Abstract
The objective of the present study was to optimise the biosynthesis of gold nanoparticles using aqueous extract of Pongamia seed cake (PSC). Proteins, amino acids and phenolic compounds present in the PSC extract are responsible for reducing the metal precursor salt to nanoparticles. The levels of factors for optimisation of nanoparticles synthesis was evaluated using UV–Vis spectrophotometer by one factor at a time studies. Further factorial design was employed to study the significant effect of various parameters on the nanoparticles synthesis. The levels of two significant factors i.e. pH and concentration of PSC extract as obtained from factorial design were optimized further for maximum nanoparticle synthesis applying central composite design. The optimized conditions were found to be pH 3.6 and PSC extract concentration of 17.07% (w/v) at which an increase (OD at 530 nm = 1.3) in nanoparticle synthesis was observed as compared to un-optimized conditions (OD at 530 nm = 0.66). The model obtained on optimization was validated and it describes the nanoparticle synthesis aptly with an error between experimental and predicted values of just 1.15% with an R2 of 0.985. It can be concluded that, RSM approach was found to be an efficient technique to optimize the reaction conditions, where a twofold increase in synthesis of small sized monodispersed gold nanoparticles was seen.
Graphic abstract

Similar content being viewed by others
1 Introduction
In the present scenario, nanoparticles with improved properties have gained a lot of importance for majority of applications related to health and environment. Properties of nanoparticles are dependent on the size of the nanoparticles, morphology and distribution [1]. Physico-chemical methods used for the synthesis of nanoparticles have a lot of drawbacks and limitations. In the case of chemical methods, toxic by-products are formed, whereas in physical methods, maintenance of high temperature and pressure increases the cost of nanoparticle production [2]. Though, microorganisms can be used for the synthesis of nanoparticles, it requires laborious work of culture maintenance, cost of media, sterility etc., hence, researchers are concentrating on the use of biological resources that are easily available in abundance for the synthesis of nanoparticles [3]. Underutilised biological resources have been explored in recent years, which help to reduce the metal ion to its corresponding nanoparticles at room temperature due to phytochemicals present in them, thereby minimizing the energy requirements for the synthesis of nanoparticles. An investigation on use of fresh bark extract of Pongamia pinnata for synthesis of silver nanoparticles for its antibacterial activity against gram positive and gram negative pathogens was carried out [4]. The use of dried leaves of Pongamia pinnata (L) pierre for silver nanoparticle synthesis has also been reported [5]. Hence the present study is aimed at synthesis of metallic gold nanoparticles by biological means using Pongamia pinatta seed cake (PSC) extract, a byproduct from the biodiesel industry. PSC is rich in phytochemicals like proteins, carbohydrates, alkaloids, flavanoids and glycosides which can help in reduction and capping of nanoparticles thereby stabilizing them [6].
Green synthesis of nanoparticles is influenced by various factors such as pH of solution, temperature, and time of synthesis, concentration of the extracts used and the protocol of synthesis [7]. The individual effect of these factors on the biological synthesis of nanoparticles has been studied extensively [3, 8,9,10]. However, the determination of interactive effects of factors on the green synthesis of nanoparticles is a difficult task and has not been exploited much. The effect of these factors on the nanoparticles synthesis varies with respect to the chemical composition of the substrate and hence the process of nanoparticles synthesis by novel substrates has to be optimised. Response surface methodology (RSM) is a collection of statistical techniques that optimises a response of interest in a process which is influenced by several factors [11]. It also describes the individual and interactive effect of factors, on the process of interest.
In the present study, the levels of factors such as time of incubation, Gold(III)chloride trihydrate (HAuCl4·3H2O) concentration, PSC extract concentration, agitation speed, temperature, Triton X-100 concentration and pH for the optimisation of synthesis of PSC mediated gold nanoparticles (PSCAuNPs) were determined by one factor at a time (OFAT) approach. The effect of significant factors on PSCAuNPs synthesis was evaluated by Factorial design and the optimum levels of factors for maximum synthesis of PSCAuNPs were elucidated by applying central composite design (CCD) and the characteristics of synthesized nanoparticles were compared at un-optimised and optimised conditions. This is the first report on optimization and response surface methodology applied for using PSC extract as a biological reducing agent for gold nanoparticles synthesis.
2 Materials and methods
HAuCl4·3H2O was purchased from Himedia. PSC required for the study was procured from Biofuel Research, Information and Demonstration Centre, NMAMIT Nitte. Double distilled water was used for all the experiments.
2.1 Preparation of cell free extract
10% (w/v) PSC powder was used for the preparation of cell free extract and the suspension was prepared in deionized water. The suspensions were kept in the water bath maintained at 80 °C for half an hour and filtered through Whatmann No. 1 filter paper and the filtrate was used for nanoparticle production.
2.2 Synthesis of PSCAuNPs
Aqueous HAuCl4·0.3H2O (1 mM) was prepared in deionized water and was added to the cell free extract of PSC in the ratio 1:9 and mixed, such that the concentration of PSC extract was 10%w/v. The mixture was maintained at a pH of 4.8 by using 0.1 N HNO3 [12] and 1 ml of Triton X-100 (1 mM) was added [13], which acts as a protective agents and controls the size of synthesized nanoparticles [14]. The mixture was incubated at 25 °C at 100 rpm agitation and UV–Vis absorbance at 360–600 nm range was measured hourly interval for 7 h till constant readings were obtained. The nanoparticles synthesized in the reaction mixture were centrifuged and the product obtained was washed thrice with deionized distilled water and stored at − 4 °C for further analysis.
2.3 Characterization of PSC AuNPs
Biosynthesized nanoparticles were characterized by various techniques like UV- visible spectroscopy, particle size analyzer, Fourier transformed infrared spectroscopy (FT-IR), Field Emission Scanning Electron Microscope (FESEM), Energy dispersive analysis X (EDAX) and X-ray powder diffraction (XRD) at optimized and un-optimized conditions. The functional groups present in the phytoconstituents of biological material and their involvement as capping agents in the biosynthesized nanoparticles was determined by recording the IR on Bruker Alpha. Capping of nanoparticles with proteins, amino acids, and polyphenols was determined by FT-IR [15]. Zetasizer nano instrument (Malvern) was used for data acquisition and for estimating the particle size distribution as well as zeta potential distribution of a colloidal solution of nanoparticles. Size and morphology of AuNPs were examined by FESEM, CARL ZEISS by sputtering. Since the mettalic gold nanoparticles were coated with plant phytochemicals, samples were sputtered and the sputtered samples were analyzed for their microstructure at 5 kV. FESEM specimen preparation was done by taking small amount of powder of nanoparticles dispersed on carbon tape. Thin gold coating over the sample was developed by passing 20 mA current for 10 min. X-ray diffraction profile was obtained using XRD (RigakuMiniflex 600, Rigaku Co., Tokyo, Japan) using cu-kα X-ray diffraction patterns of the tested material was recorded over the 2θ range of variable degrees at a scan rate of 4°/min.
2.4 Modified 4-amino antipyrine method for estimation of phenol/polyphenol coating on nanoparticles
The presence of phenols and polyphenols present on colloidal nanoparticle solution was analysed by modified amino pyrine method [16]. 0.1 to 1 ml of nanoparticle suspension is taken, to which, 0.8 ml of 0.5 N sodium hydroxide and 1.2 ml of 0.5 M sodium bicarbonate was added. To the same mixture, 1 ml each of 4-amino-antipyrine reagent (1 mg/ml) and potassium ferricyanide (5 mg/ml) was added. Phenol standard of 0.1 mg/ml was used for calculating the phenolic content of colloidal nanoparticles. Reading of the pinkish coloured complex was taken at 520 nm in Shimadzu Spectrophotometer.
2.5 Optimisation of PSCAuNPs synthesis
2.5.1 Selection of levels of parameters by OFAT
OFAT is a method of designing experiments involving the testing of factors, or causes, one at a time instead of all simultaneously. OFAT was used to determine levels of factors to be optimized for PSCAuNPs production. The experiments were performed by varying one factor at a time by keeping other factors at a fixed value. Surface plasmon resonance (SPR) Peak was recorded at 530 nm as it is evident from literature reports that blue shift is seen due to decrease in particle size owing to monodispersed nature of nanoparticles [17]. Similarly, experiments were performed by varying other factors and keeping previously varied factors at the optimized values. The factors chosen for the study were time of incubation, HAuCl4·3H2O concentration, concentration of PSC extract, agitation speed, temperature, Triton X-100 concentration and pH (Fig. 2).
2.5.2 Selection of significant factors for PSCAuNP synthesis by factorial design
The effects of PSC extract concentration (X1), agitation speed (X2), pH (X3), and Triton X-100 concentration (X4) in 2 levels on PSCAuNPs synthesis (Table 1) were studied by a 24 factorial design encompassing 16 runs. Absorbance at 530 nm (Y1), as a measure of PSCAuNPs synthesis was recorded as the response (Table 1).
2.5.3 Optimisation of significant factors by CCD
The levels of PSC extract concentration (X1), and pH (X2) were optimised for PSCAuNPs synthesis by applying CCD. The design consisted of 12 runs i.e., 4 factorial runs, 4 star runs and 4 central runs (Table 1B). Absorbance at 530 nm (Y1), as a measure of PSCAuNPs synthesis was recorded as the response.
2.5.4 Statistical analysis
The screening and optimization experiments were designed by STATISTICA software [18]. The experiments were performed and the results obtained were analysed by ANOVA (Table S2) to display the optimized conditions by the same. Also, a second order regression equation was generated by the software that modelled the PSCAuNPs synthesis.
2.5.5 Validation of the second order model
Experiments were performed at random values of parameters within levels considered for optimization for PSCAuNPs synthesis. In addition to these, experiments were also performed at optimized conditions indicated by the software (Table 3). The experimental values of absorbance at 530 nm (Y1ex) were compared with the response predicted by the model (Y1pr), to evaluate the goodness of fit of the model.
3 Results and discussion
3.1 Synthesis and characterization of PSCAuNPs at un-optimised conditions
Colour change of the solution from light yellow to deep purple within 5 h of incubation indicated the formation of gold nanoparticles at 530 nm (Fig. 1b), which was confirmed by UV–visible spectrophotometer (Fig. 1a). Synthesis of gold nanoparticles by Padina tetrastromarica took 24 h [12], compared to this present study where faster synthesis of nanoparticles is observed. Nanoparticles were centrifuged at 10,000 RPM for 30 min and washed thrice with double distilled water followed by one wash with ethanol and dried in desiccator. The dessicated nanoparticles were sent for characterization. From the FESEM analyses, biologically synthesized gold nanoparticles were found to be 20 nm in size and spherical in shape, in monodispersed aggregated form (Fig. 1c). Edax results confirmed the purity of the samples and capping of nanoparticles with proteins (Fig. 1d).
Characterization of PSCAuNPs at un-optimised conditions: UV–visible spectrum of PSCAuNPs showing maximum absorbance at 560 nm (a); visual observation confirming PSCAuNPs synthesis (PSC Extract—left, PSCAuNPs—right) (b); SEM image of PSCAuNPs (c); EDAX spectrum of PSCAuNPs (d); Zetasizer (e) and Zeta potential distribution of PSCAuNPs (f); FT-IR spectrum for PSCAuNPs and PSC extract (1—PSC extract, 2—PSCAuNPs) (g); XRD pattern of PSCAuNPs (h)
Figure 1e, f shows particle size distribution in the range of 625 nm with intensity of 173.7% with Z-average value of 651.7 nm and high polydispersity index (PDI) of 0.537 and Zeta potential value of − 5.19 mV for PSCAuNPs. This indicates that, negatively charged groups are capped on the surface of PSCAuNPs which are also responsible for moderate stability of nanoparticles which can lead to agglomeration of nanoparticles.
FTIR results (Fig. 1g) confirmed the vanishing of peak at 3351 cm−1 in nanoparticles synthesized from PSC extract, whereas the peak was present in the original extract, which confirms the fact that the phenolic groups are involved in the synthesis of nanoparticles [19]. Nanoparticles also showed a peak at 2360 cm−1, which signifies the protein coating on nanoparticles [20]. The presence of proteins in both extract and nanoparticles was confirmed by a peak at 3761 cm−1, which corresponds to stretching frequency of OH-bond arising from proteins or absorbed water in sample [21]. Reduction in peak at 2925 cm−1 was seen in nanoparticles when compared to extract which signifies the involvement of proteins as reducing and capping agent. Earlier studies of IR spectra of biologically synthesized nanoparticles have shown the presence of biomolecules on their surface [13, 22, 23]. From the present study also it can be confirmed that the nanoparticles synthesized from PSC extract are capped with organic residues like proteins and amino acids and phenolic compounds, which are mainly involved in the reduction of metallic solutions. It is a well known fact that amines of the proteins facilitate the binding to nanoparticles and therefore, stabilization of the nanoparticles occurs through surface bound proteins [22].
Figure 1h shows the XRD pattern for PSCAuNPs, which exhibited broader peaks [24], where four characteristic diffraction peaks at 2θ values of 37.76°, 43.80°, 64.32°, and 77.10°, which correspond to (1, 1, 1), (2, 0, 0), (2, 2, 0), and (3, 1, 1). The peaks revealed that the resultant gold nanoparticles were in face cantered cube structure. Further, the average crystal sizes of PSCAuNPs were calculated by applying Scherer equation and were found to be 21.4 nm. The observed XRD pattern, thus, confirmed that the nanoparticles are formed by the reduction AuCl4− ions by phytochemicals of PSC extracts and the nanoparticles are crystalline in nature. Apart from normal peaks, some additional and unidentified peaks were also noticed at the vicinity of the characteristic peaks of gold nanoparticles. These peaks may be because of some bioorganic compounds or proteins of the extract [25, 26]. The presence of phenols and polyphenols was confirmed by modified 4-amino antipyrine method (Fig. 1S).
3.2 Optimisation of PSCAuNPs synthesis
3.2.1 Selection of levels of parameters by OFAT
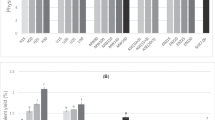
OFAT was employed to investigate the selection of levels of factors that showed considerable PSCAuNPs synthesis, so that the optimisation of factors could be performed. The conditions were varied one at a time in levels shown in Table 1, keeping others factors at a fixed level and the variation of response with respect to that of the factor were observed (Fig. 2).
From OFAT results it is clear that PSCAuNPs synthesis reached a plateau at around 5 h of incubation indicating that 5 h is sufficient for nanoparticles synthesis at conditions of 0.5 mM HAuCl4·3H2O concentration and 10%w/v PSC extract concentration. Single SPR peak was seen at 0.5 mM HAuCl4·3H2O concentration which indicates uniform sized nanoparticles, whereas at higher concentrations multiple SPR peaks were observed indicating polydispered sized nanoparticle synthesis [27, 28]. Similarly, single SPR peak was seen at 25 °C, whereas at higher temperatures multiple SPR peaks were visualised which could be due to agglomeration of particles at higher temperature [29]. Therefore, HAuCl4·3H2O concentration, temperature and time of incubation were fixed at 0.5 mM, 25 °C and 5 h respectively for further studies.
The OFAT studies showed significant effect of concentration of PSC extract (X1), agitation speed (X2), pH (X3), and Triton X-100 concentration (X4) on nanoparticle synthesis. Therefore, these factors were chosen for subsequent optimization studies. The levels of these factors for factorial design were determined on the basis of a peak obtained in the absorbance at 530 nm (Table 2). For instance, when Triton X-100 was varied between 0.5 and 2 mM, a peak in the absorbance was observed from 1.0 to 2.0 mM of Triton X-100, hence the minimum and maximum levels were set as 1.0 mM and 2 mM respectively.
3.2.2 Selection of significant factors for PSCAuNP synthesis by factorial design
The influence of the chosen independent factors on PSCAuNPs synthesis was studied by a 24 factorial design (Table 1), which also shows the absorbance values at 530 nm (Y1). The effect of the factors on the PSCAuNPs synthesis was determined by means of ANOVA (Table 1A). The factors i.e. concentration of PSC extract and pH were found to be highly significant factors that effect the PSCAuNP synthesis as they showed p value ≤ 0.05. The co-efficient of determination (R2 value) was observed to be 0.9913 which indicated that there is a strong linear relationship between the experimental values of response and values predicted by the design. The optimum levels of pH and PSC extract concentration were determined further by CCD and the values of agitation speed (X2) and Triton X-100 concentration (X4) were maintained at the centre of their levels.
3.3 Optimisation of significant factors by CCD
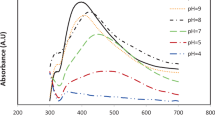
The influence of extract concentration (X1) and pH (X3) on PSCAuNPs synthesis was determined by applying CCD as indicated in Table 2, which also presents the observed values for absorbance at 530 nm for PSCAuNPs (Y1) at different combinations of the independent parameters. Absorbance values at 530 nm was found to vary from 0.233 to 1.028 in the experiments conducted which shows the strong influence of PSC extract concentration and pH on PSCAuNPs synthesis. From the ANOVA table (data not shown), it can be observed that, both the linear and quadratic effects of pH and PSC extract concentration as well the interaction between pH and concentration of PSC extract had highly significant effect on the PSCAuNPs syntheisis as indicated by the p value (≤ 0.05). The lack of fit of the model was found to be insignificant (p value > 0.05) indicating that, the model was significant. The influence of interaction of pH and concentration of PSC extract on the PSCAuNPs synthesis can be explained perfectly from the response surface plot (Fig. 3). At lower pH (3.5) there was a continous increase in synthesis of PSCAuNPs with increase in PSC extract concentration since lower pH has proven to be suitable for gold nanoparticle synthesis [14]. But at higher pH values (6.0) the PSCAuNPs synthesis decreased with increase in concentration of PSC extract since alkaline pH conditions are not favorable for synthesis [14]. At lower concentration of PSC extract (5%w/v) the PSCAuNPs synthesis was high at lower pH (3.5), decreased at intermediate pH (4.5–6.0) and continued to increase at high pH (> 6.0). However, at high concentration of PSC extract (15%w/v) the PSCAuNPs synthesis decreased with increase in pH since the extract concentration was too high which lead to faster capping therby leading to blue shift leading to agglomeration of nanoparticles resulting in precipitation [30]. At extract concentration above 15% and pH above 4, synthesis of nanoparticles were faster which lead to broader SPR leading to agglomeration of particles leading to decrease in absorbance values which is comparable with earlier reports [30].
The optimum levels of pH and concentration of PSC extract for maximum synthesis of PSCAuNPs were obtained from the desirability profiles (Fig. 4). The desirability values can vary between 0 and 1 and is a measure of closeness of response to its maximum value. When the response reaches the maximum value achieved at the given level of factors, then the desirability is 1.0 [26]. From the desirability profiles the optimised conditions were observed to be pH 3.6 at extract concentration of 17.07%w/v. This is comparable with earlier reports indicating homogenous nanoparticle synthesis under low pH conditions [31].
The second order regression equation that models the synthesis of PSCAuNPs as a function of the significant factors is given by
Co-efficient of determination for the second order equation as obtained from CCD was found to be 0.985 which indicated that there was an error of only 1.5% between the experimental and predicted absorbance values.
3.4 Validation of the second order model
The model was validated by conducting experiments at random values within the optimised level ranges of factors. The experimental absorbance values were found to be comparable with the values as predicted by the second order model obtained (Table 3). Also, experiment was performed at the optimised conditions (pH 3.58 and PSC extract concentration = 17.07% w/v). The experimental values of absorbance at 530 nm indicating PSCAuNPs synthesis (1.305) at these optimum conditions were quite close to the model predicted values (1.358). A twofold increase in the absorbance at 530 nm (1.305) was obtained by optimisation using RSM for PSCAuNPs synthesis (i.e. PSCAuNPs absorbance of 0.66 at 530 nm under unoptimized conditions).
3.5 Charecterization of PSCAuNPs synthesized post optimisation
PSCAuNPs were characterized by FESEM, EDAX, particle size analyser, FTIR and XRD after optimization studies to check if optimization helped in maximizing the synthesis of nanoparticles with better surface properties. Visual observation of optimized nanoparticle solution revealed a colour change from purple (Broader SPR leading to agglomeration) to reddish purple (Narrrow SPR) which is ascribes to blue shift. Under unoptimized condition aggregation of nanoparticles was seen due to red shift, which may be due to SPR coupling between neighboring particles [29]. Smaller size nanoparticles without agglomeration were synthesized upon optimization, which was confirmed by particle size data, where the zeta potential values improved from − 5.16 mV (un-optimized) to − 25.8 mV (optimized) which prooves the non –aggregated behavior of nanoparticles. Particle size results improved from 625 nm with 74% intensity (un-optimized) to 260.9 nm with 99% intensity (optimized) and polidispersity index (PDI) improved from 0.537 (un-optimized) to 0.279 (optimized) indicating monodisperse nature of nanoparticles under optimized conditions.
FTIR results of optimized PSCAuNPs were almost same as that of unoptimized PSCAuNPs showing the involvement of phenolic compounds as reducing agents and proteins as capping agents. Although, new peak was seen at 1674 cm−1 in optimized PSCAuNPs which could be due to bending vibrations of primary amines of proteins hence better protein coating of PSCAuNPs is acheived under optimized condition, thereby stabilizing them (Data not shown). Diffraction results for optimised PSCAuNPs showed sharper peaks compared to un-optimised conditions, owing to highly crystalline nature of nanoparticles upon optimization [29]. From EDAX spectrum for PSCAuNPs, it was seen that, the weight percent of gold increased to 76% (optimized) from 35% (un-optimized) indicating the increased purity of the nanoparticles post optimization. A slight increase in the size of PSCAuNPs was observed post optimization (25–37 nm) when compared to un-optimized condition (20 nm). This could be due to the reason that at acidic pH, proteins present in plant extract act as better reducing agents, thereby involving in more nucleation and the remaining proteins are involved in capping [9]. This was confirmed from the XRD results obtained post optimisation which shows sharp crystalline peaks indicating less protein capping on the nanoparticles (Fig. 5). Phenol and polyphenol estimation results confirmed the capping of nanoparticles with these molecules. Hence it is evident from the resullts that a refined coating was seen in optimized PSCAuNPs (0.09 mg/ml) when compared to unoptimized collidal solution (0.21 mg/ml) (Table 1C).
Characterization of PSCAuNPs at optimised conditions: UV–visible spectrum of PSCAuNPs showing maximum absorbance at 530 nm (a); visual observation confirming PSCAuNPs synthesis (PSC extract—left, PSCAuNPs—right) (b); SEM image of PSCAuNPs (c); EDAX spectrum of PSCAuNPs (d); Zetasizer (e); zeta potential distribution of PSCAuNPs (f); XRD pattern of PSCAuNPs (g)
4 Conclusions
PSC extract, an inexpensive and abundant byproduct from biodiesel industry, is rich in plant nutrients such as nitrogen and sulphur. This was found to be potential biological reducing agent for synthesis of gold nanoparticles. The gold nanoparticles synthesized were capped with proteins and polyphenols present in PSC extract. The present study showed the effect of factors on the synthesis of gold nanoparticles by PSC extract and subsequent optimisation of significant factors by RSM. Concentration of PSC extract and pH confirmed to be the most effective parameter for increased synthesis of nanoparticles which was evident from RSM studies. RSM was found to be an efficient methodology for rapid optimization of influencing significant factors and modelling of the PSCAuNPs synthesis, which led to a two fold increase in the nanoparticle synthesis. Significance of this work is that it includes the use of biodiesel byproduct as biological reducing agent for gold nanoparticle synthesis, which in turn reduces the cost involved in nanoparticle synthesis as well as reduces burden on environment by effective byproduct management. Moreover, the optimized conditions obtained from this study can be used for large-scale cost-effective production of gold nanoparticles.
Abbreviations
- RSM:
-
Response surface methodology
- PSC:
-
Pongamia seed cake
- PSCAuNPs:
-
PSC mediated gold nanoparticles
- OFAT:
-
One factor at a time
- CCD:
-
Central composite design
- FTIR:
-
Fourier transform infrared spectroscopy
- FESEM:
-
Field emission scanning electron microscope
- EDAX:
-
Energy dispersive X-ray
- XRD:
-
X-ray powder diffraction
- ANOVA:
-
Analysis of variance
References
Manimegalai G, Kumar SS, Sharma C (2011) Pesticide mineralization in water using silver nanoparticles. Int J Chem Sci 9(3):1463–1471
Shankar SS, Rai A, Ahmad A, Sastry M (2014) Rapid synthesis of Au, Ag, and bimetallic Au core–Ag shell nanoparticles using Neem (Azadirachta indica) leaf broth. J Colloid Interface Sci 275(2):496–502
Soni N, Prakash S (2011) Factors affecting the geometry of silver nanoparticles synthesis in Chrysosporium tropicum and Fusarium oxysporum. Am J Nanotechnol 2(1):112–121
Rajeshkumar S (2016) Synthesis of silver nanoparticles using fresh bark of Pongamia pinnata and characterization of its antibacterial activity against gram positive and gram negative pathogens. Resour Effic Technol 2(1):30–35
Raut RW, Kolekar NS, Lakkakula JR, Mendhulkar VD, Kashid SB (2010) Extracellular synthesis of silver nanoparticles using dried leaves of Pongamia pinnata (L) pierre. Nano-Micro Lett 2(2):106–113
Yadav RD, Jain SK, Alok S, Prajapati SK, Verma A (2011) Pongamia pinnata: an overview. Int J Pharma Sci Res 2(3):494–499
Patra JK, Baek K (2014) Green nanobiotechnology: factors affecting synthesis and characterization techniques. J Nanomater. https://doi.org/10.1155/2014/417305
Kredy HM (2018) The effect of pH, temperature on the green synthesis and biochemical activities of silver nanoparticles from Lawsonia inermis extract. J Pharm Sci Res 10(8):2022–2026
Aziz WJ, Jassi HA (2018) A novel study of pH influence on Ag nanoparticles size with antibacterial and antifungal activity using green synthesis. World Sci News 97:139–152
Elemike EE, Onwudiwe DC, Arijeh O, Nwankwo HU (2017) Plant-mediated biosynthesis of silver nanoparticles by leaf extracts of Lasienthra africanum and a study of the influence of kinetic parameters. Bull Mater Sci 40(1):129–137
Bas D, Boyaci IH (2007) Modeling and optimization II: comparison of estimation capabilities of response surface methodology with artificial neural networks in a biochemical reaction. J Food Eng 78(3):846–854
Rekavandi N, Malekzadeh A, Ghiasi E (2019) Methyl orange degradation over nano-LaMnO3 as a green catalyst under the mild conditions. Nanochem Res 4(1):1–10
Ibrahim EH, Kilany M, Ghramh HA, Khan KA, ul Islam S (2019) Cellular proliferation/cytotoxicity and antimicrobial potentials of green synthesized silver nanoparticles (AgNPs) using Juniperus procera. Saudi J Biol Sci 26(7):1689–1694
Perumal R, Casale S, De Stefano L, Spadavecchia J (2017) Synthesis and characterization of Ag-protoporphyrin nano structures using mixed co-polymer method. Front Lab Med 1(2):49–54
Edison TJI, Sethuraman MG (2012) Instant green synthesis of silver nanoparticles using Terminalia chebula fruit extract and evaluation of their catalytic activity on reduction of methylene blue. Process Biochem 47(9):1351–1357
Ettinger M, Ruchhoft C, Lishka R (1951) Sensitive 4-aminoantipyrine method for phenolic compounds. Anal Chem 23(12):1783–1788
Princy KF, Gopinath A (2018) Optimization of physicochemical parameters in the biofabrication of gold nanoparticles using marine macroalgae Padina tetrastromatica and its catalytic efficacy in the degradation of organic dyes. J Nanostruct Chem 8(3):333–342
Statsoft (1999) Statistics for Windows, Statsoft, Tulsa, Okla, USA
Vasudev DK, Pramod SK (2013) Green synthesis of copper nanoparticles using Ocimum sanctum leaf extract. Int J Chem Stud 1(3):1–4
Qutub N, Sabir S (2012) Optical, thermal and structural properties of Cds quantum dots synthesized by a route. Int J Nanosci Nanotechnol 8(2):111–120
Labhane PK, Huse VR, Patle LB, Chaudhari AL, Sonawane GH (2015) Synthesis of Cu doped ZnO nanoparticles: crystallographic, optical, FT-IR, morphological and photo catalytic study. J Mater Sci Chem Eng 3:39–51
Arulkumar S, Sabesan M (2010) Rapid preparation process of antiparkinsonian drug Mucuna pruriens silver nanoparticle by bioreduction and their characterization. Pharmacogn Res 2(4):233–236
Mubarak D, Thajuddin N, Jeganathan K, Gunasekaran M (2011) Plant extract mediated synthesis of silver and gold nanoparticles and its antibacterial activity against clinically isolated pathogens. Colloids Surf B 85(2):360–365
Chen D, Chen C (2002) Formation and characterization of Au–Ag bimetallic nanoparticles in water-in-oil microemulsions. J Mater Chem 12(5):1557–1562
Kalishwaralal K, Deepak V, Pandian SRK, Kottaisamy M, BarathManiKanth S, Kartikeyan B, Gurunathan S (2010) Biosynthesis of silver and gold nanoparticles using Brevibacterium casei. Colloids Surf B Biointerfaces 77(2):257–262
Roopan SM, Madhumitha G, Rahuman AA, Kamaraj C, Bharathi A, Surendra TV (2013) Low-cost and eco-friendly phyto-synthesis of silver nanoparticles using Cocos nucifera coir extract and its larvicidal activity. Ind Crops Prod 43:631–635
Lisha KP, Anshup PT (2009) Enhanced visual detection of pesticides using gold nanoparticles. J Environ Sci Health B 44(7):697–705
Dhumale VA, Gangwar RK, Datar SS, Sharma RB (2012) Reversible aggregation control of polyvinylpyrrolidone capped gold nanoparticles as a function of pH. Mater Exp 2(4):311–318
Suvith VS, Philip D (2014) Catalytic degradation of methylene blue using biosynthesized gold and silver nanoparticles. Spectrochim Acta A Mol Biomol Spectrosc 118:526–532
Rajan MM, Philip D (2014) Shape tailored green synthesis and catalytic properties of gold nanocrystals. Spectrochim Acta A Mol Biomol Spectrosc 118:793–799
Armendariz V, Herrera I, Jose-yacaman M, Troiani H, Santiago P, Gardea-Torresdey JL (2004) Size controlled gold nanoparticle formation by Avena sativa biomass: use of plants in nanobiotechnology. J Nanopart Res 6(4):377–382
Acknowledgements
The authors acknowledge the financial support received from Vision Group of Science and Technology (VGST) under RGS/F scheme bearing Grant Number KSTePS/VGST-RGS/F/GRD No. 705/2017-18 for carrying out this research work. Authors would like to thank Manipal College of Pharmaceutical sciences, Manipal and DST-PURSE laboratory, Mangalore University for technical support in characterizing the nanoparticles.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The authors report no conflict of interest.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Nayak, S., Sajankila, S.P., Goveas, L.C. et al. Two fold increase in synthesis of gold nanoparticles assisted by proteins and phenolic compounds in Pongamia seed cake extract: response surface methodology approach. SN Appl. Sci. 2, 634 (2020). https://doi.org/10.1007/s42452-020-2348-5
Received:
Accepted:
Published:
DOI: https://doi.org/10.1007/s42452-020-2348-5