Abstract
G-protein-biased agonism of the mu-opioid receptor (μ-OR) is emerging as a promising strategy in analgesia. A deep understanding of how biased agonists modulate and differentiate G-protein-coupled receptors (GPCR) signaling pathways and how this is transferred into the cell are open questions. Here, using extensive all-atom molecular dynamics simulations, we analyzed the binding recognition process and signaling effects of three prototype μ-OR agonists. Our suggested structural mechanism of biased signaling in μ-OR involves an allosteric sodium ion site, water networks, conformational rearrangements in conserved motifs and collective motions of loops and transmembrane helices. These analyses led us to highlight the relevance of a bias-inducing allosteric binding site in the understanding of μ-OR’s G-protein-biased signaling. These results also suggest a competitive equilibrium between the agonists and the allosteric sodium ion, where the bias-inducing allosteric binding site can be modulated by this ion or an agonist such as herkinorin. Notably, herkinorin arises as the archetype modulator of μ-OR and its interactive pattern could be used for screening efforts via protein–ligand interaction fingerprint (PLIF) studies.
Article highlights
-
Agonists and a sodium ion compete for the bias-inducing allosteric binding site that modulates signaling in mu-opioid receptors.
-
Molecular dynamics simulations of the prototype μ-OR agonist suggest a competitive equilibrium involving the agonist and an allosteric sodium ion.
-
Analysis of experimental data from the literature and molecular models provides the structural bases of biased agonism on μ-OR.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
1 Introduction
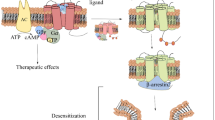
G-protein-coupled receptors (GPCRs) are seven-transmembrane (TM) helical proteins that trigger signaling pathways, mediated by interactions with heterotrimeric G-proteins and/or with β-arrestins. Preferential activation to either pathway is possible, leading to different physiological effects. Agonists that have this ability are called biased agonists, and the effect is named functional selectivity. Other modes of GPCR activation are intracellular activation, dimerization activation, transactivation, or biphasic activation [1].
Biased activation of the mu-opioid receptor (μ-OR) towards G-protein is relevant for the design of painkillers with reduced detrimental side effects. For example, activation of β-arrestin signaling pathways is suggested to lead to tolerance, desensitization, addiction, constipation, and respiratory suppression [2]. Albeit not fully understood, these effects seem to be related to β-arrestin recruitment or receptor internalization. Moreover, the coexistence of diverse and complex GPCR activation types is an active research topic. This complexity is further evidenced by the ability of GPCRs to adopt different conformations. Experimental evidence shows that different agonists can stabilize these conformations, leading to new conformational landscapes [3,4,5]. This behavior is counterintuitive, based on the paradigm that all agonists would conduct to only one active conformation [6].
Besides the biased ligands, experimental evidence shows that a conserved allosteric binding site plays a crucial role in the activation of class A GPCRs that include μ-OR [7]. Modulation of the GPCRs via this allosteric site is related to downstream signaling and physiological effects [8, 9]. The high-resolution (1.8 Å) X-ray crystal structure of the adenosine A2A receptor revealed a highly ordered cluster of atoms, composed of one sodium ion, water molecules, and conserved amino acids [8] located on TM2 and TM3, such as D2.50 and S3.39. The importance of an allosteric sodium binding site nearby D2.50 has been reviewed [7,8,9], particularly on the receptor activation process and G-protein/arrestin signaling bias. Also, the structure of the delta-opioid receptor (δ-OR), in the inactive state, showed an electron density distribution located in an allosteric site in the vicinity of D2.50, corresponding to a sodium ion and also unveiled N3.35 as an important amino acid in this binding site [9]. In agreement with these findings, mutations N3.35 V and N3.35A decreased and abolished δ-OR agonist sodium effect, respectively. Interestingly, in both cases, the δ-OR arrestin-mediated signaling pathway is increased. However, both mutations did not affect the canonical signaling, which is mediated by G-proteins. This suggests that while the allosteric sodium ion can modulate the recruitment of β-arrestin, the G-protein–receptor binding only depends on the agonist [9]. Initial experiments showed a dependence on sodium concentration for the affinity of agonists but not for antagonists [10, 11]. Antagonists/weak partial agonists are also affected by D2.50 mutations, which become β-arrestin-biased agonist [7]. Other small molecules such as amiloride can compete with the allosteric sodium site [9, 12, 13].
Mutations at D2.50 and N3.35 in δ-OR at varied concentration of Na+ affect recruitment of β-arrestin, most likely by affecting the TM7 helical bundle [9]. Similarly, the relevance of TM7 movement is suggested in arginine-vasopressin type 2 receptor (V2R) [14] using fluorescence spectroscopy; and in β2-adrenergic receptors (β2AR) [15] characterized by 19F-NMR experiments. These studies revealed that movements at TM6 helix are required for G-protein signaling. In contrast, β-arrestin-biased ligands impact the TM7-H8 region. This information could be transferable across class A GPCRs [7, 8, 16, 17] where the allosteric sodium site is preserved. To note, experimental information regarding the allosteric sodium site in μ-OR is scarce [18, 19], but many computational studies have shown the collapse of the allosteric sodium site in the activation and the egress pathways of the sodium ion in the presence of agonists and modulators [19,20,21]. These studies also suggest the lack of spontaneous release of the sodium ion from the bundle which required random accelerated molecular dynamics simulations to explore possible outlets from the allosteric site [19].
This important behavior has motivated the search for allosteric modulators in μ-OR. For example, ignavine increases agonist response for DAMGO and has positive modulatory activity in morphine and DAMGO assays. Furthermore, ignavine produces an analgesic effect in vivo through opioid response, proved by tail-flick and tail pressure tests [22]. However, the mechanistic modulation transmission in opioid receptors is still unclear. The positive allosteric modulator (PAM) BMS-986122 of μ-OR, which suppresses the β-arrestin recruitment in conjunction with agonists [23], can decrease the ability of NaCl to modulate the binding affinity of agonists and has been proposed to act as an indirect competitor of the allosteric sodium ion [18]. Additionally, molecular dynamics (MD) simulations of BMS-986122 suggested that residues on TM7 (7.53 and 7.35) are involved in the positive allosteric signal transmission [24]. Notably, the structure of BMS-986122 contains motifs related to C2-benzyloxy of herkinorin; this is in agreement with our suggested binding mode of herkinorin involving modulation of TM7 [25].
Comparing the X-ray structures of μ-OR in the active [26] and inactive [27] states allows analyzing helical rearrangements related to the activation process. These rearrangements are common to other members of the class A GPCR superfamily (β-2AR, rhodopsin, M2R) [7, 8, 16], such as helical movements at TM6 and TM7. Moreover, changes in water-mediated interactions involving amino acids inside the receptor’s cavity are observed when the activation is established [26]. The active state of the receptor is characterized by an ordered network involving water molecules. Notably, careful analysis of the water network shows that it is functionally relevant, transferring information from the extracellular side (ligand binding site) through the receptor [26], and reaching the interior of the cell. This last step is characterized by collective movements of intracellular regions of the TMs and loops. This information relates to the exquisite modulation of the receptor that results from ligand binding.
Interestingly, herkinorin, a non-nitrogenous μ-OR agonist, does not produce β-arrestin2 recruitment and μ-OR cell internalization, even when the G-protein-coupled receptor kinase 2 (GRK2) is overexpressed. This is remarkable because GRK2 is known to recognize and phosphorylate activated GPCRs [28, 29], promoting β-arrestin recruitment and internalization [30]. In turn, morphine-activated μ-OR generates limited β-arrestin2 recruitment and undergoes low μ-OR internalization under normal conditions. However, if GRK2 is overexpressed, internalization is readily promoted. In contrast, DAMGO (H-Tyr-D-Ala-Gly-N(Me)Phe-Gly-ol), a selective μ-OR peptide agonist [31], recruits β-arrestin2 and undergoes μ-OR internalization under normal conditions [29]. We previously proposed a binding mode of herkinorin into μ-OR. Our model suggests that herkinorin’s C2-benzyloxy group locates in the allosteric sodium binding site. Binding free energy calculations of the herkinorin’s proposed binding mode are in full agreement with experimental data [25]. To note, small structural modifications affect binding selectivity across opioid receptor subtypes as well as biased signaling preference. For instance, salvinorin A, a precursor of herkinorin, is a selective agonist of the kappa opioid receptor (κ-OR) and significantly recruits arrestin [32, 33].
In this work, we present the structural basis of biased agonism in μ-OR with particular emphasis on the allosteric sodium site, what we call bias-inducing allosteric binding site. Taking advantage of the signaling differences for herkinorin, morphine and DAMGO, molecular dynamic simulations were performed with these agonists, while conditions in the allosteric sodium site were studied. We observed specific conformations for the NPxxY conserved motif [9], associated with the modulation at the bias-inducing allosteric binding site. Also, we concluded that this allosteric site can modify the water distribution in the intracellular side of μ-OR, in the presence of the allosteric sodium ion. We found that herkinorin can modulate this allosteric binding site preserving one conformation of NPxxY. In contrast, morphine and DAMGO systems in the absence of allosteric sodium display two extra different conformations that can be responsible for the non-canonical signaling due to the lack of modulation in this allosteric site. Finally, the conjunction between agonist and the bias-inducing allosteric binding site can regulate intracellular loops movement and the final part of transmembrane helices that are responsible for coupling macromolecular structures.
2 Results
The μ-OR biased signaling mechanism, proposed in this work, initiated at the orthosteric binding site continues via an allosteric binding site [26], then reaches a hydrophobic barrier (framed by DRY and NPxxY conserved motifs) [34, 35] to finalize in the C-terminal helix H8, as depicted in Fig. 1. To explore different activation mechanisms, we analyzed agonists with a variety of downstream signaling effects, such as β-arrestin recruitment and receptor internalization, as well as the impact of an allosteric sodium ion. The three agonists studied here, shown in Fig. 1, are herkinorin, morphine, and DAMGO; we incorporated naloxone as the antagonist reference. Based on βarr2-GFP and confocal microscopy assays, herkinorin does not recruit β-arrestins2 and does not internalize μ-OR; DAMGO recruits β-arrestins2 and internalizes μ-OR, and morphine undergoes these processes when GRK2 is overexpressed [2, 29]. To note, based on non-imaging-based β-arrestin recruitment assays, the degree of biased agonism of herkinorin has shown dependence on experimental conditions. The presence of fusion tags directly attached to the C terminus of GPCRs may explain the differences observed in these experiments [36, 37]. For clarity, the systems proposed in this work as predominant are highlighted in italics throughout the text.
Specific regions involved in the activation of μ-OR. The blue region is the agonist binding site, yellow is the allosteric sodium pocket, brown regions are the limits of the hydrophobic barrier (framed by NPxxY and DRY motifs), and purple is the H8 helix. Chemical structures of the three agonists studied in this work are shown to the right. As reference, the trend of β-arrestin2 recruitment is shown in grayscale underneath the chemical structures. Darker color represents higher recruitment based on βarr2-GFP assays [29]
Binding poses were selected based on docking studies; see description in the Supplementary Material. MD simulations were carried out in triplicates. The allosteric sodium ion effect was studied for all systems except for herkinorin. The herkinorin C2-benzyloxy group occupies the allosteric sodium site, near N1503.35 [25]. In the absence of the allosteric sodium ion, the D2.50 amino acid was set as protonated, as suggested in previous pKa studies of GPCRs [38]. The simulation stability was assessed following the RMSD of the protein backbone and the ligand (Figures S1 and S2) clearly, all the proteins reached equilibrium. Interestingly, the conformational stability of DAMGO (+Na+) is affected by the presence of allosteric sodium.
In what follows, we present analyses of the different systems regarding conformational changes in conserved motifs, differences in water distributions, and the dependence of the hydrophobic barrier and helical motions on the ligand studied. This information is confronted with experimental information available.
2.1 Agonist and allosteric modulation affect conformations around N332
NPxxY is a highly conserved motif for class A GPCRs; it is located in the intercellular region of TM7 and is involved in the activation processes and modifications of a hydrophobic barrier [39]. Experimental data show that mutations in the NPxxY motif of β2AR or bradykinin B2 (B2R) affect coupling with G-proteins, exhibiting phosphorylation, and subsequent sequestration of these receptors [38, 40,41,42]. The conformation of NPxxY in μ-OR active crystal structure shows that N3327.49 and Y3367.53 residues interact with the backbone carbonyl group of V2856.40 through a specific water-mediated polar network (Figure S3). This interaction has also been observed in X-ray crystal structures of rhodopsin [26]. Notably, in the inactive state of μ-OR, the water-mediated interaction between Y3367.53 and V2856.40 is broken.
We analyzed the interactions involving the NPxxY motif and the side-chain dihedral angles χ1 and χ2 of N3327.49, which were sampled during the simulations. In our systems, the residues Y7.53, N7.49 (of NPxxY) and V6.40 interact similarly to those observed in crystal structures of class A GPCRs (Fig. 2 and Figure S3). Our data are also in agreement with the number and location of specific water molecules. X-ray crystal structures show water-mediated interactions between N3327.49 and V2856.40, involving two water molecules for active states and one water molecule for inactive states. Crystal structures also show different water-mediated interactions for the residue Y3367.53. While in the inactive state this residue interacts with N1.50; in the active form, it interacts with V6.40.
Networks and conformations of the conserved motif NPxxY. Comparison of the NPxxY water-mediated interaction in μ-OR and χ2 angle conformations of N3327.49. The systems are colored by agonist and the presence of the allosteric sodium ion. The probability distribution function (PDF) was calculated from three simulations of 500 ns each
We observed three possible χ2 angle conformations of N3327.49. The first one is centered at −23 degrees and was observed for agonist systems herkinorin(−Na+), morphine(+Na+), DAMGO(+Na+) and the reference antagonist system naloxone(+Na+). This value is comparable with the one observed in the X-ray crystal structure of μ-OR in the active state (PDBID: 5C1M) [26], being χ2 −13 degrees. The other two conformations populate χ2 dihedral angles at 85 and −120 degrees. These conformations were significantly populated by morphine(−Na+) and DAMGO(−Na+). The cryo-electron microscopy (cryo-EM) structure of μ-OR which bound the agonist DAMGO (PDBID: 6DDE) [43] exhibits the conformation with χ2 dihedral angle centered at −82 degrees (Figure S3).
For the dihedral angle χ1, two distributions centered at −72 and −162 degrees were observed in the systems that involve agonists (Figure S4). In the naloxone(+Na+) system only the distribution at −72 degrees is favored. Remarkably, the dihedral angle χ1 at −162 degrees was more populated in the morphine(−Na+) and DAMGO(−Na+) systems with respect to morphine(+Na+) and DAMGO(+Na+) systems which populated distributions at −72 degrees. In turn, the herkinorin(−Na+) system retained the conformation at −72 degrees as the system in the presence of the allosteric sodium ion. The experimental information discussed previously shows that χ1 angle is found at −163 degrees for the cryo-EM structure 6DDE and −76 degrees for the X-ray crystal 5C1M of μ-OR.
All in all, this information suggests that, in the absence of sodium ion, herkinorin is modulating the NPxxY motif (TM7), thus affecting downstream signaling. In contrast, morphine and DAMGO are dependent on allosteric sodium ion modulation, which is lost in the activation process. Thereby herkinorin seems to be a sodium-independent modulator of G-protein signaling. It is expected that the μ-OR in the active state, obtained bound to a G-protein mimetic nanobody (Nb39), retains NPxxY configurations, just as it does when it interacts with G-proteins. The fact that herkinorin retains the conformation observed by NPxxY in the crystal structure agrees with our model.
Let us describe the second and third conformations observed for the χ2 dihedral angle. Our simulations show that morphine(−Na+) and DAMGO(−Na+) populated χ2 at 85 or −120 degrees. Additionally, water molecules mediating N3327.49−V2856.40 interaction are relocated, and they no longer interact with Y3367.53. Furthermore, the interaction between D1142.50 and N3327.49 nitrogen was mediated by one water molecule (Fig. 2). This suggests that different conformations of NPxxY are involved in different downstream signaling pathways, such as phosphorylation in Y3367.53 when it is not participating in the water network interaction. Non-canonical signaling has been tested experimentally through Y3367.53 phosphorylation of μ-OR [44].
For the observed changes in NPxxY, we propose a dynamic equilibrium in morphine(+Na+)/morphine(−Na+) and DAMGO(+Na+)/DAMGO(−Na+) systems. The equilibrium seems to be favored towards morphine(−Na+) or DAMGO(−Na+). Moreover, experimentally DAMGO promotes internalization of the receptor more easily than morphine. The conformation that seems to be related to this signaling pathway is the one with χ2 dihedral angle at either 85 or −120 degrees. Similar conclusions can be drawn from the analysis of χ1. These configurations are favored in morphine(−Na) and DAMGO(−Na) systems. We then explored if water networks were also distinctive between agonists.
2.2 Water network is involved in activation but not in biased agonism
Crystal structures of GPCRs have shown functionally relevant water networks. Inspection of our simulations provided relevant water-mediated intercommunications in the orthostatic site. In all the systems studied, the most remarkable water-mediated interaction was primarily observed between H2976.52 and the ligand. The exception is for DAMGO(–Na+/ +Na+), which could make this contact in the presence or absence of water molecules. In the active state, the μ-OR crystal structure shows a water-mediated interaction between H2976.52 and its co-crystal agonist BU72 [26]. This interaction is stabilized by the backbone carbonyl group of K2335.39, similarly to our systems of morphine(+Na±Na+) and herkinorin. However, herkinorin(−Na+) retained this interaction through the carbonyl group of C4-methyl ester (Figure S5). Additionally, H2976.52 interconnects with D1142.50 by four water molecules on average; this interaction was observed for all the systems studied except DAMGO(−Na+), where W2936.48 disrupts the network. If the sodium ion was present, it contributed to mediate this interaction. Other different networks in the crystal structures are the interaction mediated by one or two water molecules, observed between ligands and amine group of K2335.39 or the water-mediated interactions between D1142.50- D1473.32 amino acids that were stabilized by neighbors amino acid. Thus, water networks are relevant for the activation process, but it is not clear if they are involved in the transmission of biased signaling information.
2.3 Distribution of water molecules and hydrophobic barriers through the receptors
Differences in the distribution of water molecules across the different agonists were two-fold, from one side the probability of finding voxels (cavities) inside of the receptor and on the other side on the topology of the water distribution itself, as shown in Fig. 3. Voxels are colored by the probability of finding water molecules across the simulations. Red and blue colors represent high and low probability, respectively. The topology of the volume occupied by water molecules was characterized by a narrow region (Fig. 3). This narrow region involves interactions between M1613.46, R1653.50, and Y3367.53 (Figure S6). Interestingly, R1653.50 and Y3367.53 are part of the conserved DRY and NPxxY motifs, respectively. These two structural motifs are the limits of a large hydrophobic barrier formed by approximately six amino acids on the inactive state of class A GPCRs where in the activation process, once the hydrophobic barrier opens, hydrogen bonding interactions emerge between DRY and NPxxY motifs, and the hydrophobic gap rearranges, as shown experimentally by Schertler’s group [34, 35]. In our work, for the naloxone(+Na+) system, the probability of finding water molecules on the intracellular part of μ-OR is low (no green and red regions, Fig. 3). The distance between R1653.50–Y3367.53 is larger than the one observed in the agonist systems (Table 1), supporting a hydrophobic barrier kept in the inactive state.
Interestingly, the distribution of water molecules in the intracellular part of μ-OR was denser for morphine(−Na+), DAMGO(−Na+), and herkinorin(−Na+) complexes than the system in the inactive state or morphine(+Na+) and DAMGO(+Na+). Correspondingly, our data show comparable conformations to those observed at M(L)3.46, R3.50, and Y7.53 in μ-OR and rhodopsin crystal structures in the active state (Figure S6).
2.4 The hydrophobic barrier in the herkinorin/μ-OR system
Notably, the herkinorin(−Na+) system has a short and modified hydrophobic region (enclosed by DRY and NPxxY motifs) under the parameters implemented in the trj_cavity tool. This could be explained by the close interaction between R1653.50 and Y3367.53 in the herkinorin(−Na+) system (Table 1). MD simulations of oliceridine in μ-OR (which mitigates the β-arrestin recruitment) highlight the relevance of the interaction with R1653.50 in the transmission of information initiated by biased ligands [45].
The changes in the N3327.49 conformation and the water-mediated interactions at NPxxY (involving Y3367.53) did not seem to affect the short hydrophobic barrier in all systems with agonist due to no significant differences for R1653.50–Y3367.53 distances. It is consistent with the activation processes in GPCR. Although water distribution differences were observed in the intracellular part of μ-OR with agonists, this can be attributed to the presence of the allosteric sodium ion. These observations suggest that the changes observed in the conformation of N3327.49 in the absence of the allosteric sodium ion do not affect the activation mechanism but could be responsible for alternative downstream signaling.
2.5 Structural helical and loop motions: functional mode analysis (FMA)
We performed an extended analysis of the μ-OR backbone chain’s structural collective motions and preservation of helicity over TM7 information. Typically, principal component analysis (PCA) provides compacted information about projections or principal components (PCs), which means separated motions are contributing to the mean square fluctuation (MSF) of atomic coordinates. To get a large fraction (80–90%) of atomic MSF, one should analyze the first 10–20 PCs in protein simulation [46, 47]; however, analysis of 10 PCs is demanding. In contrast, FMA builds a vector formed by PCs (up to 50), representing a collective motion (just one projection) that is directly related to a time-dependent biological function. In our work, the collective motion was calculated as ewMCM (ensemble-weighted maximal correlation motion) related to the RMSD of the backbone chain. ewMCM is more dependent on PCs and protein energetic maps than MCM (maximally correlation motion) [46]. Table 2 shows the number of selected PCs to obtain the best model (Rm) and the best cross-validation (Rc) related to the RMSD of the protein backbone of μ-OR. Figure 4 and Figure S7 show the ewMCM oscillation between extremes \(p_{a}^{min}\) and \(p_{a}^{max}\) collective motions. As mentioned earlier, our systems reached equilibrium, as shown by the steady changes in backbone-RMSD values. Thus, the collective motion is thought of as harmonic oscillation.
2.6 ICL2, ICL3, ECL2, and H8 motion
The collective motions in morphine(−Na+) and DAMGO(−Na+) presented remarkable oscillations in the intracellular loops (ICL) ICL2 and ICL3 (Fig. 4). These two intracellular loops, along with TM3, TM5, and TM6, are described as the primary interactive sites in GPCR–G-protein complexes [43]. In contrast, herkinorin(−Na+) stabilized ICL2 diminishing the oscillations in the three simulations that might help G-protein signaling. As morphine(+Na+) and DAMGO(+Na+) also restrained the movements in ICL2, the modulation in the allosteric sodium site may affect the ICL2 oscillations. Morphine(+Na+) and DAMGO(+Na+) showed oscillations at TM1 (Figure S7), which was only seen on the naloxone(+Na+) system, indicating instability of the active state of μ-OR. This information supports the idea that the equilibrium morphine(+Na+)/morphine(−Na+) favors morphine(−Na+) system and that the DAMGO(+Na+)/DAMGO(−Na+) equilibrium favors DAMGO(−Na+). The extracellular loop (ECL) ECL2 in herkinorin(−Na+) is modified concerning the other systems, leading to more oscillations. ECL2 has been linked to the positive allosteric modulation of ligand binding in μ-OR to G-protein signaling as the binding mode of the biased agonist oliceridine [45, 48].
Interestingly, the DAMGO(−Na+) system has a preponderant collective backbone motion over the H8 helix compared to morphine(−Na+) and herkinorin(−Na+) systems. Under normal experimental conditions, DAMGO induces μ-OR internalization and phosphorylation at T370, and S375 amino acid of H8 could explain recruitment by arrestins [29]. Moreover, the computational work of oliceridine suggests that F347(H8) is one of the most relevant amino acids in the transmission of information in μ-OR [45]. This means sizable and specific H8 motion would facilitate the sampling of conformations that produce phosphorylation, which can, in turn, induce receptor recruitment.
DAMGO(−Na+) system shows functionally relevant motion at H8, and the conformation at N3327.49 (in NPxxY motif) was different from DAMGO(+Na+), morphine(+Na+), and herkinorin(−Na+), suggesting that the conformation of N3327.49 would have influenced TM7 downstream signaling to affect H8 motion. All in all, systems started signaling in the orthosteric-position of agonist and the modulation of the allosteric sodium site, passing by the modification of NPxxY, which altered H8 motion. Although the herkinorin(−Na+) system had no allosteric sodium ion, it did not produce larger motion at H8, particularly noticeable in two of the simulation replicas, explained by the retention of N3327.49 conformation. This is similar to the one observed in the active state crystal structure because the C2-benzyloxy group of herkinorin modulated the allosteric sodium site preventing H8 and NPxxY changes. Interestingly, in the herkinorin(−Na+) simulation where H8 displacement was observed (simulation 2), the ECL2 did not suffer any change.
2.7 Helicity
During the MD simulations, the degree of helicity was modified (Figure S8). For example, the herkinorin(−Na+) system had lower degrees of helicity, such as morphine(−Na+) and DAMGO(−Na+), at W3187.35 (Fig. 4). Also, the herkinorin(−Na+) system showed loss of helicity at C3367.53 and L3357.52, which are more stable compared to morphine(−Na+) and DAMGO(−Na+) systems. Because of N3327.49 proximity, the unstable helicity in this region could be affected by the χ2 angle conformations of N3327.49 in morphine(−Na+) and DAMGO(−Na+). Interestingly, it has been proposed that W3187.35 and C3367.53 are involved in the TM7 transmission of mechanistic signaling, creating flexions at TM7 in the presence of a PAM [24]. By this means, herkinorin could induce positive downstream signaling over TM7.
2.8 The relation of binding mode with functional selectivity in μ-OR. Protein–ligand interactions
In previous studies, we [25] and others [49, 50] have explored protein–ligand interaction fingerprints (PLIF) to classify and identify biased vs. non-biased ligands. Here, we generated a PLIF based on the contacts obtained from the MD simulations and compared biased vs. non-biased μ-OR agonists. Protein–ligand contacts within a radius of 4.0 Å were taken into account. Analysis of the contacts for herkinorin(−Na+), morphine(−Na+), and DAMGO(–Na+) systems (Fig. 5), showed amino acids that make contacts during the entire simulation time, for example, D1473.32, I2966.51, V3006.55, W3187.35, and I3227.39. We also observed that these contacts were present for the naloxone(+Na+) system (Figure S9) during more than 90 percent of the simulation time. Interactions involving these amino acids are found in active and inactive crystal BU72-μ-OR and could be a reasonable representation of the orthosteric binding site. Prevalent and unique contacts of the herkinorin(−Na+) system corresponded to two allosteric binding sites. The first one encompasses the amino acids A1172.53, V1433.28, and C217ECL2 (TM2, TM3 and ECL2), which have been reported for the biased agonist oliceridine (TRV-130) (Q1242.60, V1433.28, and C217ECL2) [45]. Oliceridine promotes G-protein signaling [51], similar bias-agonists mitragynine and NAP also interact with these allosteric amino acids [36, 52, 53]. The second binding site for herkinorin included contacts with N1503.35 and S3297.46, corresponding to the allosteric sodium region. Consistently, a representative pose of oliceridine from the MD simulations makes contact with S3297.46 through its methoxy-thiophene moiety [45].
2.9 Binding mode is affected by the allosteric sodium ion
Morphine has the same PLIF profile in the presence or absence of the allosteric sodium ion with a small variation in L2335.39, but DAMGO did not (Figure S9). The lack of allosteric sodium ion DAMGO researched extracellular regions on ECL2 and TM5 making contacts with T218ECL2, L219ECL2, and L2325.38, whereas the DAMGO(+Na+) system makes contacts with N1272.63, W133ECL1, and V1433.28, interacting with the extracellular part of TM2-TM3 and ECL1. It is important to note that these contact differences were mostly due to placements of Phe and Gly-ol group in DAMGO. If DAMGO(−Na+) system predominates in the equilibrium DAMGO(+Na+)/DAMGO(−Na+), β − arrestin signaling will be favored because, in contrast to biased agonists, the extracellular region of TM2 and TM3 cannot be reached all time.
3 Discussion
In summary, analysis of water distribution, conformational _ in NPxxY, and water networks suggested that DRY, NPxxY motifs and intracellular part of μ-OR are affected by agonist presence and the modulation of the allosteric sodium binding site. The main alteration in the NPxxY motif (in the hydrophobic limit) was on the conformation around the χ2 angle of N3327.49 amino acid and the water-mediated network. The reason for this alteration seems to be related to the modulation of the allosteric sodium pocket but not in the orthosteric site. The binding modes observed for morphine and DAMGO did not affect the conformation at the NPxxY motif. The conformation involving N7.49 and water molecules in DAMGO(−Na+) and morphine(−Na+) systems gave a conformation at the NPxxY motif that may explain β-arrestin signaling pathways. Notably, herkinorin(−Na+) retained a conformation at the χ2 dihedral angle as the one observed in the crystallographic structure, in the active state. In the presence of herkinorin, there is no need for an allosteric sodium ion to control χ1 and χ2 angles, granted that the C2-benzyloxy group occupies this pocket. This site could be called, G-protein bias-inducing allosteric binding site since it can be modulated by Na+ or biased ligands reaching that site. The impact of information transmission on the μ-OR’s intracellular backbone was mainly observed in H8 helix and ICL3 loop movements by FMA. In this analysis of collective harmonic motion in TMs and loops, morphine(−Na+) and DAMGO(−Na+) produce changes in the ICL2 while herkinorin(−Na+) restrains the loop movement. H8 helix in DAMGO(−Na+) has larger displacement, particularly noticeable in two of the simulations. Although H8 did not have a complete sequence in our system, it can be suggested that this motion could affect the rest of the helix. We propose an equilibrium between morphine(+Na+)/morphine(−Na+) and DAMGO(+Na+)/DAMGO(−Na+). This equilibrium may alter the transduction of the signaling pathway. The main outcome is the understanding of the modulation of the allosteric sodium pocket, exemplified by herkinorin(−Na+) through the C2-benzyloxy group. New μ-OR biased ligands can be designed by maintaining the interacting pattern obtained from the PLIF studies in the allosteric sodium region.
This work highlights the relevance of the allosteric modulation in the extracellular part of class A GPCRs, in agreement with previous studies [54]. It has been proposed that, in contrast to non-biased agonists, biased agonists with specific and extended binding modes convey a closure of the binding pocket. Biased agonists could also promote an equivalent effect for the closure of the biased-inducing allosteric binding site. Our data show that the sodium ion or herkinorin induce the retention of conformations in conserved residues of class A GPCRs. The C2-benzyloxy group of herkinorin or the sodium ion participate in the closure of the allosteric site. As a result, specific water network interactions that connect the extracellular and intracellular regions of μ-OR are also induced by the allosteric modulation.
4 Materials and methods
4.1 Molecular alignments
The flexible alignment was carried out through the software molecular operating environment (MOE) developed by Chemical Computing Group Inc. This process is useful to find structural similarity features through property density functions [55, 56]. Co-crystal ligands BU72 and β-FNA (from PDBIDs 5C1M and 4DKL) [27] were used as structural ligand references in the orthosteric binding site of active and inactive μ-OR, respectively. On the other hand, agonist morphine and antagonist naloxone were placed into μ-OR with molecular alignment since they are structurally related to ligand references. The calculation parameters were adjusted by previous work [25].
4.2 Docking
Automated docking was performed for ligand agonists DAMGO and herkinorin. Processed with the software Internal Coordinate System, (ICM) 3.8–344. All non-receptor molecules were removed, and the binding site was defined using the ICM PocketFinder algorithm45,46 with previously established [25] parameters. Amino acids found in the binding site were A1172.53, D1473.32, Y1483.33, M1513.36, K2335.39, V2365.42, F2375.43, W2936.48, I2966.51, H2976.52, V3006.55, I3227.39, G3257.42, and Y3267.43. Ligand poses were selected considering structural experimental and computational information and previously generated models [25, 57,58,59].
4.3 Molecular dynamics simulations setup
Active and inactive states of μ-OR were obtained from their corresponding X-ray coordinates (PDBIDs 5C1M and 4DKL, respectively) [9, 27]. Amino acid sequence range was from M65 to F347 in the active state, whereas the range in the inactive state was from M65 to I351. First, all non-receptor molecules were removed. Then, acetyl and methylamide capping groups were placed on N-terminal and C-terminal amino acids, and the disulfide bond between the amino acids C140-C217 was inserted. The missing amino acids on ICL3 (MLSGSK sequence) in the inactive state were included with ProMod Version 3.70 implemented in SWISS-MODEL [60,61,62]. Histidine residues were simulated as the neutral tautomers. Protonation states of all amino acids were assigned at pH 7 except for D1142.50, protonated only in allosteric sodium ion absence. To study the presence of the allosteric sodium ion, the residue D1142.50 was charged negatively, and the sodium was placed around it. Conversely, in the absence of allosteric sodium ion, the residue was kept neutral [24, 26, 63]. Systems with allosteric sodium ion included: naloxone(+Na+), morphine(+Na+) and DAMGO(+Na+); and without sodium: herkinorin(−Na+), morphine(−Na+) and DAMGO(−Na+). The system of herkinorin with sodium was not considered since the C2-benzyloxy moiety of herkinorin occupies the sodium ion region.18 We used CHARMM-GUI, an online tool for membrane preparation that uses the CHARMM36 force field all-atom for protein, lipids, and salts ions [64,65,66,67,68,69,70,71]. The online tool generated systems with the receptor placed in a lipid bilayer constituted of 1-palmitoyl, 2-oleoyl-sn-glycero-3-phosphocholine (POPC, with transition phase 271), 1-palmitoyl-2-oleoyl-sn-glycero-3-phosphoethanolamine (POPE, transition phase 298 K) [72] and cholesterol (CHL), in a molar proportion 2:2:1 of POPC/POPE/CHL [73]. Na+ and Cl− ions were added to neutralize charges, reaching a final concentration of around 150 mM. The systems were solvated with type TIP3 water molecules. Also, internal solvation was performed by a simulation including the restricted receptor, water molecules, and ions (1 ns of pre-equilibrated NPT ensemble with the parameters mentioned below). The water molecules overlapping the ligand coordinates were deleted. CHARMM General Force Field (CGenFF) 1.0.0 was implemented for μ-OR ligands [74,75,76].
4.4 Molecular dynamics simulations
Simulations were carried out using GROMACS 5.0.4, and three simulations were performed for each system [77]. Energy minimizations were conducted using the steepest descent algorithm. Equilibration and MD runs were performed with the leap-frog integration algorithm at 310 K and 1 bar. The systems were equilibrated through seven steps decreasing harmonic restrictions on protein, lipid, and ligands (Table S1). Runs corresponding to production step consisted of 500 ns with 2 fs time step applying the Nose–Hoover thermostat (relaxation time 2 ps) and the semi-isotropic Parrinello-Raman barostat (relaxation time 5 ps). Bond lengths to hydrogen atoms were constrained with LINCS. Receptor–ligand, lipids, and ion–solvent sets were considered as three separated temperature-coupling groups. Non-bonded interactions used the Verlet scheme with a 12.0 Å cut-off. Long-range electrostatic interactions were computed using the particle mesh Ewald (PME) method, and the Lennard–Jones (LJ) interaction smoothly decayed to zero crossing 10 Å distances. The initial conformations of morphine and naloxone were based on the molecular alignment to the crystallized ligand coordinates in the active and inactive states, respectively. The conformation and binding mode of DAMGO and herkinorin were selected from docking studies, preceding binding models, and experimental information [25, 57, 58, 78,79,80].
4.5 Water molecules distribution in μ-OR
The analysis was performed with the tool trj_cavity [81] implemented within the GROMACS package. This tool is handy to identify and detect cavities in MD trajectories with voxels (volume pixel), including solvent analysis. The number of surrounding protein directions was set to six, ideal to find internal protein cavities. Water molecules density distribution was extracted according to occupation frequency of water in a cavity voxel through 500 ns MD.
4.6 Amino acid contact
The amino acid contact analysis was undertaken with the MDAnalysis library in the python language [82, 83]. Protein–ligand interaction contacts were calculated on 500 ns MD. First, we selected amino acids for each 0.5 ns time step with distances less than 4.0 Å from ligand atoms. Afterward, frequency contact was extracted for each amino acid over all the simulation time. Finally, contacts with less than 5% of frequency were not considered.
4.7 Functional mode analysis
We analyzed the collective atomic motions of μ-OR using the methodology known as functional mode analysis (FMA) [46]. This technique allows detecting collective motion related to a function such as a root mean square deviation (RMSD). The maximally correlated motion (MCM) extracted from FMA represents a functional state in terms of a single collective coordinate. Then, the motions that display the largest possibility to induce a change in the functional quantity are estimated from the given protein ensemble.
Hence, we diagonalized the covariance matrix \(C\) to obtain eigenvector \(e_{i}\) and eigenvalues \(\sigma_{i}^{2}\) to build principal components \(p_{i}\).
FMA uses \(e_{i}\) to establish an MCM that depends on the time by the projection \(p_{a}\) along the normalized collective vector \(a\) correlated to the change in the RMSD or some function \(f\) through Pearson’s coefficient \(R\).
The ensemble-weighted maximally correlated motion (ewMCM) is related as the most probable collective motion \(v^{*}\) generating a functional state \(p_{a}^{*}\) that is an arbitrary and fixed \(p_{a}\) value between extremes \(p_{a}^{\min }\) and \(p_{a}^{\max }\).
\(p_{a}^{*} = v^{*} \cdot a\) For each value of pa*, a set of pi* is given and related by the expression.
For each value of \(p_{a}^{*}\) a set of \(p_{i}^{*}\) is given and related by the expression
We can visualize the ewMCM for the given ensemble according to the expression
Since \(p_{a} \left( t \right)\) is a collective motion dependent on time, \(p_{a}^{\min }\) and \(p_{a}^{\max }\) are motions in the minimum or maximum of RMSD around the average. If the system reaches equilibrium (constant RMSD), an oscillatory motion will be observed. We visualized the μ-OR backbone (−N–Cα–CO–) ewMCM correlated to RMSD in all systems since this weighted collective motion is more dependent on PCs and protein energetic maps than just the MCM [46]. We used 500 ns MD to perform FMA and then selected the best PCs set based on correlation coefficients of models and cross-validation. Models were built from the first 400 ns, while the last 100 ns completed the cross-validation. Table 2 gives the PCs number used in the FMA.
4.8 Helicity
Data were extracted by GROMACS 5.0.4 inbuilt tools gmx helix, based on an ideal helix that is formed by at least three consecutive amino acids with ideal Φ and Ψ dihedral angles (axis of rotation on N-Cα and Cα-CC=O, respectively) [84, 85]. For each amino acid, both time percentage and α-helix structure on 500 ns were calculated from MD.
5 Conclusion
The molecular dynamic simulations for μ-OR performed here show that the modulation of the bias-inducing allosteric binding site influences conformational changes in the conserved NPxxY motif of class A GPCRs. Also, the water distribution in the intracellular part of μ-OR can be attributed to this allosteric site. In the absence of the allosteric sodium ion, the agonist herkinorin can modulate the bias-inducing allosteric binding site and can preserve the conformation of NPxxY (TM7). This can explain the biased agonism of herkinorin for G-protein signaling in μ-OR. In contrast, morphine and DAMGO systems in the absence of the allosteric sodium ion display two different NPxxY conformations that can be responsible for the lack of G-protein canonical signaling. Lastly, the conjunction between agonist and the modulation of the bias-inducing allosteric binding site in μ-OR appears essential for the regulation of movements at intracellular loops and transmembrane helices. These regions directly interact and recognize G-proteins or arrestins.
References
Wang W, Qiao Y, Li Z (2018) New Insights into Modes of GPCR Activation. Trends Pharmacol Sci 39(4):367–386
Bohn LM, Gainetdinov RR, Lin F-T, Lefkowitz RJ, Caron MG (2000) μ-Opioid receptor desensitization by β-arrestin-2 determines morphine tolerance but not dependence. Nature 408:720. https://doi.org/10.1038/35047086
Kenakin T (2011) Functional selectivity and biased receptor signaling. J Pharmacol Exp Ther 336(2):296–302. https://doi.org/10.1124/jpet.110.173948
Onaran HO, Costa T (1997) Agonist efficacy and allosteric models of receptor action. Ann NY Acad Sci 812(1):98–115. https://doi.org/10.1111/j.1749-6632.1997.tb48150.x
Onaran HO, Scheer A, Cotecchia S, Costa T (2000) A look at receptor efficacy. from the signalling network of the cell to the intramolecular motion of the receptor. In: Kenakin T, Angus JA (eds) The pharmacology of functional, biochemical, and recombinant receptor systems. Springer, Berlin, pp 217–259
Kenakin T, Miller LJ (2010) Seven Transmembrane receptors as shapeshifting proteins: the impact of allosteric modulation and functional selectivity on new drug discovery. Pharmacol Rev 62(2):265–304. https://doi.org/10.1124/pr.108.000992
Katritch V, Fenalti G, Abola EE, Roth BL, Cherezov V, Stevens RC (2014) Allosteric sodium in class A GPCR signaling. Trends Biochem Sci 39(5):233–244. https://doi.org/10.1016/j.tibs.2014.03.002
Liu W, Chun E, Thompson AA, Chubukov P, Xu F, Katritch V, Han GW, Roth CB, Heitman LH, Ijzerman AP, Cherezov V, Stevens RC (2012) Structural basis for allosteric regulation of GPCRs by sodium ions. Sci (NY) 337(6091):232–236. https://doi.org/10.1126/science.1219218
Fenalti G, Giguere PM, Katritch V, Huang X-P, Thompson AA, Cherezov V, Roth BL, Stevens RC (2014) Molecular control of δ-Opioid receptor signaling. Nature 506(7487):191–196. https://doi.org/10.1038/nature12944
Pert CB, Snyder SH (1973) Opiate receptor: demonstration in nervous tissue. Science 179(4077):1011–1014
Snyder SH, Pasternak GW (2003) Historical review: opioid receptors. Trends Pharmacol Sci 24(4):198–205. https://doi.org/10.1016/s0165-6147(03)00066-x
Heitman LH, Ye K, Oosterom J, Ijzerman AP (2008) Amiloride derivatives and a nonpeptidic antagonist bind at two distinct allosteric sites in the human gonadotropin releasing hormone receptor. Mol Pharmacol 73(6):1808
Gao Z-G, Melman N, Erdmann A, Kim SG, Müller CE, Ijzerman AP, Jacobson KA (2003) Differential allosteric modulation by amiloride analogues of agonist and antagonist binding at A(1) and A(3) adenosine receptors. Biochem Pharmacol 65(4):525–534. https://doi.org/10.1016/s0006-2952(02)01556-3
Rahmeh R, Damian M, Cottet M, Orcel H, Mendre C, Durroux T, Sharma KS, Durand G, Pucci B, Trinquet E, Zwier JM, Deupi X, Bron P, Banères J-L, Mouillac B, Granier S (2012) Structural insights into biased G protein-coupled receptor signaling revealed by fluorescence spectroscopy. Proc National Acad Sci 109(17):6733–6738
Liu JJ, Horst R, Katritch V, Stevens RC, Wüthrich K (2012) Biased signaling pathways in β2-adrenergic receptor characterized by 19F-NMR. Science 335(6072):1106–1110
Friedman S, Tauber M, Ben-Chaim Y (2020) Sodium ions allosterically modulate the M2 muscarinic receptor. Sci Rep 10:11177
Zhang C, Srinivasan Y, Arlow DH, Fung JJ, Palmer D, Zheng Y, Green HF, Pandey A, Dror RO, Shaw DE, Weis WI, Coughlin SR, Kobilka BK (2012) High-resolution crystal structure of human protease-activated receptor 1. Nature 492(7429):387–392
Livingston KE, Traynor JR (2014) Disruption of the Na(+) ion binding site as a mechanism for positive allosteric modulation of the mu-opioid receptor. Proc Natl Acad Sci U S A 111(51):18369–18374. https://doi.org/10.1073/pnas.1415013111
Shang Y, LeRouzic V, Schneider S, Bisignano P, Pasternak GW, Filizola M (2014) Mechanistic insights into the allosteric modulation of opioid receptors by sodium ions. Biochemistry 53:5140–5149
Selent J, Sanz F, Pastor M, De Fabritiis G (2010) Induced effects of sodium ions on dopaminergic G-protein coupled receptors. PLoS Comput Biol 6(8):e1000884. https://doi.org/10.1371/journal.pcbi.1000884
Yuan S, Vogel H, Filipek S (2013) The role of water and sodium ions in the activation of the μ-opioid receptor. Angew Chem Int Ed 52:10112–10115
Ohbuchi K, Miyagi C, Suzuki Y, Mizuhara Y, Mizuno K, Omiya Y, Yamamoto M, Warabi E, Sudo Y, Yokoyama A, Miyano K, Hirokawa T, Uezono Y (2016) Ignavine: a novel allosteric modulator of the μ opioid receptor. Sci Rep 6:31748. https://doi.org/10.1038/srep31748
Burford NT, Clark MJ, Wehrman TS, Gerritz SW, Banks M, O’Connell J, Traynor JR, Alt A (2013) Discovery of positive allosteric modulators and silent allosteric modulators of the μ-opioid receptor. Proc Natl Acad Sci 110(26):10830–10835
Bartuzi D, Kaczor AA, Matosiuk D (2016) Interplay between Two allosteric sites and their influence on agonist binding in human μ Opioid receptor. J Chem Inf Model 56(3):563–570. https://doi.org/10.1021/acs.jcim.5b00705
Marmolejo-Valencia AF, Martínez-Mayorga K (2017) Allosteric modulation model of the mu opioid receptor by herkinorin, a potent not alkaloidal agonist. J Comput Aided Mol Des 31(5):467–482. https://doi.org/10.1007/s10822-017-0016-7
Huang W, Manglik A, Venkatakrishnan AJ, Laeremans T, Feinberg EN, Sanborn AL, Kato HE, Livingston KE, Thorsen TS, Kling RC, Granier S, Gmeiner P, Husbands SM, Traynor JR, Weis WI, Steyaert J, Dror RO, Kobilka BK (2015) Structural insights into μ -opioid receptor activation. Nature 524(7565):315–321. https://doi.org/10.1038/nature14886
Manglik A, Kruse AC, Kobilka TS, Thian FS, Mathiesen JM, Sunahara RK, Pardo L, Weis WI, Kobilka BK, Granier S (2012) Crystal structure of the µ-opioid receptor bound to a morphinan antagonist. Nature 485:321–326. https://doi.org/10.1038/nature10954
Harding WW, Tidgewell K, Byrd N, Cobb H, Dersch CM, Butelman ER, Rothman RB, Prisinzano TE (2005) Neoclerodane diterpenes as a novel scaffold for μ Opioid receptor ligands. J Med Chem 48(15):4765–4771. https://doi.org/10.1021/jm048963m
Groer CE, Tidgewell K, Moyer RA, Harding WW, Rothman RB, Prisinzano TE, Bohn LM (2007) An opioid agonist that does not induce μ-opioid receptor—arrestin interactions or receptor internalization. Mol Pharmacol 71(2):549–557. https://doi.org/10.1124/mol.106.028258
Gärtner F, Seidel T, Schulz U, Gummert J, Milting H (2013) Desensitization and internalization of endothelin receptor a: impact of G protein-coupled receptor kinase 2 (GRK2)-mediated phosphorilation. J Biol Chem 288(45):32138–32148. https://doi.org/10.1074/jbc.M113.461566
Minami M, Onogi T, Nakagawa T, Katao Y, Aoki Y, Katsumata S, Satoh M (1995) DAMGO, a μ-opioid receptor selective ligand, distinguishes between μ-and κ-opioid receptors at a different region from that for the distinction between μ- and δ-opioid receptors. FEBS Lett 364(1):23–27. https://doi.org/10.1016/0014-5793(95)00340-f
DiMattio KM, Ehlert FJ, Liu-Chen L-Y (2015) Intrinsic relative activities of κ opioid agonists in activating Gα proteins and internalizing receptor: differences between human and mouse receptors. Eur J Pharmacol 761:235–244. https://doi.org/10.1016/j.ejphar.2015.05.054
Hernández-Alvarado RB, Madariaga-Mazón A, Ortega A, Martinez-Mayorga K (2020) DARK classics in chemical neuroscience: salvinorin A. ACS Chem Neurosci 11(23):3979–3992
Li J, Edwards PC, Burghammer M, Villa C, Schertler GFX (2004) Structure of bovine rhodopsin in a trigonal crystal form. J Mol Biol 343(5):1409–1438. https://doi.org/10.1016/j.jmb.2004.08.090
Trzaskowski B, Latek D, Yuan S, Ghoshdastider U, Debinski A, Filipek S (2012) Action of molecular switches in GPCRs—theoretical and experimental studies. Curr Med Chem 19(8):1090–1109. https://doi.org/10.2174/092986712799320556
Manglik A, Lin H, Aryal DK, McCorvy JD, Dengler D, Corder G, Levit A, Kling RC, Bernat V, Hübner H, Huang X-P, Sassano MF, Giguère PM, Löber S, Da D, Scherrer G, Kobilka BK, Gmeiner P, Roth BL, Shoichet BK (2016) Structure-based discovery of opioid analgesics with reduced side effects. Nature 537:185
Pedersen MH, Pham J, Mancebo H, Inoue A, Asher WB, Javitch JA (2021) A novel luminescence-based-β-arrestin recruitment assay for unmodified receptors. J Biol Chem. https://doi.org/10.1016/j.jbc.2021.100503
Vickery ON, Carvalheda CA, Zaidi SA, Pisliakov AV, Katritch V, Zachariae U (2018) Intracellular transfer of Na+ in an active-state G-protein-coupled receptor. Structure 26(1):171–180. https://doi.org/10.1016/j.str.2017.11.013
Sena DM, Cong X, Giorgetti A, Kless A, Carloni P (2017) Structural heterogeneity of the μ-opioid receptor’s conformational ensemble in the apo state. Sci Rep 7:45761. https://doi.org/10.1038/srep45761
Massink A, Louvel J, Adlere I, van Veen C, Huisman BJH, Dijksteel GS, Guo D, Lenselink EB, Buckley BJ, Matthews H, Ranson M, Kelso M, Ijzerman AP (2016) 5′-Substituted amiloride derivatives as allosteric modulators binding in the sodium ion pocket of the adenosine A2A receptor. J Med Chem 59(10):4769–4777. https://doi.org/10.1021/acs.jmedchem.6b00142
Kalatskaya I, Schüssler S, Blaukat A, Müller-Esterl W, Jochum M, Proud D, Faussner A (2004) Mutation of tyrosine in the conserved NPXXY sequence leads to constitutive phosphorylation and internalization, but not signaling, of the human B2 bradykinin receptor. J Biol Chem 279(30):31268–31276. https://doi.org/10.1074/jbc.M401796200
Schönegge A-M, Gallion J, Picard L-P, Wilkins AD, Le Gouill C, Audet M, Stallaert W, Lohse MJ, Kimmel M, Lichtarge O, Bouvier M (2017) Evolutionary action and structural basis of the allosteric switch controlling β(2)AR functional selectivity. Nat Commun 8:2169. https://doi.org/10.1038/s41467-017-02257-x
Koehl A, Hu H, Maeda S, Zhang Y, Qu Q, Paggi JM, Latorraca NR, Hilger D, Dawson R, Matile H, Schertler GFX, Granier S, Weis WI, Dror RO, Manglik A, Skiniotis G, Kobilka BK (2018) Structure of the µ-opioid receptor–Gi protein complex. Nature 558(7711):547–552. https://doi.org/10.1038/s41586-018-0219-7
Zhang L, Zhao H, Qiu Y, Loh HH, Law P-Y (2009) Src phosphorylation of μ-receptor is responsible for the receptor switching from an inhibitory to a stimulatory signal. J Biol Chem 284(4):1990–2000. https://doi.org/10.1074/jbc.M807971200
Schneider S, Provasi D, Filizola M (2016) How oliceridine (TRV-130) binds and stabilizes a μ-Opioid receptor conformational state that selectively triggers G protein signaling pathways. Biochemistry 55(46):6456–6466. https://doi.org/10.1021/acs.biochem.6b00948
Hub JS, de Groot BL (2009) Detection of functional modes in protein dynamics. PLoS Comput Biol 5(8):e1000480. https://doi.org/10.1371/journal.pcbi.1000480
Amadei A, Linssen ABM, Berendsen HJC (1993) Essential dynamics of proteins. Proteins Struct Funct Bioinf 17(4):412–425. https://doi.org/10.1002/prot.340170408
Hu X, Provasi D, Filizola M (2019) Mechanism of µ-opioid receptor-magnesium interaction and positive allosteric modulation. Biophys J 118(4):909–921
Da C, Kireev D (2014) Structural protein-ligand interaction fingerprints (SPLIF) for structure-based virtual screening: method and benchmark study. J Chem Inf Model 54(9):2555–2561. https://doi.org/10.1021/ci500319f
Madariaga-Mazón A, Marmolejo-Valencia AF, Li Y, Toll L, Houghten RA, Martinez-Mayorga K (2017) Mu-Opioid receptor biased ligands: a safer and painless discovery of analgesics? Drug Discovery Today 22(11):1719–1729. https://doi.org/10.1016/j.drudis.2017.07.002
Chen X-T, Pitis P, Liu G, Yuan C, Gotchev D, Cowan CL, Rominger DH, Koblish M, DeWire SM, Crombie AL, Violin JD, Yamashita DS (2013) Structure-Activity Relationships and Discovery of a G Protein Biased μ Opioid Receptor Ligand, [(3-Methoxythiophen-2-yl)methyl]({2-[(9R)-9-(pyridin-2-yl)-6-oxaspiro-[4.5]decan-9-yl]ethyl})amine (TRV130), for the treatment of acute severe pain. J Med Chem 56(20):8019–8031. https://doi.org/10.1021/jm4010829
Váradi A, Marrone GF, Palmer TC, Narayan A, Szabó MR, Le Rouzic V, Grinnell SG, Subrath JJ, Warner E, Kalra S, Hunkele A, Pagirsky J, Eans SO, Medina JM, Xu J, Pan Y-X, Borics A, Pasternak GW, McLaughlin JP, Majumdar S (2016) Mitragynine/corynantheidine pseudoindoxyls As opioid analgesics with Mu agonism and delta antagonism, which do not recruit β-Arrestin-2. J Med Chem 59(18):8381–8397. https://doi.org/10.1021/acs.jmedchem.6b00748
Li G, Aschenbach LC, Chen J, Cassidy MP, Stevens DL, Gabra BH, Selley DE, Dewey WL, Westkaemper RB, Zhang Y (2009) Design, synthesis, and biological evaluation of 6α- and 6β-N-heterocyclic substituted naltrexamine derivatives as μ Opioid receptor selective antagonists. J Med Chem 52(5):1416–1427. https://doi.org/10.1021/jm801272c
Bermudez M, Bock A (2019) Does divergent binding pocket closure drive ligand bias for class A GPCRs? Trends Pharmacol Sci 40(4):236–239
Molecular Operating Environment (MOE) (2021) 2019.01; Chemical Computing Group ULC, Montreal
Labute P, Williams C, Feher M, Sourial E, Schmidt JM (2001) Flexible alignment of small molecules. J Med Chem 44(10):1483–1490. https://doi.org/10.1021/jm0002634
Bartuzi D, Kaczor AA, Matosiuk D (2015) Activation and allosteric modulation of human μ Opioid receptor in molecular dynamics. J Chem Inf Model 55(11):2421–2434. https://doi.org/10.1021/acs.jcim.5b00280
Fenalti G, Zatsepin NA, Betti C, Giguere P, Han GW, Ishchenko A, Liu W, Guillemyn K, Zhang H, James D, Wang D, Weierstall U, Spence JCH, Boutet S, Messerschmidt M, Williams GJ, Gati C, Yefanov OM, White TA, Oberthuer D, Metz M, Yoon CH, Barty A, Chapman HN, Basu S, Coe J, Conrad CE, Fromme R, Fromme P, Tourwé D, Schiller PW, Roth BL, Ballet S, Katritch V, Stevens RC, Cherezov V (2015) Structural basis for bifunctional peptide recognition at human δ-Opioid receptor. Nat Struct Mol Biol 22(3):265–268. https://doi.org/10.1038/nsmb.2965
Shim J, Coop A, MacKerell AD (2013) Molecular details of the activation of the μ Opioid receptor. J Phys Chem B 117(26):7907–7917. https://doi.org/10.1021/jp404238n
Biasini M, Bienert S, Waterhouse A, Arnold K, Studer G, Schmidt T, Kiefer F, Cassarino TG, Bertoni M, Bordoli L, Schwede T (2014) SWISS-MODEL: modelling protein tertiary and quaternary structure using evolutionary information. Nucleic Acids Res 42(Web Server issue):W252–W258. https://doi.org/10.1093/nar/gku340
Arnold K, Bordoli L, Kopp J, Schwede T (2006) The SWISS-MODEL workspace: a web-based environment for protein structure homology modelling. Bioinformatics 22(2):195–201. https://doi.org/10.1093/bioinformatics/bti770
Benkert P, Biasini M, Schwede T (2011) Toward the estimation of the absolute quality of individual protein structure models. Bioinformatics 27(3):343–350. https://doi.org/10.1093/bioinformatics/btq662
Cheng J, Sun X, Li W, Liu G, Tu Y, Tang Y (2016) Molecular switches of the κ opioid receptor triggered by 6′-GNTI and 5′-GNTI. Sci Rep 6:18913. https://doi.org/10.1038/srep18913
Klauda JB, Venable RM, Freites JA, O’Connor JW, Tobias DJ, Mondragon-Ramirez C, Vorobyov I, MacKerell AD, Pastor RW (2010) Update of the CHARMM all-atom additive force field for lipids: validation on six lipid types. J Phys Chem B 114(23):7830–7843. https://doi.org/10.1021/jp101759q
MacKerell AD, Bashford D, Bellott M, Dunbrack RL, Evanseck JD, Field MJ, Fischer S, Gao J, Guo H, Ha S, Joseph-McCarthy D, Kuchnir L, Kuczera K, Lau FTK, Mattos C, Michnick S, Ngo T, Nguyen DT, Prodhom B, Reiher WE, Roux B, Schlenkrich M, Smith JC, Stote R, Straub J, Watanabe M, Wiórkiewicz-Kuczera J, Yin D, Karplus M (1998) All-atom empirical potential for molecular modeling and dynamics studies of proteins. J Phys Chem B 102(18):3586–3616. https://doi.org/10.1021/jp973084f
Jo S, Kim T, Iyer VG, Im W (2008) CHARMM-GUI: a web-based graphical user interface for CHARMM. J Comput Chem 29(11):1859–1865. https://doi.org/10.1002/jcc.20945
Wu EL, Cheng X, Jo S, Rui H, Song KC, Dávila-Contreras EM, Qi Y, Lee J, Monje-Galvan V, Venable RM, Klauda JB, Im W (2014) CHARMM-GUI Membrane Builder toward realistic biological membrane simulations. J Comput Chem 35(27):1997–2004. https://doi.org/10.1002/jcc.23702
Jo S, Lim JB, Klauda JB, Im W (2009) CHARMM-GUI membrane builder for mixed bilayers and its application to yeast membranes. Biophys J 97(1):50–58. https://doi.org/10.1016/j.bpj.2009.04.013
Jo S, Kim T, Im W (2007) Automated builder and database of protein/membrane complexes for molecular dynamics simulations. PLoS ONE 2(9):e880. https://doi.org/10.1371/journal.pone.0000880
Brooks BR, Brooks CL, MacKerell AD, Nilsson L, Petrella RJ, Roux B, Won Y, Archontis G, Bartels C, Boresch S, Caflisch A, Caves L, Cui Q, Dinner AR, Feig M, Fischer S, Gao J, Hodoscek M, Im W, Kuczera K, Lazaridis T, Ma J, Ovchinnikov V, Paci E, Pastor RW, Post CB, Pu JZ, Schaefer M, Tidor B, Venable RM, Woodcock HL, Wu X, Yang W, York DM, Karplus M (2009) CHARMM: the biomolecular simulation program. J Comput Chem 30(10):1545–1614. https://doi.org/10.1002/jcc.21287
Lee J, Cheng X, Swails JM, Yeom MS, Eastman PK, Lemkul JA, Wei S, Buckner J, Jeong JC, Qi Y, Jo S, Pande VS, Case DA, Brooks CL, MacKerell AD, Klauda JB, Im W (2016) CHARMM-GUI Input generator for NAMD, GROMACS, AMBER, OpenMM, and CHARMM/OpenMM simulations using the CHARMM36 additive force field. J Chem Theory Comput 12(1):405–413. https://doi.org/10.1021/acs.jctc.5b00935
Tieleman DP, Forrest LR, Sansom MSP, Berendsen HJC (1998) Lipid properties and the orientation of aromatic residues in OmpF, influenza M2, and alamethicin systems: molecular dynamics simulations. Biochemistry 37(50):17554–17561. https://doi.org/10.1021/bi981802y
Grossfield A, Pitman MC, Feller SE, Soubias O, Gawrisch K (2008) Internal hydration increases during activation of the G protein-coupled receptor rhodopsin. J Mol Biol 381(2):478–486. https://doi.org/10.1016/j.jmb.2008.05.036
Vanommeslaeghe K, Hatcher E, Acharya C, Kundu S, Zhong S, Shim J, Darian E, Guvench O, Lopes P, Vorobyov I, MacKerell AD (2010) CHARMM General Force Field (CGenFF): A force field for drug-like molecules compatible with the CHARMM all-atom additive biological force fields. J Comput Chem 31(4):671–690. https://doi.org/10.1002/jcc.21367
Vanommeslaeghe K, MacKerell AD (2012) Automation of the CHARMM general force field (CGenFF) I: bond perception and atom typing. J Chem Inf Model 52(12):3144–3154. https://doi.org/10.1021/ci300363c
Vanommeslaeghe K, Raman EP, MacKerell AD (2012) Automation of the CHARMM general force field (CGenFF) II: assignment of Bonded parameters and partial atomic charges. J Chem Inf Model 52(12):3155–3168. https://doi.org/10.1021/ci3003649
Abraham MJ, Murtola T, Schulz R, Páll S, Smith JC, Hess B, Lindahl E (2015) GROMACS: High performance molecular simulations through multi-level parallelism from laptops to supercomputers. SoftwareX 1–2:19–25
Seki T, Minami M, Nakagawa T, Ienaga Y, Morisada A, Satoh M (1998) DAMGO recognizes four residues in the third extracellular loop to discriminate between μ- and κ-opioid receptors. Eur J Pharmacol 350(2–3):301–310. https://doi.org/10.1016/s0014-2999(98)00240-4
Mansour A, Taylor LP, Fine JL, Thompson RC, Hoversten MT, Mosberg HI, Watson SJ, Akil H (1997) Key residues defining the μ-Opioid receptor binding pocket: a site-directed mutagenesis study. J Neurochem 68(1):344–353. https://doi.org/10.1046/j.1471-4159.1997.68010344.x
Shim J, Coop A, MacKerell AD (2011) Consensus 3D model of μ-opioid receptor ligand efficacy based on a quantitative conformationally sampled pharmacophore. J Phys Chem B 115(22):7487–7496. https://doi.org/10.1021/jp202542g
Paramo T, East A, Garzón D, Ulmschneider MB, Bond PJ (2014) Efficient characterization of protein cavities within molecular simulation trajectories: trj_cavity. J Chem Theory Comput 10(5):2151–2164. https://doi.org/10.1021/ct401098b
Gowers RJ, LM Barnoud, J, Reddy, TJ E, Melo MN, Seyler SL, Dotson DL, Domanski J, Buchoux S, Kenney IM, OB MDAnalysis: A Python package for the rapid analysis of molecular dynamics simulations. In: Rostrup SB, S (eds). In: Proceedings of the 15th Python in Science Conference, Austin, TX, 2016. Pp 102–109
Michaud-Agrawal N (2011) MDAnalysis: a Toolkit for the Analysis of Molecular Dynamics Simulations. J Comput Chem 32:2319–2327. https://doi.org/10.1002/jcc.21787
Hirst JD, Brooks CL (1994) Helicity, circular dichroism and molecular dynamics of proteins. J Mol Biol 243(2):173–178. https://doi.org/10.1006/jmbi.1994.1644
Bystroff C, Garde S (2003) Helix propensities of short peptides: Molecular dynamics versus bioinformatics. Proteins Struct Funct Bioinf 50(4):552–562. https://doi.org/10.1002/prot.10252
Acknowledgments
This work was supported by Instituto de Quimica and DGAPA-UNAM (PAPIIT IN210518). The computations were supported by the Dirección General de Cómputo y de Tecnologías de Información y Comunicación (DGTIC)-UNAM, providing Miztli computer resources under grant LANCAD-UNAM-DGTIC-367. AFMV acknowledges CONACyT (335795) for scholarship, and KM-M thanks DGAPA-PASPA for financial support. Authors thank the developers of ChemAxon, VMD/NAMD, GROMACS, and CHARMM for kindly providing open-source software and libraries.
Author information
Authors and Affiliations
Contributions
AFMV contributed to conceptualization, investigation, methodology, analysis, and writing—original draft), AMM performed analysis, investigation, visualization, and writing—review and editing, and KMM contributed to conceptualization, project administration, resources, funding acquisition, supervision, and writing—review and editing.
Corresponding author
Ethics declarations
Conflict of interest
There are no conflicts to declare.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Marmolejo-Valencia, A.F., Madariaga-Mazón, A. & Martinez-Mayorga, K. Bias-inducing allosteric binding site in mu-opioid receptor signaling. SN Appl. Sci. 3, 566 (2021). https://doi.org/10.1007/s42452-021-04505-8
Received:
Accepted:
Published:
DOI: https://doi.org/10.1007/s42452-021-04505-8