Abstract
Purpose
Based on medical reports, it is hard to find levels of different hospitalized symptomatic COVID-19 patients according to their features in a short time. Besides, there are common and special features for COVID-19 patients at different levels based on physicians’ knowledge that make diagnosis difficult. For this purpose, a hierarchical model is proposed in this paper based on experts’ knowledge, fuzzy C-mean (FCM) clustering, and adaptive neuro-fuzzy inference system (ANFIS) classifier.
Methods
Experts considered a special set of features for different groups of COVID-19 patients to find their treatment plans. Accordingly, the structure of the proposed hierarchical model is designed based on experts’ knowledge. In the proposed model, we applied clustering methods to patients’ data to determine some clusters. Then, we learn classifiers for each cluster in a hierarchical model. Regarding different common and special features of patients, FCM is considered for the clustering method. Besides, ANFIS had better performances than other classification methods. Therefore, FCM and ANFIS were considered to design the proposed hierarchical model. FCM finds the membership degree of each patient’s data based on common and special features of different clusters to reinforce the ANFIS classifier. Next, ANFIS identifies the need of hospitalized symptomatic COVID-19 patients to ICU and to find whether or not they are in the end-stage (mortality target class). Two real datasets about COVID-19 patients are analyzed in this paper using the proposed model. One of these datasets had only clinical features and another dataset had both clinical and image features. Therefore, some appropriate features are extracted using some image processing and deep learning methods.
Results
According to the results and statistical test, the proposed model has the best performance among other utilized classifiers. Its accuracies based on clinical features of the first and second datasets are 92% and 90% to find the ICU target class. Extracted features of image data increase the accuracy by 94%.
Conclusion
The accuracy of this model is even better for detecting the mortality target class among different classifiers in this paper and the literature review. Besides, this model is compatible with utilized datasets about COVID-19 patients based on clinical data and both clinical and image data, as well.
Highlights
• A new hierarchical model is proposed using ANFIS classifiers and FCM clustering method in this paper. Its structure is designed based on experts’ knowledge and real medical process. FCM reinforces the ANFIS classification learning phase based on the features of COVID-19 patients.
• Two real datasets about COVID-19 patients are studied in this paper. One of these datasets has both clinical and image data. Therefore, appropriate features are extracted based on its image data and considered with available meaningful clinical data. Different levels of hospitalized symptomatic COVID-19 patients are considered in this paper including the need of patients to ICU and whether or not they are in end-stage.
• Well-known classification methods including case-based reasoning (CBR), decision tree, convolutional neural networks (CNN), K-nearest neighbors (KNN), learning vector quantization (LVQ), multi-layer perceptron (MLP), Naive Bayes (NB), radial basis function network (RBF), support vector machine (SVM), recurrent neural networks (RNN), fuzzy type-I inference system (FIS), and adaptive neuro-fuzzy inference system (ANFIS) are designed for these datasets and their results are analyzed for different random groups of the train and test data;
• According to unbalanced utilized datasets, different performances of classifiers including accuracy, sensitivity, specificity, precision, F-score, and G-mean are compared to find the best classifier. ANFIS classifiers have the best results for both datasets.
• To reduce the computational time, the effects of the Principal Component Analysis (PCA) feature reduction method are studied on the performances of the proposed model and classifiers. According to the results and statistical test, the proposed hierarchical model has the best performances among other utilized classifiers.
Graphical Abstract

Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Severe acute respiratory syndrome virus strain reported as causing the respiratory disease COVID-19 (Gorbalenya et al. 2020). The World Health Organization (WHO) has issued a global warning about the outbreak of the coronavirus (Chan et al. 2020; WHO 2020). This disease is reported to be very aggressive and it has affected millions of people in all countries (Ivanov 2020; Rahimi et al. 2020). Attempts have been made around the world to deal with COVID-19 from a variety of medical, psychological, and engineering perspectives (Pan et al. 2020). Despite suggestions for treatment and prevention of the disease, there is still no definitive method against this disease (Di Lorenzo et al. 2020; Saghazadeh and Rezaei 2020). Quarantine and social distance have been proposed as preventive measures for cities with high prevalence (Peak et al. 2020; Thu et al. 2020). Several advanced epidemiological models are designed to provide prevention, detection, and prediction approaches and present the effects of this disease (Koolhof et al. 2020). The impact of underlying diseases such as cardiovascular disease on COVID-19 patients has been studied (Bansal 2020). The virus can also increase the risk of death in people with chronic obstructive pulmonary disease, diabetes mellitus, and kidney failure (Rahmani and Mirmahaleh 2021). Some studies have shown that pregnant women suffer worse than non-pregnant women (Fan et al. 2020). For predicting possibly unknown ncRNA-disease relationships used multi-type hierarchical clustering (Barracchia et al. 2020). Some papers aim to find possible treatments or drug/gene-disease association clustering ( Loucera et al. 2020). These articles show the high complexity of this disease for different patients. However, the exact medical method for diagnosing different levels of symptomatic COVID-19 patients has not yet been identified. On the other hand, accurate identification of relationships among patient characteristics diagnoses by physicians requires a long time. On the other hand, machine learning methods can learn many relationships among different features of asymptomatic and symptomatic COVID-19 patients based on their data. These methods present good performances for diagnosing and predicting the prevalence of COVID-19 (Rekha 2020). It is noteworthy that radiological imaging techniques, X-rays, and CT scans are great complements for the diagnosis of different levels of symptomatic and asymptomatic COVID-19 patients (Rubin et al. 2020; Shi et al. 2020). Machine learning methods could consider different features of COVID-19 patients along with their image data to detect their levels.
Therefore, a new hierarchical model is proposed in this paper to find different levels of symptomatic COVID-19 patients. Detecting the need for symptomatic COVID-19 patients to ICU and finding whether or not they are in the end-stage are important targets. In this regard, two real datasets about COVID-19 patients are selected and different features of them are studied to choose suitable features through available features. One of these datasets has both clinical and image data about COVID-19 patients. Therefore, some image processing and deep learning methods are utilized to extract meaningful features from available images. It should be noted that physicians first utilize clinical features to detect the severity level of COVID-19 patients. Some symptomatic COVID-19 patients are returned to their homes after examination and training and the rest are hospitalized based on medical diagnoses. Then, physicians decide on the need for hospitalized symptomatic COVID-19 patients to ICU and whether or not they are in the end-stage according to their clinical and image data. Accordingly, the proposed model in this paper is quite analogous to what happens in reality.
Then, different types of well-known classification methods including case-based reasoning (CBR), decision tree (DT), convolutional neural networks (CNN), K-nearest neighbors (KNN), learning vector quantization (LVQ), multi-layer perceptron (MLP), Naive Bayes (NB), radial basis function network (RBF), support vector machine (SVM), recurrent neural networks (RNN), fuzzy type-I inference system (FIS), and adaptive neuro-fuzzy inference system (ANFIS) are designed for these datasets and their results are analyzed for different random groups of the train and test data. The best classifier for these datasets is selected based on various indicators including accuracy, sensitivity, specificity, precision, F-score, and G-mean. ANFIS has most of the best performances for both datasets among other classifiers. Besides, fuzzy C-Mean (FCM) clustering method is utilized to cluster different groups of COVID-19 patients and reinforce ANFIS classification learning. It is noteworthy that there are various linear and nonlinear relationships between the features of patients and the class. It greatly increases the complexity of classification learning. Consequently, the FCM clustering method is utilized to deal with it. The divide-to-conquer approach imposed by the FCM clusters allows classifiers to be more specialized in COVID-19 subpopulations, typically leading to higher classification performance.
It is noteworthy that one of the selected datasets has image data. Therefore, some features are extracted based on image data and added to other clinical data. Then, useless features among all features are reduced using principal component analysis (PCA) method to improve the results of classification. The computational results verify the superiority of the proposed hierarchical model compared to other utilized classifiers in this paper. Moreover, the results of the Wilcoxon signed-rank test prove the effectiveness of the proposed extracted features based on images and the PCA feature reduction method. This model is compatible with both datasets about COVID-19 patients based on clinical data and both clinical and image data, as well.
The rest of the article is as follows. The “Methods” section is devoted to reviewing the literature of relevant studies. Next, the selected datasets, utilized methods, the proposed hierarchical model, and the different types of extensions are represented in the “Results” section. The clinical and image data of datasets are introduced and the relevant experimental results are presented in the “Discussion” section. The last section is about conclusions and future studies.
Literature review
Many diseases affected different people in the world. There are different linear and nonlinear relationships among features of patients that create complexities for medical treatments. Therefore, machine learning methods are widely utilized in these fields to find better diagnoses and treatment plans (Ershadi and Seifi 2020a, b). These methods are appropriate to cope with detecting infectious diseases, especially in COVID-19, a widely spread pandemic in the world. In the following, there is a quick review of the most important research studies that are merely focused on detecting different diseases using learning approaches.
Machine learning methods present good performances for differential diagnosis of COVID-19 (Dai et al. 2020; Rauschecker et al. 2020). By machine learning and using data from 29 patients at Tongji Hospital in China, another developed algorithm is proposed to find the mortality risk of infected COVID-19 patients (Yan et al. 2020). On other hand, these methods present better performances in comparison with medical diagnoses based on experts’ knowledge (Ershadi and Seifi 2020a, b). Another study focuses on the diagnosis of diabetes type-II patients and proposes a hybrid machine learning-based ensemble model for this purpose (Sarwar et al. 2020). Heart disease diagnosis is studied using a new expert system based on a fuzzy Bayesian network. This study presents the advantages of a machine learning method based on experts’ knowledge (Zarandi et al. 2017).
Another study uses medical information such as age, sex, income level, place of residence, household type, disability, respiratory symptoms, route of infection, and medical background to extract new meaningful features for COVID-19 diagnosis. The extracted features can improve classification methods learning (including RBF, SVR, and KNN) in comparison with clinical data (An et al. 2020). New extracted features are considered in another paper for COVID-19 patents, as well. This paper showed that machine learning methods can save radiologists time for diagnosis and can be more cost-effective than standard COVID-19 tests (Bullock et al. 2020). Another method is proposed for accurate and automatic diagnosis of COVID-19 patients based on advanced artificial intelligence using chest CT. This method can classify the chest image with high accuracy according to extensive computational results (Ozturk et al. 2020). A clinical study with 1014 patients in Wuhan obtained chest CT with 60% accuracy, 97% sensitivity, and 25% specificity for COVID-19 diagnosis (Ai et al. 2020). The convolutional neural network model is proposed for the automatic diagnosis of COVID-19 from chest X-ray images of patients (Elaziz et al. 2020). The diagnosis of respiratory decompensation in COVID-19 patients is studied in another article using a machine-learning approach (Burdick et al. 2020). Other most recently published papers in this area and their comparison in terms of different features including the type of utilized learning algorithms and the modality of images are reported in Table 1.
According to the literature review, the diagnosis of COVID-19 patients is helpful and most papers proposed different machine-learning methods for this purpose (Lalmuanawma et al. 2020). However, the increasing number of COVID-19 patients in different countries in the world lead to many problems in hospitalization. COVID-19 disease has new symptoms and effects in comparison with other infectious diseases. They make it difficult to find suitable medical treatments for hospitalized COVID-19 patients. Accordingly, several critical evaluations of the significant papers in the literature are in order. Although the proposed methods in the literature are applicable for disease diagnosis, finding different levels of hospitalized symptomatic COVID-19 patients is ignored as the first point (see Table 1 and Appendix). Secondly, evaluations of the proposed models are limited to accuracies in most papers, and other measurements including sensitivity and specificity. are neglected. Third, COVID-19 datasets without images and few classifiers are considered in most papers. These points lead to different research gaps. At first, the COVID-19 diagnosis cannot support physicians to find the correct levels of COVID-19 patients. Besides, the accuracies of a few classifiers are evaluated regardless of their different measurements and performances of other types of classifiers.
Contributions
According to the literature, contributions are summarized as follows to close part of mentioned gaps:
-
1-
A new hierarchical model is proposed using ANFIS classifiers and FCM clustering method in this paper. Its structure is designed based on experts’ knowledge and real medical process. FCM reinforces the ANFIS classification learning phase based on the features of COVID-19 patients
-
2-
Two real datasets about COVID-19 patients are studied in this paper. One of these datasets has both clinical and image data. Therefore, appropriate features are extracted based on its image data and considered with available meaningful clinical data. Different levels of hospitalized symptomatic COVID-19 patients are considered in this paper including the need of patients to ICU and whether or not they are in the end-stage
-
3-
Well-known classification methods including case-based reasoning (CBR), decision tree, convolutional neural networks (CNN), K-nearest neighbors (KNN), learning vector quantization (LVQ), multi-layer perceptron (MLP), Naive Bayes (NB), radial basis function network (RBF), support vector machine (SVM), recurrent neural networks (RNN), fuzzy type-I inference system (FIS), and adaptive neuro-fuzzy inference system (ANFIS) are designed for these datasets and their results are analyzed for different random groups of the train and test data
-
4-
According to unbalanced utilized datasets, different performances of classifiers including accuracy, sensitivity, specificity, precision, F-score, and G-mean are compared to find the best classifier. ANFIS classifiers have the best results for both datasets
-
5-
To reduce the computational time, the effects of the principal component analysis (PCA) feature reduction method are studied on the performances of the proposed model and classifiers. According to the results and statistical test, the proposed hierarchical model has the best performances among other utilized classifiers.
Methods
Before getting into the details of the proposed hierarchical model, some brief explanations about the scheme of research, selected datasets, data preprocessing, utilized feature reduction, classifications, FCM clustering method, and image processing are presented in the following to make this paper self-contained.
Schema of research
To illustrate the utilized schema in this research, Fig. 1 is represented as follows.
Selected datasets
Two real datasets about symptomatic COVID-19 patients are utilized in this paper as follows:
-
The first dataset about COVID-19 patients is the “COVID-19 patient pre-condition dataset.” This dataset was obtained from the Mexican government dataset and includes 566,602 patients with 23 clinical features (Mukherjee 2020) [https://www.kaggle.com/tanmoyx/covid19-patient-precondition-dataset].
-
The second dataset is “Chest Imaging with Clinical and Genomic Correlates Representing a Rural COVID-19 Positive Population (COVID-19-AR).” This dataset has clinical and image data of about 105 patients (Desai et al. 2020a, b) [https://wiki.cancerimagingarchive.net/pages/viewpage.action?pageId=70226443#70226443bcab02c187174a].
There is a collection of radiographic and CT imaging studies for patients who tested positive for COVID-19 in the second dataset. Clinical data correlates with key radiology for every patient from the same population. Detailed descriptions of the second dataset are demonstrated in Table 2.
The size of the first dataset is less than 50 MB and there is no image data about patients in this dataset. These datasets help to evaluate the performances of the proposed model for only clinical data and clinical with image data. It is noteworthy that the compatibility of classification models with image data is important for some diseases, specially COVID-19 due to its characteristics.
Data preprocessing
There are both quantitative and qualitative clinical features in selected datasets. First, the qualitative features are categorized based on related items in the attribute to find new quantitative features. Secondly, the missing features for the first dataset are deleted due to the large number of available data in this dataset. However, missing features of the second dataset are replaced with the average of available features due to the limited number of patients in the second dataset. In the third step, features with equal values (e.g., name of the hospital) and features with unique values (e.g., patient ID) for all patients are eliminated in both datasets because these columns had no meaningful information for the decision-making process. Moreover, remained features of clinical data have different dimensions and scales in both datasets. Therefore, they must be normalized and descaled such that their values state between (0, 1) in the fourth step. Finally, the prepared datasets are divided into two different sections called to train and test data in the fifth step. It is noteworthy that these sections are selected randomly for both datasets to have less dependency on the data.
Utilized feature reduction, classification, and clustering methods
Principal component analysis
PCA is one of the feature reduction methods that find eigenvalues of the dataset’s features. Then, eigenvectors are defined based on the eigenvalues to represent meaningful data features. In other words, PCA assigns principal components for each feature based on eigenvectors to make a new expressive dataset. The dimensions of the new space are usually designed based on some of the features with the most eigenvalues. Finally, all data are generated in a new space with fewer features and no correlation. Therefore, these new features may increase the performances of classifiers as the irrelevant features have been omitted. In this paper, some of the top components with the highest eigenvalues are considered to generate a new dataset such that they have more than 99% of the variations.
Case-based reasoning
CBR tries to find the class of test data based on its similarity to other data. In this paper, Euclidean distances between different features of two data are considered as the similarity. The minimum number of similar data, related weights, and selection type of CBR were selected and optimized using the design of experiments (DoE) and Taguchi method before applying this classifier in this paper.
Decision tree
This classifier considers different subsets for a dataset in the form of a tree structure. Creating decision nodes and leaf nodes in this method is obtained by combining mentioned subsets. Every decision node has two or more branches and every leaf node presents a decision. There are different algorithms to make a decision tree for a dataset. The C4.5 algorithm is utilized in this paper for this purpose. Criterion type, splitter, maximum depth, minimum samples for split, minimum samples for leaf, minimum weight fraction, maximum features, related impurities, and class weight of this classifier were selected and optimized using DoE and Taguchi method before applying it in this paper.
Convolutional neural network
CNN is a class of artificial neural networks based on the shared-weight architecture of the convolution kernels or filters that slide along input features and provide translation equivariant responses known as feature maps. CNNs take advantage of the hierarchical pattern in data and assemble patterns of increasing complexity using smaller and simpler patterns embossed in their filters. Kernel size, number of filters, filter size, activation function, pooling type, and its size of this classifier were selected and optimized using DoE and Taguchi method before applying this classifier in this paper.
K-Nearest neighbors
KNN is one of the instance-based learning methods that consider the distances between different features of test data and some train data to find the appropriate class. Therefore, different methods for calculating the distance can be used in this method. Euclidean distance is considered in this paper for this purpose. The number of neighbors, weights, learning algorithm type, leaf size, and metric of KNN were selected and optimized using DoE and Taguchi method before applying this classifier in this paper.
Learning vector quantization
LVQ is one of the artificial neural network methods based on the winner-take-all approach. The type of utilized artificial neural networks in this classification method is self-organizing maps that consider a learning algorithm to find the best winner for every test data. Margins, likelihood ratio, distance type, related kernel, dissimilarities, and learning type of LVQ were selected and optimized using DoE and Taguchi method before applying this classifier in this paper.
Multi-layer perceptron
MLP is one of the feedforward artificial neural network methods that consider different layers of perceptron with special threshold activation for them. It utilizes backpropagation as its supervised learning technique. MLP classifiers present good performances to detect data whether or not they are linearly separable. Training methods, size of hidden layers, activation type, learning type, solver function, alpha, beta, epsilon, batch size, learning rate, maximum iteration, shuffle, verbose, and validation fraction of MLP were selected and optimized using DoE and Taguchi method before applying this classifier in this paper.
Naive Bayes
NB is one of the probabilistic classifiers based on the Bayesian theorem. It considers independence assumptions between the features and finds a suitable probability associated with each feature category. This classifier presents acceptable performances with simple assumptions in different applications. Parameter estimation type, variation smoothing function, and divisions’ size of NB were selected and optimized using DoE and Taguchi method before applying this classifier in this paper.
Radial basis function network
RBF is one of the artificial neural networks with activation functions based on radial basis functions. The class of every test data is a linear combination of radial basis functions of its features in this method. The size of hidden layers, activation type, batch size, maximum iteration, shuffle, and verbose of RBF were selected and optimized using DoE and Taguchi method before applying this classifier in this paper.
Support vector machine
SVM classifiers try to find the best hyperplanes among different classes of train data of a dataset. This hyperplane has the highest distance to the nearest training-data point of different classes. Then, the class of every test data is defined based on its position and defined hyperplanes. Kernels, degree, gamma, shrinking, probability, class weight, verbose, maximum iteration, decision function shape, and penalty cost of SVM were selected and optimized using DoE and Taguchi method before applying this classifier in this paper.
Recurrent neural network
An RNN is a CNN where connections between nodes form a directed or undirected graph along a temporal sequence. In other words, RNN works on the principle of saving the output of a particular layer and feeding this back to the input in order to predict the output of the layer. Kernel size, number of filters, filter size, activation function, pooling type and the size of this classifier were selected and optimized using DoE and Taguchi method before applying this classifier in this paper.
Fuzzy type-inference system
FIS classifiers are one type of fuzzy classifiers based on fuzzy logic applications. They consider some if–then rules according to the Takagi–Sugeno method and define fuzzy membership functions for different features of a dataset. Finally, features of test data are analyzed based on extracted rules, and the best classes are selected for them. The number of fuzzy rules, type of related fuzzy membership functions, number of neurons in different layers, connection weights, and summation function type of FIS were selected and optimized using DoE and Taguchi method before applying this classifier in this paper.
Adaptive neuro-fuzzy inference system
ANFIS is one of the artificial neural networks based on the Takagi–Sugeno fuzzy inference system. It utilized the advantages of both fuzzy logic principles and artificial neural networks. The artificial neural network in ANFIS tries to find the optimal parameters for Takagi–Sugeno fuzzy inference system using a supervised-based learning approach. The number of neurons in different layers, activation type, learning type, alpha, beta, epsilon, batch size, learning rate, maximum iteration, related membership functions, connection weights, and summation function type of ANFIS were selected and optimized using DoE and Taguchi method before applying this classifier in this paper.
Fuzzy C-mean
Clustering methods attempt to find different clusters for some data such that all data in the same cluster have the most similarity (compactness measure) and data in different clusters are as dissimilar as possible (separation measure). However, FCM clustering defines the membership function to find the membership degree of each data to each cluster. Therefore, each data can belong to more than one cluster. It leads to more computational capabilities for this soft clustering method. The number of clusters, maximum iteration, minimum threshold, array exponentiation applied to the membership function, initial fuzzy c-partitioned matrix, initial seed, and stopping criterion of FCM were selected and optimized using DoE and Taguchi method before applying this classifier in this paper.
Preprocessing for images
There are two different datasets in this study. The first one is based on clinical data and the second one has both clinical and image data. Clinical data of both datasets are studied in this paper using the mentioned classifiers and the proposed hierarchical model. Besides, some features are extracted based on the image data of the second dataset and added to its available features. Therefore, a new dataset is created based on its clinical features and the new extracted features based on the chest radiograph images. Then, the performances of utilized classifiers and the proposed hierarchical model are obtained for the mentioned new dataset to find the effects of added features on their learning phase. The performances of utilized classifiers and the proposed hierarchical model can indicate whether or not the extracted features from the images are effective to find different levels of hospitalized symptomatic COVID-19 patients. In this subsection, preprocessing steps for radiology images of the second dataset are presented as follows:
-
1)
The suitable chest radiograph image for each patient is selected in the first step. Given that there are several images for each patient, the image with the greatest number of white pixels in the chest area is considered. This image is related to the most sensitive condition of any patient with the greatest number of dead cells in his/her lung. It is noteworthy that some areas in the photo, the angle, and the intensity of the images are similar for different patients
-
2)
In the second step, the additional information from the chest radiograph images including shoulder, non-lung tissues, and the separate parts of the body are eliminated using appropriate cutting operations
-
3)
All of the noises through the intensity adjustment function are deleted to make the pictures brighter. Then, all pictures changed to gray-level pictures
-
4)
An FCM clustering method is utilized to find the three clusters on the final image from the previous step. These clusters select different pixels based on their values and their centers are near 0, 0.5, and 1, respectively. The results show that the last two clusters are more suitable than the first cluster
-
5)
Make black and white segments for each image using the threshold \(=\frac{(\max_{\mathrm{lable}\;\mathrm{cluster}=2}\mathrm{gray}-\mathrm{level}\;\mathrm{value}+\min_{\mathrm{lable}\;\mathrm{cluster}=3}\mathrm{gray}-\mathrm{level}\;\mathrm{value})}2\)
After performing the above steps, some features are extracted based on prepared images as presented in the following subsection.
Extracted features based on images
Coronavirus attacks different parts of the body and causes inflammation according to physicians and experts’ knowledge. The lung is ground zero for COVID-19 and its healthy cells are affected by occurred inflammations. This causes the death of respiratory cells in different parts of the lungs. Therefore, the white parts of the chest radiograph image in the lungs of injured people are more than healthy people. As a result, it is necessary to extract features from the image that are sensitive to its white pixels. Consequently, some statistical features are extracted according to the prepared images. Let consider an image of size \(MN\) and \(p(i,j)\) to demonstrate the pixel value in point \((i,j)\). Then, the mean is calculated using Eq. (1) as the first statistical feature. It can support classifiers to find more general sensitivity in the image.
The second statistical feature is entropy to show distribution diversity as represented in Eq. (2). It defines the distribution diversity of an image based on determining the number of pixels with a special level.
In this equation, \({z}_{k}\) is the total number of pixels with the level \(k\), and \(L\) is the total number of levels.
Skewness is the third statistical feature represented in Eq. (3). It shows the degree of asymmetry of a pixel in the window specified around its distribution average. Regarding this feature, we could consider asymmetry of special scatter pixel around its distribution average.
For the fourth statistical feature, the rate of flatness of distribution relative to a normal distribution is considered as kurtosis measurement and represented in Eq. (4). This feature defines behaviors of special scatter pixel around its distribution average.
The fifth feature extracted from prepared images is standard deviation as represented in Eq. (5). This feature determines the degree of COVID-19 spread in patients’ lung.
The proposed hierarchical model
In real medical diagnosis, experts considered a special set of features for different groups of COVID-19 patients to find their treatment plans. Physicians pay attention to special features of different groups of patients and classify them. This approach is considered in the structure of the proposed hierarchical model. In the proposed model, we applied clustering methods to patients’ data to determine some clusters. Then, we learn classifiers for each cluster in a hierarchical model. FCM and ANFIS were considered to design the proposed hierarchical model due to the following details.
ANFIS is a classifier that utilizes an adaptive neural network to search optimal parameters systematically for learning the Takagi–Sugeno-type fuzzy model. This classification technique is utilized to diagnose different diseases and cancers in most cases in the literature. ANFIS consists of 5 layers and represents good performances for many applications due to using advantages of both fuzzy logic principles and artificial neural networks. Let (\({x}_{1},\dots ,{x}_{n}\)) are \(n\) inputs, \((R=m)\) are \(m\) fuzzy rules, and \({I}_{k}^{l}\) are output of \({k}^{th}\) node in \({l}^{th}\) layer. Then, the structure of utilized ANFIS in this paper is presented in Fig. 2.
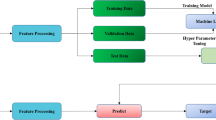
There are \((n*m)\) nodes and \({I}_{k}^{1}={\mu }_{i}\left({x}_{j}\right)={\left(1+{\left|\frac{{x}_{j}-{c}_{k}}{{a}_{k}}\right|}^{2{b}_{k}}\right)}^{-1}\forall i,j\in 1,\dots ,n , k\in 1,\dots ,n*m\) is membership degree of \({j}^{th}\) input for \({i}^{th}\) rule in layer1. Premise parameters (\({a}_{k}\),\({b}_{k}\), and\({c}_{k}\)) change the shapes of the utilized bell-shaped membership function in this layer. The number of nodes in layer2 is the same as \(m\) fuzzy rules and \({I}_{k}^{2}={w}_{i}=\prod_{j=1}^{N}{\mu }_{i}\left({x}_{j}\right) \forall k=i\in 1,\dots ,m\) is firing strength of \({i}^{th}\) fuzzy rule. The number of nodes in layer3 is the same as \(m\) fuzzy rules and \({I}_{k}^{3}=\overline{{w }_{i}}=\frac{{w}_{i}}{{\sum }_{i=1}^{m}{w}_{i}} \forall k=i\in 1,\dots ,m\) is the firing strength of \({i}^{th}\) rule after normalization. The number of nodes in layer4 is the same as \(m\) fuzzy rules and the \({k}^{th}\) node of this layer finds the contribution of \({i}^{th}\) rule towards the all output with \({I}_{k}^{4}={\overline{{w }_{i}}f}_{i}=\frac{{w}_{i}}{{\sum }_{i=1}^{m}{w}_{i}}({\sum }_{j=1}^{N}{p}_{i,j}{x}_{j}+{r}_{i}) \forall k=i\in 1,\dots ,m\) node function based on \({p}_{i,j}\) and, \({r}_{i}\) as consequent parameters. Finally, the single node in layer5 finds\({I}_{5}={\sum }_{i=1}^{m}{\overline{{w }_{i}}f}_{i}\). Therefore, the training process of ANFIS attempts to find the best values of mentioned parameters. Typical ANFIS and other classifiers have the following process in Fig. 3 for the diagnosis of different diseases and cancers in this paper.
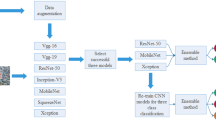
Although ANFIS has the best performances among all utilized classifications in this paper for finding appropriate levels for hospitalized symptomatic COVID-19 patients, its performances have significant weaknesses in some computational experiments. After careful examination, we found different groups of COVID-19 patients and each group has specific characteristics. However, there are a lot of common characteristics among different groups of COVID-19 patients. These variations make the decision-making process very hard to find the appropriate levels for symptomatic COVID-19 patients. To cope with this problematic issue, an FCM clustering method is applied before ANFIS classification to clustering COVID-19 patients. Other clustering methods assign each data to a single cluster, but there are a lot of common characteristics among different groups of COVID-19 patients. Therefore, we select FCM to cluster these patients. FCM determines the membership degree of each data to every cluster. Besides, the threshold equal to 0.05 is defined to eliminate some data from the cluster(s) whose membership degrees are less than 0.05. It leads to fewer complexities in computational calculations. Then, ANFIS classification should be performed separately for all COVID-19 patients belonging to each cluster. It is noteworthy that each patient is found in different clusters according to the characteristics of the FCM method. Figure 4 represents more details about it. There are 4 diverse groups of COVID-19 patients in this figure and the FCM tries to learn classifiers based on its results.
Therefore, the outputs of all ANFIS classifications for different clustering are combined using the following algorithm.
1. Find the appropriate number of clusters (or \(c\)) for FCM in each dataset using (average Euclidean distances of clusters centers or separation-average Euclidean distance of the data within the clusters or compactness) among different numbers of clusters (between 2 and \(\sqrt{\mathrm{all}\;\mathrm{record}\;\mathrm{in}\;\mathrm{each}\;\mathrm{dataset}}\)) based on train data |
2. Find the membership degree of data/patient \(i\) in cluster \(j\), \({u}_{ji}=\frac{1}{\sum_{k=1}^{c}{(\frac{{d}_{ji}}{{d}_{jk}})}^{={~}^{2}\!\left/ \!{~}_{(m-1)}\right.}}\) where \(c\) is the number of clusters, \(m\) is equal to 2, \({d}_{ji}\) is Euclidean distance of \({i}^{th}\) data from the center of \({j}^{th}\) cluster, and \({d}_{jk}\) is Euclidean distance of \({j}^{th}\) cluster of \({k}^{th}\) cluster \((k=1,\dots ,c \mathrm{except} j)\) |
3. Find all \({u}_{ji}\) less than the mentioned threshold (0.05) and replace them by zero |
4. Learn a special ANFIS classifier for each cluster based on its train data. It is noteworthy that \({i}^{th}\) data belongs to \({j}^{th}\) cluster if its membership degree (\({u}_{ji}\)) is greater than zero |
5. Find the appropriate clusters or cluster \((J)\) for each test data (\(t\in T)\) and feed its feature to learned ANFIS. The outputs of different ANFIS are stored in \({O}_{t}^{j} \forall j\in J, t\in T\) |
6. Find the class of \({t}^{th}\) test data based on \(\mathit{arg}\underset{j}{\mathit{max}}{R}_{tj}\); where \({R}_{tj}={O}_{t}^{j}*{u}_{jt}\) |
The proposed hierarchical model and its algorithm led to improve ANFIS classifiers learning. According to this model, a special ANFIS classifier is learned for each cluster of COVID-19 patients. Each cluster attempts to consider both the special and common characteristics of each group of COVID-19 patients. Figure 5 represents the processes of the proposed hierarchical model in this paper.
Results
In this section, more details about the clinical and image data of selected datasets are presented. Then, the effects of PCA on both datasets are evaluated as a feature reduction method. Finally, different groups of results and related statistical tests are presented.
Clinical data
There are different features for each patient in utilized datasets. Some of these features are removed using preprocessing steps and the obtained features are described in Table 3.
Besides, there are image data in the second dataset. More details about them are presented in the next subsection.
Image data
Each patient in the second dataset has clinical data with different types of CT, CR, and DX images. A patient may see a physician multiple times and order multiple types of images at different times. Therefore, the total record for all of these patients is greater than the number of patients (31,935 images recorded for 105 patients). We used the images in the latest updated version of this dataset on Dec 17, 2020. According to the properties of different types of features, CR images are selected for 105 patients. Then, the image preprocessing steps are applied to them. The results of four images after image preprocessing steps for three patients are presented in Figs. 6, 7, and 8 to represent the effects of utilized steps.
Applying PCA feature reduction
In this paper, some features based on image data are added to the second dataset. These features are mean, entropy, skewness, kurtosis, and standard deviation measurements of images. Considering these features along with other clinical features (see Table 3) could increase the complexity and decrease classification performances. Accordingly, the PCA feature reduction method is implemented for the second dataset before classification to eliminate meaningless features. For this purpose, PCA sorts all features based on their eigenvalues. Therefore, the first of them has the most eigenvalues among all features. Moreover, the cumulative percentage of covering data is changed based on the eigenvalues. Figure 9 presents more details to understand the process of PCA. In Fig. 9, features 1 to 5 are extracted features based on the image and features 6 to 23 are 18 clinical features of the second dataset represented in Table 3. Eigenvalues of different features and the cumulative percentage of covering data are demonstrated in Fig. 9.
According to Fig. 9, the first 18 features out of 23 cover more than 99% of the variations. Therefore, we ignore some meaningless features that cover less than 1% of the variations according to the results of PCA. It helps classifiers to learn better relationships among new features in less time.
Computational results
MATLAB R2022a 64-bit software is utilized on a computer with Intel(R) Core (TM) i5 CPU@2.30 GHz processor and 4.00 GB RAM for all runs in this research. There are two different target classes in the second dataset. Therefore, the performances of different classifiers and the proposed hierarchical classifier are obtained for the first dataset, the clinical data of the second dataset for both target classes, and both clinical features and extracted features of image data of the second dataset for both target classes.
Due to imbalanced utilized datasets, different measurements including accuracy, sensitivity, specificity, precision, F-score (\(F.\mathrm{mea}=\frac{2*\mathrm{sensitivity}*\mathrm{precision}}{\mathrm{sensitivity}+\mathrm{precision}}\)), G-mean (\(GSS=\sqrt{\mathrm{sensitivity}*\mathrm{specificity}}\)) are utilized to show the performances of classifiers. Besides, the average of these measurements for 30 different groups of train and test data based on standard tenfold cross validation is represented in this paper. While these groups are created based on 30 random seeds, they are the same for all classifiers in this paper. It is noteworthy that about 80% of data are considered as train data. Accordingly, 453,281 data of the first dataset and 84 data of the second dataset are considered as train data and the rest are test data. The number of experimental trials is considered 25 for each classifier and groups of train and test data to check its stability. Besides, the test data results corresponded to the training data results and validated the classification process.
At first, the results of classifiers and the proposed hierarchical model for clinical data of both datasets are presented in Tables 4 and Table 5. The ICU target class is considered for all results in both tables. Besides, these results are related to all features of datasets after data preparation. In other words, the PCA feature reduction is not utilized yet. The top three results for each measurement are highlighted and the best of them is determined using underline. According to these results, ANFIS has the most top results among other classifiers. Therefore, we consider ANFIS in the proposed model to improve related measurements. Besides, the proposed hierarchical model has all of the best results in both tables.
Performances of classifiers for different situations about the second dataset before applying PCA feature reduction are presented in Table 6. The average and standard deviation of three measurements (accuracy, F-score, and G-mean) for 10 different groups of train and test data are presented for this purpose. In addition, other results are presented in Table 7 for predefined situations after applying PCA feature reduction.
As is illustrated in Tables 6 and 7, the average of determined measurements of the proposed hierarchical model is the best performance among other classifiers. Besides, the ANFIS has the best performance among other classifiers according to these results. The PCA improves the performances of most of the classifiers and the proposed model, as well. Also, all of the standard deviations in Tables 6 and 7 are less than 0.005. It demonstrates the small difference among each measurement of the utilized classifier in different random groups of train and test data. Therefore, the results can be considered stable for utilized classifiers and the proposed hierarchical model in this paper.
To determine whether the proposed image features and feature reduction make significant statistical differences, the non-parametric two-sided Wilcoxon signed rank test has been carried out between different groups of results (Taheri and Hesamian 2013). It is assumed that the distribution is not a normal distribution and the outliers do not affect its performance (Derrac et al. 2011). It tests the presence of a significant statistical difference between two groups of results. The statistically significant difference between each pair of groups is evaluated based on the average of measures from classifiers on the selected dataset. Moreover, their related p-values are represented in Table 8. In this decision test, H = 1 is the logical value that indicates a statistically significant difference between the results of two groups at the given significance level. In this research, α = 0.05, 0.01, and 0.005 are considered for the levels of significance to evaluate decision tests. According to the results, the proposed image features and feature reduction method are statistically better than other groups of results for considered metrics.
According to these results, the proposed features of images and the feature reduction method enhance the performances of the proposed hierarchical model and make it more efficient for the diagnosis of different levels of symptomatic COVID-19 patients.
Discussion
Two real datasets about symptomatic COVID-19 patients are utilized in this paper including the COVID-19 patient pre-condition dataset and Chest Imaging with Clinical and Genomic Correlates Representing a Rural COVID-19 Positive Population (COVID-19-AR). The first dataset has clinical data and the second one has both clinical and image data. Performances of utilized classifiers and the proposed hierarchical model are compared based on different measurements including accuracy, sensitivity, specificity, precision, F-score, and G-mean. To have more precise results, we utilized principal component analysis (PCA) on clinical features and extracted features of image data for the second dataset. All results proved that the proposed hierarchical model has the best performances among utilized classifiers for both datasets. It achieves 92% accuracy and 98% accuracy for the first and second datasets, respectively. The proposed extracted features from image data and the utilized feature reduction method improve the performances of classifiers in comparison with other groups of results.
Different research studies utilized these datasets, as well. Dutta et al. (2020) utilized the COVID-19 patient pre-condition dataset and proposed a stacked Gated Recurrent Unit (GRU) based model to identify whether a patient can be infected by this disease or not. The accuracy of their proposed method was 66%. Different properties of the second dataset are evaluated in the paper of Santa Cruz et al. (2021). They found the second dataset appropriate for the proper assessment of the risk of bias. Desai et al. 2020a, b) explained the different features of the second dataset to guide other researchers about this dataset. They determined different features of this dataset as target classes that could be helpful for different purposes. Tang et al. (2020) proposed a segmentation model for identifying the opacity regions from the COVID-19-positive chest X-rays including haziness, ground-glass opacity, and lung consolidation. Although their results are accurate and robust, these results are not analyzed using other classifiers to make an appropriate diagnosis system. Sarv Ahrabi et al. (2021) proposed a convolutional neural network (CNN) with optimized parameters for the second dataset. They utilized deep learning (DL) paradigms for analyzing X-rays in this dataset to achieve an accuracy of 93%. According to these results and literature review, the proposed model in this paper could be supportive in medical purposes for fighting COVID-19 (Rahimi Rise et al. 2022; Rise and Ershadi 2022; Sadat et al. 2022).
Conclusions
In today’s world, there are many newly developed methods to deal with infectious diseases along with a new algorithm. Most of these methods help with disease diagnosis and other areas such as detecting levels of symptomatic patients in a hospital are neglected. Besides, most of the proposed methods in different papers considered clinical data for disease, and the image data are ignored, as well. It is noteworthy that image data plays an important role to find the level of hospitalized symptomatic COVID-19 patients. This gap becomes very serious for various types of Coronavirus disease due to its effects on patients. Timely detection of this disease in its early stages would remarkably increase the chance of complete treatment in a shorter time. Furthermore, on-time detection of the patients’ level would directly affect the service plans in the hospital.
Therefore, in this paper, for the first time and to the best of our knowledge, we used both clinical and image data to find the levels of different hospitalized COVID-19 patients. The need for these patients to ICU and whether or not they are in end-stage are considered as target classes for these patients. A new hierarchical model is proposed for this aim in this paper. This model cluster all patients based on predefined measurements and learn different ANFIS for each of the clusters. Different groups of COVID-19 patients have special and common characteristics. Therefore, we select the FCM clustering method that considers the membership function degree for each data to different clusters. Besides, ANFIS classifiers have the best performances among the 12 utilized classifiers in this paper including case-based reasoning (CBR), decision tree, convolutional neural networks (CNN), K-nearest neighbors (KNN), learning vector quantization (LVQ), multi-layer perceptron (MLP), Naive Bayes (NB), radial basis function network (RBF), support vector machine (SVM), recurrent neural networks (RNN), fuzzy type-i inference system (FIS), and adaptive neuro-fuzzy inference system (ANFIS). Consequently, ANFIS classification methods are selected to understand relationships among features of each cluster.
Accordingly, the results of the proposed models and determining features of images in this paper could be supportive for medical decisions about COVID-19 patients. The proposed hierarchical model in this paper could be supportive in hospitals to find the appropriate levels for different hospitalized symptomatic COVID-19 patients. A graphical user interface is designed based on the proposed hierarchical model and presented in Fig. 10.
The most important limitation for this work is the dependence of the proposed model on data. Therefore, the limitations of data and appropriate features have a high impact on its performance. It is noteworthy that different groups of COVID-19 patients are dissimilar in different cities/countries. Therefore, this graphical user interface has to be updated based on related datasets about hospitalized symptomatic COVID-19 patients in a special city/country.
In future studies, one can use other classification and clustering methods with non-Euclidean distances to find the performances of the used methods. Furthermore, different combinations of clustering and classification methods could be tailored based on geographical situations.
Data availability
Two real datasets about symptomatic COVID-19 patients are utilized in this paper as follows:
• The first dataset about COVID-19 patients is the “COVID-19 patient pre-condition dataset”. This dataset was obtained from the Mexican government dataset and includes 566,602 patients with 23 clinical features (https://www.kaggle.com/tanmoyx/covid19-patient-precondition-dataset.).
• The second dataset is “Chest Imaging with Clinical and Genomic Correlates Representing a Rural COVID-19 Positive Population (COVID-19-AR)” [https://wiki.cancerimagingarchive.net/pages/viewpage.action?pageId=70226443#70226443bcab02c187174a 288dbcbf95d26179e8). This dataset has clinical and image data of about 105 patients.
Code availability
Related codes are uploaded.
References
An C, Lim H, Kim DW, Chang JH, Choi YJ, Kim SW. Machine learning prediction for mortality of patients diagnosed with COVID-19: a nationwide Korean cohort study. Sci Rep. 2020;10(1):1–11. https://doi.org/10.1038/s41598-020-75767-2.
Bansal M. Cardiovascular disease and COVID-19. Diabetes Metab Syndr. 2020;14(3):247–50. https://doi.org/10.1016/j.dsx.2020.03.013.
Barracchia EP, Pio G, D’Elia D, Ceci M. Prediction of new associations between ncRNAs and diseases exploiting multi-type hierarchical clustering. BMC Bioinformatics. 2020;21(1):1–2. https://doi.org/10.1186/s12859-020-3392-2.
Bullock J, Luccioni A, Pham KH, Lam CSN, Luengo-Oroz M. Mapping the landscape of artificial intelligence applications against COVID-19. J Artif Intel Res. 2020;69:807–45. https://doi.org/10.1613/jair.1.12162.
Burdick H, Lam C, Mataraso S, Siefkas A, Braden G, Dellinger RP, ... and Das R. Prediction of respiratory decompensation in Covid-19 patients using machine learning: the READY trial. Comput Biol Med.2020; 124:103949. https://doi.org/10.1016/j.compbiomed.2020.103949
Chan JFW, Yuan S, Kok KH, To KKW, Chu H, Yang J, ... and Yuen KY. A familial cluster of pneumonia associated with the 2019 novel coronavirus indicating person-to-person transmission: a study of a family cluster. Lancet. 2020;395(10223):514–523. https://doi.org/10.1016/S0140-6736(20)30154-9
Cohen JP, Morrison P, Dao L, Roth K, Duong TQ, and Ghassemi M. Covid-19 image data collection: prospective predictions are the future. arXiv preprint arXiv:2006.11988. 2020. https://arxiv.org/abs/2006.11988.
Dai WC, Zhang HW, Yu J, Xu HJ, Chen H, Luo SP, ... and Lin F. CT imaging and differential diagnosis of COVID-19. Can Assoc Radiol J. 2020;71(2):195–200. https://doi.org/10.1177/0846537120913033
Derrac J, García S, Molina D, Herrera F. A practical tutorial on the use of nonparametric statistical tests as a methodology for comparing evolutionary and swarm intelligence algorithms. Swarm Evol Comput. 2011;1(1):3–18. https://doi.org/10.1016/j.swevo.2011.02.002.
Desai S, Baghal A, Wongsurawat T, Al-Shukri S, Gates K, Farmer P, Rutherford M, Blake GD, Nolan T, Powell T, Sexton K, Bennett W, Prior F. Data from chest imaging with clinical and genomic correlates representing a rural COVID-19 positive population. Cancer Imaging Arch. 2020a. https://doi.org/10.7937/tcia.2020.py71-5978. Available at: https://wiki.cancerimagingarchive.net/pages/viewpage.action?pageId=70226443#70226443bcab02c187174a288dbcbf95d26179e8.
Desai S, Baghal A, Wongsurawat T, Jenjaroenpun P, Powell T, Al-Shukri S, ... Prior F. Chest imaging representing a COVID-19 positive rural US population. Sci Data. 2020b;7(1):1–6. https://doi.org/10.1038/s41597-020-00741-6
Di Lorenzo G, Di Trolio R, Kozlakidis Z, Busto G, Ingenito C, Buonerba L, ... Leo E. COVID 19 therapies and anti-cancer drugs: a systematic review of recent literature. Crit Rev Oncol/Hematol. 2020;152:102991. https://doi.org/10.1016/j.critrevonc.2020.102991
Dutta S, Bandyopadhyay SK. Artificial intelligence-based study on analyzing of habits and with history of diseases of patients for prediction of recurrence of disease due to covid-19. Int J Eng Manag Res (IJEMR). 2020;10(4):106–13. https://doi.org/10.31033/ijemr.10.4.16.
Elaziz MA, Hosny KM, Salah A, Darwish MM, Lu S, Sahlol AT. New machine learning method for image-based diagnosis of COVID-19. Plos one. 2020;15(6):e0235187. https://doi.org/10.1371/journal.pone.0235187.
Ershadi MM, Seifi A. An efficient Bayesian network for differential diagnosis using experts’ knowledge. Int J Intell Comput Cybern. 2020a. https://doi.org/10.1108/IJICC-10-2019-0112.
Ershadi MM, Seifi A. An efficient multi-classifier method for differential diagnosis. Intell Decis Technol. 2020b;14(3):337–47. https://doi.org/10.3233/IDT-190060.
Fan DP, Zhou T, Ji GP, Zhou Y, Chen G, Fu H, ... Shao L. Inf-net: Automatic covid-19 lung infection segmentation from ct images. IEEE Trans Med Imaging. 2020;39(8):2626–2637. https://doi.org/10.1109/TMI.2020.2996645
Fan C, Lei D, Fang C, Li C, Wang M, Liu Y, ..., Wang S. Perinatal transmission of 2019 coronavirus disease–associated severe acute respiratory syndrome coronavirus 2: should we worry?. Clin Infect Dis. 2021;72(5):862–864. https://doi.org/10.1093/cid/ciaa226
Gorbalenya AE, Baker SC, Baric RS, de Groot RJ, Drosten C, Gulyaeva AA, ... Ziebuhr J. Severe acute respiratory syndrome-related coronavirus: the species and its viruses–a statement of the Coronavirus Study Group. BioRxiv. 2020. https://doi.org/10.1038/s41564-020-0695-z.
Ivanov D. Predicting the impacts of epidemic outbreaks on global supply chains: a simulation-based analysis on the coronavirus outbreak (COVID-19/SARS-CoV-2) case. Transp Res Part e: Logist Transp Rev. 2020;136:101922.
Khanday AMUD, Rabani ST, Khan QR, Rouf N, Din MMU. Machine learning based approaches for detecting COVID-19 using clinical text data. Int J Inf Technol. 2020;12(3):731–9. https://doi.org/10.1007/s41870-020-00495-9.
Koolhof IS, Gibney KB, Bettiol S, Charleston M, Wiethoelter A, Arnold AL, ... Firestone SM. The forecasting of dynamical Ross River virus outbreaks: Victoria, Australia. Epidemics. 2020;30:100377. https://doi.org/10.1016/j.epidem.2019.100377
Lalmuanawma S, Hussain J, Chhakchhuak L. Applications of machine learning and artificial intelligence for Covid-19 (SARS-CoV-2) pandemic: a review. Chaos Solitons Fractals. 2020;139:110059. https://doi.org/10.1016/j.chaos.2020.110059.
Li WT, Ma J, Shende N, Castaneda G, Chakladar J, Tsai JC, ... Ongkeko WM. Using machine learning of clinical data to diagnose covid-19. medRxiv. 2020. https://doi.org/10.1186/s12911-020-01266-z.
Liang H, Guo Y, Chen X, Ang KL, He Y, Jiang N, ... Zhong N. Artificial intelligence for stepwise diagnosis and monitoring of COVID-19. Eur Radiol. 2022;1–11. https://doi.org/10.1007/s00330-021-08334-6
Loucera C, Esteban-Medina M, Rian K, Falco MM, Dopazo J, Peña-Chilet M. Drug repurposing for COVID-19 using machine learning and mechanistic models of signal transduction circuits related to SARS-CoV-2 infection. Signal Transduct Target Ther. 2020;5(1):1–3. https://doi.org/10.1038/s41392-020-00417-y.
Mahmud T, Rahman MA, Fattah SA. CovXNet: A multi-dilation convolutional neural network for automatic COVID-19 and other pneumonia detection from chest X-ray images with transferable multi-receptive feature optimization. Comput Biol Med. 2020;122:103869. https://doi.org/10.1016/j.compbiomed.2020.103792.
Mukherjee T. COVID-19 patient pre-condition dataset. 2020. Available at: https://www.kaggle.com/tanmoyx/covid19-patient-precondition-dataset.
Oyelade ON, Ezugwu AE. A case-based reasoning framework for early detection and diagnosis of novel coronavirus. Inform Med Unlocked. 2020;20:100395. https://doi.org/10.1016/j.imu.2020.100395.
Pan SL, Cui M, Qian J. Information resource orchestration during the COVID-19 pandemic: a study of community lockdowns in China. Int J Inform Manag. 2020;54:102143. https://doi.org/10.1016/j.ijinfomgt.2020.102143.
Panwar H, Gupta PK, Siddiqui MK, Morales-Menendez R, Singh V. Application of deep learning for fast detection of COVID-19 in X-Rays using nCOVnet. Chaos Solitons Fractals. 2020;138:109944. https://doi.org/10.1016/j.chaos.2020.109944.
Peak CM, Kahn R, Grad YH, Childs LM, Li R, Lipsitch M, Buckee CO. Individual quarantine versus active monitoring of contacts for the mitigation of COVID-19: a modelling study. Lancet Infect Dis. 2020;20(9):1025–33. https://doi.org/10.1016/S1473-3099(20)30361-3.
Pinter G, Felde I, Mosavi A, Ghamisi P, Gloaguen R. COVID-19 pandemic prediction for Hungary; a hybrid machine learning approach. Mathematics. 2020;8(6):890. https://doi.org/10.3390/math8060890.
Rahimi Rise Z, Ershadi MM, Ershadi MJ. Multidisciplinary analysis of international environments based on impacts of Covid-19: State of art. IJIEPR. 2022;33(1):1–10. https://doi.org/10.22068/ijiepr.33.1.14.
Rahimi Rise Z, Ershadi MM, Shahabi Haghighgi SH. Scenario-based analysis about COVID-19 outbreak in Iran using systematic dynamics modeling-with a focus on the transportation system. J Transp Res . 2020;17(2):33–48. Available at: http://www.trijournal.ir/article_107879.html?lang=en.
Rahmani AM, Mirmahaleh SYH. Coronavirus disease (COVID-19) prevention and treatment methods and effective parameters: a systematic literature review. Sustain Cities Soc. 2021;64:102568. https://doi.org/10.1016/j.scs.2020.102568.
Rauschecker AM, Rudie JD, Xie L, Wang J, Duong MT, Botzolakis EJ, ... Gee JC. Artificial intelligence system approaching neuroradiologist-level differential diagnosis accuracy at brain MRI. Radiology. 2020;295(3):626–637. https://doi.org/10.1148/radiol.2020190283
Rise ZR, Ershadi MM. Socioeconomic analysis of infectious diseases based on different scenarios using uncertain SEIAR system dynamics with effective subsystems and ANFIS. J Econ Adm Sci. 2022;ahead-of-print No. ahead-of-print. https://doi.org/10.1108/JEAS-07-2021-0124
Rubin GD, Ryerson CJ, Haramati LB, Sverzellati N, Kanne J P, Raoof S, ... Leung AN. The role of chest imaging in patient management during the COVID-19 pandemic: a multinational consensus statement from the Fleischner Society. Radiology. 2020;96(1):172–180. https://doi.org/10.1016/j.chest.2020.04.003
SadaAsl AA, Ershadi MM, Sotudian S, Li X, Dick S. Fuzzy expert systems for prediction of ICU admission in patients with COVID-19. Intell Decis Technol. 2022;16(1):159–68. https://doi.org/10.3233/IDT-200220.
Saghazadeh A, Rezaei N. Towards treatment planning of COVID-19: rationale and hypothesis for the use of multiple immunosuppressive agents: anti-antibodies, immunoglobulins, and corticosteroids. Int Immunopharmacol. 2020;84:106560. https://doi.org/10.1016/j.intimp.2020.106560.
Santa Cruz BG, Bossa MN, Soelter J, Husch AD. Public Covid-19 X-ray datasets and their impact on model bias-a systematic review of a significant problem. medRxiv. 2021. https://doi.org/10.1101/2021.02.15.21251775
SarvAhrabi S, Scarpiniti M, Baccarelli E, Momenzadeh A. An accuracy vs. complexity comparison of deep learning architectures for the detection of COVID-19 disease. Computation. 2021;9:3. https://doi.org/10.3390/computation9010003.
Sarwar A, Ali M, Manhas J, Sharma V. Diagnosis of diabetes type-II using hybrid machine learning based ensemble model. Int J Inf Technol. 2020;12(2):419–28. https://doi.org/10.1007/s41870-018-0270-5.
Shi F, Wang J, Shi J, Wu Z, Wang Q, Tang Z, ... Shen D. Review of artificial intelligence techniques in imaging data acquisition, segmentation, and diagnosis for COVID-19. IEEE Rev Biomed Eng. 2020;14:4–15. https://doi.org/10.1109/RBME.2020.2987975
Suman G, Panda A, Korfiatis P, Edwards ME, Garg S, Blezek DJ, ... Goenka AH. Development of a volumetric pancreas segmentation CT dataset for AI applications through trained technologists: a study during the COVID 19 containment phase. Abdom Radiol. 2020;45(12):4302–4310. https://doi.org/10.1007/s00261-020-02741-x
Swapnarekha H, Behera HS, Nayak J, Naik B. Role of intelligent computing in COVID-19 prognosis: a state-of-the-art review. Chaos, Solitons Fractals. 2020;138:109947. https://doi.org/10.1016/j.chaos.2020.109947.
Taheri SM, Hesamian G. A generalization of the Wilcoxon signed-rank test and its applications. Stat Pap. 2013;54(2):457–70. https://doi.org/10.1007/s00362-012-0443-4.
Tang H, Sun N, Li Y. Segmentation model of the opacity regions in the chest X-rays of the Covid-19 patients in the us rural areas and the application to the disease severity. medRxiv. 2020. https://doi.org/10.1101/2020.10.19.20215483
Thu TPB, Ngoc PNH, Hai NM. Effect of the social distancing measures on the spread of COVID-19 in 10 highly infected countries. Sci Total Environ. 2020;742:140430. https://doi.org/10.1016/j.scitotenv.2020.140430.
Wang Y, Dong C, Hu Y, Li C, Ren Q, Zhang X, ... Zhou M. Temporal changes of CT findings in 90 patients with COVID-19 pneumonia: a longitudinal study. Radiology. 2020;296(2):E55-E64. https://doi.org/10.1148/radiol.2020200642
World Health Organization. Novel Coronavirus (2019-nCoV): Situation Report-3; WHO: Geneva, Switzerland, 2020; Available online: https://www.who.int/docs/default-source/coronaviruse/situation-reports/20200123-sitrep-3-2019-ncov.pdf. (accessed on 28 April 2022).
Yan L, Zhang HT, Xiao Y, Wang M, Guo Y, Sun C, ... Yuan Y. Prediction of criticality in patients with severe Covid-19 infection using three clinical features: a machine learning-based prognostic model with clinical data in Wuhan. MedRxiv. 2020. https://doi.org/10.1101/2020.02.27.20028027
Zarandi MF, Seifi A, Ershadi MM, Esmaeeli H. An expert system based on fuzzy bayesian network for heart disease diagnosis. In North American Fuzzy Information Processing Society Annual Conference (pp. 191–201). Springer, Cham. 2017. https://doi.org/10.1007/978-3-319-67137-6_21
Acknowledgements
The authors wish to thank Dr. S. Ershadi for providing expert’s knowledge used in this research.
Author information
Authors and Affiliations
Contributions
The authors of this paper are in the following order:
1. Mohammad Mahdi Ershadi 2. Zeinab Rahimi Rise
The credit role(s) of each author are as follows:
1. Basic modeling and conceptualization, investigation, data curation, software implementation, and writing the draft of the paper were done by the first author.
2. Basic modeling and conceptualization, supervision, methodology, validation of results, review, and rewriting the paper were done by the second author.
Corresponding author
Ethics declarations
Ethics approval
The authors of this paper accepted ethical standards.
Consent to participate
This statement is to certify that all authors have seen and approved the manuscript being submitted in Research on Biomedical Engineering.
Consent for publication
This statement is to certify that all authors have seen and approved the manuscript being published in Research on Biomedical Engineering. We warrant that the article is the authors’ original work. We warrant that the article has not received prior publication and is not under consideration for publication elsewhere. On behalf of all co-authors, the corresponding author shall bear full responsibility for the submission. This research has not been submitted for publication nor has it been published in whole or in part elsewhere. All authors agree that author list is correct in its content and order.
Competing interests
The authors declare no competing interests.
Research involving human participants and/or animals
This article does not contain any studies with human or animals’ participants performed by any of the authors.
Additional information
Publisher's note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Electronic supplementary material
Below is the link to the electronic supplementary material.
Appendix
Appendix
Published research of Web of Science between the years 1980 and 2022 about COVID-19 and applications of machine learning methods are analyzed using VOSviewer 1.6.10 software and related results are presented in Fig.
11 as a science map. A search with “COVID-19” or other equivalent terms founds more than about 6,500,000 results. Besides, "machine learning methods" or other equivalent terms are found in more than 850,000 results. However, less than 250 of them focused on the machine learning models for COVID-19. Less than 10 articles focus on machine learning to diagnose different levels of hospitalized COVID-19 patients. Therefore, there is a significant gap in this area and this paper presents a proposed model for this purpose. It is noteworthy that different circles show the different number of articles according to their first keywords in Fig. 11. The larger circles are associated with more articles with the same first keyword. The lines between circles show the relationships among their papers in terms of their references. Besides, VOSviewer 1.6.10 software clusters different circles in this figure into some colored clusters according to their relationships. Figure
12 shows the year of published articles presented in Fig. 11. According to Fig. 12, most papers about machine learning applications in disease diagnosis publish are published in recent years. However, there are different papers about COVID in previous years.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Ershadi, M.M., Rise, Z.R. Fusing clinical and image data for detecting the severity level of hospitalized symptomatic COVID-19 patients using hierarchical model. Res. Biomed. Eng. 39, 209–232 (2023). https://doi.org/10.1007/s42600-023-00268-w
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s42600-023-00268-w