Abstract
Coronavirus disease-19 (COVID-19) is caused by SARS-CoV-2, a highly contagious respiratory virus that has caused a global pandemic. Despite the urgent need for effective diagnostic screening technologies, ideal methods for COVID-19 detection have not yet been developed. To address this issue, we developed a Raman spectroscopy technique for rapid and sensitive on-site detection of SARS-CoV-2, utilizing the unique spectral fingerprint of molecular vibrations. The proposed technique is non-invasive and label-free that enables the detection of molecular vibrations, providing a unique spectral fingerprint for different molecules. Raman spectra from 75 positive and 75 negative swab samples were analyzed, processed by smoothening and baseline correction of spectral data. The peaks in the processed data were detected and assigned based on literature peak, with peaks specific to positive samples used for detection with minimal false positives. These peaks were attributed to various molecules, including amino acids in proteins, glycoproteins, lipids, and protein structures. Our Raman spectroscopy technique provides a reliable and non-invasive approach for the detection of SARS-CoV-2, with potential to expand to other infectious agents. This method has significant implications for global health, aiding in effective control measures against COVID-19.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
1 Introduction
Coronavirus disease-19 (COVID-19) is a respiratory illness caused by the novel coronavirus, severe acute respiratory syndrome coronavirus-2 (SARS-CoV-2). First identified in Wuhan, China in December 2019, COVID-19 has since spread to over 100 countries worldwide [1]. Compared to other coronavirus-related diseases such as SARS and MERS, COVID-19 is highly contagious [2,3,4] and can lead to severe symptoms, including fever, shortness of breath, cough, sore throat, sputum, and myalgia [5]. Patients with underlying health conditions such as heart disease, diabetes, obesity, chronic kidney disease, and chronic obstructive pulmonary disease are more susceptible to severe symptoms [6]. However, some individuals infected with SARS-CoV-2 may not exhibit any symptoms, but they can still spread the virus and are therefore required to quarantine.
To detect SARS-CoV-2 in infected patients and prevent transmission to vulnerable individuals, various methods have been developed. The most widely used method is the lateral flow immunoassay, which is cost-effective and provides relatively quick results [7]. However, some lateral flow immunoassay tests are only qualitative or partially quantitative and require extensive validation and quality [8]. Another commonly used method is the polymerase chain reaction (PCR), which is recognized as the most reliable method by government and health facilities. PCR provides accurate results with high sensitivity but requires up to 48 h to obtain results and cannot be used on patients who have recently recovered from COVID-19 [9].
Optical spectroscopy is an emerging method for detecting SARS-CoV-2 and has shown promising results in rapidly detecting other types of viruses with high accuracy [10,11,12,13,14]. Optical spectroscopy has been used more frequently in recent years due to its speed and non-destructive nature of the method [15]. Raman spectroscopy, a type of optical spectroscopy, involves the scattering of monochromatic light on biomolecules that make up a virus, resulting in a change in wavelength due to the excitation of atoms from the ground state to the virtual state [16]. The change in wavelength is quantified by Raman shift (measured in cm−1), with larger change in wavelength corresponding to larger Raman shift. Raman spectroscopy is non-invasive as the samples used typically do not require further heat and chemical treatment. Furthermore, Raman spectroscopy is superior to other methods of optical spectroscopy such as infrared or ultraviolet, mainly because the collected Raman signal is not significantly affected by the presence of water in the sample [17]. Biofingerprint detection based on Raman spectroscopy is mainly used in medical applications such as detection of various types of cancer and diagnosis of other diseases [17, 18]. This has been used in previous research studies to identify presence of viral particles in samples via scattering of monochromatic light on biomolecules that make up virus [10, 12,19, 20].
Although most countries have entered the endemic phase of the pandemic, where previously applied restrictions have been relaxed, there is still a need to contain COVID-19 to prevent a surge in cases. The public has seen the highly contagious and potentially deadly nature of COVID-19, especially for vulnerable individuals [21]. Therefore, there is a high demand for research and development of rapid and robust methods for detecting SARS-CoV-2. In this study, we analyze the Raman peaks present in negative and positive samples using spectral data obtained using a portable Raman system built in our laboratory for the purpose of SARS-CoV-2 detection. The proposed method would benefit the development of pattern recognition model by utilizing the detected peaks, consequently improving specificity and reducing false negatives of the model.
2 Data and methods
2.1 Sample preparation
The study involved the analysis of 75 nasal swab samples collected from patients infected with SARS-CoV-2 as positive samples, and 75 nasal swab samples from healthy patients as negative samples. The samples were collected from Kuala Lumpur International Healthcare Centre Kota Kinabalu (KLIHCKK) and were stored in − 18 °C after being placed in vials containing 3 mL of viral transport media (VTM). A blank VTM sample is also used in this study as control sample to compare its spectral data with positive and negative samples.
To prepare the samples for acquiring Raman spectra, 1 mL of each sample including the blank VTM sample was transferred into cuvettes using a pipette. This was done to ensure that the samples were homogenously mixed and prepared for analysis. The samples were then analyzed using Raman spectroscopy to determine their molecular composition and identify any differences between the positive and negative samples. It should be noted that all necessary safety precautions were taken during the collection and handling of the samples to prevent any potential contamination or exposure to the virus. Additionally, the study was conducted in accordance with ethical guidelines and regulations to ensure the protection of patient privacy and confidentiality.
2.2 Spectral data acquisition
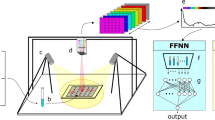
Figure 1 illustrates the simplified setup for Raman system used in this study. The system consists of a 200-mW laser diode as the excitation source, emitting a laser beam with a wavelength of 532 nm. Raman scattering light from a sample is collected using a microscope objective with a numerical aperture of 0.22 and magnification of 10X. The collected light then passes through a long-wave pass filter, which allows the transmission of Raman scattered light at longer wavelengths while rejecting Rayleigh scattered light.
To enhance the normally weak Raman signal, a charged-couple device (CCD) is used in the setup, which is cooled down for 15 min to remove noise associated with CCD. An acquisition time of 15 s is required for every sample to further enhance the signal collected at the CCD.
Before the Raman system can be used for analysis, it must be calibrated to optimize the acquisition of Raman spectra. A standard sample is used for calibration to ensure that the signal acquired by the CCD matches the required Raman shift displayed by the computer with minimal error. In this study, polystyrene is used as the standard sample for calibration as it is readily available and has numerous Raman peaks that can be compared with the datasheet for polystyrene.
Overall, the Raman system used in this study was carefully calibrated and optimized to ensure the accurate acquisition of Raman spectra from the collected samples. This setup allowed for the reliable comparison and analysis of Raman spectra between positive and negative samples.
2.3 Processing of Raman spectra
The Raman system shown in Fig. 1 is used to acquire Raman spectral data from the collected samples. The Raman shift range covered by the system is from 200 to 3800 cm−1. The acquired spectral data is processed using MATLAB software to observe the Raman peaks.
In this study, the spectral data within the range of 400 and 2000 cm−1 is of particular interest, as Raman peaks located in this range indicate the presence of functional groups found in viral particles [22]. Therefore, the acquired spectral data is trimmed to focus only on this range of high interest.
Next, background noise is removed from the spectral data using a Savitzky-Golay filter with a third order polynomial taken seven points at a time. This filter smoothens the spectral data and ensures that Raman peaks can be clearly observed. After this, the smoothened data undergoes baseline correction to remove wide peaks that might be the product of background fluorescence [12]. The baseline is regressed using a spline approximation within windows of width 150 cm−1. This ensures that sharp peaks are preserved while wider humps are removed from spectral data, leading to a clearer and more accurate representation of the Raman peaks. Figure 2 illustrates Raman spectra of a sample before and after Savitzky-Golay filter and baseline correction.
2.4 Selection and assignment of peaks from spectral data
To facilitate the selection of peaks based on a fixed prominence threshold, the intensity axis of spectral data is converted to relative intensity by setting the minimum and maximum values of the intensity axis to 0 and 1, respectively. Raman peaks are detected using the findpeaks function in MATLAB, which applies a set of criteria [12]. The findpeaks function is improved and modified in this study to identify the location at which half the prominence of each peak is taken, along with other parameters such as the peak’s prominence and width. This is important to ensure that peaks located within another peak are not counted. Peaks with a width larger than 10 cm−1 and a prominence larger than 0.2 in the Raman spectra are selected, while other peaks that may be attributed to background noise remaining after the smoothing procedure are removed.
Next, the experimental peaks of positive samples are assigned based on literature peaks obtained from previous studies [22,23,24]. Only peaks that coincide or are in proximity to the literature peaks associated with positive samples are assigned. This step is repeated for negative samples, and the presence of peaks is compared between positive and negative samples.
3 Results and discussion
3.1 Detection of peaks in positive and negative samples
All 75 positive and 75 negative samples were analyzed by detecting and analyzing their respective peaks based on specific criteria. As shown in Table 1, positive samples exhibited more detected peaks than negative samples, suggesting the presence of Raman peaks that are exclusive to positive samples. However, further assignment of these peaks is necessary to verify that they are indeed unique to positive samples. This is particularly important given the relatively low number of Raman peaks detected in positive samples (as few as 5) and the higher number of peaks detected in negative samples (up to 19). Therefore, rigorous peak assignment is crucial to ensure accurate differentiation between positive and negative samples.
3.2 Assignment of peaks from positive and negative samples
Table 2 reveals the number of positive and negative samples with the experimental peak present coinciding or close to the literature peak, obtained from previous studies [22,23,24]. Between 2 and 15 peaks in positive samples, and 1 to 6 peaks in negative samples, are assigned based on literature peaks listed in Table 2.
Figure 3a and b shows Raman spectra of both selected positive and negative samples, with the peaks detected by MATLAB are highlighted in grey. Meanwhile, Fig. 3c reveals Raman spectra for a blank VTM sample.The spectra highlighted in brown are assigned based on literature peak in Table 2. Peaks that coincide with the literature peak or deviate from literature peak not more than 10 cm−1 are assigned. By analysing all peaks in positive samples, peaks at around 501.5, 509, 577, 656.2, 716, 748, 782.5, 846.5, 951.5, 983.7, 1000–1060, 1207, 1249, 1317, 1384, 1453 and 1554 cm−1 are present in positive samples significantly more than negative samples. Since less than 10% of the samples containing these peaks are negative samples, these peaks can be used to determine positive samples with possibility of false positive detection on negative samples below 10%. Meanwhile, analysis of all peaks in negative samples shows that peaks at 1126, 1155, 1263, 1600.7 and 1665–1669 cm−1 are present in negative samples more than positive samples. These peaks can be used detect negative samples. However, some negative peaks are present in positive samples. Using these peaks in pattern recognition might cause these positive samples to be identified as negative samples, leading to false negative. It is noted that peaks located in proximity to 1340.4, 1378 and 1492.1 cm−1 are not considered as these peaks are also significant in blank VTM samples.
Raman peaks situated in close proximity to 577, 656.2, 748, 951.5, 983.7, 1048, 1207, and 1554 cm−1 are prominent in significantly more positive samples and attributed to amino acids that are present in coronavirus. Experimental peaks close to 577, 748, 951.5, 983.7 and 1554 cm−1 originate from tryptophan (Trp) found in positive samples, but not in negative samples. Another peak at 1048 cm−1 (situated between 1000 and 1060 cm−1) prevalent in positive samples correspond to aromatic amino acids, particularly phenylalanine (Phe) and Trp [23]. The environment rich in these amino acids in positive samples is due to the importance of Phe and Trp in developing viral protein structure or interaction with physiologically expressed molecules [25]. However, it is also possible that these peaks are due to other amino acids such as cysteine (Cys) in 577 cm−1 and valine (Val) in 983.7 cm−1
Some of these Raman peaks (close to 509, 922, 1155, and 1453 cm−1) are associated with glycoproteins and lipids. Many positive and negative samples have Raman peak located near 1155 cm−1, assigned with glucose, which is due to assembled and fragmented viral particles present in positive and negative samples, respectively [23]. In contrast, other Raman peaks are more prominent in positive samples, indicating that these peaks are due to assembled viral particles represented by physio/pathological modifications of lipids and glycoproteins in contact with the virus [26, 27].
Amide bands in protein structure are represented by Raman peaks of 1263 and 1665–1669 cm−1 in this experiment. Amide I and III bands present in some positive and negative samples correspond to β-sheets/coils. This is consistent with the previous study by Huang et al. which stated that amide bands in Raman spectra of SARS-CoV-2 show more obvious β-sheets/coils compared to α-helices [24]. Not all positive samples exhibit amide I and III bands based on their Raman spectra. This is a common occurrence as stated by Sanchez et al. [22] and observed in several previous studies [23, 28, 29]. This is due to dependence of absence of amide bands in Raman spectra on the size of amino acid side chain [28]. Despite that, the absence of amide bands in many positive samples is also due to deactivation of virus that might break amino acids. Also, some negative samples exhibit amide I and III bands, which are attributed to fragmented proteins that may still circulate during convalescence and weeks after infection.
Table 3 reveals the summary of Raman peaks that are shown to be significant in positive and negative samples. By analysing all 75 positive and 75 negative samples, it is found that peaks associated with presence of SARS-CoV-2 are present in the positive samples significantly more than negative samples. Because of this, these peaks can be used to distinguish positive samples from negative samples with minimal occurrence of false positives. In contrast, while peaks associated with absence of SARS-CoV-2 are present in negative samples, these peaks are also prominent in positive samples. The use of these peaks to distinguish negative samples may lead to considerable false negatives. To ensure that these peaks can be reliably incorporated into development of pattern recognition model, only peaks associated with positive samples shall be considered to minimize false negatives due to peaks in negative samples. These peaks can be used with high specificity as they are not significantly present in blank VTM samples, as revealed in Fig. 3c.
4 Conclusion
Peak assignment on Raman spectra of positive and negative samples was applied to serve as database for rapid detection of SARS-CoV-2 in samples using pattern recognition. The assigned peaks are analyzed in this study to determine the significant peaks that can be used to distinguish positive and negative samples based on Raman spectra. Peaks associated with positive samples are reliable in identifying positive samples. As these peaks are prominent in positive samples but not in negative samples nor in blank VTM sample, these peaks can be implemented in pattern recognition model with reduced false negatives and high specificity.
Data availability
The datasets analysed during the current study are not publicly available due to confidentiality but are available from the corresponding author on reasonable request.
References
Haynes BF, Corey L, Fernandes P, Gilbert PB, Hotez PJ, Rao S, …, Arvin A (2020) Prospects for a safe COVID-19 vaccine. Sci Transl Med 12:1–12. https://doi.org/10.1126/scitranslmed.abe0948
Cao Z, Zhang Q, Lu X, Pfeiffer D, Jia Z, Song H, Zeng DD (2020) Estimating the effective reproduction number of the 2019-nCoV in China. MedRxiv. https://doi.org/10.1101/2020.01.27.20018952
Shen M, Peng Z, Xiao Y, Zhang L (2020) Modeling the epidemic trend of the 2019 novel coronavirus outbreak in China. Innovation(United States) 25(1):1–2. https://doi.org/10.1016/j.xinn.2020.100048
Viceconte G, Petrosillo N (2020) Covid-19 R0: magic number or conundrum? Infect Dis Rep 12(1):1–2. https://doi.org/10.4081/idr.2020.8516
O’Connor T, Javidi B (2022) COVID-19 screening with digital holographic microscopy using intra-patient probability functions of spatio-temporal bio-optical attributes. Biomed Opt Exp 13(10):5377–5389. https://doi.org/10.1364/boe.466005
Hacker KA, Briss PA, Richardson L, Wright J, Petersen R (2021) COVID-19 and chronic disease: the Impact now and in the future. Prev Chronic Dis 18(E62):1–6
Di Nardo F, Chiarello M, Cavalera S, Baggiani C, Anfossi L (2021) Ten years of lateral flow immunoassay technique applications: trends, challenges and future perspectives. Sensors 21(5185):1–33. https://doi.org/10.3390/s21155185
Lingervelder D, Koffijberg H, Kusters R, IJzerman MJ (2019) Point-of-care testing in primary care: a systematic review on implementation aspects addressed in test evaluations. Int J Clin Pract 73(10):1–12. https://doi.org/10.1111/ijcp.13392
Tan Q, Wu S, Liu Z, Wu X, Forsberg E, He S (2022) High sensitivity detection of SARS-CoV-2 by an optofluidic hollow eccentric core fiber. Biomed Opt Exp 13(9):4592–4605. https://doi.org/10.1364/boe.465136
Bilal M, Ullah R, Khan S, Ali H, Saleem M, Ahmed M (2017) Lactate based optical screening of dengue virus infection in human sera using Raman spectroscopy. Biomed Opt Exp 8(2):1250–1256. https://doi.org/10.1364/boe.8.001250
Hasbi NH, Bade A, Chee FP, Rumaling MI (2022) Pattern recognition for human diseases classification in spectral analysis. Computation 10(96):1–24
Khan S, Ullah R, Khan A, Ashraf R, Ali H, Bilal M, Saleem M (2018) Analysis of hepatitis B virus infection in blood sera using Raman spectroscopy and machine learning. Photodiagn Photodyn Ther 23:89–93. https://doi.org/10.1016/j.pdpdt.2018.05.010
Roy S, Perez-Guaita D, Bowden S, Heraud P, Wood BR (2019) Spectroscopy goes viral: diagnosis of hepatitis B and C virus infection from human sera using ATR-FTIR spectroscopy. Clin Spectrosc 1:100001. https://doi.org/10.1016/j.clispe.2020.100001
Sakudo A, Baba K, Ikuta K (2012) Discrimination of influenza virus-infected nasal fluids by Vis-NIR spectroscopy. Clin Chim Acta 414:130–134. https://doi.org/10.1016/j.cca.2012.08.022
Hasbi NH, Bade A, Chee FP (2022) Pattern recognition for ultraviolet and fourier transform data: a walkthrough of techniques and direction. J Phys: Conf Series 2314(1):012012
Orlando A, Franceschini F, Muscas C, Pidkova S, Bartolli M, Rovere M, Tagliaferro A (2021) A Comprehensive Review on Raman Spectroscopy Applications. Chemosensors 9(262):1–28. https://doi.org/10.3390/chemosensors9090262
Kuhar N, Sil S, Taru V, Umapathy S (2018) Challenges in applications of Raman spectroscopy to biology and materials. RSC Adv 8:25888–25908. https://doi.org/10.1039/C8RA04491K
Auner GW, Koya SK, Huang C, Broadbent B, Trexler M, Auner Z, Brusatori MA (2018) Applications of Raman spectroscopy in cancer diagnosis. Cancer and Metastasis Rev 37:691–717
Desai S, Mishra SV, Joshi A, Sarkar D, Hole A, Mishra R, Dutt A (2020) Raman spectroscopy-based detection of RNA viruses in saliva: a preliminary report. J Biophotonics 13(10):1–5
Dou T, Li Z, Zhang J, Evilevitch A, Kurouski D (2020) Nanoscale structural characterization of individual viral particles using atomic force microscopy infrared spectroscopy (AFM-IR) and tip-enhanced Raman spectroscopy (TERS). Anal Chem 92:11297–11304. https://doi.org/10.1021/acs.analchem.0c01971
Rumaling MI, Chee FP, Bade A, Hasbi NH, Daim S, Juhim F, Rasmidi R (2022) Methods of optical spectroscopy in detection of virus in infected samples: a review. Heliyon 8(9):1–15
Sanchez JE, Jaramillo SA, Settles E, Velazquez Salazar JJ, Lehr A, Gonzalez J, Jose Yacaman M (2021) Detection of SARS-CoV-2 and its S and N proteins using surface enhanced Raman spectroscopy. RSC Adv 11:25788–25794. https://doi.org/10.1039/d1ra03481b
Carlomagno C, Bertazioli D, Gualerzi A, Picciolini S, Banfi PI, Lax, A.,… Bedoni, M. (2021) COVID-19 salivary Raman fingerprint: innovative approach for the detection of current and past SARS-CoV-2 infections. Sci Rep 11:1–13
Huang J, Wen J, Zhou M, Ni S, Le W, Chen G, Du K (2021) On-site detection of SARS-CoV-2 antigen by deep learning-based surface-enhanced Raman spectroscopy and its biochemical foundations. Anal Chem 93(26):9174–9182
Fan C, Hu Z, Riley LK, Purdy GA, Mustapha A, Lin M (2010) Detecting food- and waterborne viruses by surface-enhanced raman spectroscopy. J Food Sci 75(5):302–307. https://doi.org/10.1111/j.1750-3841.2010.01619.x
Azkur AK, Akdis M, Azkur D, Sokolowska M, van de Veen W, Brüggen MC, Akdis CA (2020) Immune response to SARS-CoV-2 and mechanisms of immunopathological changes in COVID-19. Allergy: Eur J Allergy Clin Immunol 75(7):1564–1581
Walls AC, Park YJ, Tortorici MA, Wall A, McGuire AT, Veesler D (2020) Structure, function, and antigenicity of the SARS-CoV-2 Spike glycoprotein. Cell 181(2):281–292. https://doi.org/10.1016/j.cell.2020.02.058
Kurouski D, Postiglione T, Deckert-Gaudig T, Deckert V, Lednev IK (2013) Amide i vibrational mode suppression in surface (SERS) and tip (TERS) enhanced Raman spectra of protein specimens. Analyst 138(6):1665–1673. https://doi.org/10.1039/c2an36478f
Zhang C, Zheng T, Wang H, Chen W, Huang X, Liang J, Tan W (2021) Rapid one-pot detection of SARS-CoV-2 based on a lateral flow assay in clinical samples. Anal Chem 93:3325–3330
Acknowledgements
The authors would like to express their gratitude towards Universiti Malaysia Sabah for supporting this research study by providing the research grant with code SDK0163-2020.
Funding
Funding is supported by Universiti Malaysia Sabah under research grant with code SDK0163-2020.
Author information
Authors and Affiliations
Contributions
FPC and AB: conceptualization. MIR, FPC, AB and LPWG: methodology. FPC and LPWG: validation. MIR and FJ: formal analysis. FPC and AB: resources. MIR and FJ: data curation. MIR, FPC, AB and LPWG: writing—original draft. FJ: visualization. FPC and AB: supervision. FPC: funding acquisition. All authors reviewed the manuscript.
Corresponding author
Ethics declarations
Conflict of interest
The authors declare no competing interests.
Ethical approval
Not applicable as this study doesn’t involve human and/ or animal studies. This study was registered at National Medical Research Register with the reference number: NMRR ID-22-02382-ZNZ.
Additional information
Publisher’s Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Rumaling, M.I., Chee, F.P., Bade, A. et al. Biofingerprint detection of corona virus using Raman spectroscopy: a novel approach. SN Appl. Sci. 5, 197 (2023). https://doi.org/10.1007/s42452-023-05419-3
Received:
Accepted:
Published:
DOI: https://doi.org/10.1007/s42452-023-05419-3