Abstract
The human gut microbiota is widely considered to be a metabolic organ hidden within our bodies, playing a crucial role in the host’s physiology. Several factors affect its composition, so a wide variety of microbes residing in the gut are present in the world population. Individual excessive imbalances in microbial composition are often associated with human disorders and pathologies, and new investigative strategies to gain insight into these pathologies and define pharmaceutical therapies for their treatment are needed. In vitro models of the human gut microbiota are commonly used to study microbial fermentation patterns, community composition, and host-microbe interactions. Bioreactors and microfluidic devices have been designed to culture microorganisms from the human gut microbiota in a dynamic environment in the presence or absence of eukaryotic cells to interact with. In this review, we will describe the overall elements required to create a functioning, reproducible, and accurate in vitro culture of the human gut microbiota. In addition, we will analyze some of the devices currently used to study fermentation processes and relationships between the human gut microbiota and host eukaryotic cells.
Graphic abstract

Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
The human gut microbiota is a trending research topic. Besides supporting the digestive process, protecting the host from endogenous and exogenous infections, and shaping the anatomy and functionality of the entire gastrointestinal tract [1,2,3,4,5], gut microorganisms are also implicated in physiological processes involving other distal organs (e.g., the microbiota-gut-brain axis) [6, 7].
In vivo experimental models (e.g., mice, rats, pigs) have been widely used to study the effects of the gut microbiota on various organs [8,9,10]. Although in vivo models are quite common, they are recognized to have low reproducibility, ethical and cost restrictions, and to represent other physiologies than human ones. In addition, due to intrinsic difficulties in taking samples frequently from animals, in vivo models do not typically allow continuous monitoring of intestinal conditions and interactions between gut microorganisms and the host. Contextually, the focus on in vitro models that mimic the gut environment has grown in recent years thanks to the advent of innovative culture techniques, materials, technologies, and screening systems that have allowed researchers to recreate, study, and understand the effects of the human gut microbiota on the host (Fig. 1). For example, artificial co-cultures made up of the human gut microbiota, intestinal components (i.e., mucus), and mono- or multi-layered tissues have become an incredibly powerful and novel tool to study this intricate tangle of cellular and ecological interactions. This is also thanks to the development of more physiologically relevant tissue models, such as organoids [11].
In this review, we aim to analyze and discuss some of the “ingredients” needed to make a reliable in vitro model of the human gut microbiota (Fig. 2) and how they have already been used to study each facet of the microbiota universe. The reliability of an in vitro model is principally related to its reproducibility over different experiments and its accuracy in recreating a physiologically relevant model (e.g., microbial composition and metabolic functions) similar to the in vivo state. The latter can be difficult to achieve for in vitro models because not all of the systems described in the next sections can promote the stability and functions of the human gut microbiota. These are clearly critical points to be evaluated when an in vitro model of the human microbiota has been designed and set up.
In particular, this review will first focus on the several choices that must be made prior to culturing the gut microbiota (e.g., selection of the appropriate sample and culture medium, choosing the environmental parameters to consider). Then, we will describe the technological advances in gut microbiota in vitro culture, especially highlighting the features of the main systems presented in the literature before now.
Fabricating an in vitro model of the human gut microbiota: setting the operation parameters
Recreating human gut microbiota composition: not only a problem of numbers
The most ambitious goal of recreating the human gut microbiota in vitro is to replicate the complex network of bacterial metabolisms ex vivo [11]. This is not necessarily associated with the precise composition of the microbial community. However, careful selection of the microbial sample to be inoculated in the in vitro model has to be considered as the first step in designing the perfect model (Fig. 2). Although the majority of studies in the literature use human stool or samples obtained through endoscopic procedures, aspiration of intestinal fluids, ileostomies, or endoscopic capsules [12,13,14] as sources of colonic and small intestine microbiota, others prefer using a few target bacteria able to synthetize metabolites and chemical compounds which reproduce the principal metabolic pathways found in the intestine [15, 16]. Recreating a complex community such as the human gut microbiota can require two approaches. The “top-down” approach is used when, starting from a large microbial community and varying environmental parameters, bacteria are selected to reproduce a certain metabolic process. Conversely, the “bottom-up” approach is used when the information derived from multi-omics technologies is employed to recreate a certain metabolic process from the microbial selection [17]. For example, Petrof et al. recreated a narrow microbial population, metabolically comparable to the gut microbiota, using 33 species of bacteria, and demonstrated that a fecal transplant with this suspension could resolve Clostridioides difficile infections [15]. From an in vitro perspective, Krause et al. described a simplified gut microbiota model (SIHUMIx) that included the eight most abundant bacterial species of the human gut microbiota [16, 18]. The aim of these approaches is not only to reduce the interconnection variables between the various species but also to perform more replicable experiments by maintaining a bacterial core able to guarantee certain metabolic pathways of the human gut microbiota. Nevertheless, the dramatic reduction of microbial richness, biodiversity, and interactions can be problematic when settings like these are designed.
Finding the best culture medium
One of the crucial points in the in vitro culture of the human gut microbiota is to identify a suitable culture medium that guarantees the survival and replication of most of the microorganisms which constitute the microbial community. Although gut microbiota medium (GMM) is known to encourage reproduction of a diverse microbial community [19], a “universal” medium to culture all the microorganisms of the human gut microbiota still does not exist. The composition of a culture medium for microbiological studies is typically water, a carbon source, a nitrogen source, and some mineral salts [20]. In addition, some demanding microorganisms need other elements to grow, such as amino acids, vitamins, purines, and pyrimidines [20]. Different media have been tested to study the final composition of in vitro-cultured microbiota [21]. Kim et al. used three different culture media, i.e., brain heart infusion broth (BHIB), high concentration carbohydrate medium (HCM), and low concentration carbohydrate medium (LCM), to culture the human gut microbiota extracted from fecal samples, demonstrating that LCM inoculated with a fecal suspension at a final concentration of 3% granted the highest microbial abundances of the principal phyla within the human gut microbiota (e.g., Firmicutes, Bacteroidetes, Actinobacteria, and Proteobacteria) after a short period culture time (i.e., 18 h of incubation) [22]. Li et al. tested the effects of inorganic salts, bile salts, short-chain fatty acids (SCFAs), and mucins on the functionality of the human gut microbiota by metaproteomics [23]. Yousi et al. tested four bacterial culture media (BHIB, GMM, fastidious anaerobe broth (FAB), and BGM) and demonstrated the differences in terms of microbial composition and SCFA production of the cultured microbiota [24].
Selecting the appropriate environmental conditions
The environmental parameters constantly change throughout the entire length of the gastrointestinal tract, with different regions of the intestine harboring microbial populations that are directly shaped by these different environmental conditions. For instance, pH, partial oxygen pressure, quality of nutrients, gut peristalsis, and concentration of hormones and bile salts are only a few of the shaping factors that characterize the ecological niche in which each microorganism is allowed to live and multiply [25,26,27]. Even in the diametral section of the intestine, environmental factors such as oxygen profoundly differ [28]. Specifically, the small intestine is mainly populated by Lactobacillaceae and Enterobacteriaceae, with a low overall bacterial load due to low pH levels and compounds secreted by intestinal, hepatic, and pancreatic cells, such as bile salts and antimicrobial peptides [27]. In the large intestine, on the other hand, one finds the highest number of microbial species, with a prevalence of Bacteroidaceae, Prevotellaceae, Rikenellaceae, Lachnospiraceae, and Ruminococcaceae [27]. Especially in this part of the intestine, the pH is widely variable from person to person or even in the same individual, depending on diet-driven fermentation processes, thus leading to changes in the bacterial clusters that inhabit this area [26, 29, 30].
From an in vitro culture perspective, several works have shown how changing one or more of these environmental parameters leads to growth inhibition or stimulation of certain bacterial taxa. Duncan et al. noted a strong decrease in Bacteroides species (spp.) when moving from a controlled condition of pH 6.5 to 5.5 [26]. In addition, a pH shift from 6 to 4 resulted in a less biodiverse microbial population with lower abundances of acetate and propionate producers but more lactate-secreting bacteria [26]. Haindl et al. showed an increase in acetate and propionate levels and in the abundance of Bacteroidetes and Verrucomicrobia, as well as a decrease in the concentration of butyrate and in the abundance of Actinobacteria and Firmicutes in parallel with the rising pH of the culture [29].
Transit time (i.e., transit of luminal content along the gastrointestinal tract) is one of the most common parameters used to describe gut motility [31]. Since gut motility is related to nutrient availability in the different intestinal regions, it is reasonable to deduce that the composition of the human gut microbiota is linked to transit time [32, 33]. Using a blue-dye method, Asnicar et al. showed that some bacterial species (i.e., Akkermansia muciniphila, Bacteroides spp., and Alistipes spp.) are linked to longer transit time [34]. Furthermore, Tottey et al. followed an in vitro approach using the 3-Stage Environmental Control System for Intestinal Microbiota (3S-ECSIM), which is a multi-compartmental reactor that simulates the physiochemical parameters of the proximal, transverse, and distal portions of the colon, to describe how human gut microbiota composition and metabolism change with transit time [35]. In particular, lower biomass was found in the distal colon compartment, where there is an increased transit time and increased fermentation activity of microorganisms.
A surface issue
Similar to eukaryotic cells in vitro, cultured microorganisms grow better on surfaces than they multiply suspended in culture medium [36]. Microbial adhesion to a surface is a complex biophysical process which can be divided into two main phases. In the first phase, a reversible adhesion occurs in which bacteria adhere to the surface due to thermodynamic phenomena (all these are well reviewed in [37]). The Derjaguin–Landau–Verwey–Overbeek (DLVO) theory of colloidal stability is typically applied to describe the different chemical/physical phenomena occurring in this phase of the adhesion process [38]. In particular, the extended DLVO theory describes the Gibbs free energy, which is the sum of the Lifshitz–van der Waals interactions, electrostatic double-layer interactions, and acid–base bonds. This model is commonly applied to microbial adhesion process to a surface because bacterial mean size fits the colloidal particle dimensions (0.5–2 µm) [39]. Following this formulation, a resulting negative Gibbs free energy promotes bacterial adhesion, while positive free energy may inhibit it. Some results have shown that this general model is accurate because bacteria, which are characterized by a hydrophobic external surface, are attracted to hydrophobic materials, and vice versa [40]. Furthermore, the presence of external appendages on the bacterial surface of certain microorganisms (e.g., flagella, pili, fimbriae) creates a connection with the surface of the material that promotes this reversible adhesion by acting as a spring [41,42,43,44].
The transition from reversible to irreversible adhesion, the second phase of the adhesion process, derives from a series of physical and chemical mechanisms involving, for example, production of extracellular polymeric substance (EPS) and consequent formation of the so-called “biofilm.” A biofilm is a structured community of microorganisms encapsulated in their EPS and attached to a surface [45]. Bacterial biofilms are well known in the field of medicine for being the leading cause of artificial-implant failure, oral-cavity disease, and bacterial resiliency after drug and disinfectant treatment [46].
Changing the properties of a surface (e.g., surface charge, wettability, roughness, topography, or stiffness) can lead to conflicting results in bacterial adhesion [47]. One example is the surface charge density of the material used as the physical support for the culture. Typically, bacterial surfaces are characterized by the presence of a net negative charge due to carboxyl, amine, and phosphate groups, which influence the initial adhesion, making them attracted to positively charged materials [42, 48, 49]. However, in some cases, some bacteria also adhere to negatively charged surfaces thanks to their appendages and surface polymers typically found in Gram-negative bacteria (e.g., lipopolysaccharides [50]). In extreme cases, both negatively and positively charged materials reduced the adhesion of Pseudomonas aeruginosa on polystyrene plates [51]. The same contradictory results can be observed for the wettability, topography, and stiffness of a certain surface (Table 1). These opposite behaviors clearly demonstrate the complexity in selecting a universally valid surface for bacterial culture. In fact, ideal physical supports for culturing the human gut microbiota are difficult to achieve due to the complex microbial composition and the remarkable differences in adhesive surface properties of various bacterial species.
Adherence of bacteria to a physical substrate is not only dependent on its mechanical and physical properties but also on its chemical characteristics. Different molecules have been studied for their effects on bacterial adhesion to material surfaces [36]. Fibronectin, for example, promotes adhesion of Staphylococcus aureus [68, 69], but inhibits that of Staphylococcus epidermidis [70]. Mucins are molecules that can be added in an in vitro model to reproduce more faithfully the gut environment and its resident microbiota. They are glycoproteins produced by the epithelial tissue of the gastrointestinal tract, and several works have already suggested their importance in the attachment process of certain microbial species and in the resulting composition of the human gut microbiota [71, 72]. In fact, the mucus layer in the gut is the home of different microbes such as Akkermansia muciniphila and Lactobacillus spp., which are able to colonize the mucosal layer under healthy conditions [72]. Lactobacillus reuteri, for example, binds the mucins thanks to the presence of mucus binding domain on its outer membrane [73]. Culturing a complex population such as the human gut microbiota may lead to different results due to the presence of microorganisms not able to adhere to mucins. For example, in our previous work, we demonstrated that biofilm production of a fecal microbiota on electrospun gelatine structures was lower in the presence of mucins [64]. Also, using the M-SHIME bioreactor, Van den Abbeele et al. demonstrated diametrically opposite differences in the composition of the human gut microbiota in the luminal and the mucosal environment, with a prevalence of Firmicutes in the mucus layer [74]. For this reason, introducing the mucus layer to an in vitro model of the human gut microbiota could be useful if coupled with non-mucins layer to enhance the adhesion and replication of both mucus- and non-mucus-adhesive microorganisms.
Shear stress studies
The effects of moving fluid on microorganisms attached to a physical substrate can be of critical importance for creation of an in vitro model of the gut microbiota. Studying the mechanical behavior of bacteria departs from the microbiological tradition, in which bacterial adhesion and proliferation is guided mainly by the chemical environment [75]. It is known, for example, that circulation of a culture medium tends to boost microbial proliferation by carrying anabolites and washing away waste products. However, the flow could also inhibit microbial proliferation, affecting the density of the formed biofilm and reducing its size, depending on the intensity of flow [76, 77]. Also in this case, there are remarkable differences in the behavior of different microbes, with adhesion forces dependent on both environment and surface properties [78]. For example, Lecuyer et al. showed with a microfluidic device that the rate of attachment of P. aeruginosa increases up to a shear stress of 3.5 Pa on different surfaces (i.e., hydrophilic glass and hydrophobic polydimethylsiloxane (PDMS)) [79]. Detachment of P. aeruginosa occurs when the shear stress suddenly decreases, thus showing that bacteria respond dynamically to shear velocity by modifying the adhesive state accordingly. The relation between the nature of surfaces and shear stress is also described by Moreira et al. [80]. In their study, Escherichia coli was exposed to different fluid flows, showing that up to a threshold level of 8–10 mL/s, the species bound better to the hydrophobic physical substrate (wall shear stress was evaluated with computational fluid dynamics and ranges from 0.05 to 0.07 Pa). Then, beyond this threshold, a diminished number of attached bacteria were observed, thus demonstrating a correlation between shear stress and adhesion of E. coli.
Other examples are reported in Table 2. Some of these highlight a correlation between increased shear stress and a higher rate of bacterial attachment on the surface, which is a diametrically opposite behavior than that of most eukaryotic cells cultured in vitro [81].
Biofilm topology is also affected by flow. Rusconi et al. found formation of long filamentous structures in E.coli culture following the direction of fluid flow [82]. The formation of these streamers is also enhanced by the presence of surface irregularities. Uncontrolled development of these streamers could be problematic in an in vitro culture, clogging the channel and consequently stopping the flow [83, 84]. For these reasons, the absence of sharp edges, which enhances these effects, could be added as a design specification for an in vitro culture device.
Culture medium for co-culture with human cells
Optimization of the culture medium is crucial for success of the co-culture between the gut microbiota and human eukaryotic cells, as it is responsible for the nourishment and viability of the different cell types involved. Still, it is laborious and time-consuming due to the huge number of possible combinations, as highlighted in the review by Vis et al. [88].
Co-culture strategy and medium composition are the first two variables to consider when designing an in vitro co-culture (Table 3). Then medium volume, waste product accumulation, and reuse of the medium itself must be taken into account.
Medium volume affects primarily the concentration and dilution of waste products and produced metabolites [98,99,100]. A higher medium volume leads to lower concentrations of cell secretomes (i.e., soluble factors and extracellular vesicles), which is particularly important for cell–cell communication, cell proliferation, and differentiation [98]; however, a lower medium volume is more cost-effective and favorable for culturing some cell types such as neuron-like cells and adipose tissue-derived mesenchymal stem cells [92].
Concerning the problem of waste accumulation, the medium must be replaced periodically to maintain proper concentrations of nutrients and growth factors and allow the removal of waste products generated by cellular metabolism. Typically, this goal is achieved with continuous reactor systems. However, changing the medium implies the removal of secretomes, whose production determines further stress for cells and has a negative impact on their viability [101]. To overcome this issue, dialysis systems have drawn the attention of experts in in vitro cultures [102,103,104,105]. Their integration in in vitro culture systems enables the selective removal of waste products and reintroduction of nutrients and vitamins into the medium, while ensuring retention of cell secretomes. This allows reuse of the culture medium and helps in creating a more physiological environment for cells [88]. In addition, dialysis membranes show promise as they are already used in indirect co-culture systems to perform physical separation of different cellular types and/or microorganisms [106,107,108].
For co-culture of microorganisms and mammalian cells, an additional issue must be considered. The majority of the microbes constituting the human gut microbiota are obligate anaerobic bacteria, which constantly crosstalk with the colonic epithelium in a mucosal anoxic–oxic interface [109]. This constraint establishes the need for two different culture media, one anoxic and the other oxygenated, to guarantee cellular survival and suitable environmental conditions. Therefore, indirect co-culture and partitioned culture environments are the most frequently implemented solutions for in vitro co-culture of the human gut microbiota in the presence of host cells, as described in the next section.
Studying an in vitro model of the human gut microbiota: what are we looking for?
Now that various preliminary steps are understood, we will move on to discuss how the human gut microbiota could be studied in an in vitro model, taking into consideration its complex physiology and richness. Fermentation studies (whose purpose is mainly to replicate as closely as possible the intestinal conditions to investigate the response of microbial fermentative pathways to the presence of specific dietary compounds, toxic molecules, pathogens, etc.) can be distinguished from interaction studies, which involve co-culture with human cells.
Fermentation models
The fermentation processes carried out by the human gut microbiota play a key role in physiological digestion of food [110]. Metabolites produced by these pathways are mostly absorbed by the intestinal mucosa and, while some have health benefits, others have harmful effects on the host [111]. The fermentation patterns associated with the intake of specific nutritional components have been widely studied through different systems described in the literature.
An initial distinction must be made between static and dynamic fermentation systems. Static systems are typically batch fermentation models built with a closed and controlled environment (i.e., flask, beaker, closed vessel) that simulates one stage at a time (e.g., mouth, stomach, small intestine, colon) [112, 113]. Different methodologies and protocols have been published in an attempt to standardize the culture conditions, such as environmental parameters and digestive fluid composition, digestion time, and operation steps, as well as the post-process (e.g., determination of enzyme activity, collection of samples during the digestive process) [114,115,116]. These models are simple and have good reproducibility but lack the absorption process by the mucosal component, and transit time between the different compartments of the gastrointestinal tract is not considered. Also, in these static systems, the culture conditions are difficult to standardize due to cell activity and resource concentration. Conversely, dynamic systems are characterized by single or multiple reactors that, thanks to the setting of compartment-specific environmental parameters such as pH, oxygen, temperature, and transit time, can more accurately recreate the intestinal environment. Most of the fermentation models, especially in dynamic conditions, are well reviewed in [117, 118]. In particular, we must consider that the stability of the microbial profile is not always guaranteed in these systems, especially in the mono-compartment models, as described by Liu and colleagues [119]. Here we aim to highlight some of the technological enhancements of these fermenters over the last 40 years.
Dynamic fermentation systems can be classified into mono- and multi-compartment models, and which of these are selected depends only on the experimental specifications. As explained by Firrman et al. [120], both models are able to develop microbial communities with different species compositions from the same initial sample. In mono-compartment fermentation models, only a single region of the gastrointestinal tract is reproduced. For example, the artificial colon (ARCOL) reproduces the colon environment of humans and animals in vitro (Table 4) [121,122,123,124]. The ARCOL bioreactor is equipped with various probes and ports and can be inoculated with fresh stool from healthy animals or human volunteers. The temperature and pH are kept constant by adding NaOH. This model is the first to allow a continuous anaerobic condition inside a fermenter solely through the metabolic activity of bacteria. The Reading Model developed in 1988 by Gibson and co-workers [125] is one of the first examples of a multistage bioreactor. The three vessels used to mimic the proximal, transverse, and distal colon are aligned in series. The pH, the chemical components inside the three vessels, and the fermentation substrate are predefined to simulate food fermentation in the gut. While the first vessel has a mildly acidic environment and high substrate concentration to induce microbial growth as in the proximal colon, the others have neutral pH and a few substrates to resemble the conditions in the transverse and distal colon. Also, the microaerophilic environment is maintained with the insertion of N2 and O2 and controlled by a dissolved oxygen sensor. Thanks to its simplicity and easy customization, this model is still used to examine the effects of prebiotic and dietary components on the human gut microbiota [126, 127]. Another example of customizable technology which has evolved during the last 30 years is the in vitro dynamic model of the gastrointestinal tract (TIM) described for the first time in 1995 [128]. The first configuration (TIM-1) comprises four compartments (i.e., stomach, duodenum, jejunum, and ileum) connected to each other by peristaltic pumps that allow chyme transport between the vessels. The TIM system has been heavily customized and improved over the years. For example, the tiny-TIM system is a smaller version of the TIM that comprises two compartments resembling the stomach and small intestine only [129], while the TIMagc simulates the specific conditions in the corpus and antrum part of the stomach [130]. In addition, the TIM-2 system operates with high-density gut microbiota samples to mimic the dynamic and metabolic conditions in the colon (Table 5) [131].
To simulate the gastrointestinal tract, most fermentation systems use working volumes similar to the physiological ones. This choice leads to an increase in costs, mainly due to the culture medium used, and in the physical space required for the overall system. The MiniBioReactor arrays (MBRAs) and the smallest intestine (TSI) (Tables 4 and 5) are two examples of dynamic mono- and multi-compartment systems, respectively, where the working volume inside the system is drastically reduced. The TSI, being constituted by five reactor units enclosed in a box where the environmental parameters are constantly controlled, replicates transit through the small intestine [132]. A dialysis chamber is used to simulate the absorption of nutrients. The results obtained from a study involving the TSI reveal that several strains of Lactobacillus have been successfully cultured inside the model [132].
A common problem among the different in vitro fermentation systems is the inoculation of the fecal sample. In fact, most systems use a liquid fecal suspension as inoculum without a physical substrate, resulting in several limitations like the absence of biofilm-associated microorganisms [133]. To overcome this issue, the PolyFermS system uses immobilized microorganisms (Table 5). The system is composed of an inoculum reactor made of micro-encapsulated microorganisms from the human gut microbiota. This reactor is used to supply other reactors disposed in parallel, which have varying environmental conditions. This model has been found to maintain a stable microbial community for 38 days [134]. Another approach in guaranteeing a stable microbial profile, even where cells are suspended, is described by Li and colleagues [135]. In fact, they demonstrated that unlike a continuous reactor system, a looped mass transfer can stabilize microbial communities over a long period of time.
Co-culture systems
One of the bioengineering challenges over the last 10 years has been to create an in vitro model explaining how the human gut microbiota interacts with eukaryotic cells from the host. Several strategies derived from tissue engineering principles have been applied to microbiology to reproduce a co-culture between microorganisms and mammalian cells. These models are well reviewed in [148,149,150,151,152,153,154]. Some examples of these co-culture approaches, with a focus on the devices, cultured cells, and bacteria, are reported in Table 6 and Fig. 3.
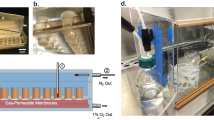
Scheme of other devices used to study interactions between the human gut microbiota and eukaryotic cells. a The HoxBan system used for static co-culture between microorganisms, F. prausnitzii, and Caco-2 cells (reproduced from [156], Copyright 2015, with permission from the authors). b The HMI module consists of two chambers (the upper one contains bacteria, while the lower one contains enterocytes) separated by a polyamide semipermeable membrane and a mucus layer that form a double functional layer (reproduced from [157], Copyright 2014, with permission from the authors). c The HuMiX module is composed of three silicone rubber gaskets, each of which defines a distinct spiral-shaped channel (200 mm in length, 4 mm in width, and 0.5 mm in height), separated by two semipermeable membranes (a microporous membrane between the perfusion chamber and the cell chamber and a nanoporous membrane between the cell chamber and the microbial chamber). The whole structure is enclosed between two polycarbonate sheets (reproduced from [158], Copyright 2016, with permission from the authors). d The anoxic–oxic interface-on-a-chip module is fabricated through soft lithography and is composed of two PDMS parts. Anoxic (blue) and oxic (red) culture media are supplied through two different microchannels, separated by a porous PDMS membrane, to recreate an oxygen gradient (reproduced from [97], Copyright 2019, with permission from the authors)
The purpose of this section is to describe some of these technologies to evaluate if and how they can be adapted to study the effects of the human gut microbiota on the host.
Transwell culture models are particularly useful for studying the interaction between bacteria and intestinal epithelial cells in aerobic conditions [159]. Typically, these wells consist of a lower compartment on which the first cell line can be placed, and a removable upper insert consisting of a microporous membrane, on which a second cell line can be seeded. The pores are large enough for the passage of growth factors and other molecules released by the cell, but too small to allow passage of the cells themselves. Recently, a variant of this Transwell culture system has been developed, allowing analysis of host-microbe interactions between Caco-2 cells and anaerobic Fecalibacterium prausnitzii, as reported in the study by Ulluwishewa et al. [160]. Caco-2 cells were grown on microporous membrane inserts. Due to the polarization of the Caco-2 monolayer (i.e., cells are arranged in an organized manner with the basal part on the bottom of the membrane and the apical part on the top), two culture media were used. In the basal compartment, an aerobic medium was used to prevent cellular death due to hypoxia. In the apical compartment, an anaerobic medium was used to culture F. prausnitzii, instead. The overall system was isolated from the external environment and the co-culture chamber was placed in an anaerobic workstation. The well was also equipped with a pair of electrodes to assess the integrity of the cell monolayer junction. It is interesting to note how Maier et al. adopted the “mixed medium” approach for their Transwell system [155]. Thus, F. prausnitzii was cultured in an anaerobic medium composed of 50% M199 (cell-culture medium) and 50% BHI (bacterial-culture medium). This combination improved not only the viability of F. prausnitzii but also attachment of HEK293-TLR2-Luc cells on the collagen-coated inserts. Conversely, the basal compartment was filled with aerobic Dulbecco’s modified eagle medium (DMEM) supplemented with 10% fetal bovine serum (FBS), to allow generation of the anoxic–oxic interface and guarantee HEK293 cell survival. Although these models are most commonly used by the pharmaceutical industry, they cannot reproduce the physiological morphology of three-dimensional (3D) intestinal cells and tissues as well as some of the key intestinal functions (i.e., mucus production and villi formation). Furthermore, being static models, they present the problem of periodic manual change of the culture media to avoid toxic waste product accumulation and cannot support culture of the human gut microbiota together with human intestinal cells for more than one day [153].
Bioreactors are another example of devices used in tissue engineering to promote cell proliferation and differentiation. These systems perform “incubator” functions, ensuring dynamic environmental conditions with a greater physiological relevance. Among these millifluidic devices, the host-microbiota interaction (HMI) [157] is designed to be connected to a SHIME system [142] (Fig. 3b). This indirect co-culture bioreactor has two compartments separated by a semi-permeable membrane and an artificial mucus layer made of mucin and agar. The lower compartment is dedicated to cell culture, while the upper compartment carries the products from the SHIME. The system was tested with fermentation products from the yeast Saccharomyces cerevisiae, and the group found that Caco-2 cells remained viable after 48 h of co-culture [157].
Although dynamic conditions promote bacterial proliferation as well as recirculation of the culture medium and elimination of waste products, these systems present some problems related to their size and feasibility. To overcome some of the limitations described, microfluidics devices have been properly implemented to study the human gut microbiota. A microfluidic device can be defined as a perfusion device that hosts one or more cell types and aims at reproducing key structures, functions, and aspects of human metabolism of a given tissue or organ in normal and pathological physiology [153]. Miniaturization of these devices still allows integration of control, sensors, imaging systems, and other analytical components [161]. There are several examples of microfluidic devices in the literature with different and peculiar features. The “gut-on-chip” described by Kim et al. was fabricated by soft lithography and composed of two parallel micro-channels [162]. These two compartments were separated by a porous PDMS membrane coated with an extracellular matrix (ECM) solution composed of rat type I collagen and Matrigel in serum-free DMEM. The culture medium was perfused through the microchannels, representing the fluid flow and shear stresses present in the human intestine, while cyclic deformations were applied to mimic peristalsis movement. Inside this device, Caco-2 cells were arranged to produce cellular monolayer and villi structures, and Lactobacillus rhamnosus was co-cultured. Results showed that after 96 h of co-culture in dynamic conditions, L. rhamnosus continued to adhere to the Caco-2 surfaces, ensuring a 95% viability rate of the monolayer, while in the control (i.e., a Transwell system in static condition) the death of the epithelial monolayer was observed after only 48 h. Jalili-Firoozinezhad et al. proposed a similar device that allowed the human intestinal epithelium to be cultured together with the human gut microbiota [163]. Using this device, it is possible to culture in vitro microorganisms in direct contact with host cells and their naturally produced mucus layer for at least 5 days. Culturing a complex microbial community together with host cells over a long period is difficult due to the high growth rate of bacteria compared to that of mammalian cells. Bacteria can also invade and kill epithelial cells, so separating microorganisms from host cells may overcome this problem. Pajoumshariati et al. described another system where different enteric bacterial species (i.e., E. coli, Enterococcus faecalis, Klebsiella pneumoniae, P. aeruginosa) and other bacterial species isolated from the ileum of patients with Crohn’s disease (CD) were incorporated within chitosan-coated alginate-based microfibers (60 µm in diameter) that spatially separated microorganisms from co-cultured cells [164]. This encapsulation in a physical substrate successfully recreated the biofilm-associated microorganisms in the intestine. To reproduce the gut environment, mucins were also incorporated into the alginate. The results showed that this model is able to retain bacteria within the microfibers for an acceptable period, thus providing a 3D microenvironment for bacterial growth and proliferation. The model also allows cells to more closely mimic their natural growth than planktonic and 2D in vitro methods.
Culturing cells and bacteria in a microfluidic system, particularly in the long term, can also lead to other problems, including fluid leakage [165], clogging [166], and unwanted accumulation of bubbles in the channels. Although some of these problems are manageable through a careful selection of materials or protocols, bubble accumulation is a frequent obstacle that is extremely difficult to avoid in most PDMS microfluidic systems [167]. Bubbles have a high probability of forming in the connection between the channel and the tube from which the fluid arrives. In addition, bubbles can gradually grow in the channel due to temperature and pressure variation. The presence of air bubbles can damage cells, rupture the cell membrane, or even wash away the cells. For this reason, some devices may involve the use of “traps” to eliminate these bubbles [168].
Ultimately, one of the new frontiers of research is to co-culture the human gut microbiota and intestinal organoids. The latter are self-assembling 3D cellular constructs made from stem cells that represent and reproduce the main physiological properties of an organ. From a purely physical and geometric point of view, an intestinal organoid can be considered as a closed 3D geometry with an internal cavity where a low level of oxygen is present. Furthermore, intestinal cells exhibit polarity and are typically arranged with the apical part toward the inside of the cavity. One of the first attempts to co-culture the gut microbiota and an organoid was made by inserting intestinal microorganisms directly into the cavity by micro-injection [169,170,171,172,173]. Although this technique is very easy and straightforward, it presents many risks related to co-culture over long periods such as disruption of the organoid membrane, a lack of medium recirculation, and a high probability of infection. Another possibility is to make organoids with the apical cellular part in contact with the external environment [174]. In this way, it becomes possible to insert the microbial suspension directly into the culture medium, preventing leakage problems during the micro-injection phase. Another method that can be applied is to linearize the structure of an organoid by switching from a 3D to a 2D construct. Indeed, standardized organoids can be fragmented and cultured to make monolayers [175, 176]. Using this method, several systems including anaerobic microorganisms have been realized through the formation of oxygen gradients or anaerobiosis chambers [177, 178]. This method can also be applied to microfluidic systems by inserting monolayers derived from organoids [97]. In this way, it is possible to use these systems for long-term cultures (i.e., 24 h for the Transwell systems above described [177, 178]) and, thanks to dynamic conditions, accumulation of a mucus layer, with a thickness similar to that found in vivo, can be observed [179].
Concluding remarks
Revealing the connections between the composition of the human gut microbiota and the consequent alteration of the normal physiological state is currently one of the most challenging research topics. Different studies have attempted to bridge this gap by exploiting in vitro models, drugs, pathogens, or highly predictive tools to show the effects of dietary components, on the activity and composition of a complex microbial community such as the gut microbiota, as well as to unravel the dense network of interactions between microorganisms and eukaryotic cells in different physiological states. Recreating a complete in vitro model of the human gut microbiota requires several initial steps, and a priority at this stage is comprehension of the ultimate purpose of the study itself. When fermentative processes and metabolic pathways carried out by the gut microbiota are investigated, more attention must be paid to the selected culture medium and maintenance of the culture parameters. On the other hand, when studying the effects of the human gut microbiota on eukaryotic cells, creation of an oxygen gradient between bacteria and cells, as well as the presence of cytotoxic molecules produced by microorganisms, becomes important priorities. Although extensive experience has been gained with in vitro systems over the last 20 years, most of the results are inconclusive and comprise single bacterial strains as opposed to a complex microbial profile like the human gut microbiota. In the future, researchers should primarily focus on designing human gut microbiota models whose activity and composition remain constant over time, or at least have only small fluctuations, and which are comparable with those of in vivo communities. Furthermore, it will be necessary for these technologies to be supported by in silico tools to create more adequate predictive models.
References
Rowland I, Gibson G, Heinken A et al (2018) Gut microbiota functions: metabolism of nutrients and other food components. Eur J Nutr 57:1–24. https://doi.org/10.1007/s00394-017-1445-8
Steer T, Carpenter H, Tuohy K et al (2000) Perspectives on the role of the human gut microbiota and its modulation by pro- and prebiotics. Nutr Res Rev 13:229–254. https://doi.org/10.1079/095442200108729089
Turnbaugh PJ, Ley RE, Mahowald MA et al (2006) An obesity-associated gut microbiome with increased capacity for energy harvest. Nature 444:1027–1031. https://doi.org/10.1038/nature05414
Rajilić-Stojanović M, Biagi E, Heilig HGHJ et al (2011) Global and deep molecular analysis of microbiota signatures in fecal samples from patients with irritable bowel syndrome. Gastroenterology 141:1792–1801. https://doi.org/10.1053/j.gastro.2011.07.043
Dicksved J, Halfvarson J, Rosenquist M et al (2008) Molecular analysis of the gut microbiota of identical twins with Crohn’s disease. ISME J 2:716–727. https://doi.org/10.1038/ismej.2008.37
Schroeder BO, Bäckhed F (2016) Signals from the gut microbiota to distant organs in physiology and disease. Nat Med 22:1079–1089. https://doi.org/10.1038/nm.4185
Cryan JF, O’riordan KJ, Cowan CSM et al (2019) The microbiota-gut-brain axis. Physiol Rev 99:1877–2013. https://doi.org/10.1152/physrev.00018.2018
Kostic AD, Howitt MR, Garrett WS (2013) Exploring host-microbiota interactions in animal models and humans. Genes Dev 27:701–718. https://doi.org/10.1101/gad.212522.112
Uzbay T (2019) Germ-free animal experiments in the gut microbiota studies. Curr Opin Pharmacol 49:6–10. https://doi.org/10.1016/j.coph.2019.03.016
Al-Asmakh M, Zadjali F (2015) Use of germ-free animal models in microbiota-related research. J Microbiol Biotechn 25(10):1583–1588. https://doi.org/10.4014/jmb.1501.01039
Pearce SC, Coia HG, Karl JP et al (2018) Intestinal in vitro and ex vivo models to study host-microbiome interactions and acute stressors. Front Physiol 9:1584. https://doi.org/10.3389/fphys.2018.01584
Jones RB, Zhu X, Moan E et al (2018) Inter-niche and inter-individual variation in gut microbial community assessment using stool, rectal swab, and mucosal samples. Sci Rep 8:4139. https://doi.org/10.1038/s41598-018-22408-4
Tang Q, Jin G, Wang G et al (2020) Current sampling methods for gut microbiota: a call for more precise devices. Front Cell Infect Microbiol 10:151. https://doi.org/10.3389/fcimb.2020.00151
Booijink CCGM, El-Aidy S, Rajilić-Stojanović M et al (2010) High temporal and inter-individual variation detected in the human ileal microbiota. Environ Microbiol 12:3213–3227. https://doi.org/10.1111/J.1462-2920.2010.02294.X
Petrof EO, Gloor GB, Vanner SJ et al (2013) Stool substitute transplant therapy for the eradication of Clostridium difficile infection: ‘RePOOPulating’ the gut. Microbiome 1:3. https://doi.org/10.1186/2049-2618-1-3
Krause JL, Schaepe SS, Fritz-Wallace K et al (2020) Following the community development of SIHUMIx—a new intestinal in vitro model for bioreactor use. Gut Microbes 11:1116–1129. https://doi.org/10.1080/19490976.2019.1702431
Lawson CE, Harcombe WR, Hatzenpichler R et al (2019) Common principles and best practices for engineering microbiomes. Nat Rev Microbiol 17:725–741. https://doi.org/10.1038/s41579-019-0255-9
Schäpe SS, Krause JL, Engelmann B et al (2019) The simplified human intestinal microbiota (SIHUMIx) shows high structural and functional resistance against changing transit times in in vitro bioreactors. Microorganisms 7:641. https://doi.org/10.3390/microorganisms7120641
Ito T, Sekizuka T, Kishi N et al (2019) Conventional culture methods with commercially available media unveil the presence of novel culturable bacteria. Gut Microbes 10:77–91. https://doi.org/10.1080/19490976.2018.1491265
Bonnet M, Lagier JC, Raoult D et al (2020) Bacterial culture through selective and non-selective conditions: the evolution of culture media in clinical microbiology. New Microbes New Infect 34:100622. https://doi.org/10.1016/j.nmni.2019.100622
Tidjani Alou M, Naud S, Khelaifia S et al (2020) State of the art in the culture of the human microbiota: new interests and strategies. Clin Microbiol Rev 34(1):e00129-19. https://doi.org/10.1128/CMR.00129-19
Kim BS, Kim JN, Cerniglia CE (2011) In vitro culture conditions for maintaining a complex population of human gastrointestinal tract microbiota. J Biomed Biotechnol 2011:838040. https://doi.org/10.1155/2011/838040
Li L, Zhang X, Ning Z et al (2018) Evaluating in vitro culture medium of gut microbiome with orthogonal experimental design and a metaproteomics approach. J Proteome Res 17:154–163. https://doi.org/10.1021/acs.jproteome.7b00461
Yousi F, Kainan C, Junnan Z et al (2019) Evaluation of the effects of four media on human intestinal microbiota culture in vitro. AMB Expr 9:69. https://doi.org/10.1186/s13568-019-0790-9
Thursby E, Juge N (2017) Introduction to the human gut microbiota. Biochem J 474:1823–1836. https://doi.org/10.1042/BCJ20160510
Duncan SH, Louis P, Thomson JM et al (2009) The role of pH in determining the species composition of the human colonic microbiota. Environ Microbiol 11:2112–2122. https://doi.org/10.1111/j.1462-2920.2009.01931.x
Donaldson GP, Lee SM, Mazmanian SK (2015) Gut biogeography of the bacterial microbiota. Nat Rev Microbiol 14:20–32. https://doi.org/10.1038/nrmicro3552
Espey MG (2013) Role of oxygen gradients in shaping redox relationships between the human intestine and its microbiota. Free Radic Biol Med 55:130–140. https://doi.org/10.1016/J.FREERADBIOMED.2012.10.554
Haindl R, Schick S, Kulozik U (2021) Influence of cultivation pH on composition, diversity, and metabolic production in an in vitro human intestinal microbiota. Fermentation 7:156. https://doi.org/10.3390/fermentation7030156
Zihler Berner A, Fuentes S, Dostal A et al (2013) Novel polyfermentor intestinal model (PolyFermS) for controlled ecological studies: validation and effect of pH. PLoS ONE 8(10):e77772. https://doi.org/10.1371/JOURNAL.PONE.0077772
Corsetti M, Costa M, Bassotti G et al (2019) First translational consensus on terminology and definitions of colonic motility in animals and humans studied by manometric and other techniques. Nat Rev Gastroenterol Hepatol 169(16):559–579. https://doi.org/10.1038/s41575-019-0167-1
Parthasarathy G, Chen J, Chen X et al (2016) Relationship between microbiota of the colonic mucosa vs feces and symptoms, colonic transit, and methane production in female patients with chronic constipation. Gastroenterology 150:367–379. https://doi.org/10.1053/J.GASTRO.2015.10.005
Vandeputte D, Falony G, Vieira-Silva S et al (2016) Stool consistency is strongly associated with gut microbiota richness and composition, enterotypes and bacterial growth rates. Gut Microbiota 65:57–62. https://doi.org/10.1136/GUTJNL-2015-309618
Asnicar F, Leeming ER, Dimidi E et al (2021) Blue poo: impact of gut transit time on the gut microbiome using a novel marker. Gut Microbiota 70:1665–1674. https://doi.org/10.1136/GUTJNL-2020-323877
Tottey W, Feria-Gervasio D, Gaci N et al (2017) Colonic transit time is a driven force of the gut microbiota composition and metabolism: in vitro evidence. J Neurogastroenterol Motil 23:124–134. https://doi.org/10.5056/jnm16042
An YH, Friedman RJ (1998) Concise review of mechanisms of bacterial adhesion to biomaterial surfaces. J Biomed Mater Res 43:338–348. https://doi.org/10.1002/(SICI)1097-4636(199823)43:3%3c338::AID-JBM16%3e3.0.CO;2-B
Carniello V, Peterson BW, van der Mei HC et al (2018) Physico-chemistry from initial bacterial adhesion to surface-programmed biofilm growth. Adv Colloid Interf Sci 261:1–14. https://doi.org/10.1016/j.cis.2018.10.005
Hermansson M (1999) The DLVO theory in microbial adhesion. Colloids Surf B Biointerf 14:105–119. https://doi.org/10.1016/S0927-7765(99)00029-6
Hori K, Matsumoto S (2010) Bacterial adhesion: from mechanism to control. Biochem Eng J 48:424–434. https://doi.org/10.1016/J.BEJ.2009.11.014
Krasowska A, Sigler K (2014) How microorganisms use hydrophobicity and what does this mean for human needs? Front Cell Infect Microbiol 4:112. https://doi.org/10.3389/FCIMB.2014.00112/BIBTEX
Sharma S, Conrad JC (2014) Attachment from flow of Escherichia coli bacteria onto silanized glass substrates. Langmuir 30:11147–11155. https://doi.org/10.1021/la502313y
Oh JK, Yegin Y, Yang F et al (2018) The influence of surface chemistry on the kinetics and thermodynamics of bacterial adhesion. Sci Rep 8:17247. https://doi.org/10.1038/s41598-018-35343-1
Kankainen M, Paulin L, Tynkkynen S et al (2009) Comparative genomic analysis of lactobacillus rhamnosus GG reveals pili containing a human-mucus binding protein. Proc Natl Acad Sci 106:17193–17198. https://doi.org/10.1073/pnas.0908876106
Sicard JF, Le Bihan G, Vogeleer P et al (2017) Interactions of intestinal bacteria with components of the intestinal mucus. Front Cell Infect Microbiol 7:387. https://doi.org/10.3389/fcimb.2017.00387
Dunne WM (2002) Bacterial adhesion: seen any good biofilms lately? Clin Microbiol Rev 15:155–166. https://doi.org/10.1128/CMR.15.2.155-166.2002
Jamal M, Ahmad W, Andleeb S et al (2018) Bacterial biofilm and associated infections. J Chin Med Assoc 81:7–11. https://doi.org/10.1016/j.jcma.2017.07.012
Zheng S, Bawazir M, Dhall A et al (2021) Implication of surface properties, bacterial motility, and hydrodynamic conditions on bacterial surface sensing and their initial adhesion. Front Bioeng Biotechnol 9:82. https://doi.org/10.3389/FBIOE.2021.643722/BIBTEX
Chen C, Petterson T, Illergard J et al (2019) Influence of cellulose charge on bacteria adhesion and viability to PVAm/CNF/PVAm-modified cellulose model surfaces. Biomacromol 20(5):2075–2083. https://doi.org/10.1021/acs.biomac.9b00297
Kovačević D, Pratnekar R, Torkar KG et al (2016) Influence of polyelectrolyte multilayer properties on bacterial adhesion capacity. Polymers 8(10):345. https://doi.org/10.3390/POLYM8100345
Rzhepishevska O, Hakobyan S, Ruhal R et al (2013) The surface charge of anti-bacterial coatings alters motility and biofilm architecture. Biomater Sci 1:589–602. https://doi.org/10.1039/C3BM00197K
Kao WK, Gagnon PM, Vogel JP et al (2017) Surface charge modification decreases pseudomonas aeruginosa adherence in vitro and bacterial persistence in an in vivo implant model. Laryngoscope 127(7):1655–1661. https://doi.org/10.1002/lary.26499
Guégan C, Garderes J, Le Pennec G et al (2014) Alteration of bacterial adhesion induced by the substrate stiffness. Colloids Surf B Biointerf 114:193–200. https://doi.org/10.1016/j.colsurfb.2013.10.010
Lichter JA, Thompson MT, Delgadillo M et al (2008) Substrata mechanical stiffness can regulate adhesion of viable bacteria. Biomacromol 9:1571–1578. https://doi.org/10.1021/bm701430y
Song F, Ren D (2014) Stiffness of cross-linked poly(dimethylsiloxane) affects bacterial adhesion and antibiotic susceptibility of attached cells. Langmuir 30:10354–10362. https://doi.org/10.1021/la502029f
Wang Y, Guan A, Isayeva I et al (2016) Interactions of Staphylococcus aureus with ultrasoft hydrogel biomaterials. Biomaterials 95:74–85. https://doi.org/10.1016/j.biomaterials.2016.04.005
Hou S, Gu H, Smith C et al (2011) Microtopographic patterns affect Escherichia coli biofilm formation on poly(dimethylsiloxane) surfaces. Langmuir 27:2686–2691. https://doi.org/10.1021/la1046194
Lu N, Zhang W, Weng Y et al (2016) Fabrication of PDMS surfaces with micro patterns and the effect of pattern sizes on bacteria adhesion. Food Contr 68:344–351. https://doi.org/10.1016/j.foodcont.2016.04.014
Perni S, Prokopovich P (2013) Micropatterning with conical features can control bacterial adhesion on silicone. Soft Matter 9:1844–1851. https://doi.org/10.1039/c2sm26828k
Xu LC, Siedlecki CA (2012) Submicron-textured biomaterial surface reduces staphylococcal bacterial adhesion and biofilm formation. Acta Biomater 8:72–81. https://doi.org/10.1016/j.actbio.2011.08.009
Ge X, Leng Y, Lu X et al (2015) Bacterial responses to periodic micropillar array. J Biomed Mater Res Part A 103:384–396. https://doi.org/10.1002/jbm.a.35182
Yang M, Ding YH, Ge X et al (2015) Control of bacterial adhesion and growth on honeycomb-like patterned surfaces. Colloid Surf B 135:549–555. https://doi.org/10.1016/j.colsurfb.2015.08.010
Kargar M, Wang J, Nain AS et al (2012) Controlling bacterial adhesion to surfaces using topographical cues: a study of the interaction of pseudomonas aeruginosa with nanofiber-textured surfaces. Soft Matter 8:10254. https://doi.org/10.1039/c2sm26368h
Biagini F, Calvigioni M, Lapomarda A et al (2020) A novel 3D in vitro model of the human gut microbiota. Sci Rep 10:21499–21510. https://doi.org/10.1038/s41598-020-78591-w
Biagini F, Calvigioni M, De Maria C et al (2022) Study of the adhesion of the human gut microbiota on electrospun structures. Bioengineering 9:96. https://doi.org/10.3390/bioengineering9030096
Verhorstert KWJ, Guler Z, de Boer L et al (2020) In vitro bacterial adhesion and biofilm formation on fully absorbable poly-4-hydroxybutyrate and nonabsorbable polypropylene pelvic floor implants. ACS Appl Mater Interf 12(48):53646–53653. https://doi.org/10.1021/acsami.0c14668
Yuan Y, Hays MP, Hardwidge PR et al (2017) Surface characteristics influencing bacterial adhesion to polymeric substrates. RSC Adv 7:14254–14261. https://doi.org/10.1039/c7ra01571b
De-la-Pinta I, Cobos M, Ibarretxe J et al (2019) Effect of biomaterials hydrophobicity and roughness on biofilm development. J Mater Sci Mater Med 30:77. https://doi.org/10.1007/S10856-019-6281-3
Vaudaux P, Suzuki R, Waldvogel FA et al (1984) Foreign body infection: role of fibronectin as a ligand for the adherence of Staphylococcus aureus. J Infect Dis 150:546–553. https://doi.org/10.1093/INFDIS/150.4.546
Kuusela P, Vartio T, Vuento M et al (1985) Attachment of staphylococci and streptococci on fibronectin, fibronectin fragments, and fibrinogen bound to a solid phase. Infect Immun 50:77. https://doi.org/10.1128/iai.50.1.77-81.1985
Herrmann M, Vaudaux P, Pittet D et al (1988) Fibronectin, fibrinogen, and laminin act as mediators of adherence of clinical staphylococcal isolates to foreign material. J Infect Dis 158:693–701. https://doi.org/10.1093/INFDIS/158.4.693
Etienne-Mesmin L, Chassaing B, Desvaux M et al (2019) Experimental models to study intestinal microbes–mucus interactions in health and disease. FEMS Microbiol Rev 43:457–489. https://doi.org/10.1093/FEMSRE/FUZ013
Paone P, Cani PD (2020) Mucus barrier, mucins and gut microbiota: the expected slimy partners? Gut 69:2232–2243. https://doi.org/10.1136/GUTJNL-2020-322260
Boekhorst J, Helmer Q, Kleerebezem M et al (2006) Comparative analysis of proteins with a mucus-binding domain found exclusively in lactic acid bacteria. Microbiology 152:273–280. https://doi.org/10.1099/MIC.0.28415-0
Van den Abbeele P, Roos S, Eeckhaut V et al (2012) Incorporating a mucosal environment in a dynamic gut model results in a more representative colonization by lactobacilli. Microb Biotechnol 5:106–115. https://doi.org/10.1111/J.1751-7915.2011.00308.X
Persat A, Nadell CD, Kim MK et al (2015) The mechanical world of bacteria. Cell 161:988–997. https://doi.org/10.1016/j.cell.2015.05.005
Liu Y, Tay JH (2002) The essential role of hydrodynamic shear force in the formation of biofilm and granular sludge. Water Res 36:1653–1665. https://doi.org/10.1016/S0043-1354(01)00379-7
Purevdorj B, Costerton JW, Stoodley P (2002) Influence of hydrodynamics and cell signaling on the structure and behavior of pseudomonas aeruginosa biofilms. Appl Environ Microbiol 68:4457–4464. https://doi.org/10.1128/AEM.68.9.4457-4464.2002
Garrett TR, Bhakoo M, Zhang Z (2008) Bacterial adhesion and biofilms on surfaces. Prog Nat Sci 18:1049–1056. https://doi.org/10.1016/j.pnsc.2008.04.001
Lecuyer S, Rusconi R, Shen Y et al (2011) Shear stress increases the residence time of adhesion of pseudomonas aeruginosa. Biophys J 100:341–350. https://doi.org/10.1016/j.bpj.2010.11.078
Moreira JMR, Araújo JDP, Miranda JM et al (2014) The effects of surface properties on Escherichia coli adhesion are modulated by shear stress. Colloids Surf B Biointerf 123:1–7. https://doi.org/10.1016/j.colsurfb.2014.08.016
Tilles AW, Baskaran H, Roy P et al (2001) Effects of oxygenation and flow on the viability and function of rat hepatocytes cocultured in a microchannel flat-plate bioreactor. Biotechnol Bioeng 73:379–389. https://doi.org/10.1002/bit.1071
Rusconi R, Lecuyer S, Guglielmini L et al (2010) Laminar flow around corners triggers the formation of biofilm streamers. J R Soc Interf 7:1293–1299. https://doi.org/10.1098/rsif.2010.0096
Stoodley P, Lewandowski Z, Boyle JD et al (1999) Structural deformation of bacterial biofilms caused by short-term fluctuations in fluid shear: an in situ investigation of biofilm rheology. Biotechnol Bioeng 65:83–92. https://doi.org/10.1002/(SICI)1097-0290(19991005)65:1%3c83::AID-BIT10%3e3.0.CO;2-B
Kim MK, Drescher K, Shun Pak O et al (2014) Filaments in curved streamlines: rapid formation of Staphylococcus aureus biofilm streamers. New J Phys 16:065024. https://doi.org/10.1088/1367-2630/16/6/065024
Thomen P, Robert J, Monmeyran A et al (2017) Bacterial biofilm under flow: first a physical struggle to stay, then a matter of breathing. PLoS ONE 12:e0175197. https://doi.org/10.1371/journal.pone.0175197
Siddiqui S, Chandrasekaran A, Lin N et al (2019) Microfluidic shear assay to distinguish between bacterial adhesion and attachment strength on stiffness-tunable silicone substrates. Langmuir 35:8840–8849. https://doi.org/10.1021/acs.langmuir.9b00803
Li ZJ, Mohamed N, Ross JM (2000) Shear stress affects the kinetics of Staphylococcus aureus adhesion to collagen. Biotechnol Prog 16:1086–1090. https://doi.org/10.1021/bp000117r
Vis MAM, Ito K, Hofmann S (2020) Impact of culture medium on cellular interactions in in vitro co-culture systems. Front Bioeng Biotechnol 8:911. https://doi.org/10.3389/FBIOE.2020.00911/BIBTEX
Zhu S, Ehnert S, Rouß M et al (2018) From the clinical problem to the basic research-co-culture models of osteoblasts and osteoclasts. Int J Mol Sci 19:2284. https://doi.org/10.3390/ijms19082284
Jones GL, Motta A, Marshall MJ et al (2009) Osteoblast: osteoclast co-cultures on silk fibroin, chitosan and PLLA films. Biomaterials 30:5376–5384. https://doi.org/10.1016/j.biomaterials.2009.07.028
Lavender MD, Pang Z, Wallace CS et al (2005) A system for the direct co-culture of endothelium on smooth muscle cells. Biomaterials 26:4642–4653. https://doi.org/10.1016/j.biomaterials.2004.11.045
Goers L, Freemont P, Polizzi KM (2014) Co-culture systems and technologies: taking synthetic biology to the next level. J R Soc Interf 11:20140065. https://doi.org/10.1098/rsif.2014.0065
Chung S, Sudo R, Mack PJ et al (2009) Cell migration into scaffolds under co-culture conditions in a microfluidic platform. Lab Chip 9:269–275. https://doi.org/10.1039/B807585A
Katagiri W, Sakaguchi K, Kawai T et al (2017) A defined mix of cytokines mimics conditioned medium from cultures of bone marrow-derived mesenchymal stem cells and elicits bone regeneration. Cell Prolif 50:e12333. https://doi.org/10.1111/cpr.12333
Bidarra SJ, Barrias CC, Barbosa MA et al (2011) Phenotypic and proliferative modulation of human mesenchymal stem cells via crosstalk with endothelial cells. Stem Cell Res 7:186–197. https://doi.org/10.1016/j.scr.2011.05.006
Klitgord N, Segrè D (2010) Environments that induce synthetic microbial ecosystems. PLoS Comput Biol 6:e1001002. https://doi.org/10.1371/journal.pcbi.1001002
Shin W, Wu A, Massidda MW et al (2019) A robust longitudinal co-culture of obligate anaerobic gut microbiome with human intestinal epithelium in an anoxic-oxic interface-on-a-chip. Front Bioeng Biotechnol 7:13. https://doi.org/10.3389/fbioe.2019.00013
Yoshimura Y, Kikuiri T, Hasegawa T et al (2017) How much medium do you use for cell culture? Medium volume influences mineralization and osteoclastogenesis in vitro. Mol Med Rep 16:429–434. https://doi.org/10.3892/MMR.2017.6611/HTML
Shimomura A, Iizuka-Kogo A, Yamamoto N et al (2016) A lower volume culture method for obtaining a larger yield of neuron-like cells from mesenchymal stem cells. Med Mol Morphol 49:119–126. https://doi.org/10.1007/s00795-015-0131-2
Simão VA, Evangelista-Ribeiro CP, Brand H et al (2019) Metabolic and proliferation evaluation of human adipose-derived mesenchymal stromal cells (ASC) in different culture medium volumes: standardization of static culture. Biologicals 62:93–101. https://doi.org/10.1016/J.BIOLOGICALS.2019.08.006
Krüger-Genge A, Fuhrmann R, Jung F et al (2015) Morphology of primary human venous endothelial cell cultures before and after culture medium exchange. Clin Hemorheol Microcirc 61:151–156. https://doi.org/10.3233/CH-151992
Büntemeyer H, Wallerius C, Lehmann J (1992) Optimal medium use for continuous high density perfusion processes. Cytotechnology 9:59–67. https://doi.org/10.1007/BF02521732
Chen G, Gulbranson DR, Hou Z et al (2011) Chemically defined conditions for human iPSC derivation and culture. Nat Methods 85(8):424–429. https://doi.org/10.1038/nmeth.1593
Shinohara M, Choi H, Ibuki M et al (2019) Endodermal differentiation of human induced pluripotent stem cells using simple dialysis culture system in suspension culture. Regen Ther 12:14–19. https://doi.org/10.1016/J.RETH.2019.05.004
Côme J, Nissan X, Aubry L et al (2008) Improvement of culture conditions of human embryoid bodies using a controlled perfused and dialyzed bioreactor system. Tissue Eng Part C Methods 14:289–298. https://doi.org/10.1089/TEN.TEC.2008.0029
Stieb M, Schink B (1987) Cultivation of syntrophic anaerobic bacteria in membrane-separated culture devices. FEMS Microbiol Lett 45:71–76. https://doi.org/10.1111/j.1574-6968.1987.tb02341.x
Ohno M, Okano I, Watsuji T et al (1999) Establishing the independent culture of a strictly symbiotic bacterium symbiobacterium thermophilum from its supporting bacillus strain. Biosci Biotechnol Biochem 63(6):1083–1090. https://doi.org/10.1271/bbb.63.1083
Kapoore RV, Padmaperuma G, Maneein S et al (2022) Co-culturing microbial consortia: approaches for applications in biomanufacturing and bioprocessing. Crit Rev Biotechnol 42:46–72. https://doi.org/10.1080/07388551.2021.1921691
Shin W, Kim HJ (2018) Intestinal barrier dysfunction orchestrates the onset of inflammatory host–microbiome cross-talk in a human gut inflammation-on-a-chip. Proc Natl Acad Sci 115:E10539–E10547. https://doi.org/10.1073/pnas.1810819115
Flint HJ (2012) The impact of nutrition on the human microbiome. Nutr Rev 70:S10–S13. https://doi.org/10.1111/j.1753-4887.2012.00499.x
Louis P, Hold GL, Flint HJ (2014) The gut microbiota, bacterial metabolites and colorectal cancer. Nat Rev Microbiol 12:661–672. https://doi.org/10.1038/nrmicro3344
Bohn T, Carriere F, Day L et al (2018) Correlation between in vitro and in vivo data on food digestion. What can we predict with static in vitro digestion models? Crit Rev Food Sci Nutr 58:2239–2261. https://doi.org/10.1080/10408398.2017.1315362
Sarbini SR, Kolida S, Naeye T et al (2011) In vitro fermentation of linear and α-1,2-branched dextrans by the human fecal microbiota. Appl Environ Microbiol 77(15):5307–5315. https://doi.org/10.1128/AEM.02568-10
Minekus M, Alminger M, Alvito P et al (2014) A standardised static in vitro digestion method suitable for food – an international consensus. Food Funct 5:1113–1124. https://doi.org/10.1039/C3FO60702J
Brodkorb A, Egger L, Alminger M et al (2019) INFOGEST static in vitro simulation of gastrointestinal food digestion. Nat Protoc 14:991–1014. https://doi.org/10.1038/s41596-018-0119-1
Pérez-Burillo S, Molino S, Navajas-Porras B et al (2021) An in vitro batch fermentation protocol for studying the contribution of food to gut microbiota composition and functionality. Nat Protoc 16:3186–3209. https://doi.org/10.1038/s41596-021-00537-x
Ji H, Hu J, Zuo S et al (2021) In vitro gastrointestinal digestion and fermentation models and their applications in food carbohydrates. Crit Rev Food Sci Nutr 62(19):5349–5371. https://doi.org/10.1080/10408398.2021.1884841
Dupont D, Alric M, Blanquet-Diot S et al (2019) Can dynamic in vitro digestion systems mimic the physiological reality? Crit Rev Food Sci 59(10):1546–1562. https://doi.org/10.1080/10408398.2017.1421900
Liu Z, Cichocki N, Hübschmann T et al (2019) Neutral mechanisms and niche differentiation in steady-state insular microbial communities revealed by single cell analysis. Environ Microbiol 21:164–181. https://doi.org/10.1111/1462-2920.14437
Firrman J, Liu L, Mahalak K et al (2021) Comparative analysis of the gut microbiota cultured in vitro using a single colon versus a 3-stage colon experimental design. Appl Microbiol Biotechnol 105:3353–3367. https://doi.org/10.1007/S00253-021-11241-X
Blanquet-Diot S, Denis S, Chalancon S et al (2012) Use of artificial digestive systems to investigate the biopharmaceutical factors influencing the survival of probiotic yeast during gastrointestinal transit in humans. Pharm Res 29:1444–1453. https://doi.org/10.1007/S11095-011-0620-5
Cordonnier C, Thévenot J, Etienne-Mesmin L et al (2015) Dynamic in vitro models of the human gastrointestinal tract as relevant tools to assess the survival of probiotic strains and their interactions with gut microbiota. Microorganisms 3:725. https://doi.org/10.3390/MICROORGANISMS3040725
Thévenot J, Etienne-Mesmin L, Denis S et al (2013) Enterohemorrhagic Escherichia coli O157:H7 survival in an in vitro model of the human large intestine and interactions with probiotic yeasts and resident microbiota. Appl Environ Microbiol 79:1058–1064. https://doi.org/10.1128/AEM.03303-12
Thévenot J, Cordonnier C, Rougeron A et al (2015) Enterohemorrhagic Escherichia coli infection has donor-dependent effect on human gut microbiota and may be antagonized by probiotic yeast during interaction with Peyer’s patches. Appl Microbiol Biotechnol 99:9097–9110. https://doi.org/10.1007/S00253-015-6704-0
Gibson GR, Cummings JH, Macfarlane GT (1988) Use of a three-stage continuous culture system to study the effect of mucin on dissimilatory sulfate reduction and methanogenesis by mixed populations of human gut bacteria. Appl Environ Microbiol 54:2750–2755. https://doi.org/10.1128/aem.54.11.2750-2755.1988
Costabile A, Walton GE, Tzortzis G et al (2015) Effects of orange juice formulation on prebiotic functionality using an in vitro colonic model system. PLoS ONE 10:e0121955. https://doi.org/10.1371/journal.pone.0121955
Healey G, Murphy R, Butts C et al (2017) Variability in gut microbiota response to an inulin-type fructan prebiotic within an in vitro three-stage continuous colonic model system. Bioact Carbohydrates Diet Fibre 11:26–37. https://doi.org/10.1016/j.bcdf.2017.07.001
Minekus M, Marteau P, Havenaar R et al (1995) A multicompartmental dynamic computer-controlled model simulating the stomach and small-intestine. Atla-Altern Lab Anim 23(2):197–209. https://doi.org/10.1177/026119299502300205
Verwei M, Minekus M, Zeijdner E et al (2016) Evaluation of two dynamic in vitro models simulating fasted and fed state conditions in the upper gastrointestinal tract (TIM-1 and tiny-TIM) for investigating the bioaccessibility of pharmaceutical compounds from oral dosage forms. Int J Pharm 498:178–186. https://doi.org/10.1016/J.IJPHARM.2015.11.048
Bellmann S, Lelieveld J, Gorissen T et al (2016) Development of an advanced in vitro model of the stomach and its evaluation versus human gastric physiology. Food Res Int 88:191–198. https://doi.org/10.1016/J.FOODRES.2016.01.030
Venema K (2015) The TNO in vitro model of the colon (TIM-2). In: Verhoeckx K, Cotter P, López-Expósito I, et al (Eds.), The impact of food bioactives on health. Springer International Publishing, Cham, pp 293–304. https://doi.org/10.1007/978-3-319-16104-4_26
Cieplak T, Wiese M, Nielsen S et al (2018) The smallest intestine (TSI)—a low volume in vitro model of the small intestine with increased throughput. Fems Microbiol Lett 365(21):231. https://doi.org/10.1093/FEMSLE/FNY231
Macfarlane S, Dillon J (2007) Microbial biofilms in the human gastrointestinal tract. J Appl Microbiol 102:1187–1196. https://doi.org/10.1111/J.1365-2672.2007.03287.X
Zihler A, Gagnon M, Chassard C et al (2011) Protective effect of probiotics on Salmonella infectivity assessed with combined in vitro gut fermentation-cellular models. BMC Microbiol 11:264. https://doi.org/10.1186/1471-2180-11-264
Li S, Abdulkadir N, Schattenberg F et al (2022) Stabilizing microbial communities by looped mass transfer. Proc Natl Acad Sci 119:e2117814119. https://doi.org/10.1073/pnas.2117814119
Thuenemann EC, Mandalari G, Rich GT et al (2015) Dynamic gastric model (DGM). In: Verhoeckx K, Cotter P, López-Expósito I, et al (Eds.), The impact of food bioactives on health. Springer International Publishing, Cham, pp 47–59. https://doi.org/10.1007/978-3-319-16104-4_6
Vardakou M, Mercuri A, Barker SA et al (2011) Achieving antral grinding forces in biorelevant in vitro models: comparing the USP dissolution apparatus II and the dynamic gastric model with human in vivo data. AAPS PharmSciTech 12(2):620–626. https://doi.org/10.1208/s12249-011-9616-z
Mercuri A, Lo Curto A, Wickham MSJ et al (2008) Dynamic gastric model (DGM): a novel in vitro apparatus to assess the impact of gastric digestion on the droplet size of self-emulsifying drug-delivery systems. J Pharm Pharmacol 60:4
O’donnell MM, Rea MC, Shanahan F et al (2018) The use of a mini-bioreactor fermentation system as a reproducible, high-throughput ex vivo batch model of the distal colon. Front Microbiol 9:1844. https://doi.org/10.3389/fmicb.2018.01844
McDonald JA, Schroeter K, Fuentes S et al (2013) Evaluation of microbial community reproducibility, stability and composition in a human distal gut chemostat model. J Microbiol Methods 95:167–174. https://doi.org/10.1016/J.MIMET.2013.08.008
McDonald JA, Fuentes S, Schroeter K et al (2015) Simulating distal gut mucosal and luminal communities using packed-column biofilm reactors and an in vitro chemostat model. J Microbiol Methods 108:36–44. https://doi.org/10.1016/J.MIMET.2014.11.007
Molly K, Vande Woestyne M, Verstraete W (1993) Development of a 5-step multi-chamber reactor as a simulation of the human intestinal microbial ecosystem. Appl Microbiol Biotechnol 39:254–258. https://doi.org/10.1007/BF00228615
Van den Abbeele P, Grootaert C, Marzorati M et al (2010) Microbial community development in a dynamic gut model is reproducible, colon region specific, and selective for Bacteroidetes and Clostridium cluster IX. Appl Environ Microb 76(15):5237–5246. https://doi.org/10.1128/Aem.00759-10
Van de Wiele T, Van den Abbeele P, Ossieur W et al (2015) The simulator of the human intestinal microbial ecosystem (SHIME®). In: Verhoeckx K, Cotter P, López-Expósito I, et al (Eds.), The impact of food bioactives on health. Springer International Publishing, Cham, pp 305–317. https://doi.org/10.1007/978-3-319-16104-4_27
Wang M, Wichienchot S, He X et al (2019) In vitro colonic fermentation of dietary fibers: fermentation rate, short-chain fatty acid production and changes in microbiota. Trends Food Sci Technol 88:1–9. https://doi.org/10.1016/J.TIFS.2019.03.005
Barroso E, Cueva C, Peláez C et al (2015) Development of human colonic microbiota in the computer-controlled dynamic SIMulator of the gastroIntestinal tract SIMGI. LWT - Food Sci Technol 61:283–289. https://doi.org/10.1016/J.LWT.2014.12.014
Poeker SA, Geirnaert A, Berchtold L et al (2018) Understanding the prebiotic potential of different dietary fibers using an in vitro continuous adult fermentation model (PolyFermS). Sci Rep 8:4318. https://doi.org/10.1038/s41598-018-22438-y
Burmeister A, Grünberger A (2020) Microfluidic cultivation and analysis tools for interaction studies of microbial co-cultures. Curr Opin Biotechnol 62:106–115. https://doi.org/10.1016/j.copbio.2019.09.001
Tan HY, Toh YC (2020) What can microfluidics do for human microbiome research? Biomicrofluidics 14:51303. https://doi.org/10.1063/5.0012185
Trujillo-de Santiago G, Lobo-Zegers MJ, Montes-Fonseca SL et al (2018) Gut-microbiota-on-a-chip: an enabling field for physiological research. Microphysiol Syst 2:7 https://doi.org/10.21037/mps.2018.09.01
von Martels JZH, Sadaghian Sadabad M, Bourgonje AR et al (2017) The role of gut microbiota in health and disease: in vitro modeling of host-microbe interactions at the aerobe-anaerobe interphase of the human gut. Anaerobe 44:3–12. https://doi.org/10.1016/j.anaerobe.2017.01.001
Roupar D, Berni P, Martins JT et al (2021) Bioengineering approaches to simulate human colon microbiome ecosystem. Trends Food Sci Technol 112:808–822. https://doi.org/10.1016/j.tifs.2021.04.035
Bein A, Shin W, Jalili-Firoozinezhad S et al (2018) Microfluidic organ-on-a-chip models of human intestine. Cell Mol Gastroenterol Hepatol 5(4):659–668. https://doi.org/10.1016/j.jcmgh.2017.12.010
Bartfeld S (2016) Modeling infectious diseases and host-microbe interactions in gastrointestinal organoids. Dev Biol 420:262–270. https://doi.org/10.1016/J.YDBIO.2016.09.014
Maier E, Anderson RC, Altermann E et al (2018) Live Faecalibacterium prausnitzii induces greater TLR2 and TLR2/6 activation than the dead bacterium in an apical anaerobic co-culture system. Cell Microbiol 20(2):e12805. https://doi.org/10.1111/CMI.12805
Sadaghian Sadabad M, von Martels JZH, Khan MT et al (2016) A simple coculture system shows mutualism between anaerobic faecalibacteria and epithelial Caco-2 cells. Sci Rep 5:17906. https://doi.org/10.1038/srep17906
Marzorati M, Vanhoecke B, De Ryck T et al (2014) The HMI™ module: a new tool to study the host-microbiota interaction in the human gastrointestinal tract in vitro. BMC Microbiol 14:133. https://doi.org/10.1186/1471-2180-14-133
Shah P, Fritz JV, Glaab E et al (2016) A microfluidics-based in vitro model of the gastrointestinal human-microbe interface. Nat Commun 7:1–15. https://doi.org/10.1038/ncomms11535
Parlesak A, Haller D, Brinz S et al (2004) Modulation of cytokine release by differentiated CACO-2 cells in a compartmentalized coculture model with mononuclear leucocytes and nonpathogenic bacteria. Scand J Immunol 60(5):477–485. https://doi.org/10.1111/J.0300-9475.2004.01495.X
Ulluwishewa D, Anderson RC, Young W et al (2015) Live Faecalibacterium prausnitzii in an apical anaerobic model of the intestinal epithelial barrier. Cell Microbiol 17(2):226–240. https://doi.org/10.1111/cmi.12360
Ramadan Q, Zourob M (2020) Organ-on-a-chip engineering: toward bridging the gap between lab and industry. Biomicrofluidics 14:041501. https://doi.org/10.1063/5.0011583
Kim HJ, Huh D, Hamilton G et al (2012) Human gut-on-a-chip inhabited by microbial flora that experiences intestinal peristalsis-like motions and flow. Lab Chip 12:2165. https://doi.org/10.1039/c2lc40074j
Jalili-Firoozinezhad S, Gazzaniga FS, Calamari EL et al (2019) A complex human gut microbiome cultured in an anaerobic intestine-on-a-chip. Nat Biomed Eng 3:520–531. https://doi.org/10.1038/s41551-019-0397-0
Pajoumshariati SR, Azizi M, Zhang SY et al (2018) A microfluidic-based model for spatially constrained culture of intestinal microbiota. Adv Funct Mater 28(48):1805568. https://doi.org/10.1002/adfm.201805568
Taeuber S, Schmitz J, Bloebaum L et al (2021) How to perform a microfluidic cultivation experiment-a guideline to success. Biosensors 11(12):485. https://doi.org/10.3390/BIOS11120485
Hassanpourfard M, Ghosh R, Thundat T et al (2016) Dynamics of bacterial streamers induced clogging in microfluidic devices. Lab Chip 16:4091–4096. https://doi.org/10.1039/C6LC01055E
Skelley AM, Voldman J (2008) An active bubble trap and debubbler for microfluidic systems. Lab Chip 8:1733–1737. https://doi.org/10.1039/B807037G
Zheng W, Wang Z, Zhang W et al (2010) A simple PDMS-based microfluidic channel design that removes bubbles for long-term on-chip culture of mammalian cells. Lab Chip 10:2906–2910. https://doi.org/10.1039/C005274D
Bartfeld S, Bayram T, van de Wetering M et al (2015) In vitro expansion of human gastric epithelial stem cells and their responses to bacterial infection. Gastroenterology 148:126-136.e6. https://doi.org/10.1053/J.GASTRO.2014.09.042
McCracken KW, Catá EM, Crawford CM et al (2014) Modelling human development and disease in pluripotent stem-cell-derived gastric organoids. Nature 516:400–404. https://doi.org/10.1038/NATURE13863
Leslie JL, Huang S, Opp JS et al (2015) Persistence and toxin production by Clostridium difficile within human intestinal organoids result in disruption of epithelial paracellular barrier function. Infect Immun 83(1):138–145. https://doi.org/10.1128/Iai.02561-14
Hill DR, Huang S, Nagy MS et al (2017) Bacterial colonization stimulates a complex physiological response in the immature human intestinal epithelium. eLife 6:e29132. https://doi.org/10.7554/ELIFE.29132
Williamson IA, Arnold JW, Samsa LA et al (2018) A high-throughput organoid microinjection platform to study gastrointestinal microbiota and luminal physiology. Cell Mol Gastroenter 6(3):301–319. https://doi.org/10.1016/j.jcmgh.2018.05.004
Co JY, Margalef-Catala M, Li XN et al (2019) Controlling epithelial polarity: a human enteroid model for host-pathogen interactions. Cell Rep 26(9):2509–2520. https://doi.org/10.1016/j.celrep.2019.01.108
Ettayebi K, Crawford SE, Murakami K et al (2016) Replication of human noroviruses in stem cell-derived human enteroids. Science 353:1387–1393. https://doi.org/10.1126/SCIENCE.AAF5211
VanDussen KL, Marinshaw JM, Shaikh N et al (2015) Development of an enhanced human gastrointestinal epithelial culture system to facilitate patient-based assays. Gut 64:911–920. https://doi.org/10.1136/GUTJNL-2013-306651
Kim R, Attayek PJ, Wang Y et al (2019) An in vitro intestinal platform with a self-sustaining oxygen gradient to study the human gut/microbiome interface. Biofabrication 12:015006. https://doi.org/10.1088/1758-5090/ab446e
Fofanova TY, Stewart CJ, Auchtung JM et al (2019) A novel human enteroid-anaerobe co-culture system to study microbial-host interaction under physiological hypoxia. bioRxiv. https://doi.org/10.1101/555755
Eain MMG, Baginska J, Greenhalgh K et al (2017) Engineering solutions for representative models of the gastrointestinal human-microbe interface. Engineering 3(1):60–65. https://doi.org/10.1016/J.Eng.2017.01.011
Acknowledgements
This work was supported by the BIOMEMBRANE project (M-ERA.net 2 project 4246), the KERAPACK project (MANUNET MNET 17/NMAT-0060), the PRA_2018_68 (grant supported by the University of Pisa), and MIT-UNIPI project (grant supported by the University of Pisa and the MIT). The authors acknowledge the support of the Additive Manufacturing Cross-Lab of the Department of Information Engineering of the University of Pisa.
Funding
Open access funding provided by Università di Pisa within the CRUI-CARE Agreement.
Author information
Authors and Affiliations
Contributions
FB and CD were involved in analysis and writing—original draft; MC, CDM, YSZ, EG, and GV contributed to supervision and writing—review and editing. All authors read and approved the final manuscript.
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that there is no conflict of interest.
Ethical approval
This article does not contain any studies with human or animal subjects performed by any of the authors.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Biagini, F., Daddi, C., Calvigioni, M. et al. Designs and methodologies to recreate in vitro human gut microbiota models. Bio-des. Manuf. 6, 298–318 (2023). https://doi.org/10.1007/s42242-022-00210-6
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s42242-022-00210-6