Abstract
Weeds cause tremendous economic and ecological damage worldwide. The number of genomes established for weed species has sharply increased during the recent decade, with some 26 weed species having been sequenced and de novo genomes assembled. These genomes range from 270 Mb (Barbarea vulgaris) to almost 4.4 Gb (Aegilops tauschii). Importantly, chromosome-level assemblies are now available for 17 of these 26 species, and genomic investigations on weed populations have been conducted in at least 12 species. The resulting genomic data have greatly facilitated studies of weed management and biology, especially origin and evolution. Available weed genomes have indeed revealed valuable weed-derived genetic materials for crop improvement. In this review, we summarize the recent progress made in weed genomics and provide a perspective for further exploitation in this emerging field.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
The Arabidopsis thaliana genome sequence was released in 2000 and represented a hallmark in plant research as the first sequenced and assembled plant genome (The Arabidopsis Genome Initiative 2000). Driven by the rapid development of sequencing technologies and bioinformatics methods, hundreds of plant genomes have since been sequenced and assembled (Sun et al. 2022a). High-quality reference genomes have provided vital resources for molecular genetics and have accelerated and improved precision crop breeding. Whole-genome genetic information for entire populations also offers accurate and plentiful molecular markers from which to infer and reconstruct the complex evolutionary histories of plant species, particularly for crop species.
Crops are not the only plants that grow in fields, however, weeds—defined here as non-crop plants growing within crop fields—can have competitive advantages over crop plants and cause yield loss (Basu et al. 2004). To date, 2847 plant species belonging to 177 families and 1118 genera have been designated as weeds (Weed Science Society of America database, http://www.wssa.net). Notably, weeds are the main contributors to yield loss for field crops, compared to pests and pathogens, and on average result in a 30% annual yield loss across the major crops (Oerke 2006). Although agricultural production is substantially affected by weeds, until recently, weed studies have not been given sufficient attention, in terms of both traditional molecular biology and genome analyses. Recent comparative genomics and population genomics analyses have revealed the effect of weeds on crop agronomic traits and the mechanisms underlying weediness, such as in barnyard grass (Echinochloa crus-galli), tall waterhemp (Amaranthus tuberculatus), and weedy rice (Oryza sativa f. spontanea) (Guo et al. 2017; Kreiner et al. 2018, 2019; Gaines et al. 2020; Qiu et al. 2020). In addition, the complex relationships among crops, weeds, humans, and abiotic environments in agricultural ecosystems, provide an ideal model for the study of biological interactions. Considering the potential of weed biology, recently, the weed research community endeavored to initiate genome sequencing of global weed species (Ravet et al. 2018).
In this review, we summarize genome sequencing of weed species over the past decade and explore future directions and potential applications in agricultural production.
Weed genome sequencing and de novo assembly
In recent years, the number of genomes released for weed species has sharply increased (Table 1), with genomes for at least 26 weed species being sequenced. Their genome sizes range from 270 Mb (Barbarea vulgaris) to 4360 Mb (Aegilops tauschii); 17 of these genomes have been assembled to the chromosome level, based on long-read sequencing technologies. Meanwhile, a significant improvement in sequence quality for weed genomes was achieved along with the development of new sequencing technologies. For instance, the genomes of the barnyard grass species, E. crus-galli and E. oryzicola, which grow in paddy fields and compete with rice, were assembled into draft genomes and later anchored to chromosomes by incorporating data from chromosome conformation capture (Hi-C) (Guo et al. 2017; Ye et al. 2020; Wu et al. 2022b). The genome of weedy rice (Oryza sativa f. spontanea) was also sequenced and assembled, at the chromosome level, in 2019 (Sun et al. 2019). In addition, the genomes of tetraploid Chinese sprangletop (Leptochloa chinensis) were assembled (Wang et al. 2022). An invasive weed in wheat fields, field pennycress (Thlaspi arvense), had its genome assembled in 2015 and independently anchored to chromosomes in 2021 and 2022 (Dorn et al. 2015; Geng et al. 2021; Nunn et al. 2022). The genomes for other agronomically important weeds have also been sequenced. For example, chromosome-level genomes of highly heterozygous Amaranthus species (A. tuberculatus, A. hybridus, and A. palmeri) have been developed (Montgomery et al. 2020). Of the 26 weed species, 16 are dicots from six different families, with the remaining nine species being monocots from only one family (Poaceae) (Fig. 1). Polyploid species usually exhibit more dominant advantages in their adaptation (te Beest et al. 2012), and the genomes of four polyploid weed species, comprising three tetraploid (L. chinensis, E. oryzicola, and Capsella bursa-pastoris) and one hexaploid (E. crus-galli) species, have been sequenced. Notably, several weeds are very closely related to crop species (i.e., they represent different subspecies or accessions of the same species), and the corresponding crop genome can therefore be used as a reference genome for weeds. For example, the available barnyard millet (E. colona var. frumentacea) genome provided an important reference for barnyard grass (E. colona var. colona) (Wu et al. 2022b), as did the crop sorghum (Sorghum bicolor) for Johnsongrass (Sorghum halepense), cultivated pearl millet (Pennisetum glaucum) for wild pearl millet (Pennisetum violaceum), rye (Secale cereale) for weedy rye (S. cereale subsp. segetale), sugar beet (Beta vulgaris) for sea beet (Beta vulgaris ssp. maritima), and rice for weedy rice.
List of weed species sequenced and their phylogenetic relationships. For detailed information about all genome sequencing results, please see Table 1
Whole-genome sequencing of weed populations
Whole-genome sequencing, which provides excellent tools for mining genetic mechanisms and evolutionary studies, has been widely used in crop genomics (Jia et al. 2021). Since 2017, this method has also been applied to a limited number of weed species, mainly for paddy weeds, such as weedy rice and barnyard grass (Table 2).
Weedy rice was the first weed species to be used for genomic investigation, via whole-population sequencing. As weedy rice can be considered a wild-like rice ecotype, the genome of cultivated rice provides a good reference for calling single-nucleotide polymorphisms (SNPs) in individuals. Over 650 accessions of weedy rice have been sequenced, being derived from global rice production areas, which has deepened our understanding of weedy rice origins and adaptation strategies (Li et al. 2017; Qiu et al. 2017, 2020; Imaizumi et al. 2021; Wedger et al. 2022).
Other weeds affecting paddy fields have also been studied, at the genomic level. For barnyard grass, the release of its genome (Guo et al. 2017) heralded the beginning of population genomics in this species, with over 700 genomes of accessions collected from all over the world being re-sequenced for studies on evolutionary history and typical weed adaptation syndromes (Ye et al. 2019, 2020; Wu et al. 2022b). Similarly, genome resequencing of 89 Chinese accessions revealed that sprangletop originated from a local population in tropical areas of South Asia and Southeast Asia and that the geographical range of individuals with herbicide resistance genes expanded, likely due to field management practices (Wang et al. 2022).
In recent years, significant efforts have been made to explore the adaptation and evolutionary dynamics of field pennycress. For example, 40 field pennycress lines from different altitude regions were re-sequenced, resulting in the identification of one SNP responsible for the adaptation to latitude, via constructing ultra-high-density linkage maps (Geng et al. 2021). In another example, a genomic region located on scaffold 6 was identified as causing the seedling color phenotype in field pennycress by bulk-sequencing of DNA pools from 20 wild-type and 20 pale plants (Nunn et al. 2022).
Genomic insights into weed biology
Environmental adaptation
Weeds have great potential as model systems in which to understand plant responses to biotic and abiotic stresses (Vigueira et al. 2013). They can survive in disrupted environments and persist under multiple challenges, in particular escaping from control measures in the field, including targeted tillage practices, herbicide use, and hand-weeding (Sharma et al. 2021; Neve and Caicedo 2022). In addition, weeds are not distributed in limited ecological niches, but rather, they often exhibit a widespread distribution, even among areas with distinct conditions, exemplifying their strong environmental plasticity (Sharma et al. 2021).
Genomic studies have significantly improved our understanding of weed environmental adaptations to biotic and abiotic stresses. For example, T. arvense is an annual weed from the Brassicaceae family that lives at different altitudes, ranging from sea level to 4500 m above sea level. Genomic analyses of populations from different ecological conditions identified a SNP that led to a loss-of-function allele in FLOWERING LOCUS C on chromosome 1, which contributed to the early flowering trait that was key to the success of high-elevation populations (Geng et al. 2021).
Another conspicuous trait related to environmental adaptation in weeds is herbicide resistance (Hawkins et al. 2019; Gaines et al. 2020). Comparative genomics between herbicide-susceptible and -resistant individuals, from the same species, and between species, can offer glimpses into innovations in herbicide resistance pathways (Kreiner et al. 2018). Waterhemp (A. tuberculatus), which is troublesome in maize (Zea mays) and soybean (Glycine max) fields, is notorious for exhibiting multiple herbicide-resistant (MHR) traits. Recently, a reduction–dehydration–glutathione (GSH) conjugation system was discovered as a possible pathway for MHR (Concepcion et al. 2021). In palmer amaranth (Amaranthus palmeri), genomic analysis helped determine that herbicide resistance is conferred by an extrachromosomal circular DNA (eccDNA) of about 400 kb in length that harbors 5-ENOYLPYRUVYLSHIKIMATE-3-PHOSPHATE SYNTHASE (EPSPS), which encodes the enzyme targeted by the herbicide glyphosate (Gaines et al. 2010; Molin et al. 2020). Although the amplification of genes and gene clusters, via eccDNAs or other structures, is a common stress-avoidance mechanism in plants (Nandula et al. 2014; Singh et al. 2020), it is usually transient and not stably inherited (Lanciano et al. 2017; Gaines et al. 2019).
As the most dominant weed in rice fields, barnyard grass has also evolved global resistance to major herbicides. Genome resequencing of barnyard grass individuals from Brazil, Italy, and China revealed four mutations in the gene encoding aceto-lactate synthase (ALS), which conferred herbicide resistance, namely Ala-122-Thr, Trp-574-Leu, Ser-653-Asn, and a Gly-654-Cys substitution identified for the first time, with a tendency to occur in sub-genome A (barnyard grass is a hexaploid). Moreover, after comparing the genomes of resistant and susceptible individuals from Brazil, an Arg-86-Gln mutation in the conserved degron tail region of Echinochloa AUXIN-INDUCED (AUX)/INDOLE-3-ACETIC ACID INDUCIBLE 12 (IAA12) was identified, which has since been confirmed to confer resistance to other auxin-like herbicides (LeClere et al. 2018; Figueiredo et al. 2021; Wu et al. 2022b).
Great progress has also been made in understanding the responses of weeds to biotic stresses. Before herbicides were used in agriculture, the direct interaction between weeds and human beings was through hand-weeding, which placed high pressure on weed morphology, especially plant architecture. One example is the Vavilovian mimicry or crop mimicry seen in barnyard grass (at least in E. crus-galli and E. oryzicola), an unintentional human selection (UHS) resulting from human action (Fig. 2).
Crop mimicry describes the adaptation of a weed through its acquiring some of the morphological characteristics of neighboring domesticated crops, at a specific stage of their life history, to escape their removal by hand-weeding (Barrett 1983; Ye et al. 2019). The preadapted plants, or wild species that were first to colonize in cultivated fields, during the early agricultural stage (so-called ancient weeds), gradually became mimic weeds under strong artificial (weeding) selection. Genomic signatures of human selection on crop mimicry were elucidated by comparing the genomes of mimetic and non-mimicry lines of barnyard grass collected from paddy fields in the Yangtz River basin, China (Ye et al. 2019). Several genes underlying plant architecture (e.g., tiller angle) were identified, including LAZY1, a gene responsible for plant tiller angles, which was also under selection during rice domestication. The genomic study of mimicry of rice seedlings, by barnyard grass, is an example of how weeds can adapt to disturbed environments with selective pressure from human beings, via a genomic approach.
Allelopathic secondary metabolites also are a representative response of weeds to biotic stress. Benzoxazinoids, which acted against microbial pathogens and neighboring plants, were identified in a multitude of species of the family Poaceae, such as maize, wheat (Triticum aestivum), and barnyardgrass (Frey et al. 2009; Wu et al. 2022a). As a predominant representative of benzoxazinoids in plants, DIMBOA is present in barnyard grass with multiple copies and inhibits plant height and fresh weight of neighboring rice (Guo et al. 2017). Another example is momilactone A, which has similar functions to Benzoxazinoids in rice. Based on the momilactone A biosynthesis genes of rice, a syntenic gene cluster was identified in barnyard grass. Up-regulated expression of MAS and KSL4, within this cluster, under fungal infection indicated its contribution to resistance to blast infection in the paddy environment (Guo et al. 2017).
Origins of weeds
Understanding the origin of agricultural weeds is crucial to their proper management. Weed origins can be via several routes. Preadapted plants or wild species can colonize cultivated fields in human-made ecological niches (Larson et al. 2014). With the expansion of cultivated fields, the emergence and diversification of weeds may have resulted from hybridization between crop and wild groups, along with other routes (Iriondo et al. 2018; Janzen et al. 2019).
Recent genomic studies focused on paddy weeds revealed many interesting insights about their possible origin(s) and evolution (Fig. 2). Weedy rice (Oryza sativa f. spontanea) has attracted much attention for its origin of de-domestication, i.e., the conversion of a domesticated form to a wild-like form (Wu et al. 2021). Weedy rice mimics rice cultivars, at the seedling stage, while retaining wild phenotypes, such as strong seed dormancy and shattering. De-domestication from cultivated rice (including cultivars and landraces) is the main route for rice feralization, along with introgressions from wild rice, which is commonly seen in Southeast Asia and South China, where wild rice is distributed, as well as inter-subspecies hybridization (Stewart 2017; Sun et al. 2019; Qiu et al. 2020; Wu et al. 2021). Genomic mining, aided by comparisons between the genomes of weedy, wild, and cultivated rice populations, has revealed distinct differentiation regions on chromosomes during de-domestication compared to those resulting from domestication, with the identification of a genomic island possibly underlying feralization traits on chromosome 7. This genomic region harbors Rc, controlling red pericarp and seed dormancy (Sweeney et al. 2006), and several tandem-duplicated genes encoding seed storage proteins (Li et al. 2017; Qiu et al. 2020).
A similar process was also described for the origins of E. crus-galli var. oryzoides, which is currently regarded as a paddy weed (Fig. 2). The significantly lower nucleotide diversity, longer linkage disequilibrium decay, more immune response genes, larger grains, and non-shattering spikelets in this species, compared to weed populations, indicate that var. oryzoides is an abandoned crop (Wu et al. 2022b).
Perspectives in weed genomics
We need complete, contiguous, and accurate genome assemblies for many more weed species. Indeed, in notable contrast to the massive increase in sequenced crop genomes, only 26 weeds have been decoded thanks to the sequencing and assembly of their genomes. The enormous gap between crops and weeds underscores how much weeds are currently being overlooked. For example, Commelinales, with about 750 extant species, including pickerel weed (Monochoria vaginalis), are important weeds growing in paddy fields. Likewise, common water hyacinth (Eichhornia crassip) is the most common invasive plant according to a survey by the Weed Science Society of America database (WSSA, http://www.wssa.net). Yet, these two species still lack a representative genome. Several sedges (e.g., Cyperus, Scirpus, and Fimbristilis) are found worldwide and exhibit particular weediness traits, but very little genomic information is currently available.
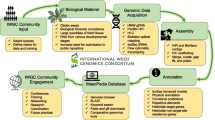
We even lack a thorough understanding and characterization of notorious weeds affecting croplands, such as hairy crabgrass (Digitaria sanguinalis), a typical upland weed growing in maize and soybean fields. Moreover, a higher-quality genome of weeds is required to shed light on related biological topics. The gap-free genomes of many plants, such as Arabidopsis, rice, and watermelon (Citrullus lanatus), have recently been assembled, providing the first complete genome structure of any plant (Song et al. 2021; Wang et al. 2021a; Deng et al. 2022). With the incorporation of sequences from highly repetitive regions and centromeres into genome assemblies at the chromosome scale, a greater understanding of the global pattern of weed polymorphisms and the genetic basis of their weedy traits and high adaptability is finally within reach, but only if more genomes are sequenced or improved upon. These issues were also noted by the International Weed Science Consortium, which has designated Plantae (www.plantae.org) as a platform for community collaboration efforts and has developed a weed genomics website (www.weedgenomics.org) (Ravet et al. 2018).
We expect and anticipate more studies exploring the population genomics of weeds, which will be helpful for the understanding of their evolutionary strategies and evolutionary ecology, while offering more options for weed management. Current evolutionary patterns tend to highlight pressure imposed by the natural environment, perhaps neglecting the role that human activities play in a novel ecosystem labeled with specific species assemblages and environmental factors. Studying weed populations with complex evolutionary trajectories of traits will enhance our ability to decode their distinct evolutionary strategies under different conditions. In addition, a better understanding of the evolution of agricultural weeds will be crucial to weed management. Given the increasing number of rapid weed adaptations, such as herbicide resistance, ongoing selection for other weedy traits should be a driving force to adjust all weed management practices to mitigate the spread and success of weeds.
With the advantage of more available genomes, weed functional genomics will step to the front stage. Our understanding of the mechanisms by which multiple weed species acquire herbicide resistance (particularly non-target resistance) to the same class of herbicide has considerably improved with released genomic information (Devine and Shukla 2000; Yuan et al. 2007; Délye 2013; Kreiner et al. 2018). For example, the availability of the barnyard grass genome made it possible to identify, for the first time, a significant increase in copy number for cytochrome P450 genes in the weed genomes, as well as a Gly-654-Cys substitution, with both strategies contributing to ALS resistance. Another example resulting from the comparative analysis of waterhemp genomes was the report of a possible pathway for MHR, via reduction–dehydration–glutathione. We anticipate that, along with the development of weed genomics, additional discoveries about gene functions and their interactions will be forthcoming.
More valuable genetic resource of weeds will be revealed with the sequencing of more weed genomes, which will have benefits for the genetic improvement of crops and even their de novo domestication. Crops, particular orphan crops, are genetically very closely related to weeds. For example, orphan crops usually have a notorious weed species in the same genus (Ye and Fan 2021). Given their strong environmental plasticity and high level of genetic variation, weeds are an untapped genetic resource for domestication. For example, mutating the orthologs for qSH1 (Shattering QTL 1) and Sh4 (Shattering 4) genes in weeping rice grass (Microlaena stipoides), an Australian wild relative of rice, caused the loss of shattering in this species (Shapter et al. 2013). Historically, some weeds have been domesticated into crops, such as rye (Rye secale) (Sun et al. 2022b). Presently, de novo domestication of new crops is an option being considered to mitigate the effects of climate change on global crop production. We propose that some weeds, in particular those mimicking crops, are ideal targets for de novo domestication.
In addition to crop improvement, weed management will also benefit from the advances in weed genomes. Gene silencing techniques are offer a promising approach to manipulate the expression level of weed traits genes to reduce their impact with improved understanding of characteristic regulated pathways (Neve 2018). For example, if genomics can identify the basis of allelopathy, weeds could be modified with low levels of allelopathic compounds, thereby reducing their competitive ability in paddy fields. However, major challenges remain to be overcome; e.g., the designation of highly specific gene silencing triggers with high heritability (Patterson et al. 2019b).
Post-transcriptional silencing, using exogenous application of RNA, known as spray-induced gene silencing (SIGS), is a promising technology that may revolutionize weed control. Several limitations and opportunities are associated with the development of this technology. The main requirement for SIGS is selective gene silencing in weeds and the absence of effects on crops and non-target organisms. Therefore, the development of this non-transgenic, and environmentally safe, technology depends largely on genome sequencing, chromosome-level assemblies, and deep knowledge of gene function for all weed species, which affect food production, and the crops whose fields they invade.
Data availability
The datasets generated during and/or analyzed during the current study are available from the corresponding author on reasonable request.
References
Abrouk M, Ahmed HI, Cubry P, Šimoníková D, Cauet S, Pailles Y et al (2020) Fonio millet genome unlocks African orphan crop diversity for agriculture in a changing climate. Nat Commun 11:4488. https://doi.org/10.1038/s41467-020-18329-4
Barrett SH (1983) Crop mimicry in weeds. Econ Bot 37:255–282. https://doi.org/10.1007/BF02858881
Basu C, Halfhill MD, Mueller TC, Stewart CN (2004) Weed genomics: new tools to understand weed biology. Trends Plant Sci 9:391–398. https://doi.org/10.1016/j.tplants.2004.06.003
Bieker VC, Battlay P, Petersen B, Sun X, Wilson J, Brealey JC et al (2022) Uncovering the genomic basis of an extraordinary plant invasion. Sci Adv 8:eabo5115
Byrne SL, Erthmann PØ, Agerbirk N, Bak S, Hauser TP, Nagy I et al (2017) The genome sequence of Barbarea vulgaris facilitates the study of ecological biochemistry. Sci Rep 7:40728. https://doi.org/10.1038/srep40728
Concepcion JCT, Kaundun SS, Morris JA, Hutchings S-J, Strom SA, Lygin AV et al (2021) Resistance to a nonselective 4-hydroxyphenylpyruvate dioxygenase-inhibiting herbicide via novel reduction–dehydration–glutathione conjugation in Amaranthus tuberculatus. New Phytol 232:2089–2105. https://doi.org/10.1111/nph.17708
de Figueiredo MRA, Küpper A, Malone JM, Petrovic T, de Figueiredo ABTB, Campagnola G et al (2021). An in-frame deletion mutation in the degron tail of auxin co-receptor IAA2 confers resistance to the herbicide 2,4-D in Sisymbrium orientale. https://doi.org/10.1101/2021.03.04.433944
Délye C (2013) Unravelling the genetic bases of non-target-site-based resistance (NTSR) to herbicides: a major challenge for weed science in the forthcoming decade. Pest Manag Sci 69:176–187. https://doi.org/10.1002/ps.3318
Deng Y, Liu S, Zhang Y, Tan J, Li X, Chu X et al (2022) A telomere-to-telomere gap-free reference genome of watermelon and its mutation library provide important resources for gene discovery and breeding. Mol Plant 15:1268–1284. https://doi.org/10.1016/j.molp.2022.06.010
Devine MD, Shukla A (2000) Altered target sites as a mechanism of herbicide resistance. Crop Prot 19:881–889. https://doi.org/10.1016/S0261-2194(00)00123-X
Dorn KM, Fankhauser JD, Wyse DL, Marks MD (2015) A draft genome of field pennycress (Thlaspi arvense) provides tools for the domestication of a new winter biofuel crop. DNA Res 22:121–131. https://doi.org/10.1093/dnares/dsu045
Frey M, Schullehner K, Dick R, Fiesselmann A, Gierl A (2009) Benzoxazinoid biosynthesis, a model for evolution of secondary metabolic pathways in plants. Phytochemistry 70:1645–1651. https://doi.org/10.1016/j.phytochem.2009.05.012
Gaines TA, Zhang W, Wang D, Bukun B, Chisholm ST, Shaner DL et al (2010) Gene amplification confers glyphosate resistance in Amaranthus palmeri. Proc Natl Acad Sci 107:1029–1034. https://doi.org/10.1073/pnas.0906649107
Gaines TA, Patterson EL, Neve P (2019) Molecular mechanisms of adaptive evolution revealed by global selection for glyphosate resistance. New Phytol 223:1770–1775. https://doi.org/10.1111/nph.15858
Gaines TA, Duke SO, Morran S, Rigon CAG, Tranel PJ, Küpper A et al (2020) Mechanisms of evolved herbicide resistance. J Biol Chem 295:10307–10330. https://doi.org/10.1074/jbc.REV120.013572
Geng Y, Guan Y, Qiong L, Lu S, An M, Crabbe MJC et al (2021) Genomic analysis of field pennycress (Thlaspi arvense) provides insights into mechanisms of adaptation to high elevation. BMC Biol 19:143. https://doi.org/10.1186/s12915-021-01079-0
Guo L, Qiu J, Ye C, Jin G, Mao L, Zhang H et al (2017) Echinochloa crus-galli genome analysis provides insight into its adaptation and invasiveness as a weed. Nat Commun 8:1031. https://doi.org/10.1038/s41467-017-01067-5
Hawkins NJ, Bass C, Dixon A, Neve P (2019) The evolutionary origins of pesticide resistance. Biol Rev 94:135–155. https://doi.org/10.1111/brv.12440
He Q, Kim K-W, Park Y-J (2017) Population genomics identifies the origin and signatures of selection of Korean weedy rice. Plant Biotechnol J 15:357–366. https://doi.org/10.1111/pbi.12630
Imaizumi T, Ebana K, Kawahara Y, Muto C, Kobayashi H, Koarai A et al (2021) Genomic divergence during feralization reveals both conserved and distinct mechanisms of parallel weediness evolution. Commun Biol 4:1–11. https://doi.org/10.1038/s42003-021-02484-5
Iriondo JM, Milla R, Volis S, Rubio de Casas R (2018) Reproductive traits and evolutionary divergence between Mediterranean crops and their wild relatives. Plant Biol 20:78–88. https://doi.org/10.1111/plb.12640
Janzen GM, Wang L, Hufford MB (2019) The extent of adaptive wild introgression in crops. New Phytol 221:1279–1288. https://doi.org/10.1111/nph.15457
Jia J, Zhao S, Kong X, Li Y, Zhao G, He W et al (2013) Aegilops tauschii draft genome sequence reveals a gene repertoire for wheat adaptation. Nature 496:91–95. https://doi.org/10.1038/nature12028
Jia L, Xie L, Lao S, Zhu Q-H, Fan L (2021) Rice bioinformatics in the genomic era: status and perspectives. Crop J 9:609–621. https://doi.org/10.1016/j.cj.2021.03.003
Kasianov AS, Klepikova AV, Kulakovskiy IV, Gerasimov ES, Fedotova AV, Besedina EG et al (2017) High-quality genome assembly of Capsella bursa-pastoris reveals asymmetry of regulatory elements at early stages of polyploid genome evolution. Plant J 91:278–291. https://doi.org/10.1111/tpj.13563
Kreiner JM, Stinchcombe JR, Wright SI (2018) Population genomics of herbicide resistance: adaptation via evolutionary rescue. Annu Rev Plant Biol 69:611–635. https://doi.org/10.1146/annurev-arplant-042817-040038
Kreiner JM, Giacomini DA, Bemm F, Waithaka B, Regalado J, Lanz C et al (2019) Multiple modes of convergent adaptation in the spread of glyphosate-resistant Amaranthus tuberculatus. Proc Natl Acad Sci 116:21076–21084. https://doi.org/10.1073/pnas.1900870116
Lanciano S, Carpentier M-C, Llauro C, Jobet E, Robakowska-Hyzorek D, Lasserre E et al (2017) Sequencing the extrachromosomal circular mobilome reveals retrotransposon activity in plants. PLOS Genet 13:e1006630. https://doi.org/10.1371/journal.pgen.1006630
Larson G, Piperno DR, Allaby RG, Purugganan MD, Andersson L, Arroyo-Kalin M et al (2014) Current perspectives and the future of domestication studies. Proc Natl Acad Sci 111:6139–6146. https://doi.org/10.1073/pnas.1323964111
LeClere S, Wu C, Westra P, Sammons RD (2018) Cross-resistance to dicamba, 2,4-D, and fluroxypyr in Kochia scoparia is endowed by a mutation in an AUX/IAA gene. Proc Natl Acad Sci 115:E2911–E2920. https://doi.org/10.1073/pnas.1712372115
Li L-F, Li Y-L, Jia Y, Caicedo AL, Olsen KM (2017) Signatures of adaptation in the weedy rice genome. Nat Genet 49:811–814. https://doi.org/10.1038/ng.3825
Liu B, Yan J, Li W, Yin L, Li P, Yu H et al (2020) Mikania micrantha genome provides insights into the molecular mechanism of rapid growth. Nat Commun 11:340. https://doi.org/10.1038/s41467-019-13926-4
Lu H, Xue L, Cheng J, Yang X, Xie H, Song X et al (2020) Polyploidization-driven differentiation of freezing tolerance in Solidago canadensis. Plant Cell Environ 43:1394–1403. https://doi.org/10.1111/pce.13745
Luo M-C, Gu YQ, Puiu D, Wang H, Twardziok SO, Deal KR et al (2017) Genome sequence of the progenitor of the wheat D genome Aegilops tauschii. Nature 551:498–502. https://doi.org/10.1038/nature24486
Mamidi S, Healey A, Huang P, Grimwood J, Jenkins J, Barry K et al (2020) A genome resource for green millet Setaria viridis enables discovery of agronomically valuable loci. Nat Biotechnol 38:1203–1210. https://doi.org/10.1038/s41587-020-0681-2
Molin WT, Yaguchi A, Blenner M, Saski CA (2020) The EccDNA replicon: a heritable, extranuclear vehicle that enables gene amplification and glyphosate resistance in Amaranthus palmeri. Plant Cell 32:2132–2140. https://doi.org/10.1105/tpc.20.00099
Montgomery JS, Giacomini D, Waithaka B, Lanz C, Murphy BP, Campe R et al (2020) Draft genomes of Amaranthus tuberculatus, Amaranthus hybridus, and Amaranthus palmeri. Genome Biol Evol 12:1988–1993. https://doi.org/10.1093/gbe/evaa177
Nandula VK, Wright AA, Bond JA, Ray JD, Eubank TW, Molin WT (2014) EPSPS amplification in glyphosate-resistant spiny amaranth (Amaranthus spinosus): a case of gene transfer via interspecific hybridization from glyphosate-resistant Palmer amaranth (Amaranthus palmeri). Pest Manag Sci 70:1902–1909. https://doi.org/10.1002/ps.3754
Neve P (2018) Gene drive systems: do they have a place in agricultural weed management? Pest Manag Sci 74:2671–2679. https://doi.org/10.1002/ps.5137
Neve P, Caicedo AL (2022) Weed adaptation as a driving force for weed persistence in agroecosystems. Persistence strategies of weeds. Wiley, US, pp 302–324. https://doi.org/10.1002/9781119525622.ch15
Nunn A, Rodríguez-Arévalo I, Tandukar Z, Frels K, Contreras-Garrido A, Carbonell-Bejerano P et al (2022) Chromosome-level Thlaspi arvense genome provides new tools for translational research and for a newly domesticated cash cover crop of the cooler climates. Plant Biotechnol J 20:944–963. https://doi.org/10.1111/pbi.13775
Oerke E-C (2006) Crop losses to pests. J Agric Sci 144:31–43. https://doi.org/10.1017/S0021859605005708
Paril J, Pandey G, Barnett E B, Rane RV, Court L, Walsh T et al (2022) Rounding up the annual ryegrass genome: high-quality reference genome of Lolium rigidum. https://doi.org/10.1101/2022.07.18.499821.
Patterson EL, Saski CA, Sloan DB, Tranel PJ, Westra P, Gaines TA (2019a) The draft genome of Kochia scoparia and the mechanism of glyphosate resistance via transposon-mediated EPSPS tandem gene duplication. Genome Biol Evol 11:2927–2940. https://doi.org/10.1093/gbe/evz198
Patterson EL, Saski C, Küpper A, Beffa R, Gaines TA (2019b) Omics potential in herbicide-resistant weed management. Plants 8:607. https://doi.org/10.3390/plants8120607
Peng Y, Lai Z, Lane T, Nageswara-Rao M, Okada M, Jasieniuk M et al (2014) De novo genome assembly of the economically important weed horseweed using integrated data from multiple sequencing platforms. Plant Physiol 166:1241–1254. https://doi.org/10.1104/pp.114.247668
Qiu J, Zhou Y, Mao L, Ye C, Wang W, Zhang J et al (2017) Genomic variation associated with local adaptation of weedy rice during de-domestication. Nat Commun 8:15323. https://doi.org/10.1038/ncomms15323
Qiu J, Jia L, Wu D, Weng X, Chen L, Sun J et al (2020) Diverse genetic mechanisms underlie worldwide convergent rice feralization. Genome Biol 21:70. https://doi.org/10.1186/s13059-020-01980-x
Ravet K, Patterson EL, Krähmer H, Hamouzová K, Fan L, Jasieniuk M et al (2018) The power and potential of genomics in weed biology and management. Pest Manag Sci 74:2216–2225. https://doi.org/10.1002/ps.5048
Shapter FM, Cross M, Ablett G, Malory S, Chivers IH, King GJ et al (2013) High-throughput sequencing and mutagenesis to accelerate the domestication of Microlaena stipoides as a new food crop. PLoS ONE 8:e82641. https://doi.org/10.1371/journal.pone.0082641
Sharma G, Barney JN, Westwood JH, Haak DC (2021) Into the weeds: new insights in plant stress. Trends Plant Sci 26:1050–1060. https://doi.org/10.1016/j.tplants.2021.06.003
Singh V, Etheredge L, McGinty J, Morgan G, Bagavathiannan M (2020) First case of glyphosate resistance in weedy sunflower (Helianthus annuus). Pest Manag Sci 76:3685–3692. https://doi.org/10.1002/ps.5917
Song J-M, Xie W-Z, Wang S, Guo Y-X, Koo D-H, Kudrna D et al (2021) Two gap-free reference genomes and a global view of the centromere architecture in rice. Mol Plant 14:1757–1767. https://doi.org/10.1016/j.molp.2021.06.018
Stein JC, Yu Y, Copetti D, Zwickl DJ, Zhang L, Zhang C et al (2018) Genomes of 13 domesticated and wild rice relatives highlight genetic conservation, turnover and innovation across the genus Oryza. Nat Genet 50:285–296. https://doi.org/10.1038/s41588-018-0040-0
Stewart CN (2017) Becoming weeds. Nat Genet 49:654–655. https://doi.org/10.1038/ng.3851
Sun G, Xu Y, Liu H, Sun T, Zhang J, Hettenhausen C et al (2018) Large-scale gene losses underlie the genome evolution of parasitic plant Cuscuta australis. Nat Commun 9:2683. https://doi.org/10.1038/s41467-018-04721-8
Sun J, Ma D, Tang L, Zhao M, Zhang G, Wang W et al (2019) Population genomic analysis and de novo assembly reveal the origin of weedy rice as an evolutionary game. Mol Plant 12:632–647. https://doi.org/10.1016/j.molp.2019.01.019
Sun Y, Shang L, Zhu Q-H, Fan L, Guo L (2022a) Twenty years of plant genome sequencing: achievements and challenges. Trends Plant Sci 27:391–401. https://doi.org/10.1016/j.tplants.2021.10.006
Sun Y, Shen E, Hu Y, Wu D, Feng Y, Lao S et al (2022b) Population genomic analysis reveals domestication of cultivated rye from weedy rye. Mol Plant 15:552–561. https://doi.org/10.1016/j.molp.2021.12.015
Sweeney MT, Thomson MJ, Pfeil BE, McCouch S (2006) Caught red-handed: Rc encodes a basic helix-loop-helix protein conditioning red pericarp in rice. Plant Cell 18:283–294. https://doi.org/10.1105/tpc.105.038430
te Beest M, Le Roux JJ, Richardson DM, Brysting AK, Suda J, Kubešová M et al (2012) The more the better? The role of polyploidy in facilitating plant invasions. Ann Bot 109:19–45. https://doi.org/10.1093/aob/mcr277
The Arabidopsis Genome Initiative (2000) Analysis of the genome sequence of the flowering plant Arabidopsis thaliana. Nature 408:796–815. https://doi.org/10.1038/35048692
Thielen PM, Pendleton AL, Player RA, Bowden KV, Lawton TJ, Wisecaver JH (2020) Reference genome for the highly transformable Setaria viridis ME034V. G3 GenesGenomesGenetics 10:3467–3478. https://doi.org/10.1534/g3.120.401345
Vigueira CC, Olsen KM, Caicedo AL (2013) The red queen in the corn: agricultural weeds as models of rapid adaptive evolution. Heredity 110:303–311. https://doi.org/10.1038/hdy.2012.104
Vogel A, Schwacke R, Denton AK, Usadel B, Hollmann J, Fischer K et al (2018) Footprints of parasitism in the genome of the parasitic flowering plant Cuscuta campestris. Nat Commun 9:2515. https://doi.org/10.1038/s41467-018-04344-z
Wang B, Yang X, Jia Y, Xu Y, Jia P, Dang N et al (2021a) High-quality Arabidopsis thaliana genome assembly with Nanopore and HiFi long reads. Genom Proteom Bioinform. https://doi.org/10.1016/j.gpb.2021.08.003
Wang L, Zhu T, Rodriguez JC, Deal KR, Dubcovsky J, McGuire PE et al (2021b) Aegilops tauschii genome assembly Aet v5.0 features greater sequence contiguity and improved annotation. G3 GenesGenomesGenetics 11:jkab325. https://doi.org/10.1093/g3journal/jkab325
Wang L, Sun X, Peng Y, Chen K, Wu S, Guo Y et al (2022) Genomic insights into the origin, adaptive evolution and herbicide resistance of Leptochloa chinensis, a devastating tetraploid weedy grass in rice fields. Mol Plant. https://doi.org/10.1016/j.molp.2022.05.001
Wedger MJ, Roma-Burgos N, Olsen KM (2022) Genomic revolution of US weedy rice in response to 21st century agricultural technologies. Commun Biol 5:1–9. https://doi.org/10.1038/s42003-022-03803-0
Wu D, Lao S, Fan L (2021) De-domestication: an extension of crop evolution. Trends Plant Sci 26:560–574. https://doi.org/10.1016/j.tplants.2021.02.003
Wu D, Jiang B, Ye C-Y, Timko MP, Fan L (2022a) Horizontal transfer and evolution of the biosynthetic gene cluster for benzoxazinoids in plants. Plant Commun 3:100320. https://doi.org/10.1016/j.xplc.2022.100320
Wu D, Shen E, Jiang B, Feng Y, Tang W, Lao S et al (2022b) Genomic insights into the evolution of Echinochloa species as weed and orphan crop. Nat Commun 13:689. https://doi.org/10.1038/s41467-022-28359-9
Xu Y, Zhang J, Ma C, Lei Y, Shen G, Jin J et al (2022) Comparative genomics of orobanchaceous species with different parasitic lifestyles reveals the origin and stepwise evolution of plant parasitism. Mol Plant 15:1384–1399. https://doi.org/10.1016/j.molp.2022.07.007
Ye C-Y, Fan L (2021) Orphan crops and their wild relatives in the genomic era. Mol Plant 14:27–39. https://doi.org/10.1016/j.molp.2020.12.013
Ye C-Y, Tang W, Wu D, Jia L, Qiu J, Chen M et al (2019) Genomic evidence of human selection on Vavilovian mimicry. Nat Ecol Evol 3:1474–1482. https://doi.org/10.1038/s41559-019-0976-1
Ye C-Y, Wu D, Mao L, Jia L, Qiu J, Lao S et al (2020) The genomes of the allohexaploid Echinochloa crus-galli and its progenitors provide insights into polyploidization-driven adaptation. Mol Plant 13:1298–1310. https://doi.org/10.1016/j.molp.2020.07.001
Yuan JS, Tranel PJ, Stewart CN (2007) Non-target-site herbicide resistance: a family business. Trends Plant Sci 12:6–13. https://doi.org/10.1016/j.tplants.2006.11.001
Zhang J, Zhang X, Tang H, Zhang Q, Hua X, Ma X et al (2018) Allele-defined genome of the autopolyploid sugarcane Saccharum spontaneum L. Nat Genet 50:1565–1573. https://doi.org/10.1038/s41588-018-0237-2
Zhang H, Hall N, Goertzen LR, Bi B, Chen CY, Peatman E et al (2019) Development of a goosegrass (Eleusine indica) draft genome and application to weed science research. Pest Manag Sci 75:2776–2784. https://doi.org/10.1002/ps.5389
Zhang X, Liu T, Wang J, Wang P, Qiu Y, Zhao W et al (2021) Pan-genome of Raphanus highlights genetic variation and introgression among domesticated, wild, and weedy radishes. Mol Plant 14:2032–2055. https://doi.org/10.1016/j.molp.2021.08.005
Zhao G, Zou C, Li K, Wang K, Li T, Gao L et al (2017) The Aegilops tauschii genome reveals multiple impacts of transposons. Nat Plants 3:946–955. https://doi.org/10.1038/s41477-017-0067-8
Zhou Y, Bai S, Li H, Sun G, Zhang D, Ma F et al (2021) Introgressing the Aegilops tauschii genome into wheat as a basis for cereal improvement. Nat Plants 7:774–786. https://doi.org/10.1038/s41477-021-00934-w
Acknowledgements
This work is supported by National Natural Science Foundation of China (31971865) to LF.
Author information
Authors and Affiliations
Contributions
LF and LB designed the review. YH performed the investigation and wrote the first manuscript. AMJ, DW, ZH, XL, LB and LF revised the manuscript, which all authors edited and approved.
Corresponding authors
Ethics declarations
Conflict of interest
The authors declare no competing interest.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Huang, Y., Wu, D., Huang, Z. et al. Weed genomics: yielding insights into the genetics of weedy traits for crop improvement. aBIOTECH 4, 20–30 (2023). https://doi.org/10.1007/s42994-022-00090-5
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s42994-022-00090-5