Abstract
Treatment of Proteus mirabilis infections is a challenge due to the high abundance of virulence factors and the high intrinsic resistance to antimicrobials. Multidrug resistance (MDR) and extensive drug resistance (XDR) further challenge the control of P. mirabilis infection. This study aimed to investigate the correlation between virulence determinants and multidrug resistance in 100 clinical isolates of P. mirabilis collected in Alexandria from December 2019 to June 2021. Susceptibility to antimicrobials was tested by the Kirby Bauer method. Detection of swarming, urease, protease, hemolysin, and biofilm formation was performed phenotypically and by PCR amplification of zapA, flaA, ureC, mrpA, atfA, ucaA, hpmA, and luxS. MDR and XDR were detected in 34% and 5%, respectively. All isolates were positive for motility, swarming, urease, and protease production. Ninety percent were positive for hemolysin production, while 73% formed biofilm. All isolates possessed the ureC and zapA genes. The luxS, flaA, ucaA, hpmA, mrpA, and atfA genes were detected in 99%, 98%, 96% 90%, 89%, and 84%, respectively. The presence of a single biofilm-related gene was statistically correlated with non-biofilm production (P= 0.018). It was concluded that P. mirabilis isolates from catheterized-urine samples were significantly associated with biofilm formation. MDR and virulence were not statistically correlated. A significant positive correlation was detected between some virulence genes in P. mirabilis. Non-MDR isolates of P. mirabilis had a high abundance of virulence factors with no statistically significant difference from MDR. Most of the MDR and all XDR isolates could produce biofilm.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Proteus spp. belong to the order Enterobacterales and to the family Morganellaceae [1]. Clinically important Proteus spp. include P. mirabilis, P. vulgaris, and P. penneri [2]. P. mirabilis is a common causative agent of a diversity of clinical infections such as urinary tract infections (UTI), wound and burn infections, prostatitis, meningitis, otitis media, and rarely respiratory tract infections [3].
Treatment of P. mirabilis infections is a challenge due to intrinsic and acquired antimicrobial resistance. P. mirabilis is characterized by its intrinsic resistance to many antimicrobial agents: colistin, polymyxin, nitrofurans, tigecycline, and tetracycline [4].
With the extensive and unrestricted use of antimicrobial agents, acquired multidrug resistance (MDR) and extensive drug resistance (XDR) have been commonly encountered among clinical isolates of P. mirabilis, posing marked challenges to the control of infection by this bacterial species [5].
The high abundance of virulence factors in Proteus spp. further augments its impact as a potential threat to public health. P. mirabilis possesses a variety of virulence determinants such as fimbriae, flagellae, urease enzyme, hemolysin production, protease enzyme production, biofilm production, and quorum sensing [6].
Fimbriae are responsible for adherence to uroepithelial cells or medical devices, causing urinary tract infection [6] as well as adhesion to wound extracellular matrix proteins such as collagen and fibronectin causing wound infections. The most common fimbriae are mannose-resistant Proteus-like fimbriae (MR/P), uroepithelial cell adhesion (UCA/NAF), ambient temperature fimbriae (ATF), P. mirabilis fimbriae (PMF), and P. mirabilis P-like fimbria (PMF) [7]. Flagellae are utilized for spread to other sites as the migration to the upper urinary tract causing pyelonephritis and the dispersal of biofilm from catheters to the urinary tract [8, 9].
Urease enzymes hydrolyze urea into carbon dioxide and ammonia, which renders the pH of the environment alkaline [10]. In UTI, the alkaline pH causes the precipitation of polyvalent cations such as calcium and magnesium resulting in stone formation [11]. In wound infections, the alkaline pH contributes to delayed wound healing [12].
Hemolysins are secreted toxins produced by P. mirabilis, which insert into host cell membranes, causing pore formation, and cytotoxicity, hence facilitating the invasion [13]. In addition, P. mirabilis produces ZapA metalloproteases which protect the organism from the host defense by cleaving immunoglobulins, IgA, and IgG [8].
P. mirabilis forms biofilms on chronic wound infections [14] and in urinary tract infections especially on catheters [6]. P. mirabilis has a unique ability to form biofilms of crystalline nature, owing to the urease activity. This leads to encrustation and obstruction in many cases [8].
Quorum sensing is the main regulator of many virulence factors. It is of particular importance in regulating the multicellular and coordinated processes of swarming and biofilm formation. Quorum sensing in P. mirabilis involves autoinducer-1 which is controlled by luxR genes and autoinducer-2 which is controlled by luxS gene [15].
This study aimed to investigate the correlation between virulence determinants and multidrug resistance in clinical isolates of P. mirabilis.
Materials and methods
Sample collection
The study was performed on 100 isolates of P. mirabilis, collected from clinical samples submitted at the Microbiology laboratory of the Medical Research Institute, Alexandria University. Isolates were collected along the period from December 2019 to June 2021. The isolates were collected from mid-stream urine samples, catheter-collected urine, and wound swabs.
Identification of isolates
Colonies were presumptively identified as Proteus species by observing the formation of swarming on blood agar and the growth of smooth, non-lactose fermenting colonies on MacConkey agar after incubation at 37°C under aerobic conditions for 16–24 h. Standard biochemical tests were used for further species identification of P. mirabilis. The detection of ureC gene by PCR was employed for further genotypic confirmation of P. mirabilis species identification, as previously described [16].
Antimicrobial susceptibility testing
Kirby Bauer disk diffusion method for susceptibility testing of the isolates was performed, and the results were interpreted as per the CLSI 2021 recommendations [17]. Nineteen antibiotic disks (Oxoid, UK) were tested: ampicillin (10 μg), amoxicillin-clavulanate (20/10 μg), ampicillin-sulbactam (10/10 μg), piperacillin-tazobactam (100/10 μg), cefepime (30 μg), cefotaxime (30 μg), ceftriaxone (30 μg), ceftazidime (30 μg), aztreonam (30 μg), ertapenem (10 μg), imipenem (10 μg), meropenem (10 μg), gentamicin (10 μg), tobramycin (10 μg), amikacin (30 μg), ciprofloxacin (5 μg), levofloxacin (5 μg), ofloxacin (5 μg), trimethoprim-sulfamethoxazole (1.25/23.75 μg). Acquired resistance to antibiotics (at least one) belonging to three or more categories of antimicrobial agents was described as multidrug resistance (MDR). Isolates sensitive to only one or two categories of antimicrobials are classified as extensive drug-resistant (XDR) isolates [18].
Detection of virulence factors of P. mirabilis
Phenotypic detection of virulence factors
Urease production
The isolates were cultured on urea agar medium by stab. The tubes were incubated at a temperature of 37°C for 24 h. The change of color from yellow to magenta was a sign of a positive result [19].
Protease production
Skim milk agar (Himedia) was inoculated with the test isolates and incubated at 37°C for 24 h. A positive reaction appeared in the form of a clear zone that developed around the colonies [20].
Hemolysin production
Tube hemolysis assay was performed by inoculating 2 mL of nutrient broth with bacteria. One hundred microliters of 1% washed human red blood cells (RBCs) was added, and the media were incubated for 24 h at 37 °C. Non-hemolyzed red blood cells settled to the bottom of the test tube and formed a button. No button was observed if the cells were lysed by hemolysin [21].
Formation of biofilm
The microtiter plate (MTP) assay was performed to test the isolates for biofilm formation. Briefly, a bacterial suspension was prepared in MHB supplemented with 1% glucose. Following the adjustment of the suspension to 5×107 CFU/mL, 200 μL was used to inoculate the wells of a 96-well MTP. After overnight incubation at 37°C, the content of the wells was discarded, and the wells were washed with normal saline. Methanol (99%) was added to the biofilms for fixation. The biofilms were then stained with 1% crystal violet for 20 min. The plate was washed with normal saline to get rid of excess dye. Finally, 200 μL of ethanol (99%) was added to release the bound crystal violet. The optical density (OD) of each well was measured at 620 nm using an MTP reader. The strength of biofilm formation was calculated in relation to the OD of negative control wells [22].
Genotypic detection of virulence factors
DNA extraction
Genomic DNA was extracted from P. mirabilis isolates by boiling method as previously described. In brief, several colonies from a fresh overnight culture of the isolates were washed with Tris-EDTA (TE) buffer and the pelleted cells were resuspended in 250 μL TE buffer by vortexing. The bacterial suspension was incubated in a boiling water bath for 10–15 min, immediately chilled on ice for 2 min, then centrifuged at 14000 rpm for 15 min. The supernatant was transferred into a new tube and was used as the stock DNA extract, which was 10 folds diluted and used as a template for PCR [23].
Amplification of virulence genes by multiplex PCR
Conventional PCR was used for the amplification of 8 virulence genes using specific primers: zapA [24] encoding extracellular metalloprotease, flaA [25] for flagellae, ureC [26] for urease enzyme large subunit, mrpA [27] for mannose-resistant Proteus-like fimbria, atfA [28] for ambient-temperature fimbriae, ucaA [29] for uroepithelial cell adhesin fimbriae, hpmA [25] for hemolysin, luxS [15] for quorum sensing.
Six genes were detected in 3 multiplex PCR reactions for amplification of (atfA + zapA), (hpmA + luxS), and (ureC+ ucaA). Each of the 2 genes flaA and mrpA was amplified in a single PCR reaction. The PCR reactions contained 12.5 μL 2x PCR master mix (Dream Taq™ Hot Start Green DNA Polymerase Master Mix (Thermo Fisher), 1 μL of each primer (10 pmol/μL), 3 μL of DNA extract, and PCR-grade water to a final volume of 25 μL.
The primers’ sequences, annealing temperatures, and the expected sizes of the amplicons are illustrated in Table S1, supplementary material. The thermal cycling conditions were 4 min of initial denaturation at 95 °C, 35 cycles of denaturation at 95 C for 30 s, annealing at the primers’ specific annealing temperature for 30 s, and extension for 72 °C for 1 min, followed by 10 min of final extension at 72 °C.
Amplification products were visualized by electrophoresis using 1.5% (w/v) agarose gel in TAE buffer stained with 5 μL of ethidium bromide solution (10 mg/mL). Gene-specific bands were observed in comparison with the bands of a 100-bp DNA ladder (Thermo Fisher) using a 302-nm UV transilluminator.
Statistical analysis
Data were analyzed using IBM SPSS software package version 20.0. (Armonk, NY: IBM Corp). The significance of the obtained results was judged at the 5% alpha level. For categorical variables, the chi-square test was used to compare different groups while Fisher’s exact or Monte Carlo correction for chi-square when more than 20% of the cells have an expected count of less than 5. Correlation analysis by the Spearman rank method was done using RStudio.
Results
The majority of P. mirabilis isolates were isolated from mid-stream urine samples 55 (55%) followed by wound swabs 31 (31%), and catheter-associated urinary tract infection 14 (14%).
Results of antimicrobial susceptibility testing
The highest percentage of resistance was 62% and 61% to ampicillin and trimethoprim-sulfamethoxazole, respectively, while the lowest resistance was 3%, 3%, 6%, and 8% to piperacillin-tazobactam, ertapenem, meropenem, and imipenem respectively (Fig. 1). Overall, 61% of isolates were non-MDR, 34% were MDR, and 5% were XDR.
Phenotypic detection of virulence factors
All isolates (100%) were positive for motility, swarming, urease, and protease production. Ninety isolates (90%) were positive for hemolysin production by tube hemolysis test. Seventy- three isolates (73%) were positive for biofilm formation: including 40 isolates forming a weak biofilm, 30 forming a moderate biofilm, and only 3 isolates forming a strong biofilm.
Genotypic detection of virulence factors
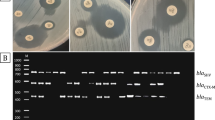
All P. mirabilis isolates (100%) were positive for ureC and zapA gene, encoding for urease and extracellular metalloprotease, respectively. As for the quorum sensing gene luxS, the flagellar gene flaA, and the fimbrial adhesin gene ucaA, they were detected in 99%, 98%, and 96% of the isolates, respectively. The hemolysin gene hpmA and the fimbrial genes mrpA and atfA genes were positive in 90%, 89%, and 84% of the isolates, respectively (Figs. S1–S5 supplementary material).
There was a statistically significant agreement (of 100%) between phenotypic and genotypic results in the detection of urease, protease, and hemolysin (P <0.001).
Among the 100 isolates, 72% carried all 8 studied virulence genes, 17% carried 7/8 of the studied virulence genes, and 8% carried 6/8 of the studied virulence genes, while 1% carried 5/8 and 2% carried 4/8 of the studied virulence genes.
There was a statistically significant association between the number of biofilm genes and biofilm formation (P= 0.03). The proportions did not differ between biofilm producers and non-biofilm producers except for 1 gene; P= 0.017, indicating that the presence of a single biofilm-related gene in the isolates (mainly luxS gene or ucaA gene) was statistically significantly associated with non-biofilm production (Table 1).
Association between virulence factors and the type of clinical infection
There was a statistically significant difference regarding biofilm formation among isolates from different types of clinical samples (P = 0.034). All isolates from catheterized urine samples formed biofilm (100%), followed by wound isolates (74.2%), and then mid-stream urine samples (65.5%) (Table 2).
Association between antimicrobial resistance and virulence factors in P. mirabilis isolates
There was no statistically significant difference in analyzing the distribution of virulence factors phenotypically and genotypically among isolates with different antimicrobial resistance patterns (Tables 3 and 4).
Only 70.5% of the non-MDR and 73.5% of MDR P. mirabilis isolates were biofilm producers, while all 5 XDR isolates produced biofilm. Most of the isolates formed weak to moderate biofilm, regardless of their resistance patterns. The 73 biofilm-forming isolates included 43 non-MDR isolates, 25 MDR, and 5 XDR. Among the 43 non-MDR isolates, 21 formed weak biofilm, 20 formed moderate, and 2 formed strong biofilm. As for the 25 MDR biofilm-forming isolates, 17 formed weak biofilm, 7 formed moderate biofilm, and 2 formed strong biofilm. Two of the 5 XDR isolates formed weak biofilm, while 3 formed moderate biofilm. Among the 27 isolates that were unable to form a biofilm, 18 were non-MDR and 9 were MDR (Table 3).
The quorum sensing gene luxS was detected in all the non-MDR P. mirabilis isolates, followed by the flagellar gene the flaA which was detected in 98.4% of these isolates. The hemolysin gene hpmA was positive in 85.2%. The prevalence of fimbrial adhesin genes (ucaA, mrpA, and atfA) was 95.1%, 90.2%, and 80.3%, respectively (Table 4).
As for the MDR isolates, the quorum sensing gene, luxS, was positive in 97.1% of the isolates. Both the flagellar gene, flaA, and hemolysin gene, hpmA, were positive in 97.1% of them. The prevalence of fimbrial adhesin genes (ucaA, atfA, and mrpA) was 97.1%, 88.2%, and 85.3%, respectively. All the XDR isolates were positive for all the aforementioned genes.
Correlation between different virulence genes, virulence factors, and resistance patterns among all P. mirabilis isolates
There was a significant positive correlation between the following pairs of virulence genes: mrpA and ucaA (rho= 0.42, P= 0.0007), atfA and hpmA (rho=0.4, P= 0.002), atfA and ucaA (rho= 0.33, P= 0.03). A perfect positive significant correlation between hpmA virulence gene and the hemolysis virulence factor was observed (rho= 1, P<0.000001). Moreover, a statistically significant positive correlation was noticed between the atfA virulence gene and hemolysis (rho= 0.4, P=0.002). Despite the fact that the rho coefficients between virulence genes are small ranging 0.33 to 0.42 indicating a fair positive correlation, the correlation is statistically significant and cannot be ignored (Fig. 2). No correlation was detected between virulence factors and antibiotic resistance patterns (MDR and XDR).
Heat map of correlation analysis by Spearman rank method between different virulence genes, virulence factors, and resistance patterns among all P. mirabilis isolates (n= 100). Blue color indicates a positive correlation while red color indicates a negative correlation. This figure was produced by the Corrplot package in RStudio. A value of +1 or −1 indicates a perfect correlation, 0.8 to 0.9 (−0.8 to −0.9) indicates a very strong correlation, 0.6 to 0.7 (−0.6 to −0.7) indicates a moderate correlation, and 0.3 to 0.5 (−0.3 to −0.5) indicates a fair correlation
Discussion
Multidrug resistance in P. mirabilis is increasingly observed. The high level of intrinsic resistance and the high virulence of this bacterial species further augment its impact on public health. Multidrug-resistant P. mirabilis has been previously isolated from a variety of human clinical specimens and food animals [30]. In this study, we determined the antibiotic resistance profiles of 100 clinical isolates of P. mirabilis, collected from urine samples and wound infection. In addition, we examined the isolates phenotypically and genotypically for the presence of some important virulence determinants including motility, swarming, urease and protease production, biofilm formation, and quorum sensing. We aimed to explore any potential correlations between virulence determinants and multidrug resistance among clinical isolates of P. mirabilis.
Multidrug resistance in P. mirabilis has been reported to be gained by means of horizontal gene transfer from other bacterial species such as K. pneumoniae, with the eventual formation of a hybrid (mosaic) plasmid that carries resistance and virulence genes from both species [31]. In our study, we detected multidrug resistance and extensive drug resistance in 34% and 5% of our isolates, respectively. This percentage is less than that reported in Israel in 2010, where the authors reported the isolation of MDR P. mirabilis from 50% of their patients who were all hospitalized with UTI [32]. A more recent study in India also revealed a much higher prevalence of MDR (85%) among UTI isolates of P. mirabilis [33]. Lower prevalence, however, was reported in Europe and Taiwan [34, 35].
Four out of the six studied virulence factors were invariably detected phenotypically among all our isolates of P. mirabilis: motility, swarming, urease, and protease production. However, at the molecular level, only ureC and zapA genes were amplified in all isolates. The flagellar gene flaA was amplified in 98% of the isolates.
With regard to our two motile isolates with non-amplified flaA gene, this result could be explained by the recombination of flaA and flaB with the formation of flaAB hybrids that have different nucleotide sequence that is not amplifiable by the flaA primers used in this study [36]. Another possible explanation is that the swarming was controlled by other genes such as fliL gene [37].
Similar results for the detection of virulence factors in urine isolates of P. mirabilis were also encountered by Filipiak et al. who found that all isolates of P. mirabilis did produce two of the studied virulence determinants phenotypically: swarming and urease production. They also reported the amplification of ureC and zapA genes in all isolates [38]. Likewise, Abd Al-Mayahi and Al-Dulaimi et al. detected flaA gene in all the swarming isolates [39, 40]. On the contrary, Ali et al. reported a lower percentage of flaA gene amplification (86.66%) among swarming isolates [25]. On the other hand, Al-Dulaimi et al. detected ureC gene in only (85.7%) of their urease-producing isolates [40]. In addition, Alsherees et al. detected zapA gene in only (39.28%) of their isolates [24].
Hemolysis was detected phenotypically and genotypically (hpmA amplified) in 90% of our isolates. On the other hand, Filipiak et al. detected hpmA gene in all isolates although hemolytic activity was phenotypically observed in 84% of their isolates, with only 16% showing typical appearance of beta hemolysis [38]. Similarly, Mirzaei et al. reported that all their P. mirabilis isolates produced hemolysin and had hpmA gene [33]. On the contrary, Jaber et al. detected hpmA gene in only 50% of their isolates [41].
Seventy-three percent of our isolates were able to form biofilm on microtiter plates, which was mostly of weak (40%) to moderate (30%) intensity. At the molecular level, biofilm-related genes, luxS quorum sensing gene, and the fimbrial genes: ucaA, mrpA, and atfA were amplified in of 99%, 96%, 89%, and 84% of the isolates respectively.
The sole presence of one biofilm-related gene in the isolates (mainly luxS gene or ucaA gene) was statistically associated with non-biofilm production (P=0.017). A statistically significant fair positive correlation was detected between some of the biofilm genes: mrpA and ucaA (P= 0.0007), as well as atfA and ucaA (P= 0.03). The fimbrial gene atfA was also found to be of a fair positive correlation with hemolysis (P=0.002) and hpmA gene (P= 0.002).
The relation between luxS gene and biofilm production was previously reported by Abd Albagar et al. [42]. However, other studies suggested that MR/P and ATF fimbriae rather than UCA fimbriae have a major role in biofilm formation [43, 44]. Based on the results of the current study, no significant correlation was detected between luxS gene and ucaA genes and they were not found to be significantly correlated with biofilm production.
The association between the high abundance of biofilm-related genes among the isolates and their ability to form biofilm phenotypically was also observed by Filipiak et al. who reported that the biofilm-genes ucaA and mrpA were amplified in all of their P. mirabilis isolates and 96% of them were able to form biofilm on polyurethane [38]. Several studies also reported a high abundance of biofilm genes among clinical isolates of P. mirabilis. For instance, Hussein et al. detected luxS and atfA genes in 100% and 98.4% of their isolates, respectively [45]. In addition, Kamel et al. detected mrpA gene in all tested isolates [46]. On the other hand, Abbas et al. reported that only 47% of their isolates carried luxS gene, while 35% carried mrpA gene [47]. Sun et al. reported a lower percentage of ucaA (33%) and atfA (64.77%) genes among their isolates of P. mirabilis [43].
Catheterized urine isolates are more liable to form biofilm than mid-stream urine isolates, since catheter surfaces facilitate adherence of bacteria without clearance, in the absence of host defense mechanisms which occur in the bladder upon binding of bacteria to epithelia, and also in the absence of the flushing mechanism of urine which also has a role [6, 48]. From this work, we observed that all the isolates from catheterized urine samples were able to form a biofilm, followed by wound swab isolates (74.2%). A statistically significant difference (P=0.034) was observed regarding the distribution of biofilm-positive isolates among different types of clinical samples.
Similarly, Jacobsen et al. proved that all P. mirabilis isolates from urinary catheters formed a biofilm [8]. Also, Hola et al. reported that catheterized urine P. mirabilis isolates had a higher ability to form biofilm than those isolated from feces as a control group [49]. Another similar study reported by Abdallah et al. found that P. mirabilis isolates from catheterized patients had a higher percentage (43.3%) of biofilm-forming ability than those of non-catheterized samples (30%) but did not reach the level of significance [50]. On the contrary, Kwiecinska-Piróg et al. reported no difference in biofilm formation ability between catheterized and non-catheterized urine isolates [22].
Our results revealed that the existence of virulence factors, including the ability to form a biofilm, is not correlated with the antimicrobial resistance profile of P. mirabilis clinical isolates. All of our isolates were positive to 4/6 of the studied virulence factors, regardless of being MDR or non-MDR. Nevertheless, it was observed that all the XDR isolates were positive to all the six studied virulence factors phenotypically and at the molecular level.
On the other hand, a negative correlation between virulence and multidrug resistance was reported by Rodulfo et al. who found that non-MDR P. mirabilis possessed a higher number of virulence factors compared to MDR, yet this was statistically significant only with 2 virulence factors that are swarming and twitching motility [51].
The hpmA hemolysin of P. mirabilis is accountable for pore formation in host cells with subsequent tissue damage [52]. This hemolysin was not detected phenotypically and genotypically in 10 of our 100 isolates, among which 9 were non-MDR. Likewise, Mishu et al. reported that among their 44 clinical isolates of P. mirabilis, 3 isolates were negative for hpmA, among which 2 were non-MDR and 1 was MDR [53].
According to Sun et al., the formation of moderate-intensity biofilm is correlated with increased virulence [43]. This agrees with our findings, as most of our isolates that were positive for biofilm formation, which was of moderate to weak intensity, were also positive for most of the studied virulence determinants.
We did not find any significant association between biofilm formation and multidrug resistance. Our results showed that biofilm formation was slightly higher among non-MDR isolates (58.9%). At the same time, most of the non-biofilm-forming isolates were also non-MDR (66.7%). However, all the XDR isolates were biofilm producers. Moreover, most of the MDR isolates (73.5%) were also biofilm producers. This high biofilm-forming ability of XDR and MDR isolates of P. mirabilis complicates infection and renders treatment more difficult.
Similar findings were reported by Rodulfo et al., who found that biofilm formation was slightly higher in the non-MDR isolates of P. mirabilis (84.8%), compared to 76.1% in the MDR group [51]. Meanwhile, Ghaima et al. and Sun et al. reported that biofilm-forming isolates of P. mirabilis significantly displayed more resistance to antimicrobial agents compared with non-biofilm-forming isolates [43, 54]. Also, Filipiak et al. reported that strong biofilm formation is correlated with multidrug resistance, and they attributed this to the blockade of antimicrobial penetration by the extracellular matrix of biofilm [38].
Conclusion
From this work, we concluded that P. mirabilis isolates collected from catheterized-urine samples are associated with a high ability of biofilm formation. A significant positive correlation was detected between some pairs of virulence genes in P. mirabilis: mrpA and ucaA, in addition to atfA and ucaA, as well as atfA and hpmA. The ability of biofilm formation and the high abundance of virulence factors were not found to be correlated with multidrug resistance. The non-MDR isolates of P. mirabilis have a large repository of virulence factors with no statistically significant difference from MDR isolates. Most of the MDR and all XDR isolates were biofilm producers, which represents a serious challenge in the management of infection by these isolates.
This study reveals that most clinical isolates of P. mirabilis, regardless of their resistance pattern, are fully equipped with a large number of virulence factors, the co-existence of many of which is significantly correlated.
References
Bonnin RA, Girlich D, Jousset AB, Gauthier L, Cuzon G, Bogaerts P et al (2020) A single Proteus mirabilis lineage from human and animal sources: a hidden reservoir of OXA-23 or OXA-58 carbapenemases in Enterobacterales. Sci Rep 10(1):1–9. https://doi.org/10.1038/s41598-020-66161-z
O’Hara CM, Brenner FW, Miller JM (2000) Classification, identification, and clinical significance of Proteus, Providencia, and Morganella. Clin Microbiol Rev 13(4):534–546. https://doi.org/10.1128/CMR.13.4.534
Drzewiecka D (2016) Significance and roles of Proteus spp. bacteria in natural environments. Microb Ecol 72(4):741–758. https://doi.org/10.1007/s00248-015-0720-6
Stock I (2003) Natural antibiotic susceptibility of Proteus spp., with special reference to P. mirabilis and P. penneri strains. J Chemother 15(1):12–26. https://doi.org/10.1179/joc.2003.15.1.12
Li Z, Peng C, Zhang G, Shen Y, Zhang Y, Liu C et al (2022) Prevalence and characteristics of multidrug-resistant Proteus mirabilis from broiler farms in Shandong Province, China. Poult Sci 101(4):101710. https://doi.org/10.1016/j.psj.2022.101710
Armbruster CE, Mobley HL, Pearson MM (2018) Pathogenesis of Proteus mirabilis infection. EcoSal Plus 8(1):10–128. https://doi.org/10.1128/ecosalplus.ESP-0009-2017
Hasan TH, Alasedi KK, Jaloob AA (2021) Proteus mirabilis virulence factors. Int J Pharm Res 13(1):2145–2149. https://doi.org/10.31838/ijpr/2021.13.01.169
Jacoben S, Stickler D, Mobley H, Shirtliff M (2008) Complicated catheter-associated urinary tract infections due to Escherichia coli and Proteus mirabilis. Clin Microbiol Rev 21:26–59. https://doi.org/10.1128/CMR.00019-07
Schaffer JN, Norsworthy AN, Sun T-T, Pearson MM (2016) Proteus mirabilis fimbriae and urease-dependent clusters assemble in an extracellular niche to initiate bladder stone formation. Proc Natl Acad Sci U S A 113(16):4494–4499. https://doi.org/10.1073/pnas.1601720113
Coker C, Poore CA, Li X, Mobley HL (2000) Pathogenesis of Proteus mirabilis urinary tract infection. Microbes Infect 2(12):1497–1505. https://doi.org/10.1016/s1286-4579(00)01304-6
Yuan F, Huang Z, Yang T, Wang G, Li P, Yang B et al (2021) Pathogenesis of Proteus mirabilis in catheter-associated urinary tract infections. Urol Int 105(5-6):354–361. https://doi.org/10.1159/000514097
Schneider LA, Korber A, Grabbe S, Dissemond J (2007) Influence of pH on wound-healing: a new perspective for wound-therapy? Arch Dermatol Res 298(9):413–420. https://doi.org/10.1007/s00403-006-0713-x
Braun V, Focareta T (1991) Pore-forming bacterial protein hemolysins (cytolysins). Crit Rev Microbiol 18(2):115–158. https://doi.org/10.3109/10408419109113511
Rajpaul K (2015) Biofilm in wound care. Br J Community Nurs 20(Sup3):S6–S11. https://doi.org/10.12968/bjcn.2015.20.Sup3.S6
Stankowska D, Czerwonka G, Rozalska S, Grosicka M, Dziadek J, Kaca W (2012) Influence of quorum sensing signal molecules on biofilm formation in Proteus mirabilis O18. Folia Microbiol (Praha) 57(1):53–60. https://doi.org/10.1007/s12223-011-0091-4
Zhang W, Niu Z, Yin K, Liu P, Chen L (2013) Quick identification and quantification of Proteus mirabilis by polymerase chain reaction (PCR) assays. Ann Microbiol 63(2):683–689. https://doi.org/10.1007/s13213-012-0520-x
Humphries R, Bobenchik AM, Hindler JA, Schuetz AN (2021) Overview of changes to the clinical and laboratory standards institute performance standards for antimicrobial susceptibility testing, M100. J Clin Microbiol 59(12):e00213. https://doi.org/10.1128/JCM.00213-21
Magiorakos A-P, Srinivasan A, Carey RB, Carmeli Y, Falagas M, Giske C et al (2012) Multidrug-resistant, extensively drug-resistant and pandrug-resistant bacteria: an international expert proposal for interim standard definitions for acquired resistance. Clin Microbiol Infect 18(3):268–281. https://doi.org/10.1111/j.1469-0691.2011.03570.x
Brink B (2010) Urease test protocol. American Society for Microbiology, Washington, DC, pp 1–7
Senior B (1999) Investigation of the types and characteristics of the proteolytic enzymes formed by diverse strains of Proteus species. J Med Microbiol 48(7):623–628. https://doi.org/10.1099/00222615-48-7-623
Mobley HL, Chippendale GR (1990) Hemagglutinin, urease, and hemolysin production by Proteus mirabilis from clinical sources. J Infect Dis 161(3):525–530. https://doi.org/10.1093/infdis/161.3.525
Kwiecinska-Piróg J, Bogiel T, Skowron K, Wieckowska E, Gospodarek E (2014) Proteus mirabilis biofilm-qualitative and quantitative colorimetric methods-based evaluation. Braz J Microbiol 45:1423–1431. https://doi.org/10.1590/s1517-83822014000400037
Peng X, Yu K-Q, Deng G-H, Jiang Y-X, Wang Y, Zhang G-X et al (2013) Comparison of direct boiling method with commercial kits for extracting fecal microbiome DNA by Illumina sequencing of 16S rRNA tags. J Microbiol Methods 95(3):455–462. https://doi.org/10.1016/j.mimet.2013.07.015
Alsherees HAA, Ali J, Rana T (2016) Molecular study of Proteus mirabilis bacteria isolated from urine and wounds in Hospitals Al-Najaf Province. Int J Adv Res Biol Sci 3(6):99–105 SOI: http://s-o-i.org/1.15/ijarbs-2016-3-6-13
Ali HH, Yousif MG (2015) Detection of some virulence factors genes of Proteus mirablis that isolated from urinary tract infection. IJAR 3(1):156–163 ISSN 2320-5407
Takeuchi H, Yamamoto S, Terai A, Kurazono H, Takeda Y, Okada Y et al (1996) Detection of Proteus mirabilis urease gene in urinary calculi by polymerase chain reaction. Int J Urol 3(3):202–206. https://doi.org/10.1111/j.1442-2042.1996.tb00517.x
Ali AS, Abid AJ, Abbas FM (2018) Molecular assessments of Proteus mirabilis virulence factors isolated from urinary tract infection patients. Int J Pharm Res 10(4):523–527. https://doi.org/10.31838/ijpr/2018.10.04.084
Zunino P, Geymonat L, Allen AG, Legnani-Fajardo C, Maskell DJ (2000) Virulence of a Proteus mirabilis ATF isogenic mutant is not impaired in a mouse model of ascending urinary tract infection. FEMS Immunol Med Microbiol 29(2):137–143. https://doi.org/10.1111/j.1574-695X.2000.tb01516.x
Mohammed GJ, Abdul-Razaq MS (2014) Molecular detection of fimbrial genes of Proteus vulgaris isolated from patients with urinary tract infection. Photon J Microbiol 107:226–235 ISJN53497294D711620112014
Algammal AM, Hashem HR, Alfifi KJ, Hetta HF, Sheraba NS, Ramadan H et al (2021) atpD gene sequencing, multidrug resistance traits, virulence-determinants, and antimicrobial resistance genes of emerging XDR and MDR-Proteus mirabilis. Sci Rep 11(1):1–15. https://doi.org/10.1038/s41598-021-88861-w
Shelenkov A, Petrova L, Fomina V, Zamyatin M, Mikhaylova Y, Akimkin V (2020) Multidrug-resistant Proteus mirabilis strain with cointegrate plasmid. Microorganisms 8(11):1775. https://doi.org/10.3390/microorganisms8111775
Cohen-Nahum K, Saidel-Odes L, Riesenberg K, Schlaeffer F, Borer A (2010) Urinary tract infections caused by multi-drug resistant Proteus mirabilis: risk factors and clinical outcomes. Infect 38(1):41–46. https://doi.org/10.1007/s15010-009-8460-5
Mirzaei A, Nasr Esfahani B, Raz A, Ghanadian M, Moghim S (2021) From the urinary catheter to the prevalence of three classes of integrons, β-lactamase genes, and differences in antimicrobial susceptibility of Proteus mirabilis and clonal relatedness with Rep-PCR. Biomed Res Int 2021:9952769. https://doi.org/10.1155/2021/9952769
Critchley IA, Cotroneo N, Pucci MJ, Jain A, Mendes RE (2020) Resistance among urinary tract pathogens collected in Europe during 2018. J Glob Antimicrob Resist 23:439–444. https://doi.org/10.1016/j.jgar.2020.10.020
Lin M-F, Liou M-L, Kuo C-H, Lin Y-Y, Chen J-Y, Kuo H-Y (2019) Antimicrobial susceptibility and molecular epidemiology of Proteus mirabilis isolates from three hospitals in Northern Taiwan. Microb Drug Resist 25(9):1338–1346. https://doi.org/10.1089/mdr.2019.0066
Manos J, Artimovich E, Belas R (2004) Enhanced motility of a Proteus mirabilis strain expressing hybrid FlaAB flagella. Microbiol 150(5):1291–1299. https://doi.org/10.1099/mic.0.26727-0
Lee Y-Y, Belas R (2015) Loss of FliL alters Proteus mirabilis surface sensing and temperature-dependent swarming. J Bacteriol 197(1):159–173. https://doi.org/10.1128/JB.02235-14
Filipiak A, Chrapek M, Literacka E, Wawszczak M, Głuszek S, Majchrzak M et al (2020) Pathogenic factors correlate with antimicrobial resistance among clinical Proteus mirabilis strains. Front Microbiol 11:579389. https://doi.org/10.3389/fmicb.2020.579389
Al-Mayahi F (2017) Phenotypic and molecular detection of virulence factors in Proteus mirabilis isolated from different clinical sources. Bas J Vet Res 16(1):369–388
Al-Dulaimi MTS, Al-Taai HRR (2020) Detection of some virulence factors and antibiotics susceptibility of Proteus mirabilis isolated from different clinical sources in Baquba. Biochem Cell Arch 20(1):803–809. https://doi.org/10.35124/bca.2020.20.1.803
Jaber AH, Almiyah SAF (2022) Molecular detection of some virulence genes for Proteus mirabilis bacteria isolated from diabetic foot ulcers. Eurasian Med Res Period 8:59–67 ISSN: 2795-7624 (E)
Abd Albagar FA, Baqer BA (2020) Phenotype and genotype detection of producing biofilm in some species of pathogenic bacteria and studying the inhibitory effect of plant extract against biofilm. Eurasia J Biosci 14(2):7193–7196
Sun Y, Wen S, Zhao L, Xia Q, Pan Y, Liu H et al (2020) Association among biofilm formation, virulence gene expression, and antibiotic resistance in Proteus mirabilis isolates from diarrhetic animals in Northeast China. BMC Vet Res 16(1):1–10. https://doi.org/10.1186/s12917-020-02372-w
Scavone P, Iribarnegaray V, Caetano AL, Schlapp G, Härtel S, Zunino P (2016) Fimbriae have distinguishable roles in Proteus mirabilis biofilm formation. Pathog Dis 74(5):ftw033. https://doi.org/10.1093/femspd/ftw033
Hussein EI, Al-Batayneh K, Masadeh MM, Dahadhah FW, Al Zoubi MS, Aljabali AA et al (2020) Assessment of pathogenic potential, virulent genes profile, and antibiotic susceptibility of Proteus mirabilis from urinary tract infection. Int J Microbiol 2020:1231807. https://doi.org/10.1155/2020/1231807
Kamel A, Al-Yasseen AK (2009) Phenotypic and genotypic characterization of some virulence factors among Proteus mirabilis isolated from clinical samples in Al-Najaf Al-Ashraf/Iraq. JGPT 11(5):471–478
Abbas KF, Al Khafaji JK, Al-Shukri MS (2015) Molecular detection of some virulence genes in Proteus mirabilis isolated from Hillaprovince. Int J Res Stud Biosci 3:85–89 ISSN 2349-0357 (Print) & ISSN 2349-0365 (Online)
Mater H, Hussein B, Mahdi L, Musafer H, Ghaeb L, Mijbel B et al (2019) Role of biofilm in reinfection in catheter-associated urinary tract infection in Iraqi women. J Glob Pharma Technol 11(2):32–37 ISSN: 0975 -8542
Hola V, Peroutkova T, Ruzicka F (2012) Virulence factors in Proteus bacteria from biofilm communities of catheter-associated urinary tract infections. FEMS Immunol Med Microbiol 65(2):343–349. https://doi.org/10.1111/j.1574-695X.2012.00976.x
Abdallah NMA, Elsayed SB, Mostafa MMY, El-gohary GM (2011) Biofilm-forming bacteria isolated from urinary tract infection, relation to catheterization and susceptibility to antibiotics. Int J Biotechnol Mol Biol Res 2(10):172–178. https://doi.org/10.5897/IJBMBR.9000008
Rodulfo H, Horta M, Mata G, Gutiérrez R, González Y, Michelli E et al (2021) Negative correlation between virulence and multidrug resistance in intrahospital and community-acquired infections by Proteus mirabilis, in Eastern Venezuela. Investig Clin 62(1):37–51. https://doi.org/10.22209/IC.v62n1a04
Cestari SE, Ludovico MS, Martins FH, da Rocha SPD, Elias WP, Pelayo JS (2013) Molecular detection of HpmA and HlyA hemolysin of uropathogenic Proteus mirabilis. Curr Microbiol 67(6):703–707. https://doi.org/10.1007/s00284-013-0423-5
Mishu NJ, Shamsuzzaman S, Khaleduzzaman H, Nabonee MA (2022) Association between biofilm formation and virulence genes expression and antibiotic resistance pattern in Proteus mirabilis, isolated from patients of Dhaka Medical College Hospital. Arch Clin Biomed Res 6(3):418–434. https://doi.org/10.26502/acbr.50170257
Ghaima KK, Hamid HH, Hasan S (2017) Biofilm formation, antibiotic resistance and detection of mannose-resistant Proteus-like (MR/P) fimbriae genes in Proteus mirabilis isolated from UTI. Int J ChemTech Res 10(5):964–971 ISSN: 0974-4290, ISSN(Online):2455-9555
Acknowledgements
The authors would like to thank Rasha Emad Mansour, MSc, Alexandria Main University Hospital, Alexandria University, for performing correlation analysis using RStudio.
Funding
Open access funding provided by The Science, Technology & Innovation Funding Authority (STDF) in cooperation with The Egyptian Knowledge Bank (EKB).
Author information
Authors and Affiliations
Contributions
Mai Elhoshi: methodology, formal analysis and investigation, original draft preparation. Eglal El-Sherbiny: conceptualization, writing — review and editing, supervision. Amel Elsheredy: conceptualization, methodology, formal analysis and investigation, writing — review and editing, supervision. Aliaa Gamaleldin Aboulela: methodology, formal analysis and investigation, original draft preparation, writing — review and editing, supervision.
Corresponding author
Ethics declarations
Ethics approval
The research design has been approved by the Ethics Committee of the Medical Research Institute, Alexandria University. The research was performed on bacterial isolates collected from clinical samples that were already cultured as part of the routine work in the Microbiology laboratory of the Medical Research Institute, Alexandria University. No human participants, their data nor biological material from them was utilized in the research.
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
ESM 1
(DOCX 1649 kb)
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Elhoshi, M., El-Sherbiny, E., Elsheredy, A. et al. A correlation study between virulence factors and multidrug resistance among clinical isolates of Proteus mirabilis. Braz J Microbiol 54, 1387–1397 (2023). https://doi.org/10.1007/s42770-023-01080-5
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s42770-023-01080-5