Abstract
With the development of modern molecular genetics, the original “one gene-one enzyme” hypothesis has been outdated. For protein coding genes, the discovery of alternative splicing and RNA editing provided the biochemical background for the RNA repertoire of a single locus, which also serves as an important pillar for the enormous protein variability of the genomes. Non-protein coding RNA genes were also revealed to produce several RNA species with distinct functions. The loci of microRNAs (miRNAs), encoding for small endogenous regulatory RNAs, were also found to produce a population of small RNAs, rather than a single defined product. This review aims to present the mechanisms contributing to the astonishing variability of miRNAs revealed by the new sequencing technologies. One important source is the careful balance of arm selection, producing sequentially different 5p- or 3p-miRNAs from the same pre-miRNA, thereby broadening the number of regulated target RNAs and the phenotypic response. In addition, the formation of 5', 3' and polymorphic isomiRs, with variable end and internal sequences also leads to a higher number of targeted sequences, and increases the regulatory output. These miRNA maturation processes, together with other known mechanisms such as RNA editing, further increase the potential outcome of this small RNA pathway. By discussing the subtle mechanisms behind the sequence diversity of miRNAs, this review intends to reveal this engaging aspect of the inherited “RNA world”, how it contributes to the almost infinite molecular variability among living organisms, and how this variability can be exploited to treat human diseases.
Similar content being viewed by others
Explore related subjects
Discover the latest articles, news and stories from top researchers in related subjects.Avoid common mistakes on your manuscript.
Introduction
One of the fundamental ideas that revolutionized biological science and was also considered to lay down the bases of molecular biology was the famous “one gene—one enzyme” hypothesis formulated by George Beadle and Edward Tatum in the beginning of the 1940s (Beadle and Tatum 1941). They proposed that enzymes carry out the metabolic functions in the cells, and all enzymes are determined by unique DNA segments called genes. With the advancement of biochemistry and genetics, and after the formulation of the “central dogma” of molecular biology (Crick 1958), the hypothesis was refined as the “one gene—one polypeptide” statement; yet, it still turned out to be a simplified version of genetic complexity. Soon it was revealed that at the RNA level, the “message” is often a combinatorial output of the DNA sequence of a particular gene. The discovery of splicing and alternative splicing provided the first evidence that several mRNA isoforms can be generated from a single gene, potentially coding for several protein species (Berget et al. 1977; Chow et al. 1977; Baralle and Giudice 2017; Shenasa and Hertel 2019). Another important discovery was the phenomenon of RNA editing, showing that the nucleotide sequence of the transcribed mRNA can be functionally modified, which also results in the alteration of the encoded polypeptide sequence (Benne et al. 1986; Powell et al. 1987). Since these original observations, follow-up investigations revealed several other RNA modification pathways, further supporting the view that gene activity indeed produces a complex pool of transcripts, rather than a single RNA with a single determined function (reviewed in (Li and Mason 2014)).
The next important breakthrough in the RNA field was the discovery of small regulatory RNAs, later identified as microRNAs (miRNAs) (Lee et al. 1993; Wightman et al. 1993). Since their initial description, miRNAs were found to be important posttranscriptional molecular modulators, and apart from fungi, they represent an intracellular regulatory network of the RNA interference (RNAi) machinery in all other organisms examined so far (Lee et al. 2004; Ameres and Zamore 2013). Considering their transcription and maturation, the original assumption again was that one miRNA gene produces a single small RNA product of a discrete length. However, especially with the unforeseen developmental pace of next generation sequencing (NGS) technologies, more and more experimental data suggested that this is not the case and the miRNA repertoire is far more complex than previously thought. Once again in the history of RNA biochemistry, the “one gene—one RNA product” hypothesis needed to be revised by the data showing that the activity of miRNA loci produce numerous small RNAs, often a large population of isomiRs (Gebert and MacRae 2019; Trabucchi and Mategot 2019; Bofill-De Ros et al. 2020). In this review I attempt to present the molecular mechanisms behind this complexity: I will address the flexibility of the once underestimated pre-miRNA arm selection regulation, as well as the mechanisms resulting in 5' and 3' heterogeneity of mature miRNAs, producing numerous 5' and 3' isomiRs from a single miRNA gene. I will also discuss the functional consequences and the medical relevance of these carefully regulated miRNA maturation processes, and how the better understanding of these could contribute to the solution of current public health problems. The aim is to give an appreciative view on the unprecedented sequence diversity of small RNA species produced from a single miRNA gene, and through that, to demonstrate the complexity that is present in the existing “RNA world” of today.
The life of miRNAs—maturation and a “wobbling” mechanism of target recognition
miRNAs are the key actors in one of the RNA interference pathways, and have been evolved in parallel with the short interfering RNAs (siRNAs) and the Piwi-associated small RNAs (piRNAs) (Ghildiyal and Zamore 2009). The latter two represent two pathways that have still kept the common ancient function of protecting the organisms from invasive genetic elements such as viruses and transposons. siRNAs are the main defense guards against outside invaders, namely viruses, and represent a “cellular immune system” which is the major security line in organisms that lack an adaptive immune system (such as lower invertebrates or most plants) (Rosendo Machado et al. 2021; Kong et al. 2022). piRNAs, however, are found exclusively in the animal kingdom, and control the “resident aliens” of the genomes, eliminating endogenous viruses and transposons, especially in the germline (Onishi et al. 2021). Although recent studies revealed various functions of piRNAs in somatic cells, they are still predominantly considered as guardians of the germline, protecting stem cells and especially the gametes (Peng and Lin 2013; Wang et al. 2022). miRNAs, on the other hand, have evolved for a different functionality: they represent an elaborate network of posttranscriptional regulators, controlling endogenous mRNAs and thereby fine-tuning cellular gene expression patterns (Gebert and MacRae 2019). They are present in most eukaryotic groups of the tree of life, having a common origin in plants and animals, and current molecular evidence suggest that the loss of this pathway in some species (such as in fungi) rather represents a secondary loss during evolution (Moran et al. 2017).
microRNAs are short, ~ 20–24 nucleotide long, single-stranded RNA molecules encoded by the genome, and they may be relatively numerous in a given organism: in the human genome, for instance, approximately 2000 miRNAs have already been described, although some still require further validation (http://mirbase.org/). miRNAs can be regulated by their own promoter or they can be located in introns (or even in exons) of other, predominantly protein coding genes (Bartel 2018). Interestingly, in mammalian genomes they often form clusters where several miRNAs can be transcribed simultaneously (Altuvia et al. 2005; Kim and Kim 2007; Chan et al. 2012; Ramalingam et al. 2014; Kabekkodu et al. 2018). After transcription of the primary-miRNAs (pri-miRNA), their maturation requires two subsequent endonucleolytic cleavage steps (Fig. 1A). In animal cells, the first one occurs in the nucleus by the “Microprocessor” complex, containing the RNase III type enzyme Drosha, and its partner protein DGCR8 (‘DiGeorge Syndrome Critical Region 8’). It generates a precursor-miRNA (pre-miRNA) with a hairpin-like structure that is transported to the cytoplasm where the second cleavage takes place by another endonuclease called Dicer. The so formed, usually mismatch containing short double-stranded RNAs are incorporated into an Argonaute (Ago) protein containing, RNA-induced silencing complex (RISC), where further processing eliminates one strand (the “passenger” strand), resulting in a single stranded miRNA-containing functional RISC (Iwakawa and Tomari 2022). The effector function of these ribonucleoprotein complexes is then elicited by the Watson–Crick base pairing complementarity between the miRNA and the target RNA, initiating RNA decay or translation inhibition (Huntzinger and Izaurralde 2011). However, in contrast to siRNAs and piRNAs, often only a small portion of the miRNA has complementarity to the target RNA. In most cases, the 2–8 nucleotides of the 5' sequence, the so-called “seed region” provides the base-pairing interactions but several validated examples have been shown where varying portions from the 3' end of the miRNA also contribute to target recognition which makes the mere bioinformatic target prediction a very difficult task (Berezikov 2011; Pasquinelli 2012). The lack of a long and strict sequence constraint also explains why one miRNA can often bind to several target mRNA molecules (in animals, usually in the 3' untranslated regions, 3' UTRs), and why a given mRNA may contain multiple miRNA-binding sites. The result of this is that similarly to the regulation by transcription factors, miRNAs and their targets represent a complex network of interactions, resulting in a post-transcriptional fine-tuning of gene expression (Arora et al. 2013; Gebert and MacRae 2019; Trabucchi and Mategot 2019).
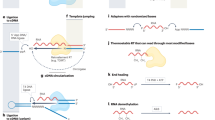
MicroRNA maturation processes A Structure and processing of the pre-miRNA formed on the primary transcript (pri-miRNA). The staggered lines indicate the two consecutive endonucleolytic cleavages: the first is performed by the Drosha/DGCR8 (Microprocessor) complex in the nucleus or substituted by the spliceosome for the Drosha-independent mirtrons. The second cleavage occurs in the cytoplasm by Dicer, the other RNase III type enzyme involved in the maturation. Red and blue lines indicate the 5p or 3p arms, green rectangles depicting the seed sequences. B If they are functionally selected, the different arms are incorporated into distinct RNA-induced Silencing Complexes (RISCs). The ratio of such complexes may vary in cell types, or in different developmental or physiological states, and they target different sets of RNA molecules, although individual targets may overlap. C RISCs formed from a given miRNA locus may differ in their isomiR contents. As examples, distinct 5' and 3' isomiR-containing RISCs are shown above the one with the “canonical” or “reference” miRNA-containing complex. Note that for 5' isomiRs, a difference in target selection may occur due to “seed shifting” (depicted by an arrow)
For many years, it was believed that miRNAs are produced exclusively by the above outlined, well-defined “canonical” biogenesis pathway, with occasional differences only in certain organisms. However, later scientific data challenged this paradigm by the discovery of the Drosha- and Dicer-independent alternative maturation pathways (Winter et al. 2009; Miyoshi et al. 2010). The most prominent of these is certainly the mirtron pathway, with the spliceosome substituting for the Drosha/DGCR8 complex (Fig. 1A) (Berezikov et al. 2007; Okamura et al. 2007; Ruby et al. 2007). Although it was first considered to be a speciality of invertebrate genomes with a short average length of introns, later studies confirmed the existence of mirtrons also in higher organisms, including mammals (Havens et al. 2012; Schamberger et al. 2012; Sibley et al. 2012). The emergence of alternative pathways increased the potential sources of miRNA precursors; however, it soon turned out that the diversity of miRNA sequences is further increased by previously underestimated mechanisms acting not only on the precursors but also on the mature miRNA species.
miRNA arm selection—the “guide” or the “passenger”?
After cleaving the terminal loop of pre-miRNA by Dicer, an important step is the selection of the appropriate strand (referred to as “arm”) of the short double-stranded RNA which will be acting as a guide for the Ago protein in the RISC (Fig. 1A, B). A similar process occurs during siRNA generation (also catalyzed by Dicer) therefore in early studies, the selection mechanism was investigated in parallel in the two RNAi pathways. Analyzing a high number of siRNA or miRNA containing complexes, a clear bias between the strands was found and was shown to be associated with the thermodynamic properties of the arm sequences. It was revealed that the strand having the less stable 5' end is favorable over the other, and it is selected for incorporating into the Ago-complex by unknown mechanisms (Khvorova et al. 2003; Schwarz et al. 2003). This “thermodynamic asymmetry rule” seemed to explain the formation of several mature miRNAs, and the selected arm was named as the “guide” strand, as opposed to the other “passenger” strand (or miRNA*) which was considered to be functionless and therefore destined to be degraded (Hutvagner 2005). Although this rule was clearly helpful in designing effective siRNAs or shRNAs for genetic knock-down experiments, large-scale sequencing data already indicated the functional presence of miRNA* species for some miRNA loci (Ruby et al. 2006), including their presence in human embryonic stem cells (Morin et al. 2008). Meanwhile, a systematic study on miRNA expression profiles from several species indicated that the thermodynamic selection rule is not universal and for various loci, both arms can be expressed simultaneously. Moreover, if expression abundance is considered for defining the dominant arm, than a tissue-specific total switch from miRNA to miRNA* was also revealed for a number miRNA genes (such as for mouse mmu-miR-30e-5p and -3p arms) (Ro et al. 2007). Analyzing NGS data, another group also identified an “outlier” from the rule: in some cancer samples, they detected an almost 1000-fold expression difference of the hsa-miR-223* species, and they proposed that this pattern is likely to be used as a biomarker for leukemia (Kuchenbauer et al. 2008). Nevertheless, the thermodynamic-based selection rule seemed to be working for most examined miRNA loci, still supporting the “one miRNA gene—one mature small RNA product” hypothesis.
But why would one consider the importance of the passenger strand at all? The first answer would be that even if the two strands in the pre-miRNA were fully complementary to each other, they would still be sequentially different: selecting the miRNA* would open up the possibility of targeting other mRNA species by the RISC, and thereby increasing the regulatory repertoire of a given miRNA locus. On the other hand, as shown in Fig. 1A, the two arms of the pre-miRNA stem-loop structure are very often not fully complementary and contain one or more bulges with varying length. This structural feature, however, provides an excellent platform for distinct sequence selection and evidence were soon given for the evolutionary independence of the two arms. In David Bartel’s laboratory, when investigating the origin and the evolution of miRNAs, they found that both arms of the aqu-miR-2015 in sponges are expressed in a cell type-dependent manner, and this regulated arm switching was found to be important not only in sponge development but also in the evolution of sponge species (Grimson et al. 2008). In a parallel study, in Eric Lai’s laboratory they revealed the high conservation and therefore the evolutionary importance of several miRNA* species, and their connection to the evolution of 3' UTR sequences of the targeted protein coding genes (Okamura et al. 2008). Several subsequent studies showed that the guide and the passenger arms show tissue-specific expression patterns in various vertebrate species, including humans, and these all indicated the active regulation of arm selection, despite the thermodynamic stability differences between the two strands (Cloonan et al. 2011; Yang et al. 2011; Marco et al. 2012; Schamberger et al. 2012; Zhou et al. 2012).
The widespread occurrence of arm switching was followed by studies where the phenomenon was connected to physiological regulations and to medical importance. An early example was the work on the expression from the hsa-miR-9/hsa-miR-9* locus in neuronal development: both strands were detected in the samples and the misregulation of their expression profiles was not only associated with Huntington disease but they were also found to target mRNAs from the same biochemical pathway, even sharing common targets (Packer et al. 2008). This was followed by various examples where the miRNA guide/passenger ratio was shown to influence pathways of medical relevance in human diseases, such as the association of the miRNA arm usage pattern with the prognosis and the therapy of macroglobulinemia (Roccaro et al. 2009). There were particularly numerous examples in different cancer types (Zhang et al. 2013; Kuo et al. 2015; Mitra et al. 2015a, 2015b), indicating that the functional expression of the miRNA* strand is much more frequent than previously thought (Table 1).
But if not only the thermodynamic rule then what else regulates pre-miRNA arm selection? Can those other mechanisms override the stability-based assortment? The first indication for a distinct and elaborate mechanism was revealed when the loading of miRNA duplexes into different Ago proteins was studied in Drosophila. In that model organism, two proteins, Ago1 and Ago2 can incorporate these double-stranded small RNA species: it was shown that Ago2 is sensitive to central bulges which can take the priority over thermodynamics and leads to the active selection of the formerly considered passenger strands (Okamura et al. 2009). Similarly in human cells, where four Agos compete for the targets, Ago3 preferentially selects for the passenger strands of certain miRNA duplexes, such as the let-7a-3p arm. It was also shown that hemin can selectively induce Ago3 expression in the K562 cell line, a widely used cellular model of myeloid leukemia, so the selective incorporation of let-7a-3p into this Argonaute protein may have relevance in that particular cancer type (Winter and Diederichs 2013). In line with these observations, an earlier study revealed that the human Ago2 has a clear bias for the first 5'-nucleotide identity, selecting mature miRNAs with either uracil (U) or adenine (A) base in that position (Frank et al. 2010). In a systematic follow-up study, this U and A preference was also shown for all human Agos and the authors proposed that the miRNA arm selection asymmetry is determined by a combination of an “analog pattern” (the thermodynamic instability preference) and a “digital code” (the U or A preference at the 5' end) (Suzuki et al. 2015). Nevertheless, this model still does not give a reassuring answer for all experimental results, nor does it explain the difference in Ago3 selectivity described above (Nakanishi 2022).
In parallel with investigating the roles of Agos in arm selection, various studies revealed that asymmetry “sensors” already influence the process before the double-stranded small RNAs are loaded into the pre-RISC. In Drosophila, Dicer-2 and its binding partner, R2D2 are responsible not only for loading the siRNA duplexes into the Ago2-containing complex but also influencing strand selection; however, it was also shown that human Dicer is dispensable for the asymmetry (Betancur and Tomari 2012). Quite contrary, in Jennifer Doudna’s lab, they provided evidence that human Dicer, in complexed with either the TRBP or the PACT protein, strongly influence miRNA arm selection of Agos (Noland and Doudna 2013), which aspect was further supported by the crystal structure of the Dicer-TRBP complex (Wilson et al. 2015). In addition, a recent study revealed that 3' uridylation of the pre-miRNA could also actively influence arm selection: in case of the pre-miR-324, it causes an alternative (re-positioned) Dicer cleavage which results in a switch from 5p to 3p usage in miRNA maturation (Kim et al. 2020). On the other hand, other mechanisms also exist which can ensure the strict usage of one miRNA arm. One prominent example is the Dicer-independent alternative maturation pathway: in such cases, illustrated by miR-451, the short stem length of the pre-miRNA excludes Dicer binding, and the Ago2 cleavage of the stem-loop structure already determines the exclusive presence of the 5p arm in the RISC-loading complex (Cheloufi et al. 2010; Cifuentes et al. 2010). Interestingly, this alternative pathway is also utilized for shRNA vector design to eliminate off-target effects caused by the expression of the passenger strand (Herrera-Carrillo et al. 2015).
Thorough investigations in recent years eventually changed our original model of pre-miRNA maturation and the exclusive arm usage determined by the thermodynamic stability differences between the two arms. From sequence constraints to the discriminating action of regulatory proteins, several mechanisms were uncovered which can influence miRNA strand selection and although the molecular details are not always completely understood, it is clear that this process is actively regulated in a cell-type dependent manner, presumably by at least the cell-type specific expression level of the identified regulatory factors. This shift in the model is also noticeable in the nomenclature (https://mirbase.org/), as we are not talking about guide and passenger (*) miRNAs any more but rather 5p or 3p species matured from a given miRNA locus. Although understanding this aspect already challenged the ‘one locus—one miRNA’ belief, the real paradigm shift came with the discovery of miRNA isomiRs.
Formation of isomiRs—substantially increasing the miRNA repertoire
After the discoveries that miRNAs are wide-spread posttranscriptional regulators in most eukaryotic genomes, scientist started annotating and placing miRNA gene sequences into databases (such as “mirbase”, https://mirbase.org/), analogously to protein coding genes with their mature mRNA sequences. However, these annotation very often ran into discrepancies concerning the mature miRNA end sequences, detecting not only alternative ends and internal sequences but also non-templated 5' or 3' extensions, the latter ones being end nucleotides different from those present in the flanking genomic sequences (Azuma-Mukai et al. 2008). When several studies reported such miRNA sequence variations in various plant and animal species, the idea of these representing merely sequencing errors were becoming less and less favorable (Ebhardt et al. 2009; Wu et al. 2009; Lee et al. 2010; Zhang et al. 2010; Wyman et al. 2011). It was even revealed that the tissue-specific distributions of these miRNA variations are significantly different, again arguing for the functional presence of these species, which were subsequently referred to as “isomiRs” (Cloonan et al. 2011; Li et al. 2011; Zhou et al. 2012). Their presence was first connected to the miRNA arm switching or the RNA editing phenomena but they were soon shown to represent a distinct miRNA modifying pathway (Ebhardt et al. 2009; Cloonan et al. 2011; Zhou et al. 2012).
Following many years of uncertainty, it finally became accepted that isomiRs exist and are functional, and they were eventually considered as “the hitherto overlooked repertoire of the microRNAome” which potentially influence target selection, miRNA stability, or affect miRNA loading into the RISCs (Burroughs et al. 2011; Neilsen et al. 2012). This again lead to a “paradigm shift”, since from that on, the “reference miRNA” sequence became very hard to define, and a miRNA gene could not be considered as a locus producing a single RNA product with a defined length and sequence, but rather a locus producing a population of related small RNAs with considerable sequence variability (Fig. 1C). According to a recent classification, at least the following categories of isomiRs can be distinguished beyond the canonical (the “reference”) miRNA species: (i) 5' isomiRs, having a different length at the 5' end; (ii) 3' isomiRs, having a different length at the 3' end; (iii) polymorphic isomiRs, with the same length but having internal sequence variations; and (iv) “mixed type” isomiRs, containing differences in any combinations of the first 3 types (Tomasello et al. 2021). However, several questions and problems arose, including the mechanisms leading to isomiR formation, their functional versatility, as well as the problem of how to reliably detect and measure the level of a single isomiR.
Concerning their production, templated and non-templated isomiRs should be discussed separately. Templated 5' and 3' isomiRs can be considered as alternative or imprecise cleavage products from the precursor transcripts, sometimes referred to as Drosha or Dicer “promiscuity” (Wu et al. 2009; Burroughs et al. 2011). As discussed above for miRNA arm selection, theses enzymes are sensitive to sequential and structural differences, and especially for Dicer, structural motifs in the precursor hairpins influence cleavage site selection and therefore the length diversity of the small dsRNA products (Starega-Roslan et al. 2011, 2015a, 2015b). For miR-203, for example, it was found that a “sliding-bulge structure” can cause the formation of two differently folded pre-miRNA structures which are then processed differently by Dicer (Ma et al. 2016). In addition, as TRBP and PACT proteins are important regulatory partners of Dicer, they were also shown to influence cleavage site selection, thereby the production of isomiRs (Lee et al. 2013; Wilson et al. 2015). Finally, it was shown that miRNA arm selection and isomiR production are interconnected: systematically analyzing NGS data it was revealed that the more stable the 5p or the 3p arm of a miRNA is, the more stable its corresponding isomiR profile is (Guo and Chen 2014). It is overall not surprising, as similar sequence motifs and the same processing enzymes are responsible for both miRNA maturation processes.
Analyzing the constantly growing volume of NGS datasets, a significant amount of non-templated isomiRs were also discovered. The majority of them represent 3' uridylated or adenylated species of various lengths which were connected to miRNA stability processes and decay intermediates: it was suggested that 3' uridylation marks miRNAs for degradation, whereas 3' adenylation is a sign for stability (Scheer et al. 2016). On the other hand, there were indications that 3' mono- and oligo-uridylation of pre-miRNAs could play opposing roles in the biogenesis of certain miRNAs (Heo et al. 2012), whereas another study revealed that terminal uridylation of mature miRNAs is physiologically significant and it is especially pervasive in the neonatal period (Jones et al. 2012). But this raised an important question: can an isomiR with a seemingly genomic 3' end sequence be in fact a non-templated isomiR? In principle yes, it could be a mono-uridylated miRNA, even if by coincidence the Dicer-cleaved mature miRNA 3' end is followed by a U nucleotide in the genomic sequence. For certain miRNA groups, such as mirtrons, these cases may be easier to test: the 5' and the 3' ends of a mirtronic pre-miRNA is determined the stringency of splicing, therefore they represent better candidates for studying posttranscriptional modifications, such as non-templated 3' uridylation (Westholm et al. 2012). However, for general miRNA molecules, these cases are both technically and biologically challenging to verify. And this also leads to a very important aspect of isomiR investigations: how can functional isomiRs be reliably identified?
The first problem is the methodology: although carefully optimized NGS methods can detect all isomiRs in a sample (Smith and Hutvagner 2022), this approach is rather expensive and cumbersome when a single locus is to be studied. Traditional RNA detection methods may be applied—but Northern blots, although detecting all isomiRs expressed above a certain threshold, cannot be used to identify new species. The more sensitive real-time qPCR methodologies can increase detectability of single known miRNA variants but the canonical assays cannot reliably distinguish among closely related isomiR sequences (Lee et al. 2010; Schamberger and Orban 2014). However, newly developed and optimized qPCR platforms could provide the desired selectivity in isomiR detection but they still require preliminary NGS data to start with, followed by careful adjustments when a new miRNA locus is to be investigated (Honda and Kirino 2015; Franco et al. 2022).
With the technological improvements, the detection problem seems to be settled. Nevertheless, the real biological problem still remains: can the functional role of a detected isomiR be verified? How can we distinguish miRNA degradation products from bona fide isomiRs? As a hint, there are examples where global changes in the NGS-determined isomiR profiles are associated with a certain phenotype or a disease. As examples, these include the drastic change in the mosquito vector isomiR distribution after Dengue virus infection (Etebari et al. 2015), the dysregulation of 5'-isomiR levels in connection to the Alzheimer disease (Wang et al. 2016), or the global changes in isomiR profiles in certain cancer types (Li et al. 2012; Kuo et al. 2015; Saiselet et al. 2015). However, these results cannot be attributed to single miRNA variants and the real challenge is to assign distinct function(s) to different individual isomiRs. For animal miRNAs, where target selection is strongly based on the seed sequence, the shift at the 5' terminus can clearly influence silencing function (Fig. 1C); however, the potential effect of changes in the length or the sequence of the 3' end could be more problematic to validate. If a new population of isomiRs is to be examined, only a rigorous pipeline of analysis could exclude potential decay intermediates and provide evidence for functional importance of the detected miRNA variants. These include investigating at least (i) the tissue-specific expression patterns, (ii) the association with Ago proteins, (iii) in vitro functional activities (in luciferase assays or on predicted targets), and (iv) the potential evolutionary conservations, although this latter one may exclude species-specific isomiRs (Tan et al. 2014). Despite such methodological challenges, many examples are already known where the concrete functions of selected isomiRs have been proven, which provides evidence that this process of miRNA regulation is actively regulated and physiologically important (Table 2).
These examples underline the relevance isomiR formation and distribution in human physiology and diseases, and the most prominent examples come from cancer biology. An earlier attempt to classify 32 cancer types using isomiR profiling was surprisingly successful: even if a “binary” (present or absent) approach was applied, this method could label different tumor types with high (90%) sensitivity and with a very low (3%) false discovery rate (Telonis et al. 2017). A recent meta-analysis of available cancer datasets and publications not only supported this approach but could identify specific miRNA isomiRs that can be used as biomarkers for certain tumor types (Zelli et al. 2021). A further deep analysis of isomiR datasets could differentiate even among breast cancer subtypes (Nersisyan et al. 2022), whereas a recent study provided evidence that the heterogeneous nuclear ribonucleoprotein C (hnRNPC) induced 5' isomiR shift of miR-21-5p could lead to liver cancer (Park et al. 2022). All these studies show that isomiR-based tumor profiling seems to be a very encouraging approach for cancer diagnostics but isomiR-based tumor targeting also holds promise for future therapy applications.
Nowadays, after many years of intensive research, it is widely accepted that miRNA isomiRs represent an actively regulated and physiologically important layer of gene regulation. The mechanisms behind isomiR formation have, in general, been revealed: beside the subtle regulation of the main miRNA processing enzymes (Drosha and Dicer), the action of terminal nucleotidyl transferases, exoribonucleases and RNA editing enzymes are also responsible for the existence of certain isomiR groups (Tomasello et al. 2021). On the other hand, the regulation of cell-type specific isomiR distribution is still not completely understood and this aspect is currently in the focus of intensive investigations (Zelli et al. 2021; Aparicio-Puerta et al. 2023). Nevertheless, isomiRs should definitely be considered in miRNA target predictions (Ahmed et al. 2014), as well as in the development of new diagnostic tools and in planned therapy applications (Nikolova et al. 2021; Scheper et al. 2022).
Conclusions for future biology
After more than a decade of intensive research, RNA scientists started to appreciate the unforeseen complexity in the hidden layers of small RNA regulation. From the once described “interesting but seemingly unique examples”, we have moved to discover the complex regulation of miRNAs and certainly reached “beyond the one-locus-one-miRNA” paradigm (Telonis et al. 2015). The development and the continuous refinement of new sequencing technologies provide an unprecedented resolution to reveal regulatory mechanisms even at the level of single autonomous cells (Smith and Hutvagner 2022). By revealing the subtle molecular mechanisms of miRNA arm selection and isomiR formation, we have started to understand both the evolutionary and the physiological importance of the elicited posttranscriptional fine-tuning of gene regulation. The discussed phenomena influencing the formation of dynamic miRNA populations have become important for diagnostic tools in several human diseases, presenting useful biomarkers especially in various cancer types, and furthermore also providing potential therapeutic targets for future biomedical research. In addition to the already described functional roles, with the constantly developing detection technologies, new important aspects of miRNA arm selection and isomiR formation can and should also be closely investigated, such as the population- and gender-specific expression profiles which were already described earlier (Loher et al. 2014). Moreover, with the current challenges of new viral threats to human populations, this unexplored population-dependency of miRNA and isomiR repertoire may have a strong influence on the response to viral infections, and therefore this medical aspect of small RNAs should not be neglected either (Rotival et al. 2020).
And the journey has certainly not stopped yet, as similar complexities are expected to be deciphered not only in the other RNAi pathways but most likely in other, RNA-related regulatory mechanisms. All these provide evidence that members of the former “RNA world” are not simple relics of our evolutionary past but they are still active contributors to the current life on Earth. Moreover, they should be considered and utilized also for human medical applications, especially in times of unexpected threats of new potent human infecting viruses.
References
Ahmed F, Senthil-Kumar M, Lee S, Dai X, Mysore KS, Zhao PX (2014) Comprehensive analysis of small RNA-seq data reveals that combination of miRNA with its isomiRs increase the accuracy of target prediction in Arabidopsis thaliana. RNA Biol 11(11):1414–1429. https://doi.org/10.1080/15476286.2014.996474
Altuvia Y, Landgraf P, Lithwick G, Elefant N, Pfeffer S, Aravin A, Brownstein MJ, Tuschl T, Margalit H (2005) Clustering and conservation patterns of human microRNAs. Nucleic Acids Res 33(8):2697–2706. https://doi.org/10.1093/nar/gki567
Ameres SL, Zamore PD (2013) Diversifying microRNA sequence and function. Nat Rev Mol Cell Biol 14(8):475–488. https://doi.org/10.1038/nrm3611
Aparicio-Puerta E, Hirsch P, Schmartz GP, Fehlmann T, Keller V, Engel A, Kern F, Hackenberg M, Keller A (2023) isomiRdb: microRNA expression at isoform resolution. Nucleic Acids Res 51(D1):D179–D185. https://doi.org/10.1093/nar/gkac884
Arora S, Rana R, Chhabra A, Jaiswal A, Rani V (2013) miRNA-transcription factor interactions: a combinatorial regulation of gene expression. Mol Genet Genom 288(3–4):77–87. https://doi.org/10.1007/s00438-013-0734-z
Azuma-Mukai A, Oguri H, Mituyama T, Qian ZR, Asai K, Siomi H, Siomi MC (2008) Characterization of endogenous human Argonautes and their miRNA partners in RNA silencing. Proc Natl Acad Sci U S A 105(23):7964–7969. https://doi.org/10.1073/pnas.0800334105
Baralle FE, Giudice J (2017) Alternative splicing as a regulator of development and tissue identity. Nat Rev Mol Cell Biol 18(7):437–451. https://doi.org/10.1038/nrm.2017.27
Bartel DP (2018) Metazoan MicroRNAs. Cell 173(1):20–51. https://doi.org/10.1016/j.cell.2018.03.006
Beadle GW, Tatum EL (1941) Genetic control of biochemical reactions in Neurospora. Proc Natl Acad Sci U S A 27(11):499–506. https://doi.org/10.1073/pnas.27.11.499
Benne R, Van den Burg J, Brakenhoff JP, Sloof P, Van Boom JH, Tromp MC (1986) Major transcript of the frameshifted coxII gene from trypanosome mitochondria contains four nucleotides that are not encoded in the DNA. Cell 46(6):819–826. https://doi.org/10.1016/0092-8674(86)90063-2
Berezikov E (2011) Evolution of microRNA diversity and regulation in animals. Nat Rev Genet 12(12):846–860. https://doi.org/10.1038/nrg3079
Berezikov E, Chung WJ, Willis J, Cuppen E, Lai EC (2007) Mammalian mirtron genes. Mol Cell 28(2):328–336. https://doi.org/10.1016/j.molcel.2007.09.028
Berget SM, Moore C, Sharp PA (1977) Spliced segments at the 5’ terminus of adenovirus 2 late mRNA. Proc Natl Acad Sci U S A 74(8):3171–3175. https://doi.org/10.1073/pnas.74.8.3171
Betancur JG, Tomari Y (2012) Dicer is dispensable for asymmetric RISC loading in mammals. RNA 18(1):24–30. https://doi.org/10.1261/rna.029785.111
Bofill-De Ros X, Yang A, Gu S (2020) IsomiRs: Expanding the miRNA repression toolbox beyond the seed. Biochim Biophys Acta Gene Regul Mech 1863(4):194373. https://doi.org/10.1016/j.bbagrm.2019.03.005
Burroughs AM, Ando Y, de Hoon MJ, Tomaru Y, Suzuki H, Hayashizaki Y, Daub CO (2011) Deep-sequencing of human Argonaute-associated small RNAs provides insight into miRNA sorting and reveals Argonaute association with RNA fragments of diverse origin. RNA Biol 8(1):158–177. https://doi.org/10.4161/rna.8.1.14300
Chan WC, Ho MR, Li SC, Tsai KW, Lai CH, Hsu CN, Lin WC (2012) MetaMirClust: discovery of miRNA cluster patterns using a data-mining approach. Genomics 100(3):141–148. https://doi.org/10.1016/j.ygeno.2012.06.007
Cheloufi S, Dos Santos CO, Chong MM, Hannon GJ (2010) A dicer-independent miRNA biogenesis pathway that requires ago catalysis. Nature 465(7298):584–589. https://doi.org/10.1038/nature09092
Chow LT, Gelinas RE, Broker TR, Roberts RJ (1977) An amazing sequence arrangement at the 5’ ends of adenovirus 2 messenger RNA. Cell 12(1):1–8. https://doi.org/10.1016/0092-8674(77)90180-5
Cifuentes D, Xue H, Taylor DW, Patnode H, Mishima Y, Cheloufi S, Ma E, Mane S, Hannon GJ, Lawson ND, Wolfe SA, Giraldez AJ (2010) A novel miRNA processing pathway independent of Dicer requires Argonaute2 catalytic activity. Science 328(5986):1694–1698. https://doi.org/10.1126/science.1190809
Cloonan N, Wani S, Xu Q, Gu J, Lea K, Heater S, Barbacioru C, Steptoe AL, Martin HC, Nourbakhsh E, Krishnan K, Gardiner B, Wang X, Nones K, Steen JA, Matigian NA, Wood DL, Kassahn KS, Waddell N, Shepherd J, Lee C, Ichikawa J, McKernan K, Bramlett K, Kuersten S, Grimmond SM (2011) MicroRNAs and their isomiRs function cooperatively to target common biological pathways. Genome Biol 12(12):R126. https://doi.org/10.1186/gb-2011-12-12-r126
Crick FH (1958) On protein synthesis. Symp Soc Exp Biol 12138–12163
Ebhardt HA, Tsang HH, Dai DC, Liu Y, Bostan B, Fahlman RP (2009) Meta-analysis of small RNA-sequencing errors reveals ubiquitous post-transcriptional RNA modifications. Nucleic Acids Res 37(8):2461–2470. https://doi.org/10.1093/nar/gkp093
Etebari K, Osei-Amo S, Blomberg SP, Asgari S (2015) Dengue virus infection alters post-transcriptional modification of microRNAs in the mosquito vector Aedes aegypti. Sci Rep. https://doi.org/10.1038/srep15968
Franco S, Pluvinet R, Sanchez-Herrero JF, Sumoy L, Martinez MA (2022) Rapid and accurate quantification of isomiRs by RT-qPCR. Sci Rep 12(1):17220. https://doi.org/10.1038/s41598-022-22298-7
Frank F, Sonenberg N, Nagar B (2010) Structural basis for 5’-nucleotide base-specific recognition of guide RNA by human AGO2. Nature 465(7299):818–822. https://doi.org/10.1038/nature09039
Gebert LFR, MacRae IJ (2019) Regulation of microRNA function in animals. Nat Rev Mol Cell Biol 20(1):21–37. https://doi.org/10.1038/s41580-018-0045-7
Ghildiyal M, Zamore PD (2009) Small silencing RNAs: an expanding universe. Nat Rev Genet 10(2):94–108. https://doi.org/10.1038/nrg2504
Grimson A, Srivastava M, Fahey B, Woodcroft BJ, Chiang HR, King N, Degnan BM, Rokhsar DS, Bartel DP (2008) Early origins and evolution of microRNAs and Piwi-interacting RNAs in animals. Nature 455(7217):1193–1197. https://doi.org/10.1038/nature07415
Guo L, Chen F (2014) A challenge for miRNA: multiple isomiRs in miRNAomics. Gene 544(1):1–7. https://doi.org/10.1016/j.gene.2014.04.039
Havens MA, Reich AA, Duelli DM, Hastings ML (2012) Biogenesis of mammalian microRNAs by a non-canonical processing pathway. Nucleic Acids Res 40(10):4626–4640. https://doi.org/10.1093/nar/gks026
Heo I, Ha M, Lim J, Yoon MJ, Park JE, Kwon SC, Chang H, Kim VN (2012) Mono-uridylation of pre-microRNA as a key step in the biogenesis of group II let-7 microRNAs. Cell 151(3):521–532. https://doi.org/10.1016/j.cell.2012.09.022
Herrera-Carrillo E, Harwig A, Berkhout B (2015) Toward optimization of AgoshRNA molecules that use a non-canonical RNAi pathway: variations in the top and bottom base pairs. RNA Biol 12(4):447–456. https://doi.org/10.1080/15476286.2015.1022024
Honda S, Kirino Y (2015) Dumbbell-PCR: a method to quantify specific small RNA variants with a single nucleotide resolution at terminal sequences. Nucleic Acids Res 43(12):e77. https://doi.org/10.1093/nar/gkv218
Huntzinger E, Izaurralde E (2011) Gene silencing by microRNAs: contributions of translational repression and mRNA decay. Nat Rev Genet 12(2):99–110. https://doi.org/10.1038/nrg2936
Hutvagner G (2005) Small RNA asymmetry in RNAi: function in RISC assembly and gene regulation. FEBS Lett 579(26):5850–5857. https://doi.org/10.1016/j.febslet.2005.08.071
Iwakawa HO, Tomari Y (2022) Life of RISC: formation, action, and degradation of RNA-induced silencing complex. Mol Cell 82(1):30–43. https://doi.org/10.1016/j.molcel.2021.11.026
Jazdzewski K, Liyanarachchi S, Swierniak M, Pachucki J, Ringel MD, Jarzab B, de la Chapelle A (2009) Polymorphic mature microRNAs from passenger strand of pre-miR-146a contribute to thyroid cancer. Proc Natl Acad Sci USA 106(5):1502–1505. https://doi.org/10.1073/pnas.0812591106
Jones MR, Blahna MT, Kozlowski E, Matsuura KY, Ferrari JD, Morris SA, Powers JT, Daley GQ, Quinton LJ, Mizgerd JP (2012) Zcchc11 uridylates mature miRNAs to enhance neonatal IGF-1 expression, growth, and survival. PLoS Genet 8(11):e1003105. https://doi.org/10.1371/journal.pgen.1003105
Kabekkodu SP, Shukla V, Varghese VK, Jeevitha DS, Chakrabarty S, Satyamoorthy K (2018) Clustered miRNAs and their role in biological functions and diseases. Biol Rev Camb Philos Soc 93(4):1955–1986. https://doi.org/10.1111/brv.12428
Karali M, Persico M, Mutarelli M, Carissimo A, Pizzo M, Singh Marwah V, Ambrosio C, Pinelli M, Carrella D, Ferrari S, Ponzin D, Nigro V, di Bernardo D, Banfi S (2016) High-resolution analysis of the human retina miRNome reveals isomiR variations and novel microRNAs. Nucleic Acids Res 44(4):1525–1540. https://doi.org/10.1093/nar/gkw039
Khvorova A, Reynolds A, Jayasena SD (2003) Functional siRNAs and miRNAs exhibit strand bias. Cell 115(2):209–216
Kim YK, Kim VN (2007) Processing of intronic microRNAs. Embo J 26(3):775–783. https://doi.org/10.1038/sj.emboj.7601512
Kim S, Lee UJ, Kim MN, Lee EJ, Kim JY, Lee MY, Choung S, Kim YJ, Choi YC (2008) MicroRNA miR-199a* regulates the MET proto-oncogene and the downstream extracellular signal-regulated kinase 2 (ERK2). J Biol Chem 283(26):18158–18166. https://doi.org/10.1074/jbc.M800186200
Kim H, Kim J, Yu S, Lee YY, Park J, Choi RJ, Yoon SJ, Kang SG, Kim VN (2020) A mechanism for microRNA arm switching regulated by uridylation. Mol Cell 78(6):1224–1236. https://doi.org/10.1016/j.molcel.2020.04.030
Kong X, Yang M, Le BH, He W, Hou Y (2022) The master role of siRNAs in plant immunity. Mol Plant Pathol 23(10):1565–1574. https://doi.org/10.1111/mpp.13250
Kuchenbauer F, Morin RD, Argiropoulos B, Petriv OI, Griffith M, Heuser M, Yung E, Piper J, Delaney A, Prabhu AL, Zhao Y, McDonald H, Zeng T, Hirst M, Hansen CL, Marra MA, Humphries RK (2008) In-depth characterization of the microRNA transcriptome in a leukemia progression model. Genome Res 18(11):1787–1797. https://doi.org/10.1101/gr.077578.108
Kuo WT, Su MW, Lee YL, Chen CH, Wu CW, Fang WL, Huang KH, Lin WC (2015) Bioinformatic Interrogation of 5p-arm and 3p-arm specific miRNA expression using TCGA datasets. J Clin Med 4(9):1798–1814. https://doi.org/10.3390/jcm4091798
Lee RC, Feinbaum RL, Ambros V (1993) The C elegans heterochronic gene lin-4 encodes small RNAs with antisense complementarity to lin-14. Cell 75(5):843–854. https://doi.org/10.1016/0092-8674(93)90529-Y
Lee R, Feinbaum R, Ambros V (2004) A short history of a short RNA. Cell 116(2 Suppl):S89-92
Lee LW, Zhang S, Etheridge A, Ma L, Martin D, Galas D, Wang K (2010) Complexity of the microRNA repertoire revealed by next-generation sequencing. RNA 16(11):2170–2180. https://doi.org/10.1261/rna.2225110
Lee HY, Zhou K, Smith AM, Noland CL, Doudna JA (2013) Differential roles of human Dicer-binding proteins TRBP and PACT in small RNA processing. Nucleic Acids Res 41(13):6568–6576. https://doi.org/10.1093/nar/gkt361
Li S, Mason CE (2014) The pivotal regulatory landscape of RNA modifications. Annu Rev Genom Hum Genet. https://doi.org/10.1146/annurev-genom-090413-025405
Li SC, Liao YL, Chan WC, Ho MR, Tsai KW, Hu LY, Lai CH, Hsu CN, Lin WC (2011) Interrogation of rabbit miRNAs and their isomiRs. Genomics 98(6):453–459. https://doi.org/10.1016/j.ygeno.2011.08.008
Li SC, Liao YL, Ho MR, Tsai KW, Lai CH, Lin WC (2012) miRNA arm selection and isomiR distribution in gastric cancer. BMC Genom 13(Suppl):1S13. https://doi.org/10.1186/1471-2164-13-S1-S13
Llorens F, Banez-Coronel M, Pantano L, del Rio JA, Ferrer I, Estivill X, Marti E (2013) A highly expressed miR-101 isomiR is a functional silencing small RNA. BMC Genom. https://doi.org/10.1186/1471-2164-14-104
Loher P, Londin ER, Rigoutsos I (2014) IsomiR expression profiles in human lymphoblastoid cell lines exhibit population and gender dependencies. Oncotarget 5(18):8790–8802. https://doi.org/10.18632/oncotarget.2405
Ma H, Wu Y, Niu Q, Zhang J, Jia G, Manjunath N, Wu H (2016) A sliding-bulge structure at the Dicer processing site of pre-miRNAs regulates alternative Dicer processing to generate 5’-isomiRs. Heliyon 2(9):e00148. https://doi.org/10.1016/j.heliyon.2016.e00148
Ma M, Yin Z, Zhong H, Liang T, Guo L (2019) Analysis of the expression, function, and evolution of miR-27 isoforms and their responses in metabolic processes. Genomics 111(6):1249–1257. https://doi.org/10.1016/j.ygeno.2018.08.004
Manzano M, Forte E, Raja AN, Schipma MJ, Gottwein E (2015) Divergent target recognition by coexpressed 5’-isomiRs of miR-142-3p and selective viral mimicry. RNA 21(9):1606–1620. https://doi.org/10.1261/rna.048876.114
Marco A, Macpherson JI, Ronshaugen M, Griffiths-Jones S (2012) MicroRNAs from the same precursor have different targeting properties. Silence 3(1):8. https://doi.org/10.1186/1758-907X-3-8
Mercey O, Popa A, Cavard A, Paquet A, Chevalier B, Pons N, Magnone V, Zangari J, Brest P, Zaragosi LE, Ponzio G, Lebrigand K, Barbry P, Marcet B (2017) Characterizing isomiR variants within the microRNA-34/449 family. FEBS Lett 591(5):693–705. https://doi.org/10.1002/1873-3468.12595
Mitra R, Lin CC, Eischen CM, Bandyopadhyay S, Zhao Z (2015a) Concordant dysregulation of miR-5p and miR-3p arms of the same precursor microRNA may be a mechanism in inducing cell proliferation and tumorigenesis: a lung cancer study. RNA 21(6):1055–1065. https://doi.org/10.1261/rna.048132.114
Mitra R, Sun J, Zhao Z (2015b) microRNA regulation in cancer: one arm or two arms? Int J Cancer 137(6):1516–1518. https://doi.org/10.1002/ijc.29512
Miyoshi K, Miyoshi T, Siomi H (2010) Many ways to generate microRNA-like small RNAs: non-canonical pathways for microRNA production. Mol Genet Genom 284(2):95–103. https://doi.org/10.1007/s00438-010-0556-1
Moran Y, Agron M, Praher D, Technau U (2017) The evolutionary origin of plant and animal microRNAs. Nat Ecol Evol 1(3):27. https://doi.org/10.1038/s41559-016-0027
Morin RD, O’Connor MD, Griffith M, Kuchenbauer F, Delaney A, Prabhu AL, Zhao Y, McDonald H, Zeng T, Hirst M, Eaves CJ, Marra MA (2008) Application of massively parallel sequencing to microRNA profiling and discovery in human embryonic stem cells. Genome Res 18(4):610–621. https://doi.org/10.1101/gr.7179508
Musiyenko A, Bitko V, Barik S (2008) Ectopic expression of miR-126*, an intronic product of the vascular endothelial EGF-like 7 gene, regulates prostein translation and invasiveness of prostate cancer LNCaP cells. J Mol Med (berl) 86(3):313–322. https://doi.org/10.1007/s00109-007-0296-9
Nakanishi K (2022) Anatomy of four human Argonaute proteins. Nucleic Acids Res 50(12):6618–6638. https://doi.org/10.1093/nar/gkac519
Neilsen CT, Goodall GJ, Bracken CP (2012) IsomiRs–the overlooked repertoire in the dynamic microRNAome. Trends Genet 28(11):544–549. https://doi.org/10.1016/j.tig.2012.07.005
Nersisyan S, Zhiyanov A, Engibaryan N, Maltseva D, Tonevitsky A (2022) A novel approach for a joint analysis of isomiR and mRNA expression data reveals features of isomiR targeting in breast cancer. Front Genet. https://doi.org/10.3389/fgene.2022.1070528
Nikolova M, Naydenov M, Glogovitis I, Apostolov A, Saare M, Boggavarapu N, Salumets A, Baev V, Yahubyan G (2021) Coupling miR/isomiR and mRNA expression signatures unveils new molecular layers of endometrial receptivity. Life (basel). 11(12):1319. https://doi.org/10.3390/life11121391
Noland CL, Doudna JA (2013) Multiple sensors ensure guide strand selection in human RNAi pathways. RNA 19(5):639–648. https://doi.org/10.1261/rna.037424.112
Okamura K, Hagen JW, Duan H, Tyler DM, Lai EC (2007) The mirtron pathway generates microRNA-class regulatory RNAs in Drosophila. Cell 130(1):89–100. https://doi.org/10.1016/j.cell.2007.06.028
Okamura K, Phillips MD, Tyler DM, Duan H, Chou YT, Lai EC (2008) The regulatory activity of microRNA* species has substantial influence on microRNA and 3’ UTR evolution. Nat Struct Mol Biol 15(4):354–363. https://doi.org/10.1038/nsmb.1409
Okamura K, Liu N, Lai EC (2009) Distinct mechanisms for microRNA strand selection by Drosophila Argonautes. Mol Cell 36(3):431–444. https://doi.org/10.1016/j.molcel.2009.09.027
Onishi R, Yamanaka S, Siomi MC (2021) PiRNA- and siRNA-mediated transcriptional repression in Drosophila mice and yeast: new insights and biodiversity. EMBO Rep 22(10):e53062. https://doi.org/10.15252/embr.202153062
Packer AN, Xing Y, Harper SQ, Jones L, Davidson BL (2008) The bifunctional microRNA miR-9/miR-9* regulates REST and CoREST and is downregulated in Huntington’s disease. J Neurosci 28(53):14341–14346. https://doi.org/10.1523/JNEUROSCI.2390-08.2008
Park S, Yang HD, Seo JW, Nam JW, Nam SW (2022) hnRNPC induces isoform shifts in miR-21-5p leading to cancer development. Exp Mol Med 54(6):812–824. https://doi.org/10.1038/s12276-022-00792-2
Pasquinelli AE (2012) MicroRNAs and their targets: recognition, regulation and an emerging reciprocal relationship. Nat Rev Genet 13(4):271–282. https://doi.org/10.1038/nrg3162
Pass HI, Goparaju C, Ivanov S, Donington J, Carbone M, Hoshen M, Cohen D, Chajut A, Rosenwald S, Dan H, Benjamin S, Aharonov R (2010) hsa-miR-29c* is linked to the prognosis of malignant pleural mesothelioma. Cancer Res 70(5):1916–1924. https://doi.org/10.1158/0008-5472.CAN-09-3993
Peng JC, Lin H (2013) Beyond transposons: the epigenetic and somatic functions of the Piwi-piRNA mechanism. Curr Opin Cell Biol 25(2):190–194. https://doi.org/10.1016/j.ceb.2013.01.010
Powell LM, Wallis SC, Pease RJ, Edwards YH, Knott TJ, Scott J (1987) A novel form of tissue-specific RNA processing produces apolipoprotein-B48 in intestine. Cell 50(6):831–840. https://doi.org/10.1016/0092-8674(87)90510-1
Ramalingam P, Palanichamy JK, Singh A, Das P, Bhagat M, Kassab MA, Sinha S, Chattopadhyay P (2014) Biogenesis of intronic miRNAs located in clusters by independent transcription and alternative splicing. RNA 20(1):76–87. https://doi.org/10.1261/rna.041814.113
Ro S, Park C, Young D, Sanders KM, Yan W (2007) Tissue-dependent paired expression of miRNAs. Nucleic Acids Res 35(17):5944–5953. https://doi.org/10.1093/nar/gkm641
Roccaro AM, Sacco A, Chen C, Runnels J, Leleu X, Azab F, Azab AK, Jia X, Ngo HT, Melhem MR, Burwick N, Varticovski L, Novina CD, Rollins BJ, Anderson KC, Ghobrial IM (2009) microRNA expression in the biology, prognosis, and therapy of Waldenstrom macroglobulinemia. Blood 113(18):4391–4402. https://doi.org/10.1182/blood-2008-09-178228
Rosendo Machado S, van der Most T, Miesen P (2021) Genetic determinants of antiviral immunity in dipteran insects—compiling the experimental evidence. Dev Comp Immunol. https://doi.org/10.1016/j.dci.2021.104010
Rotival M, Siddle KJ, Silvert M, Pothlichet J, Quach H, Quintana-Murci L (2020) Population variation in miRNAs and isomiRs and their impact on human immunity to infection. Genome Biol 21(1):187. https://doi.org/10.1186/s13059-020-02098-w
Ruby JG, Jan C, Player C, Axtell MJ, Lee W, Nusbaum C, Ge H, Bartel DP (2006) Large-scale sequencing reveals 21U-RNAs and additional microRNAs and endogenous siRNAs in C. Elegans. Cell 127(6):1193–1207. https://doi.org/10.1016/j.cell.2006.10.040
Ruby JG, Jan CH, Bartel DP (2007) Intronic microRNA precursors that bypass Drosha processing. Nature 448(7149):83–86. https://doi.org/10.1038/nature05983
Saiselet M, Gacquer D, Spinette A, Craciun L, Decaussin-Petrucci M, Andry G, Detours V, Maenhaut C (2015) New global analysis of the microRNA transcriptome of primary tumors and lymph node metastases of papillary thyroid cancer. BMC Genom. https://doi.org/10.1186/s12864-015-2082-3
Salem O, Erdem N, Jung J, Munstermann E, Worner A, Wilhelm H, Wiemann S, Korner C (2016) The highly expressed 5’isomiR of hsa-miR-140–3p contributes to the tumor-suppressive effects of miR-140 by reducing breast cancer proliferation and migration. BMC Genom. https://doi.org/10.1186/s12864-016-2869-x
Schamberger A, Orban TI (2014) 3’ IsomiR species and DNA contamination influence reliable quantification of microRNAs by stem-loop quantitative PCR. PLoS ONE 9(8):e106315. https://doi.org/10.1371/journal.pone.0106315
Schamberger A, Sarkadi B, Orban TI (2012) Human mirtrons can express functional microRNAs simultaneously from both arms in a flanking exon-independent manner. RNA Biol 9(9):1177–1185. https://doi.org/10.4161/rna.21359
Scheer H, Zuber H, De Almeida C, Gagliardi D (2016) Uridylation earmarks mRNAs for degradation… and more. Trends Genet 32(10):607–619. https://doi.org/10.1016/j.tig.2016.08.003
Scheper M, Romagnolo A, Besharat ZM, Iyer AM, Moavero R, Hertzberg C, Weschke B, Riney K, Feucht M, Scholl T, Petrak B, Maulisova A, Nabbout R, Jansen AC, Jansen FE, Lagae L, Urbanska M, Ferretti E, Tempes A, Blazejczyk M, Jaworski J, Kwiatkowski DJ, Jozwiak S, Kotulska K, Sadowski K, Borkowska J, Curatolo P, Mills JD, Aronica E, Consortium ME (2022) MiRNAs and isomiRs: serum-based biomarkers for the development of intellectual disability and autism spectrum disorder in tuberous sclerosis complex. Biomedicines. 10(8): 1838. https://doi.org/10.3390/biomedicines10081838
Schwarz DS, Hutvagner G, Du T, Xu Z, Aronin N, Zamore PD (2003) Asymmetry in the assembly of the RNAi enzyme complex. Cell 115(2):199–208
Shenasa H, Hertel KJ (2019) Combinatorial regulation of alternative splicing. Biochim Biophys Acta Gene Regul Mech. 11–12:194392. https://doi.org/10.1016/j.bbagrm.2019.06.003
Sibley CR, Seow Y, Saayman S, Dijkstra KK, El Andaloussi S, Weinberg MS, Wood MJ (2012) The biogenesis and characterization of mammalian microRNAs of mirtron origin. Nucleic Acids Res 40(1):438–448. https://doi.org/10.1093/nar/gkr722
Smith CM, Hutvagner G (2022) A comparative analysis of single cell small RNA sequencing data reveals heterogeneous isomiR expression and regulation. Sci Rep 12(1):2834. https://doi.org/10.1038/s41598-022-06876-3
Starega-Roslan J, Krol J, Koscianska E, Kozlowski P, Szlachcic WJ, Sobczak K, Krzyzosiak WJ (2011) Structural basis of microRNA length variety. Nucleic Acids Res 39(1):257–268. https://doi.org/10.1093/nar/gkq727
Starega-Roslan J, Galka-Marciniak P, Krzyzosiak WJ (2015a) Nucleotide sequence of miRNA precursor contributes to cleavage site selection by Dicer. Nucleic Acids Res 43(22):10939–10951. https://doi.org/10.1093/nar/gkv968
Starega-Roslan J, Witkos TM, Galka-Marciniak P, Krzyzosiak WJ (2015b) Sequence features of Drosha and Dicer cleavage sites affect the complexity of isomiRs. Int J Mol Sci 16(4):8110–8127. https://doi.org/10.3390/ijms16048110
Suzuki HI, Katsura A, Yasuda T, Ueno T, Mano H, Sugimoto K, Miyazono K (2015) Small-RNA asymmetry is directly driven by mammalian argonautes. Nat Struct Mol Biol 22(7):512–521. https://doi.org/10.1038/nsmb.3050
Tan GC, Chan E, Molnar A, Sarkar R, Alexieva D, Isa IM, Robinson S, Zhang S, Ellis P, Langford CF, Guillot PV, Chandrashekran A, Fisk NM, Castellano L, Meister G, Winston RM, Cui W, Baulcombe D, Dibb NJ (2014) 5’ isomiR variation is of functional and evolutionary importance. Nucleic Acids Res 42(14):9424–9435. https://doi.org/10.1093/nar/gku656
Telonis AG, Loher P, Jing Y, Londin E, Rigoutsos I (2015) Beyond the one-locus-one-miRNA paradigm: microRNA isoforms enable deeper insights into breast cancer heterogeneity. Nucleic Acids Res 43(19):9158–9175. https://doi.org/10.1093/nar/gkv922
Telonis AG, Magee R, Loher P, Chervoneva I, Londin E, Rigoutsos I (2017) Knowledge about the presence or absence of miRNA isoforms (isomiRs) can successfully discriminate amongst 32 TCGA cancer types. Nucleic Acids Res 45(6):2973–2985. https://doi.org/10.1093/nar/gkx082
Tomasello L, Distefano R, Nigita G, Croce CM (2021) The microRNA family gets wider: the isomiRs classification and role. Front Cell Dev Biol. 9:668648. https://doi.org/10.3389/fcell.2021.668648
Trabucchi M, Mategot R (2019) Subcellular heterogeneity of the microRNA machinery. Trends Genet 35(1):15–28. https://doi.org/10.1016/j.tig.2018.10.006
Tsang WP, Kwok TT (2009) The miR-18a* microRNA functions as a potential tumor suppressor by targeting on K-Ras. Carcinogenesis 30(6):953–959. https://doi.org/10.1093/carcin/bgp094
Wang S, Xu Y, Li M, Tu J, Lu Z (2016) Dysregulation of miRNA isoform level at 5’ end in Alzheimer’s disease. Gene 584(2):167–172. https://doi.org/10.1016/j.gene.2016.02.020
Wang X, Ramat A, Simonelig M, Liu MF (2022) Emerging roles and functional mechanisms of PIWI-interacting RNAs. Nat Rev Mol Cell Biol. https://doi.org/10.1038/s41580-022-00528-0
Westholm JO, Ladewig E, Okamura K, Robine N, Lai EC (2012) Common and distinct patterns of terminal modifications to mirtrons and canonical microRNAs. RNA 18(2):177–192. https://doi.org/10.1261/rna.030627.111
Wightman B, Ha I, Ruvkun G (1993) Posttranscriptional regulation of the heterochronic gene lin-14 by lin-4 mediates temporal pattern formation in C. elegans. Cell 75(5):855–862. https://doi.org/10.1016/0092-8674(93)90530-4
Wilson RC, Tambe A, Kidwell MA, Noland CL, Schneider CP, Doudna JA (2015) Dicer-TRBP complex formation ensures accurate mammalian microRNA biogenesis. Mol Cell 57(3):397–407. https://doi.org/10.1016/j.molcel.2014.11.030
Winter J, Diederichs S (2013) Argonaute-3 activates the let-7a passenger strand microRNA. RNA Biol 10(10):1631–1643. https://doi.org/10.4161/rna.26424
Winter J, Jung S, Keller S, Gregory RI, Diederichs S (2009) Many roads to maturity: microRNA biogenesis pathways and their regulation. Nat Cell Biol 11(3):228–234. https://doi.org/10.1038/ncb0309-228
Wu H, Ye C, Ramirez D, Manjunath N (2009) Alternative processing of primary microRNA transcripts by Drosha generates 5’ end variation of mature microRNA. PLoS ONE 4(10):e7566. https://doi.org/10.1371/journal.pone.0007566
Wyman SK, Knouf EC, Parkin RK, Fritz BR, Lin DW, Dennis LM, Krouse MA, Webster PJ, Tewari M (2011) Post-transcriptional generation of miRNA variants by multiple nucleotidyl transferases contributes to miRNA transcriptome complexity. Genome Res 21(9):1450–1461. https://doi.org/10.1101/gr.118059.110
Yamane D, Selitsky SR, Shimakami T, Li Y, Zhou M, Honda M, Sethupathy P, Lemon SM (2017) Differential hepatitis C virus RNA target site selection and host factor activities of naturally occurring miR-122 3 variants. Nucleic Acids Res 45(8):4743–4755. https://doi.org/10.1093/nar/gkw1332
Yang JS, Phillips MD, Betel D, Mu P, Ventura A, Siepel AC, Chen KC, Lai EC (2011) Widespread regulatory activity of vertebrate microRNA* species. RNA 17(2):312–326. https://doi.org/10.1261/rna.2537911
Zelli V, Compagnoni C, Capelli R, Corrente A, Cornice J, Vecchiotti D, Di Padova M, Zazzeroni F, Alesse E, Tessitore A (2021) Emerging role of isomiRs in cancer: state of the art and recent advances. Genes (basel) 12(9):1447. https://doi.org/10.3390/genes12091447
Zhang W, Gao S, Zhou X, Xia J, Chellappan P, Zhou X, Zhang X, Jin H (2010) Multiple distinct small RNAs originate from the same microRNA precursors. Genome Biol 11(8):R81. https://doi.org/10.1186/gb-2010-11-8-r81
Zhang Y, Yang P, Sun T, Li D, Xu X, Rui Y, Li C, Chong M, Ibrahim T, Mercatali L, Amadori D, Lu X, Xie D, Li QJ, Wang XF (2013) miR-126 and miR-126* repress recruitment of mesenchymal stem cells and inflammatory monocytes to inhibit breast cancer metastasis. Nat Cell Biol 15(3):284–294. https://doi.org/10.1038/ncb2690
Zhou H, Arcila ML, Li Z, Lee EJ, Henzler C, Liu J, Rana TM, Kosik KS (2012) Deep annotation of mouse iso-miR and iso-moR variation. Nucleic Acids Res 40(13):5864–5875. https://doi.org/10.1093/nar/gks247
Acknowledgements
The author would like to thank Eszter Horváth for critical reading of the manuscript. MicroRNA-related research in the author’s lab is supported by the PC-II-12/2022 grant from the Hungarian Academy of Sciences.
Funding
Open access funding provided by ELKH Research Centre for Natural Sciences.
Author information
Authors and Affiliations
Contributions
Conceptualization and manuscript preparation were done by T.I.O.
Corresponding author
Ethics declarations
Conflict of interest
The author declares that he has no conflict of interest.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Orbán, T.I. One locus, several functional RNAs—emerging roles of the mechanisms responsible for the sequence variability of microRNAs. BIOLOGIA FUTURA 74, 17–28 (2023). https://doi.org/10.1007/s42977-023-00154-7
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s42977-023-00154-7