Abstract
Molecular methodologies providing data on viral concentration and infectivity have been successfully used in environmental virology, supporting quantitative risk assessment studies. The present study aimed to assess human mastadenovirus (HAdV) intact particles using a derivative of propidium monoazide associated with qPCR (PMAxx-qPCR) in aquatic matrices. Initially, different concentrations of PMAxx were evaluated to establish an optimal protocol for treating different naturally contaminated matrices, using 10 min incubation in the dark at 200 rpm at room temperature and 15 min of photoactivation in the PMA-Lite™ LED photolysis device. There was no significant reduction in the quantification of infectious HAdV with increasing concentration of PMAxx used (20 μM, 50 μM, and 100 μM), except for sewage samples. In this matrix, a reduction of 5.01 log of genomic copies (GC)/L was observed from the concentration of 50 μM and revealed 100% HAdV particles with damaged capsids. On the other hand, the mean reduction of 0.51 log in stool samples using the same concentration mentioned above demonstrated 83% of damaged particles eliminated in the stool. Following, 50 μM PMAxx-qPCR protocol revealed a log reduction of 0.91, 0.67, and 1.05 in other samples of raw sewage, brackish, and seawater where HAdV concentration reached 1.47 × 104, 6.81 × 102, and 2.33 × 102 GC/L, respectively. Fifty micrometers of PMAxx protocol helped screen intact viruses from different matrices, including sea and brackish water.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Diagnostic methods for viral detection and quantification have been investigated over the years, stimulating the development of innovative tools and technologies in virology [1]. Evaluation of infectious and non-infectious viruses is essential in environmental virology [2, 3] since the quantification of infectious viruses is a determining factor for calculating the quantitative risk assessment of exposure of susceptible individuals to contaminated environmental matrices [4, 5].
Viral quantification methodologies to estimate values of viral particles as the viral plaque assay using plaque-forming units (PFU) per ml, median tissue culture infectious dose (TCID50), or quantitative polymerase chain reaction (qPCR) have been used. The last one cannot distinguish between infectious and non-infectious viruses [6,7,8,9,10].
Cell culture has been the standard gold method in virology to assess viral infectivity; however, integrated cell culture quantitative PCR (ICC-qPCR) protocols are commonly used when dealing with environmental matrices and enteric viruses considered fastidious [11]. The ICC-qPCR method is laborious and costly but widely used for human mastadenovirus (HAdV) due to its ability to replicate in cell culture. However, cultivation in different cell types to increase the chances of isolation of serotypes is still needed [12]. HAdV has been described as a potential indicator of fecal contamination in environmental matrices mainly due to their high concentration and prevalence in sewage samples, resistance to environmental conditions, and absence of seasonality [13, 14].
In this scenario, a dye propidium monoazide (PMA) or its derivative (PMAxx) combined with qPCR (PMA(xx)-qPCR) has been pointed to as a faster, specific, and sensitive alternative approach to assessing capsid-integrity of enteric viruses in the environment, thus quantifying potentially infectious viral particles [15,16,17,18,19,20,21]. PMA is a nucleic acid intercalating dye with a photoinducible azide group that covalently binds to RNA/DNA from virion with damaged capsid, thus inhibiting genomic target amplification during qPCR [22, 23]. Factors such as viral species, dye concentration, incubation conditions, and light source for dye photoactivation influence PMAxx-qPCR efficacy and applicability [24]. In this context, this study aimed to assess PMAxx-qPCR for quantifying HAdV intact particles from different environmental matrices such as raw sewage, brackish water, and seawater.
Materials and methods
Experimental design and HAdV-positive controls
To quantify HAdV protocols using different concentrations of PMAxx (20, 50, and 100 μM) were first evaluated in three independent experiments using clinic and environmental HAdV-positive samples with known quantification. Stool samples were obtained from the Rotavirus Reference Service/Laboratory of Comparative and Environmental Virology (LVCA), Oswaldo Cruz Institute (IOC/Fiocruz) in Rio de Janeiro, Brazil, with approval of the Research Ethics Committee of the Oswaldo Cruz Foundation (CAAE number: 94144918.3.0000.5248). For analysis, stool suspensions (10% w/v) were prepared in Tris/calcium buffer (0.1 M Tris/HCl; 1.5 mM CaCl2; pH 7.2). Raw sewage, brackish, and seawater samples previously concentrated by skimmed milk flocculation method were obtained at LVCA. All clinical and environmental samples were obtained and analysed and kept at − 70 °C until used in this study.
Sampling and viral concentration method
Posteriorly, another 30 environmental samples were collected, concentrated, and analysed to quantify the presence of HAdV intact capsids using the PMAxx-qPCR established protocol. These environmental samples were collected in the Barra da Tijuca, a coastal Metropolitan Region of Rio de Janeiro City, Brazil, from 2016 to 2020 (Fig. 1). Raw sewage, brackish, and seawater samples, were concentrated by skimmed milk flocculation method described for wastewater and recreational water samples [25, 26].
PMAxx treatment
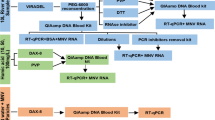
Stock solutions of PMAxx (Biotium Inc., Hayward, CA) at a concentration of 20 mM were diluted in ultrapure water to a working concentration of 500 μM and stored at – 20 °C. Different samples from each matrix were aliquoted into four fractions of 140 μL, three of them treated with PMAxx at concentrations of 20, 50, and 100 μM and one without treatment. Based on a previous study using HAdV PMAxx qPCR, we designed a protocol incubating all treated and untreated samples at room temperature for 10 min at 200 rpm in the dark, followed by photoactivation for 15 min using the blue light of the PMA-Lite™ LED Photolysis Device (Biotium) [19, 22, 24, 27]. During PMAxx treatment, the samples were protected from light. The optimal PMAxx concentration was measured by comparing the results of the treated and untreated fractions of each matrix by reducing HAdV DNA amplification in the qPCR in genomic copies (GC)/L.
For interpretation of potentially infectious HAdV by matrix, PMAxx reduction log value was defined by the formula below:
The percentage of HAdV with damaged capsids was calculated using the following formula accordingly [16]:
Results of quantification in GC/L of samples treated with PMAxx and untreated samples were compared, numerically estimating the intact and potentially infectious HAdV per matrix. The same incubation and photoactivation conditions mentioned above were applied with the 30 samples tested for the PMAxx protocol to establish after choosing the appropriate concentration.
DNA extraction and quantification
According to the manufacturer’s instructions, nucleic acids were manually extracted after the PMAxx treatment using the commercial QIAmp viral RNA mini kit (Qiagen™, Valencia, USA). The DNA extraction assay included negative control composed of DNase/RNase free water and positive controls of each matrix.
For detection and quantification of HAdV by qPCR, 25 μL reaction mixtures consisting of 12.5 μL of TaqMan Universal Master Mix II with UNG (Applied Biosystems™, Foster City, CA), 1 μL of each primer (22.5 μM), 0.5 μL of probe and 10 μL of nucleic acid. Specific HAdV primers and probes were used according to Hernroth et al. [28]. The target genomic fragments were amplified using ABI PRISM 7500 Real-Time TaqMan System. The qPCR conditions following activation of the uracil N-glycosylase (2 min, 50 °C) and activation of the AmpliTaq Gold for 10 min at 95 °C, 40 cycles (15 s at 95 °C and 1 min at 60 °C) and submitted to an infinite process at 4 °C. Negative and positive controls from nucleic acid assay and NTC were included in all procedures. All undiluted and 1:10 to 1:100 diluted nucleic acid samples were tested in triplicate to avoid inference with possible qPCR inhibitors. Standard curves were designed using gBlock gene fragments (Integrated DNA Technologies™, Coralville, Iowa, USA) with HAdV sequence fragments. Nucleic acid was quantified by Qubit Fluorometer (Thermo Fisher Scientific ™), and ten-fold dilutions were prepared and stored at – 70 °C until use. Serial dilutions (100 to 107 GC/reaction) presented the values of the slope, squared regression coefficient, and reaction efficiencies of − 3.332, 0.997, and 99.7%, respectively. Values of Ct ≤ 38 (presenting the characteristic sigmoidal curve) were considered positive.
Statistical analysis
Results were plotted using GraphPad Prism software version 8.0.1 (GraphPad Software, La Jolla, CA). A log transformation normalized data related to the number of GC/L. Differences between HAdV quantification in PMAxx-treated and untreated in different matrices were analysed using a parametric one-way ANOVA model, followed by Tukey’s multiple comparison tests (p < 0.05).
Results
Figure 2 shows the quantification by GC/L of HAdV obtained by different concentrations of PMAxx used according to the matrices evaluated. Comparing PMAxx concentrations, no significant reduction when 20, 50, or 100 μM (P > 0.986) were used as treatment independent of the matrices studied was observed, except for sewage samples. A mean value reduction of 5.01 logs GC/L was observed in this matrix when treated with 50 μM of PMAxx.
HAdV viral particles’ mean percentage revealed 99% and 95% of defective capsids in raw sewage and seawater, respectively (Table 1). In contrast, stool samples showed the lowest percentage (78%) of damaged capsid virus, followed by brackish water samples with 81%. The absence of intact HAdV particles in raw sewage was observed after the PMAxx treatment of 50 or 100 μM.
Following, 30 environmental samples, 10 from each matrix, were processed using [50 μM] PMAxx established protocol. A reduction of 0.91, 0.67, and 1.05 for raw sewage, brackish, and seawater, was observed with a mean concentration of 1.47 × 104, 6.81 × 102, and 2.33 × 102 HAdV infectious GC/L, respectively (Fig. 3). The average value percentage of HAdV viral particles revealed 88%, 79%, and 91% of defective capsids in those matrices, respectively.
Discussion
Treatment with PMA proved helpful in quantifying intact HAdV particles in all matrices evaluated, regardless of the concentration used. However, 50 μM of PMAxx presented an intermediate effect for inhibiting the signal amplification by qPCR compared to the other two concentrations, furthermore presented the best cost-benefit ratio with higher dye yield. According to recent systematic literature, many authors also used azo dyes at 50 μM in their experiments [29]. Until the completion of the present study, PMA and PMAxx were not applied to brackish and seawater matrices [29] and few studies have quantified intact HAdV particles using these dyes in environmental matrices [21, 30,31,32]. Only one study used PMAxx dye to analyse HAdV inoculated in phosphate-buffered saline (PBS) solution in thermal and free-chlorine disinfection processes [20]. The percentage of damaged capsid observed in this study was high, ranging from 69 to 100%, regardless of the concentration of PMAxx and matrix evaluated. Faeces and brackish samples showed the lowest amount of HAdV particles with damaged capsids. This quantification was confirmed when new brackish samples were evaluated, showing a variation of 157 to 1 million intact viruses, thus evidencing the high contamination of domestic sewage discarded in natura in this Lagoon used for recreational activities [33]. In seawater samples, despite the low concentration (11.3 to 1160), it was possible to detect intact HAdV. Although we cannot determine the origin of this contamination, the high concentration of those viruses in adjacent lagoons that exchange water with the sea is noteworthy. In this study, we did not assess the presence of marine currents or point and diffuse sources of faecal contamination, such as rainwater runoff and outfalls that may influence the increase or reduction of the spread and concentration of these infectious agents [34, 35]. So far, there is no study assessing HAdV PMAxx-qPCR in brackish or seawater.
The detection of HAdV intact particles with mean quantification of 103 log of HAdV GC/L in the sample treated with PMAxx, indicates the need for risk assessment studies for the exposed population to these matrices. In addition, annual risk calculations for HAdV infection in recreational waters are up to 1/1000 for a single exposure [36]. Previously, our group evidenced viral contamination on other beaches on the coast of the city of Rio de Janeiro, showing concentrations as high as 104 log of GC/L [37], thus highlighting the need for further analyses in environmental samples. It is important to note that concentrations of infectious HAdV in environmental matrices are influenced by several factors capable of changing viral stability including the concentration method and the peculiarity of each matrix. The organic flocculation method based on skimmed milk has been described as a low-cost methodology, although it presents inhibition [38], which in this study was bypassed by the dilution of the samples. The number of polluting sources of raw sewage discharge; the geomorphology and hydrology of the collection site; the physicochemical elements (temperature, pH, oxidation-reduction potential, salinity, organic, and inorganic components, among others), the presence of microorganisms with virucidal or predation action; suspended solid particles (viral aggregation), and ultraviolet light irradiation should be considered [13, 39,40,41,42].
Our results using PMAxx showed a reduction ranging from 0.91 to 5 log, unlike the result obtained in a previous study carried out for HAdV in sewage samples, where a logarithmic reduction of 0.74 was observed [22]. However, regarding the quantification of infectious HAdV, our results were similar to those found in the same study (2.69 × 104 TCID50/L and 2.27 × 105 GC/L). It is important to emphasize that although PMAxx-qPCR provides more reliable data than qPCR, it may overestimate the real numbers of infectious viruses, as described in studies of viral inactivation after heat, chlorine and UV treatments [16, 24, 31, 43, 44].
Studies using PMAxx evaluated infectious HAdV particles (serotypes 2, 5, 12, and 40) in samples of phosphate-buffered saline, food (mollusc and sausage), and environmental (soil, sewage, and freshwater from the urban river), thus demonstrating a reduction ranging from 0.42 to 4 log [19, 20, 22, 24, 27, 45]. PMA or PMAxx were also efficient in detecting another viruses, enteric or not, including SARS-CoV-2 with intact capsids in clinical, food, and environmental matrices [16, 31, 32, 43, 46,47,48,49,50,51,52,53,54] providing valuable data for environmental surveillance.
Conclusion
The presence of potentially infectious viral particles depends on each sampling, regardless of the matrix, although a higher concentration of damaged particles was evident in sewage and seawater samples. Association of this genomic intercalant with qPCR can become a quick analytical tool for screening for infectious HAdV in environmental samples, including sea and brackish water.
References
Reta DH, Tessema TS, Ashenef AS et al (2020) Molecular and immunological diagnostic techniques of medical viruses. Int J Microbiol. 2020:1–19. https://doi.org/10.1155/2020/8832728
Ding C, Liu X, Yang S (2021) The value of infectious disease modeling and trend assessment: a public health perspective. Expert Rev Anti Infect Ther. 19(9):1135–1145. https://doi.org/10.1080/14787210.2021.1882850
Larivé O, Brandani J, Dubey M, Kohn T (2021) An integrated cell culture reverse transcriptase quantitative PCR (ICC-RTqPCR) method to simultaneously quantify the infectious concentrations of eight environmentally relevant enterovirus serotypes. J Virol Methods. 296:114225. https://doi.org/10.1016/j.jviromet.2021.114225
O’Brien E, Xagoraraki I (2019) A water-focused one-health approach for early detection and prevention of viral outbreaks. One Health. 7:100094. https://doi.org/10.1016/j.onehlt.2019.100094
Sims N, Kasprzyk-Hordern B (2020) Future perspectives of wastewater-based epidemiology: Monitoring infectious disease spread and resistance to the community level. Environ Int. 139:105689. https://doi.org/10.1016/j.envint.2020.105689
Chin AWH, Chu JTS, Perera MRA et al (2020) Stability of SARS-CoV-2 in different environmental conditions. Lancet Microbe. 1(1):e10. https://doi.org/10.1016/S2666-5247(20)30003-3
Goh SG, Liang L, Gin KYH (2021) Assessment of human health risks in tropical environmental waters with microbial source tracking markers. Water Res. 207:117748. https://doi.org/10.1016/j.watres.2021.117748
da Rocha LC, Estofolete CF, Milhim BHG de A, et al. (2021) Enteric viruses circulating in undiagnosed central nervous system infections at tertiary hospital in São José do Rio Preto, São Paulo, Brazil. J Med Virol. 93(6):3539–3548. https://doi.org/10.1002/jmv.26216
Sano D, Watanabe R, Oishi W, Amarasiri M, Kitajima M, Okabe S (2021) Viral interference as a factor of false-negative in the infectious adenovirus detection using integrated cell culture-PCR with a BGM cell line. Food Environ Virol. 13(1):84–92. https://doi.org/10.1007/s12560-020-09453-x
Xagoraraki I, O’Brien E (2020) Wastewater-based epidemiology for early detection of viral outbreaks. In: O’Bannon DJ (ed) Women in Water Quality. Women in Engineering and Science. Springer International Publishing, pp 75–97. https://doi.org/10.1007/978-3-030-17819-2_5
Hamza IA, Bibby K (2019) Critical issues in application of molecular methods to environmental virology. J Virol Methods. 266:11–24. https://doi.org/10.1016/j.jviromet.2019.01.008
Greber UF (2020) Adenoviruses – infection, pathogenesis and therapy. FEBS Lett. 594(12):1818–1827. https://doi.org/10.1002/1873-3468.13849
Allard A, Vantarakis A (2019) Adenoviruses. In: Michigan State University, Rose JB, Jiménez Cisneros B, UNESCO - International Hydrological Programme (eds) Water and Sanitation for the 21st Century: Health and Microbiological Aspects of Excreta and Wastewater Management (Global Water Pathogen Project). Michigan State University. 10.14321/waterpathogens.11
Hidalgo P, González RA (2021) Adenoviruses. In: Reference Module in Biomedical Sciences. Elsevier, p B9780128187319001000. https://doi.org/10.1016/B978-0-12-818731-9.00066-5
Cuevas-Ferrando E, Randazzo W, Pérez-Cataluña A et al (2021) Viability RT-PCR for SARS-CoV-2: A Step Forward to Solve the Infectivity Quandary. Infectious Diseases (except HIV/AIDS). https://doi.org/10.1101/2021.03.22.21253818
Fuster N, Pintó RM, Fuentes C, Beguiristain N, Bosch A, Guix S (2016) Propidium monoazide RTqPCR assays for the assessment of hepatitis A inactivation and for a better estimation of the health risk of contaminated waters. Water Res. 101:226–232. https://doi.org/10.1016/j.watres.2016.05.086
Oristo S, Lee HJ, Maunula L (2018) Performance of pre-RT-qPCR treatments to discriminate infectious human rotaviruses and noroviruses from heat-inactivated viruses: applications of PMA/PMAxx, benzonase and RNase. J Appl Microbiol. Published online March 12, 2018. https://doi.org/10.1111/jam.13737
Puente H, Randazzo W, Falcó I, Carvajal A, Sánchez G (2020) Rapid selective detection of potentially infectious porcine epidemic diarrhea coronavirus exposed to heat treatments using viability RT-qPCR. Front Microbiol. 11:1911. https://doi.org/10.3389/fmicb.2020.01911
Quijada NM, Fongaro G, Barardi CRM, Hernández M, Rodríguez-Lázaro D (2016) Propidium monoazide integrated with qPCR enables the detection and enumeration of infectious enteric RNA and DNA viruses in clam and fermented sausages. Front Microbiol. 7. https://doi.org/10.3389/fmicb.2016.02008
Shirasaki N, Matsushita T, Matsui Y, Koriki S (2020) Suitability of pepper mild mottle virus as a human enteric virus surrogate for assessing the efficacy of thermal or free-chlorine disinfection processes by using infectivity assays and enhanced viability PCR. Water Res. 186:116409. https://doi.org/10.1016/j.watres.2020.116409
Randazzo W, Khezri M, Ollivier J et al (2018) Optimization of PMAxx pretreatment to distinguish between human norovirus with intact and altered capsids in shellfish and sewage samples. Int J Food Microbiol. 266:1–7. https://doi.org/10.1016/j.ijfoodmicro.2017.11.011
Leifels M, Hamza IA, Krieger M, Wilhelm M, Mackowiak M, Jurzik L (2016) From lab to lake – evaluation of current molecular methods for the detection of infectious enteric viruses in complex water matrices in an urban area. Tang P (ed) PLoS ONE 11(11):e0167105. 10.1371/journal.pone.0167105
Nocker A, Cheung CY, Camper AK (2006) Comparison of propidium monoazide with ethidium monoazide for differentiation of live vs. dead bacteria by selective removal of DNA from dead cells. J Microbiol Methods. 67(2):310–320. https://doi.org/10.1016/j.mimet.2006.04.015
Leifels M, Jurzik L, Wilhelm M, Hamza IA (2015) Use of ethidium monoazide and propidium monoazide to determine viral infectivity upon inactivation by heat, UV- exposure and chlorine. Int J Hyg Environ Health. 218(8):686–693. https://doi.org/10.1016/j.ijheh.2015.02.003
Calgua B, Rodriguez-Manzano J, Hundesa A et al (2013) New methods for the concentration of viruses from urban sewage using quantitative PCR. J Virol Methods. 187(2):215–221. https://doi.org/10.1016/j.jviromet.2012.10.012
Calgua B, Mengewein A, Grunert A et al (2008) Development and application of a one-step low cost procedure to concentrate viruses from seawater samples. J Virol Methods. 153(2):79–83. https://doi.org/10.1016/j.jviromet.2008.08.003
Fongaro G, Hernández M, García-González MC, Barardi CRM, Rodríguez-Lázaro D (2016) Propidium monoazide coupled with PCR predicts infectivity of enteric viruses in swine manure and biofertilized soil. Food Environ Virol. 8(1):79–85. https://doi.org/10.1007/s12560-015-9225-1
Hernroth BE, Conden-Hansson AC, Rehnstam-Holm AS, Girones R, Allard AK (2002) Environmental factors influencing human viral pathogens and their potential indicator organisms in the blue mussel, Mytilus edulis: the First Scandinavian Report. Appl Environ Microbiol. 68(9):4523–4533. https://doi.org/10.1128/AEM.68.9.4523-4533.2002
Leifels M, Cheng D, Sozzi E et al (2021) Capsid integrity quantitative PCR to determine virus infectivity in environmental and food applications – a systematic review. Water Res X. 11:100080. https://doi.org/10.1016/j.wroa.2020.100080
López-Gálvez F, Randazzo W, Vásquez A et al (2018) Irrigating lettuce with wastewater effluent: does disinfection with chlorine dioxide inactivate viruses? J Environ Qual. 47(5):1139–1145. https://doi.org/10.2134/jeq2017.12.0485
Randazzo W, Piqueras J, Rodríguez-Díaz J, Aznar R, Sánchez G (2018) Improving efficiency of viability-qPCR for selective detection of infectious HAV in food and water samples. J Appl Microbiol. 124(4):958–964. https://doi.org/10.1111/jam.13519
Randazzo W, Piqueras J, Evtoski Z et al (2019) Interlaboratory comparative study to detect potentially infectious human enteric viruses in influent and effluent waters. Food Environ Virol. 11(4):350–363. https://doi.org/10.1007/s12560-019-09392-2
Pedrosa de Macena LG, Castiglia Feitosa R, Vieira CB, Araújo IT, Taniuchi M, Miagostovich MP (2020) Microbiological assessment of an urban lagoon system in the coastal zone of Rio de Janeiro, Brazil. Environ Sci Pollut Res 28(1):1170–1180. https://doi.org/10.1007/s11356-020-10479-8
Ahmed W, Hamilton K, Toze S, Cook S, Page D (2019) A review on microbial contaminants in stormwater runoff and outfalls: Potential health risks and mitigation strategies. Sci Total Environ. 692:1304–1321. https://doi.org/10.1016/j.scitotenv.2019.07.055
Jiang S, Noble R, Chu W (2001) Human adenoviruses and coliphages in urban runoff-impacted coastal waters of Southern California. Appl Environ Microbiol. 67(1):179–184. https://doi.org/10.1128/AEM.67.1.179-184.2001
Crabtree KD, Gerba CP, Rose JB, Haas CN (1997) Waterborne adenovirus: a risk assessment. Water Sci Technol. 35(11-12):1–6. https://doi.org/10.2166/wst.1997.0700
Victoria M, Fumian TM, Rocha MS et al (2014) Gastroenteric virus dissemination and influence of rainfall events in urban beaches in Brazil. J Appl Microbiol. 117(4):1210–1218. https://doi.org/10.1111/jam.12592
Assis ASF, Fumian TM, Miagostovich MP, Drumond BP (2018) da Rosa e Silva ML. Adenovirus and rotavirus recovery from a treated effluent through an optimized skimmed-milk flocculation method. Environ Sci Pollut Res. 25(17):17025–17032. https://doi.org/10.1007/s11356-018-1873-x
Carratalà A, Rodriguez-Manzano J, Hundesa A et al (2013) Effect of temperature and sunlight on the stability of human adenoviruses and MS2 as fecal contaminants on fresh produce surfaces. Int J Food Microbiol. 164(2-3):128–134. https://doi.org/10.1016/j.ijfoodmicro.2013.04.007
Rexroad J, Evans RK, Middaugh CR (2006) Effect of pH and ionic strength on the physical stability of adenovirus type 5. J Pharm Sci. 95(2):237–247. https://doi.org/10.1002/jps.20496
Rigotto C, Hanley K, Rochelle PA, De Leon R, Barardi CRM, Yates MV (2011) Survival of adenovirus types 2 and 41 in surface and ground waters measured by a plaque assay. Environ Sci Technol. 45(9):4145–4150. https://doi.org/10.1021/es103922r
Ye Y, Ellenberg RM, Graham KE, Wigginton KR (2016) Survivability, partitioning, and recovery of enveloped viruses in untreated municipal wastewater. Environ Sci Technol. 50(10):5077–5085. https://doi.org/10.1021/acs.est.6b00876
Canh VD, Torii S, Yasui M, Kyuwa S, Katayama H (2021) Capsid integrity RT-qPCR for the selective detection of intact SARS-CoV-2 in wastewater. Sci Total Environ. 791:148342. https://doi.org/10.1016/j.scitotenv.2021.148342
Prevost B, Goulet M, Lucas FS, Joyeux M, Moulin L, Wurtzer S (2016) Viral persistence in surface and drinking water: Suitability of PCR pre-treatment with intercalating dyes. Water Res. 91:68–76. https://doi.org/10.1016/j.watres.2015.12.049
Bae S, Wuertz S (2012) Survival of host-associated bacteroidales cells and their relationship with Enterococcus spp., Campylobacter jejuni, Salmonella enterica Serovar Typhimurium, and Adenovirus in freshwater microcosms as measured by propidium monoazide-quantitative PCR. Appl Environ Microbiol. 78(4):922–932. https://doi.org/10.1128/AEM.05157-11
Chen NT, Cheong NS, Lin CY, Tseng CC, Su HJ (2021) Ambient viral and bacterial distribution during long-range transport in Northern Taiwan. Environ Pollut Barking Essex 1987 270:116231. https://doi.org/10.1016/j.envpol.2020.116231
Fittipaldi M, Rodriguez NJP, Codony F, Adrados B, Peñuela GA, Morató J (2010) Discrimination of infectious bacteriophage T4 virus by propidium monoazide real-time PCR. J Virol Methods. 168(1-2):228–232. https://doi.org/10.1016/j.jviromet.2010.06.011
Guix S, Fuentes C, Pintó RM et al (2020) Infectivity of norovirus GI and GII from bottled mineral water during a waterborne outbreak, Spain. Emerg Infect Dis. 26(1):134–137. https://doi.org/10.3201/eid2601.190778
Ho J, Seidel M, Niessner R, Eggers J, Tiehm A (2016) Long amplicon (LA)-qPCR for the discrimination of infectious and noninfectious phix174 bacteriophages after UV inactivation. Water Res. 103:141–148. https://doi.org/10.1016/j.watres.2016.07.032
Hong W, Xiong J, Nyaruaba R et al (2021) Rapid determination of infectious SARS-CoV-2 in PCR-positive samples by SDS-PMA assisted RT-qPCR. Sci Total Environ. 797:149085. https://doi.org/10.1016/j.scitotenv.2021.149085
Karim MR, Fout GS, Johnson CH, White KM, Parshionikar SU (2015) Propidium monoazide reverse transcriptase PCR and RT-qPCR for detecting infectious enterovirus and norovirus. J Virol Methods. 219:51–61. https://doi.org/10.1016/j.jviromet.2015.02.020
Kodama D, Tanaka M, Matsuzaki T et al (2020) Inhibition of ABL1 tyrosine kinase reduces HTLV-1 proviral loads in peripheral blood mononuclear cells from patients with HTLV-1-associated myelopathy/tropical spastic paraparesis. PLoS Negl Trop Dis. 14(7):e0008361. https://doi.org/10.1371/journal.pntd.0008361
Parshionikar S, Laseke I, Fout GS (2010) Use of propidium monoazide in reverse transcriptase PCR to distinguish between infectious and noninfectious enteric viruses in water samples. Appl Environ Microbiol. 76(13):4318–4326. https://doi.org/10.1128/AEM.02800-09
Wei X, Ge T, Wu C et al (2021) T4-like phages reveal the potential role of viruses in soil organic matter mineralization. Environ Sci Technol. 55(9):6440–6448. https://doi.org/10.1021/acs.est.0c06014
Acknowledgements
The authors thank the technical and field staff Márcia Maria Araújo Pimenta and Sérgio de Silva e Mouta Júnior (Oswaldo Cruz Foundation, RJ, Brazil) for helping with the environmental sample collection and processing. Management of Genetic Heritage and Associated Traditional Knowledge-SisGen”, in compliance with the Brazilian Law N. 13123/2015 and its regulations, under registration number ACD04AA. PDTIS DNA Sequence Platform staff at FIOCRUZ for technical support in sequencing reactions. This research study is under the scope of the activities of the Oswaldo Cruz Foundation (FIOCRUZ) as a Collaborating Centre of PAHO/WHO of Public and Environmental Health.
Funding
This work was supported by Instituto Oswaldo Cruz (PAEF), Fundação Carlos Chagas Filho de Amparo à Pesquisa do Estado do Rio de Janeiro (Faperj-E-26/202.821/2018), Conselho de Desenvolvimento Científico e Tecnológico (CNPq-Universal-406414/2016-5), Coordenação de Aperfeiçoamento de Pessoal de Nível Superior-Brasil (CAPES)-Finance Code 001.
Author information
Authors and Affiliations
Contributions
Conceptualization: Lorena da Graça Pedrosa de Macena, and Marize Pereira Miagostovich. Data curation: Lorena da Graça Pedrosa de Macena, Natália Maria Lanzarini and Adriana Gonçalves Maranhão. Formal analysis: Lorena da Graça Pedrosa de Macena, Natália Maria Lanzarini, and Marize Pereira Miagostovich. Investigation: Lorena da Graça Pedrosa de Macena, and Joseane Simone de Oliveira Pereira. Methodology: Lorena da Graça Pedrosa de Macena, Joseane Simone de Oliveira Pereira, Jansen Couto da Silva, Adriana Gonçalves Maranhão, and Fernando César Ferreira. Supervision: Marize Pereira Miagostovich. Visualization: Lorena da Graça Pedrosa de Macena and Marize Pereira Miagostovich. Writing—original draft: Lorena da Graça Pedrosa de Macena. Writing—review and editing: Lorena da Graça Pedrosa de Macena and Marize Pereira Miagostovich. Funding acquisition: Marize Pereira Miagostovich. All authors read and approved the final manuscript.
Corresponding author
Ethics declarations
Ethics approval and consent to participate
Not applicable.
Consent for publication
Not applicable.
Conflict of interest
The authors declare that they have no conflict of interest.
Additional information
Responsible Editor: Jônatas Abrahão
Publisher’s note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
About this article
Cite this article
Pedrosa de Macena, L.d., Pereira, J.S.d., da Silva, J.C. et al. Quantification of infectious Human mastadenovirus in environmental matrices using PMAxx-qPCR. Braz J Microbiol 53, 1465–1471 (2022). https://doi.org/10.1007/s42770-022-00775-5
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s42770-022-00775-5