Abstract
Myotis emarginatus seems fond of spiders and flies, a unique feeding style among European bats. The importance of each prey type varies among studies, so this paper aims to expand on the knowledge to unveil the trophic niche of M. emarginatus by studying its diet changes throughout the maternity season. We sampled five maternity colonies in the Basque Country every 2 weeks for the whole duration of the maternity season and studied their diet using DNA metabarcoding methods. We observed significant changes in diet diversity and composition. At the beginning of the season, M. emarginatus consumed a variety of prey orders, Diptera being the most abundant; but as the season progressed, the relative consumption of Araneae increased. By August, 80% of the weighted percentage of occurrences (wPOO) corresponded to the orb-web-building spider Araneus diadematus. Orb-web-building spiders need habitats of high vertical complexity to grow in abundance. Therefore, conserving them is essential to preserve attractive hunting grounds for colonies of M. emarginatus. Besides, diurnal flies, mainly Stomoxys calcitrans, were also considerably consumed in the colonies studied, which can become detrimental to the well-being of cattle, and can act as a vector of many diseases.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Since the beginning of the twenty-first century, the diet of insectivorous bats has been widely studied, especially in Holarctic regions (Taweesub et al. 2021). This is undoubtedly linked to the rise in popularity of DNA-Metabarcoding and High Throughput Sequencing techniques, which offer an easy and cost-effective way to comprehensively analyse the diet of bat individuals and colonies (Pompanon et al. 2012). Although limitations of molecular techniques are still under discussion (Alberdi et al. 2018; Littleford-Colquhoun et al. 2022), they have provided valuable insight into hidden aspects of the foraging ecology of bats (Andriollo et al. 2021; Arrizabalaga-Escudero et al. 2019) and the ecosystem services they provide (Baroja et al. 2019; Garin et al. 2019; Taylor et al. 2018). The basic knowledge elicited by dietary studies has also paved the way to understand ecological processes beyond mere prey consumption (Alberdi et al. 2012; Arrizabalaga-Escudero et al. 2015) serving as a valuable tool for the successful management of human and natural resources (Cuff et al. 2022).
Both molecular and previous morphologic studies have found that the dietary range of bats can be highly diverse and variable. Each species is constrained by morpho-ecological characteristics (Emrich et al. 2014), which limit habitat use (Starik et al. 2021) and prey detectability (Siemers and Swift 2006). Nevertheless, the diet of each bat species has been reported to vary according to intrinsic characteristics of individuals like sex (Mata et al. 2016) or age (Arrizabalaga-Escudero et al. 2019), as well as factors affecting prey availability, such as seasonality, weather, landscape, or the presence of competing species (e.g. Goiti et al. 2008; Napal et al. 2013; Novella-Fernandez et al. 2020; Razgour et al. 2011; Tournayre et al. 2020a; but see Wray et al. 2021).
This paper aims to provide insight into the diet of the Notch-Eared bat (Myotis emarginatus) in a temperate-oceanic region throughout the whole duration of the maternity season. Previous studies on the diet of M. emarginatus have shown unique characteristics among European bat species because of the high prevalence of orb-weaving spiders (Araneidae) and diurnal flies (Muscidae) (Bauerova 1986; Beck 1995; Goiti et al. 2011; Kervyn et al. 2012; Krull et al. 1991; Vallejo et al. 2019). While spiders are common prey for bats of the gleaning guild (e.g. Andreas et al. 2012a; Roswag et al. 2018), only three species have been described as spider specialists among chiropterans: Kerivoula papuensis (Schulz 2000), Myotis evotis (Burles et al. 2008) and Myotis emarginatus (Goiti et al. 2011); but see Novella-Fernandez et al. (2020). Spiders are the primary food source for these bat species, who consume them consistently. Molecular analyses have identified 45 different spider species in the M. emarginatus faeces in the Iberian Peninsula, and vertical orb-web weaving spiders of the family Araneidae are the most common spider prey item in all the studied locations (Vallejo et al. 2019). On the other hand, the consumption of the blood-sucking diurnal cattle-fly Stomoxys calcitrans is especially noteworthy. This fondness for flies has been reported to account for over half of M. emarginatus’ diet in Central Europe (Beck 1995; Kervyn et al. 2012; Steck and Brinkmann 2006) and some colonies in the Iberian Peninsula (Vallejo et al. 2019). It seems to be linked to the presence of cattle sheds in the foraging grounds, where M. emarginatus has commonly been reported to forage (Dekker et al. 2013; Dietz and Pir 2021; Krull et al. 1991).
Generalist predators, like insectivorous bats, shift from selective to opportunistic behaviour depending on the abundance of their preferred prey (Emlen 1966). For example, the Mediterranean horseshoe bat (Rhinolophus euryale) consumes a diverse diet in spring. However, it shifts to feed almost exclusively on moths during the summer, when flying insects reach their yearly peak (Hails 1982). This prey item fits better into the detection and hunting features of the bat (Goiti et al. 2008). Similarly, Myotis myotis consumes more ground dwelling spiders during the summer, when the overall abundance of its preferred prey, carabids and crickets, drops (Ramos-Pereira et al. 2002).
Kervyn et al. (2012) found a higher proportion of moths in the diet of Myotis emarginatus studied in May than in the rest of the breeding season but found no significant changes in diet composition otherwise. Vallejo et al. (2019) suggested the importance of spiders decreases from spring to summer, in favour of cattle flies and other occasional prey. Nonetheless, the understanding of how the diet of M. emarginatus can change seasonally is still scarce.
In this research, we aim to describe temporal changes in the diet of M. emarginatus in five maternity colonies in the Basque Country (Northern Iberian Peninsula). To do so, we investigated the changes in diet composition and diversity of this bat species every fortnight in maternity colonies. The diet of the notch-eared bat has previously been studied in the region by Vallejo et al. (2019) and Goiti (unpublished data) and found that both spiders and stable flies were abundant, the latter, especially during the summer months. Based on these results, we expected the diet composition and diversity to vary seasonally and precisely predict that (1) the diet would be more diverse in spring than in summer, (2) the proportion of spiders consumed will be high overall, and (3) the consumption of diurnal flies would be highest during the summer months. A decrease in the dietary diversity during the summer months, and an increase in the proportion of cattle flies consumed would indicate that cattle flies are an essential food source for M. emarginatus in this area, as it focuses on them in times of higher overall resource abundance, while it maintains a more diverse diet in times of lower resource abundance (Emlen 1966).
Materials and methods
Sample collection
The five maternity colonies studied are within a 27 km radius (the Basque Country, Southwestern Europe) and all are roosting in artificial constructions. The climate in the region is temperate oceanic, with mild temperatures throughout the year (mean: 14ºC) and abundant precipitation (1200–2000 mm). The landscape around all colonies is highly modified by human activity, as pine and eucalyptus plantations are common in the region.
Baranbio (BA)
A farm house, where around 100 M. emarginatus share the roost with 40 Rhinolophus ferrumequinum. The colony is surrounded by coniferous and eucalyptus plantations on a 5 km radius, but further away (up to 10 km) broad-leaved forests, pastures, herbaceous crops and scrubland are available.
Ereño (ER)
An empty building in an abandoned quarry gives roost to up to 50 M. emarginatus. Around 15 R. ferrumequinum and occasionally some Rhinolophus euryale are also present. It is surrounded by a diverse landscape composed of broad-leaved forests (including evergreen oak forests), pine and eucalyptus plantations, and wetland.
Lauro (LA)
Around 20 M. emarginatus roost in an abandoned unfinished two-storey construction. The area is surrounded by agricultural land of various types and a few patches of broad-leaved forests, eucalyptus plantations, pastures and scrublands.
Mañaria (MA)
A church vault where up to 200 M. emarginatus share the space with R. ferrumequinum. Mostly broad-leaved forests and pine plantations surround the area.
Zestoa (ZE)
The biggest colony of M. emarginatus in the region, with more than 500 individuals, sharing the roost with around 100 R. ferrumequinum and 30 Miniopterus schreibersii in an empty building, right next to a river bed. Pastures, meadows, broad-leaved forests and a few pine plantations surround the area.
We sampled these five colonies in the year 2020. Bats at ZE arrived the earliest (May 4th, Week 19), and left the latest (September 3rd, Week 36). On the other hand, bats from LA spent the shortest time in the roost, arriving the latest (May 29th, Week 22) and leaving the earliest (August 5th, Week 32).
We took special care to reduce disturbance to the colonies. Every fortnight, a collector was placed under the bats and faeces were collected after no more than 2 days to minimize DNA degradation. We collected up to 20 samples of 4–6 pellets each. In the cases where the risk of contamination from the faeces of non-target bat species was high, we collected up to 40 samples containing one or two pellets each. Sampling was always conducted in even weeks. Exceptionally, four collections were taken in the next week, although they were analysed as if they were collected in the week before: ZE in week 18, ER in week 20, LA in week 28 and BA in week 30.
Samples from each colony and sampling date were processed and sequenced to describe 75% of the diet diversity per colony and day (Chao et al. 2014). Unfortunately, we could only reach such sequencing depth in some of the colonies and dates due to budgetary reasons. We performed inter- and extrapolation of dietary richness using package iNEXT (Hsieh et al. 2020). In total, 595 samples were sequenced. One extraction blank was added every 23 samples, and one library blank was included in every MiSeq Run performed.
DNA extraction, PCR amplification and sequencing
The faeces were weighted, and the DNA was extracted using DNeasy PowerSoil Kit and DNeasy PowerSoil Pro Kit (Qiagen) following manufacturer’s instructions with some modifications. For the amplification process, we used primer set FWH1 (Vamos et al. 2017), which targets a 180 bp region of the COI gene. This primer set was chosen because: first, it amplifies a broad range of prey taxa, and also amplifies bat DNA (Tournayre et al. 2020b) which is helpful to detect contamination from non-target bat species in our samples; and second, it is longer than other popular primer sets used in bat dietary studies (e.g. Gillet et al. 2015; Zeale et al. 2011), which reduces the chance to amplify DNA from unwanted sources. Shorter markers can capture greater diversity (Elbrecht et al. 2019; Tournayre et al. 2020b) but are also prone to amplify very degraded and unwanted DNA coming, for example, from secondary predation (Galan et al. 2018). Usually this is not concerning, but given the fact that spiders are expected to be abundant in the diet of M. emarginatus, the probability of detecting DNA from secondary predation increases, and it would hamper its discrimination from positive data.
PCR amplification was performed following Tournayre et al. (2020b), with modifications. Libraries were built using Illumina’s Nextera XT kit, and samples were sequenced in Illumina MiSeq. PCR amplification, DNA library construction and sequencing processes were done at the Genomics and Proteomics General Service (SGIker) of the University of the Basque Country (UPV/EHU).
Details on all laboratory procedures can be found in Online Resource 1.
Sequence analysis and taxonomic assignment
Bioinformatic analyses were done using vsearch (Rognes et al. 2016) and Cutadapt (Martin 2011), following the steps described in Esnaola et al. (2018) with the necessary modifications. Details are explained in Online Resource 1.
The 595 samples (Online Resource 2) analysed yielded 7851 OTUs (Online Resource 3) which were assigned to their appropriate taxonomy using the blastn function in BLAST + (Camacho et al. 2009) to access the GenBank dataset, and Boldigger-cline (Buchner and Leese 2020) to access the BOLD dataset. Only matches over 98% similarity were considered. The output of both datasets was curated manually and using a custom script so that each OTU was only assigned to a single taxon. Each assigned taxon was classified into one of the following categories: predator, prey, not-in-study-area, environmental contamination and unassigned.
Selection of samples and OTUs
Samples were checked for contamination from potential prey species by studying the sample blanks. We found no clear contamination event from species that could be mistaken for prey. Nonetheless, an abundance threshold was applied to remove taxa with low read abundances, so that OTUs with less than 0.5% of reads were removed in each sample. While far from perfect, this method has proven to be quite effective in limiting contamination risk of multiple sources without eliminating too many rare prey taxa (Drake et al. 2022); therefore, the data were interpreted having this decision in mind.
We also checked for contamination from co-occurring bat species. Samples were discarded from the analysis if more than 10% of all the reads identified as bat species belonged to others than M. emarginatus. We obtained at least 8–12 samples eligible for analysis in most situations. Nevertheless, we removed samples from MA in weeks 20, 32 and 34, and of ZE in week 36 groups entirely from the analysis, because we did not manage to obtain more than one viable sample for the analyses. More information on the selection of samples is given in Online Resource 1.
Description of dietary metrics
All data analysis was performed in R version 4.0.4 (R Core Team 2021). Only those OTUs classified as potential prey were used for the diet analysis. Due to the lack of proven correlation between relative read abundances (RRA) and biomass of prey consumed (Paula et al. 2022), we converted our data to weighted percentages of occurrences (wPOO), a metric based on presence/absence data that provides a good proxy of consumption (Cuff et al. 2022; Deagle et al. 2018). We calculated wPOO values at the prey species, genus, family and order levels.
Multivariate analysis of diet variability and homogeneity
We explored diet composition and variability at the species, genus, family and order level of prey items. We calculated Bray–Curtis dissimilarities between all samples using function vegdist in package vegan (Oksanen et al. 2020). We explored relationships between diet composition and Week, Colony and their interaction through a Permutational Multivariate Analysis of Variance, or PERMANOVA (Anderson 2001), using function adonis2 in package vegan.
We calculated distances of samples to the centroid of their respective group in non-Euclidean space using function betadisper in package vegan to test for homogeneity of multivariate dispersion (Anderson and Walsh 2013). We performed the analysis by defining groups of samples by the combination of Week and Colony. We used the results of these analyses to explore changes in diet variability, that is, beta diversity (Anderson et al. 2006). To avoid the creation of negative eigenvalues in the process, we calculated square root transformed distances, instead of standard ones.
The distances calculated by betadisper were extracted and modelled against factors Week, Colony and their interaction in a conventional linear regression, to explore differences in diet variability. We visualized the marginal effects of each factor in the model and their interaction using package margins (Leeper 2021). In addition, we represented each group's centroid position calculated by betadisper in a PCoA to analyse underlying patterns in the overall composition of the sample groups and their relationship with each other.
Analysis of seasonal diet composition
Finally, the effect of sampling date on the diet composition was analysed at the ordinal level using a Multinomial Logit model, which is used to model discrete choices between mutually exclusive alternatives (Croissant 2017). They have been recently used to model changes in diet composition in bats (Tiede et al. 2020) and they can theoretically define the probability that an alternative—a prey item in our case—is chosen (see Croissant 2017). To run the model, we collapsed all orders found in less than 15 samples in a new category called “Others”, resulting in six different prey orders.
We chose to use package brms (Bürkner 2017), which uses Markov chain Monte Carlo (MCMC) algorithms to fit various multilevel models, including multinomial logit models, under a Bayesian framework. We chose the category “Others” as a baseline for the model. We calculated the estimated probabilities of consumption for each order every week and the marginal effect of the week on the consumption of each order, without the effect of the baseline category.
Results
Selection of samples and OTUs
After removing non-eligible samples (Online Resource 2), 359 samples were left for the diet analysis, which yielded a total of 6314 OTUs, of which only 664 had an abundance higher than 0.5% in any of the samples (Online Resource 3). Seven OTUs (accounting for 50% of total reads) belonged to the predator, M. emarginatus; 73 (4% of reads) were classified as environmental contamination; 251 (24% of reads) were identified as potential prey items; four belonged to potential prey species that are not found in the study area (0.5% of reads); and the remaining 329 (21% of reads) did not match with any sequence in the databases at least at the 98% identity level. In total, 155 unique prey species and 21 unique genera were identified. In addition, seven genera also had OTUs identified both at the genus and the species level. In these cases, a genus level occurrence was added to the dietary data. All in all, we analysed 183 prey taxa, identified at the genus or species levels, belonging to 92 families and 14 orders.
The most frequent prey species was the orb-weaving spider Araneus diadematus. It was identified in 249 samples (Frequency of occurrence (FOO) 69%) and accounted for 42% RRA and 33.9% wPOO. The next two most frequent prey items were Stomoxys calcitrans and Musca domestica, identified in 81 and 68 samples, with 9.8% and 6.6% RRA and 7.8% and 6.7% wPOO, respectively. Most prey items appeared occasionally in faeces: 108 prey species were only identified in a single sample each, 22 were identified in two samples, and 37 in between three and ten samples. Therefore, only 16 species occurred in more than ten samples out of 359 (Table 1). Half of them were orb-web weaving spiders.
Araneae and Diptera were the two most consumed prey orders, accounting for 67% (Weeks 18–20) to 88% (Weeks 34–36) of the total wPOO. During May, the consumption of other orders is noteworthy: Caenis luctuosa (Ephemeroptera) had the highest wPOO (37%) in ZE Week 18; Pharmacis fusconebulosa (Lepidoptera) was also the highest consumed prey item (wPOO: 27%) in BA Week 20; Melolontha melolontha (Coleoptera) was the third most consumed prey item (wPOO 13%) in ER Week 20.
Multivariate analysis of diet composition and variability
PERMANOVA showed significant difference in diet composition across Week, Colony and the interaction of both factors at all taxonomical levels (α = 0.05). Figure 1 shows a PCoA representing the centroid position of every group of samples in a non-Euclidean space, as calculated by betadisper. The first two PCoA axes represent 53% of the total variability at the order level, but a lower percentage at other taxonomic levels. The first axis separates groups of samples similarly regardless of the taxonomic level: it separates samples from the first four sampling weeks (18–24) and the last four sampling weeks (30–36). Mid-season samples appear all along the first PCoA axis, closer to early or late season samples depending on Colony (Fig. 1). The second PCoA axis separates certain colonies more and shows different patterns depending on the taxonomical rank used.betadisper also showed significant differences in the diet variability of Colonies, Week, and a combination of both factors at all taxonomical levels (α = 0.05). The distances to centroid calculated by betadisper were negatively correlated to Week at all taxonomical levels (Order: F1 = 62.161, p < 0.001—Family: F1 = 86.5237, p < 0.001—Genus: F1 = 90.024, p < 0.001—Species: F1 = 71.262, p < 0.001, Fig. 2a), meaning that diet variability decreased with Week. Distances to centroid were also significantly different between colonies at all taxonomical levels (Order: F4 = 14.864, p < 0.001—Family: F4 = 25.277, p < 0.001—Genus: F4 = 31.367, p < 0.001—Species: F4 = 31.000, p < 0.001). Colonies LA and ER had significantly lower average distances to the centroid (Fig. 2b). An interaction between Week and Colony showed that the temporal pattern might vary in different ways across Colonies (Order: F4 = 5.647, p < 0.001—Family: F4 = 7.429, p < 0.001—Genus: F4 = 11.335, p < 0.001—Species: F4 = 6.967, p < 0.001). ER and BA presented a sharper decrease in diet variability as the season progressed (Fig. 2c, d), compared to the rest of the colonies (Fig. 2e–g).
PCoA ordination of the position of the centroids of groups of samples collected in the same day and Colony, with prey items classified to the a Order level, b Family level, c Genus level and d Species level. The proportion of the variability explained by each of the axes is indicated in the axis labels
Marginal effects of Predicted Average Distance to Centroid (PADG) and 95% confidence intervals calculated by the linear model of distances to group centroids (at prey species level) against Week, Colony and their interaction. Plot a shows the marginal effect of Week PADG without the effect of Colony, plot b shows PADG per colony, without the effect of seasonality, and plots c–g show the colony PADG: c BA, d ER, e LA, f MA, g ZE. The effects for the models at different taxonomical levels are similar
Seasonal changes in diet composition
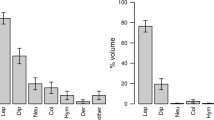
Given that most variability happened through time (Fig. 1), we modelled the changes in the proportion of six prey orders simultaneously in relation to Week using Multinomial logit models (Fig. 3). The proportion of Araneae increased significantly with Week, compared to the baseline (95% CI 9–12%). The frequency of Araneae was estimated to increase by 3.1% every week (95% CI 2.9–3.1%). On the other hand, Diptera, Lepidoptera and Ephemeroptera, and the baseline Others decreased significantly, albeit the estimated decrease for Diptera is the highest, at 1.9% points every week (%95 CI Diptera − 2 to − 1.8%, Lepidoptera − 0.7 to − 0.6%, Ephemeroptera − 0.3 to − 0.2%, Others − 0.13 to − 0.06%).
Overall, the model estimated that the consumption of Araneae would increase from a wPOO value of 28% (95% CI 10–49%) at Week 18, to a wPOO of 84% at Week 36 (95% CI 63–93%). Similarly, Diptera decreased from 46% (95% CI 19–74%) to 9% (95% CI 2.6%–27%) (Fig. 3). The confidence intervals of all orders overlapped at the beginning of the season. While Diptera showed the highest wPOO then, we cannot say that it was consumed significantly more than any other order. That situation quickly changed, as the consumption of Araneae became increasingly common. It reached up to 80% wPOO at the end of the season. As its confidence intervals did not overlap with the rest of the orders, Araneae was consumed significantly more than the rest of the orders at the end of the summer.
Discussion
Seasonal changes in diet variability and composition
Our results showed that both the variability and the composition of the diet of Myotis emarginatus changed throughout the season in the five colonies studied at multiple taxonomical levels. At the begging of the season, Diptera was the most consumed prey order. However, they were not consumed significantly more than any other order (Fig. 3). As the season progressed, the relative importance of Araneae in the diet gradually increased, and by the end of the season, spiders, particularly Araneus diadematus, constituted the main prey item of Myotis emarginatus in the colonies studied. This suggests that orb-web building spiders are the preferred prey for Myotis emarginatus in this area. Additionally, some prey items were also consumed only locally, and in abundance, during the first weeks of the bat’s stay in the breeding roosts, like C. luctuosa (Ephemeroptera) in ZE, P. fusconebulosa (Lepidoptera) in BA or M. melolontha (Coleoptera) in ER. Kervyn et al. (2012) also reported higher proportions of Lepidoptera in the diet of M. emarginatus at the beginning of the season. Colony ZE is located next to a river, which might explain the occasional high consumption of C. luctuosa.
As we predicted, the diet of M. emarginatus was more diverse at the beginning of the season, which may be a response to overall lower availability of prey, as both flying insects and spiders show a peak of abundance in late summer (Hails 1982; Hsieh and Linsenmair 2012). In the study region, the abundance of the stable fly Stomoxys calcitrans peaks during late summer-early autumn (Valbuena-Lacarra and Saloña-Bordas 2010). The consumption of a more diverse diet at times of a priori lower resource availability has also been reported in other bat species (Andreas et al. 2012b; Presetnik and Aulagnier 2013, see Introduction). However, this is not the norm, as some studies show that bats opportunistically feed on available resources, and thus the diversity of their diet mirrors that of available prey (Divoll et al. 2022; Tiede et al. 2020).
Being highly mobile, bats can respond quickly to changes in resource availability, such as those created by pest species of patchy distribution and short flying stages (Aizpurua et al. 2018; Baroja et al. 2021). This behaviour underlines insectivorous bats' plasticity and capacity to exploit various resources (Faure and Barklay 1994; Tournayre et al. 2020a).
Diet diversity did not change equally in all colonies studied; it decreased noticeably (Fig. 2c, d) in BA and ER, but not in the rest. Since the time series for colonies MA and LA were shorter, we might have been prevented from observing a significant decrease in diversity in these colonies too. Additionally, local differences in the arthropod communities could result in the observed differences regarding diet variability and composition. Finally, we did not measure local arthropod abundance and diversity in this study, which would be essential to fully understand whether and how M. emarginatus, and other insectivorous bats, adjust their foraging ecology and responds to changes in prey availability.
Consumption of spiders
In the 5 colonies studied here, out of the 16 prey species detected at least in 3% of the samples, 8 were orb-web building spiders of families Araneidae, Tetragnathidae and Uloboridae. Given how unusually other European bats prey on orb-weaving spiders (Whitaker 1988), such a consumption during the summer may alleviate the interspecific competition pressure. Bats can alter their foraging behaviour in the presence of competing species to avoid competition (e.g. Andriollo et al. 2021; Novella-Fernandez et al. 2020; Salsamendi et al. 2012; reviewed in Salinas-Ramos et al. 2019). Spiders have been rarely regarded as bat prey until recently.
In Europe, several different bat species can species consume spiders. Myotis nattereri sensu latto consumes them, as well as diurnal flies, during the autumn (Andreas et al. 2012a; Siemers and Swift 2006) and winter months (Hope et al. 2014). In the Iberian Peninsula, their sister species M. crypticus and M. escalerai also prey consistently on spiders (Novella-Fernandez et al. 2020). Spider remains have also been found in the faeces of Myotis bechsteinii and Plecotus auritus, for example (Andreas et al. 2012a). Additionally, with the use of multiple markers (Aldasoro et al. 2019), or generalist markers (Tournayre et al. 2020a) for DNA amplification, spiders are expected to appear as an occasional or regular prey item for other insectivorous bats as well. In fact, spiders have been recently reported to be an important food source for other bats of the genus Myotis in Europe and North America using molecular techniques for dietary studies, all of which forage in forests and catch their prey by gleaning (Hope et al. 2014; Maucieri and Barklay 2021; O’Rourke et al. 2021; Roswag et al. 2018).
In accordance with previous work on M. emarginatus’ diet near the study area (Vallejo et al. 2019), we predicted that the consumption of spiders would be highest in May. However, the results presented in this paper show the opposite trend. Intrinsic differences between the colonies sampled could explain the different responses detected. For example, one of the colonies from the previous study is in an area where cattle farming the most abundant in the Basque Country. Therefore, alternate food sources like diurnal flies are likely to be much more abundant there than in the colonies from this study. On the other hand, for the present study, we significantly improved the sampling effort to focus on the seasonality of the diet, highlighting the importance of an adequate study design in the results, and their correct interpretation.
In the present study, A. diadematus was the most common prey species, and it was detected in 69% of the samples. It was mostly consumed in the late season, and in some weeks, it accounted for over half the wPOO of the total diet. Contrastingly, it was rarely consumed during May (wPOO %1.6). At this time, Nuctenea umbratica and Metellina merianae, other orb-web building spiders, were the most consumed prey species. Many orb-web building species show an increase in their abundance during late summer months or autumn, while others mature in the spring (Hsieh and Linsenmair 2012). A. diadematus, specifically, reaches maturity during late summer-early autumn in temperate zones, and at this time they would be most available to bats like M. emarginatus (Olive 1981). Given their similarity in morphology and lifestyle, it would be interesting to study whether all orb-web building spiders are functionally indistinguishable by M. emarginatus besides perhaps size (Arrizabalaga-Escudero et al. 2019; Divoll et al. 2022; Spitz et al. 2014). Therefore, we hypothesize that changes in diet variability and composition within this functional group should mirror the local availability of resources.
The wing morphology and echolocation of M. emarginatus suggest that orb-web-building spiders would be gleaned over orb webs or other surfaces. Dense habitats, mainly native broad-leaved and mixed forests, are the preferred foraging habitat of M. emarginatus in some colonies (Dekker et al. 2013; Zahn et al. 2010). However, it can also use conifer plantations and mixed forests in some areas of its distribution (Flaquer et al. 2008; Goiti et al. 2011). Additionally, the use of open grasslands, riparian forests and especially cattle sheds has also been reported in the northernmost part of the distribution area (Dietz et al. 2013; Zahn et al. 2010). Accordingly, flies are abundant in the M. emarginatus diet in these areas (e. g. Krull et al. 1991), which shows that the geographic differences reported in diet composition most likely correlate with differences in habitat use for foraging (Dietz and Pir 2021).
The affinity of M. emarginatus to dense foraging habitats in its meridional distribution area is usually linked to the predation on orb-web-building spiders (Goiti et al. 2011). Cluttered habitats, especially those with a high degree of vertical complexity, are essential for web-building spider communities to thrive, because they offer plenty of web-building opportunities (Rubio and Moreno 2010; Ward and Lublin 1992). Additionally, diverse habitats also favour the diversity and abundance of spider prey (Dennis et al. 2015). Therefore, maintaining forests and other land patches with a complex vertical vegetation structure and a high degree of structural and vegetational heterogeneity is crucial to preserve attractive hunting grounds for colonies of M. emarginatus, and potentially other insectivorous bat species that, albeit to a lesser degree, also exploit this food resource.
Consumption of flies
The second and third most consumed prey species were Stomoxys calcitrans and Musca domestica (Muscidae). Together, they made 20% wPOO between the end of May and June (Weeks 22–26). Their importance in the diet decreased as the season progressed, and the consumption of A. diadematus increased. Flies are the most important food source for M. emarginatus in Central Europe (Kervyn et al. 2012; Steck and Brinkmann 2006). In contrast, studies from Mediterranean areas report lower proportions of flies and instead describe a spider-based diet (Goiti et al. 2011; Vallejo et al. 2019).
Both Stomoxys calcitrans and Musca domestica are synanthropic species, associated with cattle farms. Stomoxys calcitrans, especially, is frequently found around bovine farms and cattle farms (González 2022). Stomoxys calcitrans is considered a significant pest for livestock and domestic animals. Both sexes are hematophagous, and their bite causes physical distress and injuries to animals, particularly when their abundance increases, and can act as a vector of severe diseases and pathogens (González 2022; González et al. 2022; Patra et al. 2018). Nowadays, no specific control measures are taken against S. calcitrans by farmers, as the basic knowledge about this species is still scarce (González et al. 2022).
Besides, both are diurnal flies and remain active only in the warmest hours of the day (González 2022), so M. emarginatus should glean them directly from surfaces. In Central Europe, radio tracking data has confirmed that M. emarginatus frequently visits cattle sheds at night to forage (Dekker et al. 2013; Krull et al. 1991; Zahn et al. 2010). These structures are key foraging habitats for M. emarginatus, as they provide a reliable food source, especially on colder nights (Dekker et al. 2013). Krull et al. (1991) did not report such a correlation between weather and the use of cattle sheds as foraging grounds, however. Besides M. emarginatus, other bat species that use forests as their foraging areas have occasionally been reported hunting flies inside cattle sheds (Ancilloto et al. 2021; Siemers et al. 2012). Additionally, the activity of open foragers increases around free-ranging cattle (Ancilloto et al. 2021).
DNA metabarcoding has allowed us to confirm that the consumption of S. calcitrans by M. emarginatus is frequent in the Basque Country, especially at the beginning of the summer. Therefore, it is very likely that it also forages in cattle farms in the Basque Country. Nonetheless, specific studies are needed to confirm this fact in the area. More research is needed to explore under what conditions bats exploit diurnal flies in cattle farms, especially in the southern area of the distribution of M. emarginatus, further away from the limit of their distribution, where more examples of a spider-dominated diet have been reported (Goiti et al. 2011; Vallejo et al. 2019). On the other hand, DNA Metabarcoding does not offer accurate quantitative data (Deagle et al. 2018), so estimating the potential of M. emarginatus colonies for biological control is challenging. Methods targeted to identify a single prey species (e.g. Baroja et al. 2021) are an interesting option to overcome some of the pitfalls of DNA metabarcoding in this regard.
Data availability
All data generated or analysed during this study are included in this published article in supplementary files 2 and 3. Additional raw data will be available from the corresponding author on reasonable request.
References
Aizpurua O, Budinski I, Georgiakakis P, Gopalakrishnan S, Ibañez C, Mata V, Rebelo H, Russo D, Szodoray-Parádi F, Zhelyazkova V, Zrncic V, Gilbert MTP, Alberdi A (2018) Agriculture shapes the trophic niche of a bat preying on multiple pest arthropods across Europe: evidence form DNA metabarcoding. Mol Ecol 27:815–825. https://doi.org/10.1111/mec.14474
Alberdi A, Garin I, Aizpurua O, Aihartza J (2012) The foraging ecology of the Mountain Long-eared bat Plecotus macrobullaris revealed with DNA mini-barcodes. PLoS ONE 7(4):e35692. https://doi.org/10.1371/journal.pone.0035692
Alberdi A, Aizpurua O, Gilbert MTP, Bohmann K (2018) Scrutinizing key steps for reliable metabarcoding of environmental samples. Methods Ecol Evol 9:134–147. https://doi.org/10.1111/2041-210X.12849
Aldasoro M, Garin I, Vallejo N, Baroja U, Arrizabalaga-Escudero A, Goiti U, Aihartza J (2019) Gaining ecological insight on dietary allocation among horseshoe bats through molecular primer combination. PLoS ONE 14(7):e0220081. https://doi.org/10.1371/journal.pone.0220081
Ancilloto L, Festa F, De Benedetta F, Cosentino F, Pejic B, Russo D (2021) Free-ranging livestock and a diverse landscape structure increase bat foraging in mountainous landscapes. Agroforest Syst 95:407–418. https://doi.org/10.1007/s10457-021-00591-0
Anderson MJ (2001) A new method for non-parametric multivariate analysis of variance. Austral Ecol 26:32–46. https://doi.org/10.1111/j.1442-9993.2001.01070.pp.x
Anderson MJ, Walsh DCI (2013) PERMANOVA, ANOSIM, and the mantel test in the face of heterogeneous dispersion: what null hypothesis are you testing? Ecol Monog 83(4):557–574. https://doi.org/10.1890/12-2010.1
Anderson MJ, Ellingsen KE, McArdle BH (2006) Multivariate dispersion as a measure of beta diversity. Ecol Lett 9:683–693. https://doi.org/10.1111/j.1461-0248.2006.00926.x
Andreas M, Reiter A, Benda P (2012a) Dietary composition, resource partitioning and trophic niche overlap in three forest foliage-gleaning bats in Central Europe. Acta Chiropterologica 14(2):335–345. https://doi.org/10.3163/150811012X661657
Andreas M, Reiter A, Benda P (2012b) Prey selection and seasonal diet changes in the western barbastelle bat (Barbastella barbastellus). Acta Chiropterologica 14(1):81–92. https://doi.org/10.3161/150811012X654295
Andriollo T, Michaux JR, Ruedi M (2021) Food for everyone: differential feeding habits of cryptic bat species inferred from DNA metabarcoding. Mol Ecol 00:1–17. https://doi.org/10.1111/mec.16073
Arrizabalaga-Escudero A, Garin I, García-Mudarra JL, Alberdi A, Aihartza J, Goiti U (2015) Trophic requirements beyond foraging habitats: the importance of prey source habitats in bat conservation. Biol Conserv 191:512–519. https://doi.org/10.1016/j.biocon.2015.07.043
Arrizabalaga-Escudero A, Merckx T, García-Baquero G, Wahlberg N, Aizpurua O, Garin I, Goiti U, Aihartza J (2019) Trait-based functional dietary analysis provides a better insight into the foraging ecology of bats. J Anim Ecol 88(10):1587–1600. https://doi.org/10.1111/1365-2656.13055
Baroja U, Garin I, Aihartza J, Arrizabalaga-Escudero A, Vallejo N, Aldasoro M, Goiti U (2019) Pest consumption in a vineyard system by the lesser horseshoe bat (Rhinolophus hipposideros). PLoS ONE 14(7):e0219265. https://doi.org/10.1371/journal.pone.0219265
Baroja U, Garin I, Vallejo N, Aihartza J, Rebelo H, Goiti U (2021) Bats actively track and prey on grape pest populations. Ecol Indic 126:107718. https://doi.org/10.1016/j.ecolind.2021.107718
Bauerová Z (1986) Contribution to the trophic bionomics of Myotis emarginatus. Folia Zool 35(4):305–310
Beck A (1995) Fecal analyses of European bat species. Myotis 32(33):109–119
Buchner D, Leese F (2020) BOLDigger—a python package to identify and organise sequences with the barcode of life data systems. Metabarcoding Metagenom 4:e53535. https://doi.org/10.3897/mbmg.4.53535
Bürkner PC (2017) brms: an R package for bayesian multilevel models using stan. J Stat Softw 80(1):1–28. https://doi.org/10.18637/jss.v080.i01
Burles DW, Brigham RM, Ring RA, Reimchen TE (2008) Diet of two insectivorous bats, Myotis lucifugus and Myotis keenii, in relation to arthropod abundance in a temperate Pacific Northwest rainforest environment. Can J Zool 86(12):1367–1375. https://doi.org/10.1139/Z08-125
Camacho C, Coulouris G, Avagyan V, Ma N, Papadopoulos J, Bealer K, Madden TL (2009) BLAST+: architecture and applications. BMC Bioinform 10:421. https://doi.org/10.1186/1471-2105-10-421
Chao A, Gotelli NJ, Hsieh TC, Sander EL, Ma KH, Colwell RK, Ellison AM (2014) Rarefaction and extrapolation with hill numbers: a framework for sampling and estimation in species diversity studies. Ecol Monog 84(1):45–67. https://doi.org/10.1890/13-0133.1
Croissant Y (2017) Estimation of random utility models in R: the mlogit package. J Stat Softw 95(11):1–41. https://doi.org/10.18637/jss.v095.i11
Cuff JP, Windsor FM, Tercel MPTG, Kitson JJN, Evans DM (2022) Overcoming the pitfalls of merging dietary metabarcoding into ecological networks. Methods Ecol Evol 13:545–559. https://doi.org/10.1111/2041-210X.13796
Deagle BE, Thomas AC, McInnes JC, Clarke LJ, Vesterinen EJ, Clare EL, Kartzinel TR, Eveson JP (2018) Counting with DNA in metabarcoding studies: How should we convert sequence reads to dietary data? Mol Ecol 28:391–406. https://doi.org/10.1111/mec.14734
Dekker JJ, Regelink JR, Jansen EA, Brinkmann R, Limpens HJ (2013) Habitat use by Myotis emarginatus. Lutra 56(2):111–120
Dennis P, Skartveit J, Kunaver A, McCracken DI (2015) The response of spider (Araneae) assemblages to structural heterogeneity and prey abundance in sub-montane vegetation modified by conservation grazing. GECCO 107:1–14. https://doi.org/10.1016/j.gecco.2015.03.007
Dietz M, Pir JB (2021) Geoffroy’s Bat Myotis emarginatus (Geoffroy, 1806). In: Hackländer K, Zachos FE (eds) Handbook of the mammals of Europe. Springer Nature, Cham
Dietz M, Pir JB, Hillen J (2013) Does the survival of greater horseshoe bats and Geoffroy’s bats in Western Europe depend on traditional cultural landscapes? Biodivers Conserv 22:3007–3025
Divoll TJ, Brown VA, McCracken G, O’Keefe JM (2022) Prey size is more representative than prey taxa when measuring dietary overlap in sympatric forest bats. Environ DNA 00:1–13. https://doi.org/10.1002/edn3.354
Drake LE, Cuff JP, Young RE, Marchbank A, Chadwick EA, Symondson WOC (2022) An assessment of minimum sequence copy thresholds for identifying and reducing the prevalence of artefacts in dietary metabarcoding data. Methods Ecol Evol 13(3):694–710. https://doi.org/10.1111/2041-210X.13780
Elbrecht V, Braukmann TWA, Ivanova NV, Prosser SWJ, Hajibabaei M, Wright M, Evgeny VZ, Hebert PDN, Steinke D (2019) Validation of COI metabarcoding primers for terrestrial arthropods. PeerJ 7:e7745. https://doi.org/10.7717/peerj.7745
Emlen JM (1966) The role of time and energy in food preference. Am Nat 100(916):611–617
Emrich MA, Clare EL, Symondson WOC, Koenig SE, Fenton MB (2014) Resource partitioning by insectivorous bats in Jamaica. Mol Ecol 23(15):3648–3656. https://doi.org/10.1111/mec.12504
Esnaola A, Arrizabalaga-Escudero A, González-Esteban J, Elosegi A, Aihartza J (2018) Determining diet from faeces: selection of metabarcoding primers for the insectivore Pyrenean desman (Galemys pyrenaicus). PLoS ONE 13(12):e0208986. https://doi.org/10.1371/journal.pone.0208986
Faure PA, Barclay RMR (1994) Substrate-gleaning versus aerial-hawking: plasticity in the foraging and echolocation behaviour of the long-eared bat Myotis evotis. J Comp Physiol A 174(5):651.660. https://doi.org/10.1007/BF00217386
Flaquer C, Puig-Montserrat X, Burgas A, Russo D (2008) Habitat selection by Geoffroy’s bats (Myotis emarginatus) in a rural Mediterranean landscape: implications for conservation. Acta Chiropterologica 10(1):61–67. https://doi.org/10.3161/150811008X331090
Galan M, Pons JB, Tournayre O, Pierre E, Leuchtmann M, Pontier D, Charbonell N (2018) Metabarcoding for the parallel identification of several hundred predators and their prey: application to bat species diet analysis. Mol Ecol Resour 18:474–489. https://doi.org/10.1111/1755-0998.12749
Garin I, Aihartza J, Goiti U, Arrizabalaga-Escudero A, Nogueras J, Ibáñez C (2019) Bats from different foraging guilds prey upon the pine processionary moth. PeerJ 7:e7169. https://doi.org/10.7717/peerj.7169
Gillet F, Tiouchichine ML, Galan M, Blanc F, Némoz M, Aulagnier S, Michaux JR (2015) A new method to identify the endangered Pyrenean desman (Galemys pyrenaicus) and to study its diet, using next generation sequencing from faeces. Mamm Biol 80(6):505–509. https://doi.org/10.1016/j.mambio.2015.08.002
Goiti U, Garin I, Almenar D, Salsamendi E, Aihartza J (2008) Foraging by Mediterranean horseshoe bats (Rhinolophus euryale) in relation to prey distribution and edge habitat. J Mammal 89(2):493–502. https://doi.org/10.1644/07-MAMM-A-054R2.1
Goiti U, Aihartza J, Guiu M, Salsamendi E, Almenar D, Napal M, Garin I (2011) Geoffroy’s bat, Myotis emarginatus, preys preferentially on spiders in multistratified dense habitats: a study of foraging bats in the Mediterranean. Folia Zool 60(1):17–24. https://doi.org/10.25225/fozo.v60.i1.a3.2011
González MA (2022) Insectos de importancia Médico-Veterinaria en el Norte de España. Publicación Independiente. 240 pp
González MA, Bravo-Barriga D, Barrio-Fernández E, Frontera E, Ruiz-Arrondo I (2022) Severe Skin lesions caused by persistent bites of the stable fly Stomoxys calcitrans (Diptera: Muscidae) in a donkey sanctuary of Western Spain. J Equine Vet Sci 116:104056
Hails CJ (1982) A comparison of tropical and temperate aerial insect abundance. Biotropica 14(4):310–313. https://doi.org/10.2307/2388092
Hope PR, Bohmann K, Gilbert MTP, Zepeda-Mendoza ML, Razgour O, Jones G (2014) Second generation sequencing and morphological faecal analysis reveal unexpected foraging behaviour by Myotis nattereri (Chiroptera, Vespertilionidae) in winter. Front Zool 11:39. https://doi.org/10.1186/1742-9994-11-39
Hsieh YL, Linsenmair KE (2012) Seasonal dynamics of arboreal spider diversity in a temperate forest. Ecol Evol 2(4):768–777. https://doi.org/10.1002/ece3.221
Hsieh TC, Ma KH, Chao A (2020) iNEXT: iNterpolation and EXTrapolation for species diversity. R package version 2.0.20 URL: http://www.chao.stat.nthu.edu.tw/wordpress/software-download/. Accessed 8 June 2022
Kervyn T, Godin MC, Jocqué R, Grootaert P, Libois R (2012) Web-building spiders and blood-feeding flies as prey of the notch-eared bat (Myotis emarginatus). Belg J Zool 142(1):59–67
Krull D, Schumm A, Metzner W, Neuweiler G (1991) Foraging areas and foraging behavior in the notch- eared bat, Myotis emarginatus (Vespertilionidae). Behav Ecol Sociobiol 28:247–253
Leeper TJ (2021) margins: Marginal Effects for Model Objects. R package version 0.3.26
Littleford-Colquhoun BL, Freeman PT, Sackett VI, Tulloss CV, McGarvey LM, Geremia C, Kartzinel TR (2022) The precautionary principle and dietary DNA metabarcoding: commonly used abundance thresholds change ecological interpretation. Mol Ecol 00:1–12. https://doi.org/10.1111/mec.16352
Martin M (2011) Cutadapt removes adapter sequences from high-throughtput sequencing reads. Embnet J 17(1):10–12. https://doi.org/10.14806/ej.17.1.200
Mata VA, Amorim F, Corley MFV, McCracken GF, Rebelo H, Beja P (2016) Female dietary bias towards large migratory moths in the European free-tailed bat (Tadarida teniotis). Biol Lett 12(3):20150988. https://doi.org/10.1098/rsbl.2015.0988
Maucieri DG, Barclay RMR (2021) Consumption of spiders by the little brown bat (Myotis lucifugus) and the long-eared myotis (Myotis evotis) in the rocky mountains of Alberta. Canada Can J Zool 99(3):221–226. https://doi.org/10.1139/cjz-2020-0160
Napal M, Garin I, Goiti U, Salsamendi E, Aihartza J (2013) Past deforestation of Mediterranean Europe explains the present distribution of the strict forest dweller Myotis bechsteinii. For Ecol Manage 293:161–170. https://doi.org/10.1016/j.foreco.2012.12.038
Novella-Fernandez R, Ibañez C, Juste J, Clare EL, Doncaster CP, Razgour O (2020) Trophic resource partitioning drives fine-scale coexistence in cryptic bat species. Ecol Evol 10(24):14122–14136. https://doi.org/10.1002/ece3.7004
O’Rourke DR, Mangan MT, Mangan KE, Bokulich NA, MacManes MD, Foster JT (2021) Lord of the Diptera (and moths and a spider): molecular diet analyses and foraging ecology of Indiana Bats in Illinois. Front Ecol Evol 9:623655. https://doi.org/10.3389/fevo.2021.623655
Oksanen J, Guillaume B, Friendly M, Kindt R, Legendre P, McGlinn D, Minchin PR, O’Hara RB, Simpson GL, Solymos P, Stevens MHH, Szoecs E, Wagner H (2020) vegan: Community Ecology Package. R package version 2.5–7. https://www.CRAN.R-project.org/package=vegan. Accessed 20 Oct 2022
Olive CW (1981) Optimal phenology and body-size of orb-weaving spiders: foraging constraints. Oecologia 49:83–87. https://doi.org/10.1007/BF00376902
Patra G, Behera P, Das SK, Saikia B, Ghosh S, Biswas P, Kumar A, Alam SS, Kawlni L, Lalnunpuia C, Lalchhandama C, Bacham M, Debbarma A (2018) Stomoxys calcitrans and its importance in livestock: a review. Int J Adv Agric Res 6:30–37
Paula DP, Barros SKA, Pitta RM, Barreto MR, Togawa RC, Andow DA (2022) Metabarcoding versus mapping unassembled shotgun reads for identification of prey consumed by arthropod epigeal predators. GigaScience 11(1):1–13. https://doi.org/10.1093/gigascience/giac020
Pompanon F, Deagle BE, Symondson WOC, Brown DS, Jarman SN, Taberlet P (2012) Who is eating what: Diet assessment using next generation sequencing. Mol Ecol 21(8):1931–1950. https://doi.org/10.1111/j.1365-294X.2011.05403.x
Presetnik P, Aulagnier S (2013) The diet of Schrieber’s bent-winged bat, Miniopterus schreibersii (Chiroptera: Miniopteridae), in northeastern Slovenia (Central Europe). Mammalia 77(3):297–305. https://doi.org/10.1515/mammalia-2012-0033
R Core Team (2021) R: A language and environment for statistical computing. R Foundation for Statistical Computing, Vienna, Austria. https://www.R-project.org/. Accessed 20 Oct 2022
Ramos-Pereira MJ, Rebelo H, Rainho A, Palmereim JM (2002) Prey selection by Myotis myotis (Vespertilionidae) in a Mediterranean region. Acta Chiropterologica 4(2):183–193. https://doi.org/10.3161/001.004.0207
Razgour O, Clare EL, Zeale MRK, Hanmer J, Schnell IB, Rasmussen M, Gilbert TP, Jones G (2011) High-throughput sequencing offers insight into mechanisms of resource partitioning in cryptic bat species. Ecol Evol 1(4):556–570. https://doi.org/10.1002/ece3.49
Rognes T, Flouri T, Nichols B, Quince C, Mahé F (2016) VSEARCH: a versatile open source tool for metagenomics. PeerJ 4:e2584. https://doi.org/10.7717/peerj.2584
Roswag A, Becker NI, Encarnação JA (2018) Isotopic and dietary niches as indicators for resource partitioning in the gleaner bats Myotis bechsteinii, M. nattereri, and Plecotus auritus. Mammal Biol 89:62–70. https://doi.org/10.1016/j.mambio.2017.12.006
Rubio GD, Moreno CE (2010) Orb-weaving spider diversity in the Iberá Marshlands Argentina. Neotrop Entomol 39(4):496–505. https://doi.org/10.1590/S1519-566X2010000400006
Salinas-Ramos VB, Ancillotto L, Bosso L, Sanchez-Cordero V, Russo D (2019) Interspecific competition in bats: state of knowledge and research challenges. Mamm Rev 50:68–81. https://doi.org/10.1111/mam.12180
Salsamendi E, Garin I, Arostegui I, Goiti U, Aihartza J (2012) What mechanism of niche segregation allows the coexistence of sympatric sibling rhinolophid bats? Front Zool 9:30. https://doi.org/10.1186/1742-9994-9-30
Schulz M (2000) Diet and foraging behavior of the golden-tipped bat, Kerivoula papuensis: a spider specialist? J Mammal 81(4):948–957. https://doi.org/10.1644/1545-1542(2000)081%3c0948:DAFBOT%3e2.0.CO;2
Siemers BM, Swift S (2006) Differences in sensory ecology contribute to resource partitioning in the bats Myotis bechsteinii and Myotis nattereri (Chiroptera: Vespertilionidae). Behav Ecol Sociobiol 59:373–380. https://doi.org/10.1007/s00265-005-0060-5
Siemers BM, Kriner E, Kaipf I, Simon M, Greif S (2012) Bats eavesdrop on the sound of copulating flies. Curr Biol 22:563–564. https://doi.org/10.1016/j.cub.2012.06.030
Spitz J, Ridoux V, Brind’Amour A (2014) Let’s go beyond taxonomy in diet description: testing a trait-based approach to prey-predator relationships. J Anim Ecol 83:1137–1148. https://doi.org/10.1111/1365.2656.12218
Starik N, Göttert T, Zeller U (2021) Spatial behavior and habitat use of two sympatric bat species. Animals 11:3460. https://doi.org/10.3390/ani11123460
Steck CE, Brinkmann R (2006) The trophic niche of the Geoffroy’s bat (Myotis emarginatus) in south-western Germany. Acta Chiropterologica 8(2):445–450. https://doi.org/10.3161/1733-5329(2006)8[445:TTNOTG]2.0.CO;2
Taweesub C, Tanalgo KC, Sritongchuay T, Huges AC (2021) Understanding global patterns of insectivorous bat dietary research. J Bat Res Conserv 14(1):134–144. https://doi.org/10.14709/BarbJ.14.1.2021.12
Taylor PJ, Grass I, Alberts AJ, Joubert E, Tscharntke T (2018) Economic value of bat predation services—a review and new estimates from macadamia orchards. Ecosyst Serv 30:372–381. https://doi.org/10.1016/j.ecoser.2017.11.015
Tiede J, Diepenbruck M, Gadau J, Wemheuer B, Daniel R, Scherber C (2020) Seasonal variation in the diet of the serotine bat (Eptesicus serotinus): a high-resolution analysis using DNA metabacoding. Basic Appl Ecol 49:1–12. https://doi.org/10.1016/j.baae.2020.09.004
Tournayre O, Leuchtmann M, Galan M, Trillat M, Piry S, Pinaud D, Filippi-Codaccioni O, Pontier D, Charbonnel N (2020a) eDNA metabarcoding reveals a core and secondary diets of the greater horseshoe bat with strong spatio-temporal plasticity. Environ DNA 3(1):277–296. https://doi.org/10.1002/edn3.167
Tournayre O, Leuchtmann M, Filippi-Codaccioni O, Trillat M, Piry S, Pontier D, Charbonnel N, Galan M (2020b) In silico and empirical evaluation of twelve metabarcoding primer sets for insectivorous diet analyses. Ecol Evol 00:1–23. https://doi.org/10.1002/ece3.6362
Valbuena-Lacarra P, Saloña-Bordas MI (2010) Nuevos datos sobre la presencia de Stomoxys calcitrans (Linnaeus, 1758) (Diptera, Muscidae) en explotaciones ganaderas de Vizcaya (País Vasco, Norte de España). Boln Asoc Esp Ent 34(1–2):199–205
Vallejo N, Aihartza J, Goiti U, Arrizabalaga-Escudero A, Flaquer C, Puig X, Aldasoro M, Baroja U, Garin I (2019) The diet of the notch-eared bat (Myotis emarginatus) across the Iberian Peninsula analysed by amplicon metabarcoding. Hystrix 30(1):59–64. https://doi.org/10.4404/hystrix-00189-2019
Vamos EE, Elbrecht V, Leese F (2017) Short COI markers for freshwater macroinvertebrate metabarcoding. Metabarcoding Metagenom 1:e14625. https://doi.org/10.3897/mbmg.1.14625
Ward D, Lublin Y (1992) American arachnological society temporal and spatial segregation of web-building in a community of orb-weaving spiders. J Arachnol 20(2):73–87
Whitaker JO Jr (1988) Food habits analysis of insectivorous bats. In: Kunz TH (ed) Ecological and behavioral methods for the study of bats. Smithsonian Institution Press, Washington, pp 171–189
Wray AK, Peery MZ, Jusino MA, Kochanski JM, Banik MT, Palmer JM, Lindner DL, Gratton C (2021) Predator preferences shape the diets of arthropodivorous bats more than quantitative local prey abundance. Mol Ecol 30(3):855–873. https://doi.org/10.1111/mec.15769
Zahn A, Bauer S, Kriner E, Holzhaider J (2010) Foraging habitats of Myotis emarginatus in Central Europe. Eur J Wildl Res 56:395–400. https://doi.org/10.1007/s10344-009-0331-y
Zeale MR, Butlin RK, Barker GL, Lees DC, Jones G (2011) Taxon-specific PCR for DNA barcoding arthropod prey in bat faeces. Mol Ecol Resour 11(2):236–244. https://doi.org/10.1111/j.1755-0998.2010.02920.x
Acknowledgements
We want to thank everyone who collaborated in collecting samples, especially Ane Caro and the owners who allowed us to sample their properties and homes. Two anonymous reviewers notably improved a previous version of the manuscript. We are also grateful to the Sequencing and Genotyping Unit—Genomic Facilities—SGIker (UPV/EHU/ERDF, EU) for the technical support provided. This study was funded by research grants IT1163-19, IT1571-22, CGL-2015-69069-P and 108123GB-I00. The Spanish Ministry of Universities granted NV (FPU18/02701). Open Access funding provided by University of Basque Country.
Funding
Open Access funding provided thanks to the CRUE-CSIC agreement with Springer Nature.
Author information
Authors and Affiliations
Contributions
NV and IG conceived the ideas and designed the methodology; NV and IG collected the data; NV and LO performed DNA extractions; NV analysed the sequences and performed statistical analyses; NV led the writing of the manuscript. All authors contributed critically to the drafts and gave final approval for publication.
Corresponding author
Ethics declarations
Conflict of interest
The authors have no competing interests to declare that are relevant to the content of this article.
Additional information
Handling editor: Danilo Russo.
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
42991_2023_353_MOESM1_ESM.pdf
Supplementary file1 ESM_1: Details of the methodology for sample analysis, including DNA extraction, PCR amplification, library building and sequencing, bioinformatic analysis, taxonomic assignment of OTUs and selection of OTUs and samples for the analysis (PDF 224 KB)
42991_2023_353_MOESM2_ESM.csv
Supplementary file2 ESM_2: Samples included in the analysis and information regarding colony and date of collection, and quality of the sample regarding contamination from co-occurring bat species (see Online Resource 1) (CSV 46 KB)
42991_2023_353_MOESM3_ESM.csv
Supplementary file3 ESM_3: List of all OTUs found in the samples studied. It includes sequence, taxonomic assignation and number of reads in each sample (CSV 11446 KB)
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Vallejo, N., Aihartza, J., Olasagasti, L. et al. Seasonal shift in the diet of the notched-eared bat (Myotis emarginatus) in the Basque Country: from flies to spiders. Mamm Biol 103, 419–431 (2023). https://doi.org/10.1007/s42991-023-00353-8
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s42991-023-00353-8