydra viridis. Virol 113:704–711
Voories AA, Eisenlord SD, Marcus DN, Duhaime MB, Cavalcoli JD, Dick GJ (2015) Ecological and genetic interactions between cyanobacteria and viruses in a low-oxygen mat community inferred through metagenomics and metatranscriptomics. Environ Microbiol 18:358–371
Wilhelm SW, Suttle CA (1998) Viruses and nutrient cycles in the sea-viruses play critical roles in the structure and function of aquatic food webs. Bioscience 49:781–788
Winter C, Bouvier T, Weinbauer MG, Thingstad TF (2010) Trade-offs between competition and defense specialists among unicellular planktonic organisms: the “Killing the Winner” hypothesis revisited. Microbiol Mol Biol Rev 74:42–57
Zhao Y, Temperton B, Thrash JC, Schwalbach MS, Vergin KL, Landry ZC, Ellisman M, Deerinck T, Sullivan MB, Giovannoni SJ (2013) Abundant SAR11 viruses in the ocean. Nature 494:357–360
Zhao Z, Gonsior M, Luek J, Timko S, Ianiri H, Hertkorn N, Schmitt-Kopplin P, Fang X, Zeng Q, Jiao N, Chen F (2017) Picocyanobacteria and deep-ocean fluorescent dissolved organic matter share similar optical properties. Nat Commun 8:15284
Zhao Z, Gonsior M, Schmitt-Kopplin P, Zhan Y, Zhang R, Jiao N, Chen F (2019) Microbial transformation of virus-induced dissolved organic matter from picocyanobacteria: coupling of bacterial diversity and DOM chemodiversity. ISME J 13:2551–2565
Acknowledgements
This study was funded by the Marine S&T Fund of Shandong Province for Pilot National Laboratory for Marine Science and Technology (Qingdao) (No. 2018SDKJ0406-6), National Key Research and Development Program of China (2018YFC1406704), The Fundamental Research Funds for the Central Universities (201812002) and the Natural Science Foundation of China (Nos. 41976117 and 41606153). We thank Wenjing Zhang for drafting Fig. 1.
Author information
Authors and Affiliations
Contributions
AM and YL made an equal contribution. The ideas and substance of the review were conceived equally by AM, YL and MW. The manuscript was written by AM with content and editorial advice from YL.
Corresponding author
Ethics declarations
Conflict of interest
The authors declare no conflict of interest.
Animal and human rights statement
This article does not contain any studies with human participants or animals performed by any of the authors.
Additional information
Edited by Jiamei Li.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
McMinn, A., Liang, Y. & Wang, M. Minireview: The role of viruses in marine photosynthetic biofilms. Mar Life Sci Technol 2, 203–208 (2020). https://doi.org/10.1007/s42995-020-00042-2
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s42995-020-00042-2