Abstract
Biophysical restrictions regulate protein diffusion, nucleus deformation, and cell migration, which are all universal and important processes for cells to perform their biological functions. However, current technologies addressing these multiscale questions are extremely limited. Herein, through two-photon polymerization (TPP), we present the precise, low-cost, and multiscale microstructures (micro-fences) as a versatile investigating platform. With nanometer-scale printing resolution and multiscale scanning capacity, TPP is capable of generating micro-fences with sizes of 0.5–1000 μm. These micro-fences are utilized as biophysical restrictions to determine the fluidity of supported lipid bilayers (SLB), to investigate the restricted diffusion of Src family kinase protein Lck on SLB, and also to reveal the mechanical bending of cell nucleus and T cell climbing ability. Taken together, the proposed versatile and low-cost micro-fences have great potential in probing the restricted dynamics of molecules, organelles, and cells to understand the basics of physical biology.
Graphic abstract

Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Biophysical restrictions commonly regulate cell functions at molecular (microscale), subcellular (mesoscale) and cellular (macroscale) levels [1,2,3,4]. At the molecular level, such restrictions are frequently formed by protein clusters or cytoskeleton networks or lipid microdomains, regulating membrane protein diffusion to ensure the proper signaling initiation and transduction in the cell [5,6,7]. For example, the Src family kinase Lck is a key membrane-anchored kinase for triggering T cell receptor (TCR) signaling and activation of T cells [6, 8,9,10]. The diffusion of Lck is not only restricted by two-dimensional (2D) cell membrane [11, 12] but also by various neighboring clustered proteins (e.g., integrin, CD45, and LAT) and cytoskeleton networks [5, 10, 13, 14]. More importantly, the restricted diffusion of Lck on the plasma membrane is essential for Lck to stay with and phosphorylate the TCR-CD3 complex upon TCR activation [15]. At the subcellular level, biophysical restrictions are usually formed by the extracellular matrix or intracellular cytoskeleton networks, which can mechanically deform the cell nucleus (especially bending deformation), and also regulate mitochondrial fission [2, 16, 17]. Although the compressed deformation of cell nucleus has been reported to affect gene transcription, epigenetic modification, DNA damage and repair [18,19,20,21], the mechanism of bending deformation regulating those activities is still unclear. At the cellular level, biophysical restrictions are commonly formed by cell aggregation or cell–matrix agglutination (e.g., platelet, red blood cells and collagen form thrombosis), or cell–cell tight junctions (e.g., endothelial cells in blood vessel tightly connect with each other), and they have also been reported to hinder cell migration [3, 22, 23]. For example, T cells always migrate through extracellular matrix networks to search for targeted cancer cells and initiate their adaptive immune response [24], but it is still unclear how extracellular biophysical restrictions and cell signaling couple together to regulate T cell migration.
Answering the aforementioned multiscale questions is essential to understand the basic physical biology of a cell. Photoetching and imprinting methods have been utilized to manufacture micro-patterns (e.g., micro-grids, pillars, and scaffolds) to investigate how these biophysical patterns and structures affect membrane protein signal transduction [25,26,27,28], cell morphology [2, 29], and cell migration [3, 22], but these methods are extremely limited by resolution and working range, and are costly and tedious. Therefore, to match the multiscale cellular dynamic investigation, an effective fabricating method with the resolution down to the nanometer scale and whole working range up to several hundred micrometers is urgently desired. Due to the advantages of high spatial resolution and true three-dimensional (3D) direct writing ability, TPP based on femtosecond manufacturing laser is regarded as the preferred option. The working mechanism behind TPP is the nonlinear optical absorption that limits the polymerization reaction inside the focal zone. Thus, the printing resolution can reach 200 nm with a high numerical aperture objective, and 3D direct writing can be achieved by the relative movement of the focal spot inside the photoresist. Compared with the photoetching and imprinting methods, specified masks and molds are not essential for TPP, which brings about a much more flexible fabrication method [30, 31].
In this work, we applied TPP to directly print low-cost and versatile micro-fences for probing the multiscale restricted cellular dynamics. By using a pair of galvo mirrors, a piezo nano-positioning stage, and a large translation platform, the laser printing system based on TPP demonstrated the feature size of 0.5 μm and a working range up to 1000 μm. The TPP-manufactured micro-fences possessed stable and coherent boundaries as biophysical restrictions, which perfectly separated supported lipid bilayers (SLB) for lipid fluidity quantification and the high-throughput real-time tracking of the restricted diffusion of Lck on SLB. More importantly, TPP expanded the micro-fences to probe the cell nucleus mechanical bending and cell climbing ability in response to 3D biophysical restrictions.
Materials and methods
Materials
TPP and EBL
Coverslip (Φ18, Thermo Fisher Scientific, USA), O2 plasma cleaner (Diener, Germany), 3-(methacryloyloxy)propyltrimethoxysilane (MPS, 97%, Alfa Aesar, China), photoresist for TPP (IP-L 780, Nanoscribe GmbH, Germany), TPP software (Describe, Nanoscribe GmbH, Germany), oil immersion objective (NA = 1.4, 63 × , Zeiss, Germany), propylene glycol methyl ether acetate (PGMEA, 99%, Aladdin, China), isopropyl alcohol (Sinopharm, China), E-beam photoresist for EBL (AR-P 6200.09, ALLRESIST, Germany), conductive polymer (AR-PC 5090.02, ALLRESIST, Germany), E-beam lithography (EBPG 5150, Raith BV, Germany), developer solvent for EBL (AR 600–546, ALLRESIST, Germany), stopper solvent for EBL (AR 600–60, ALLRESIST, Germany), Multi-chamber Vacuum Coating System (ASB-EPI-C6, Syskey Technology, China), DEKTAK XT Stylus Profiler (DXT-18–1935, Bruker, Malaysia).
SLB and protein
1-palmitoyl-2-oleoyl-glycero-3-phosphocholine (POPC, Avanti Polar Lipids, USA), 1,2-dioleoyl-sn-glycero-3-[(N-(5-amino-1-carboxypentyl)iminodiacetic acid)succinyl] (nickel salt) (DGS-NTA(Ni), Avanti Polar Lipids, USA), PBS (Sangon Biotech, China), Octadecyl Rhodamine B Chloride (R18, Thermo Fisher Scientific, USA), fluorescence microscope (Ti-E, Nikon, Japan), polymerase chain reaction kit (Vazyme, China), HisSep Ni-NTA Agarose Resin 6FF (Yeasen, China), HiTrap Q HP column (GE Healthcare, USA), gel filtration (Superdex 75 10/300 GL, GE Healthcare, USA), SiO2 bead (Thermo Fisher Scientific, USA), Flow Cytometry (Beckman, USA), BSA (LEE BIOSOLUTIONS, USA), C5-Maleimide-Alexa Fluor 488 (Thermo Fisher Scientific, USA), zeba spin desalting column (Thermo Fisher Scientific, USA).
Nucleus bending and cell climbing
Paraformaldehyde (Aladdin, China), Triton X-100 (Sangon Biotech, China), DAPI (Thermo Fisher Scientific, USA), Cytochalasin D (GlpBio, USA), DMEM culture medium (Basal Media, China), FBS (Yeasen, China), PS (Basal Media, China), trypsin (Yeasen, China), 1640 culture medium (Basal Media, China), microscope (Ti-E, Nikon, Japan), camera (iXon Ultra 897, Andor, UK).
Methods
Manufacture of micro-fences by two-photon polymerization (TPP)
A piece of coverslip was sequentially immersed in acetone, isopropanol and DI water at room temperature (RT) for 5 min of sonication in individual steps to remove organic matters and particulates on its surface. The coverslip was dried up and further cleaned with O2 Plasma cleaner for 5 min followed by immersion in MPS ethanol solution (2%, v/v) for 30 min to enhance the adhesion of micro-fences onto the coverslip surface, and then dried up again with N2. Subsequently, a droplet of the photoresist was cast on the coverslip, and 780 nm femtosecond laser pulses with a repetition rate of 80 MHz were focused on the droplet through the oil immersion objective. The laser was controlled to move along the path based on the designed and edited scanning path file for micro-fences. The laser power and scan speed were set at 50 mW and 1000 μm/s, respectively. The time for TPP manufacturing depends on the micro-fence design; in the current study, normally 12 min was enough for printing micro-fences on a piece of coverslip. The coverslip was then developed in the PGMEA for 5 min and finally rinsed in isopropyl alcohol for 5 min to develop the micro-fences. The size and height of micro-fences were verified through a scanning electron microscope (SEM) and a step profiler.
To achieve a smooth structure with the designed size, the slicing and hatching distances were set at 0.5 and 0.15 μm, respectively. For the micro-fences that were beyond the range of the galvo scanning mode under the 63 × objective, the block size was 150 μm, and the overlap was 1 μm.
Manufacture of micro-fences by E-beam lithography (EBL)
To manufacture micro-fences through EBL, a piece of coverslip was first immersed sequentially in acetone, isopropanol and DI water at RT for 5 min with sonication in each immersion step to remove organic matters and particulates from the surface. The coverslip was thoroughly dried up by baking it on a hot plate at 140 °C for 1 min to prevent any surface moisture from interfering resist adhesion in the subsequent steps. Secondly, the E-beam photoresist (AR-P 6200.09) was dropped onto the coverslip, and spin-coating was performed at RT for 2 min at 4000 rpm to obtain an about 200 nm layer on the coverslip surface. Thirdly, the coverslip was baked at 150 °C for 1 min to remove any extra solvent. To dissipate the surface charge during E-beam exposure, a layer of conductive polymer was further spin-coated onto the surface for 2 min at 4000 rpm, and the coverslip was baked at 90 °C for 1 min to remove any extra solvent. Fourthly, the coverslip was exposed to E-beam lithography with 50 kV acceleration voltage at 1500 µC/cm2 exposure dose according to the designed pattern for micro-fences, the coverslip was rinsed with DI water for 1 min to remove the conductive polymer layer, the exposed resist was developed with the developer solvent (AR 600–546) for 2 min, the developing process was stopped by the stopper solvent (AR 600–60) for 30 s, the coverslip was rinsed twice with DI water, and finally dried up with pressurized nitrogen. Fifthly, vacuum pumping was performed for about 170 min to meet the vacuum degree requirement, then a 2 nm chromium (Cr) seed layer and a 8 nm gold layer were evaporated onto the developed coverslip in the multi-chamber vacuum coating system for about 10 min, with the latter step depending on the evaporation rate (0.2 A/s in our set-up). Sixthly, all E-beam resist was removed through incubating the coverslip in acetone for 10 min. It is worthy to note that after the above operations, only the Cr/Au layer was left on the coverslip surface to constitute the micro-fences. The size and height of micro-fences were verified through a SEM and a step profiler.
Comparison of TPP- and EBL-manufactured micro-fences
A comparison of TPP and EBL for manufacturing micro-fences was performed on a piece of coverslip from five aspects (size: 15 μm × 15 μm × 1 μm × 1 μm, 2500 micro-fence units in total), including the number of procedure steps, material types, instruments, financial cost, and time cost (also see Fig. 1d and Table S1). For “instruments,” the prices of certain instruments were markedly different, especially the TPP ($585,460) being much cheaper than the EBL ($2,188,430) system, making it more affordable and practical to many laboratories. EBL also requires special cleaning rooms, making it extremely expensive and not affordable to many research institutes and small laboratories. For “time cost,” the most time in TPP (50 min) was spent on cleaning and modifying the coverslip rather manufacturing micro-fences (12 min), but the most time in EBL was spent on vacuum pumping and evaporating Cr and Au onto micro-fences (170 + 10 min), indicating that TPP was much more time-saving. When performing mass production, the time for manufacturing micro-fences on average would show further sharp decrease. Considering “financial cost,” the usage fees for TPP ($11.68) and EBL ($26.68 and $48.18) were estimated according to the charge standard ($/h) in the TPP and EBL manufacturing center. In addition, $75.92 was the financial costs of Cr and Au. From all the above five aspects, the manufacturing of TPP-based micro-fences was much simpler, more time-saving and less expensive, thus it possesses several advantages over EBL.
Micro-fences for the investigation of multiscale cellular dynamics. a Chemical information and schematic diagram of steps of manufacturing micro-fences by TPP. b Multiscale micro-fences. c Schematic diagram of steps of manufacturing micro-fences by EBL. d Comparison of TPP and EBL methods. e Multiscale molecular and cellular dynamics investigation by using micro-fences
Supported lipid bilayer (SLB) preparation
To prepare the SLB, 2 μmol of lipid (90% POPC and 10% DGS-NTA(Ni) by molar ratio or 100% POPC) was mixed in 1 mL of chloroform in a pre-cleaned glass tube, the lipid was dried up by steady-flowing N2 at 37 °C for at least 5 min, and the dried lipid was kept under vacuum for 2 h to remove the extra chloroform. Then, 500 μL of PBS buffer (pH 7.4) was added to redissolve the lipid by vortexing for 1 min. The lipid was frozen in liquid nitrogen for 1 min and then immediately thawed in a 37 °C water bath for 1 min. This freeze-thawing process was repeated 30 times to make the liposome solution clear, thus could generate uniform liposomes. The liposome solution was stored at − 80 °C for more than 24 h before the fluorescence recovery after photo-bleaching (FRAP) experiment. It should be noted that the glass tube had to be carefully pre-cleaned and buffer must be pre-filtered, as this was highly critical for preparing suitable liposomes.
Diffusion coefficient quantification for SLB and Lck
After repeating the freeze-thawing process, the liposome solution was centrifuged at 15,000 g for 40 min, and the supernatant containing large unilamellar vesicles (LUV), which are commonly below 500 nm in size, was carefully dissociated from the sediment that was full of cracked liposomes [32, 33]. LUV solution (10 μL) was diluted into 200 μL of PBS buffer, the buffer was spread onto the plasma pre-cleaned coverslip with manufactured micro-fences for 30 min at RT to form SLB [34, 35], gently washed the SLB twice, and the SLB was stained with R18 dye for 3 min followed by washing with PBS twice to remove any extra R18. Next, FRAP experiment was performed to measure the diffusion coefficients of lipid. After focusing the R18-stained SLB in micro-fences, the selected region of interest (ROI) was photo-bleached with 561 nm laser (20 mW) for 15 s. After photo-bleaching, the fluorescence of ROI was immediately recorded every 5 or 10 s until the fluorescence intensity in ROI no longer recovered. Based on the FRAP curve, the diffusion coefficient (D, unit: μm2/s) was calculated by the following equation [36]:
where r represents the radius of ROI (μm) and τD represents the half-recovery time (s) from the FRAP curve.
To calculate the diffusion coefficient (D) of Lck on SLB, the following equation was used [15]:
where MSD represents the mead square displacement of Lck (μm2) and t denotes the time (s).
If the molecule did not touch the micro-fences boundary, we defined it as non-restricted diffusion. If the molecule touched and got stuck to the micro-fence boundary, we defined it as restricted diffusion.
Construction, expression, purification of Lck and ICAM-1, and labeling of Lck
The Lck construct was obtained by inserting full length human Lck cDNA (amino acid sequence: G3-P509) flanked with a C-terminal 10 × His tag into a pHAGE vector with PCR kit. The Lck-expression HEK 293F cell line was built by introducing Lck gene-contained lentivirus into cells. HEK 293F cells were cultured with SMM 293-Ti medium containing 1% PS for 5–7 days to express the Lck protein. The 293F cells were then lysed with NP40 solution, Lck was enriched by HisSep Ni–NTA Agarose Resin 6FF from 293F cell lysate, and purified by HiTrap Q HP column followed by gel filtration for final purification. The purity of Lck protein was verified by SDS-PAGE. The ICAM-1-Fc construct was obtained by inserting the extracellular domain of human ICAM-1 cDNA (amino acid sequence: Q28-E480) followed by the human IgG1 Fc region into a pHAGE vector with a PCR kit. The ICAM-1-Fc construct was transfected into 500 mL of HEK 293F cells in SMM 293-Ti medium with PEI. Five days later, the medium was harvested and concentrated. The ICAM-1-Fc protein was enriched by protein G beads and eluted by elution buffer (Glycine–HCl, pH 3.0). The eluted ICAM-1-Fc protein was neutralized with neutralization buffer (Tris–HCl, pH 9.5) and exchanged into PBS for storage. The purity of ICAM-1-Fc was verified by SDS-PAGE.
To label Lck with fluorescence dye, the Lck solution was first changed into the labeling buffer (1 × PBS, pH 7.4, 1 mM TCEP) and then incubated with C5-Maleimide-Alexa Fluor 488 (Lck-488, 1 mM) in a rotation mixer in the dark at RT for 2 h. The molar ratio of Lck and dye was controlled to be 1:10 to prevent too much dye labeling from changing the properties of Lck. After incubation, the Lck-488 solution was passed through a zeba spin desalting column to remove the unlabeled dye, and the final Lck-488 solution was stored at − 80 °C before further use.
Detection of Lck-488 on lipid bilayers by flow cytometry
SiO2 beads (10 μL, 108/mL) were added into NTA(Ni) or PC liposome solution, the bead solution was shocked at RT for 1 min and rotated at 37 °C for 30 min. Next, beads were gently washed (< 3000 rpm) twice with PBS buffer to avoid damage to lipid bilayers. After lipid bilayers were successfully coated onto the bead surface, these beads were incubated with 1% BSA solution for 30 min to block the potential non-specific interaction between proteins and lipid bilayers. Then, 1 μg of Lck-488 was incubated with NTA(Ni) or PC lipid-coated beads in 1% BSA solution while rotating for 30 min, washed twice, and detected the attachment of Lck-488 to lipid bilayer-coated beads through flow cytometry.
U2OS cell nucleus staining and imaging
The coverslip with micro-fences was packed into a chamber and immersed in 75% alcohol for 5 min, cleaned with O2 plasma cleaner for 2 min, and the chamber was then kept under UV light at least for 4 h. The U2OS cells were digested with trypsin, and an appropriate number of cells were added into the chamber, allowing cells to drop onto the micro-fences. Then, cells were cultured in DMEM medium with 10% FBS and 1% PS at 37 °C overnight. Cells were then fixed with 4% paraformaldehyde in PBS solution for 20 min at RT, and washed with PBS three times. Subsequently, cells were slotted with 0.2% TritonX-100 for 20 min, and then blocked with 0.1% TritonX-100, 5% FBS solution for 1 h at RT. Then, cells were incubated with 10 μg/mL DAPI for 20 min, washed, and imaged under confocal microscope in Z-stack scanning mode with a step size of 0.15–0.5 μm. For the inhibition experiments, cells were incubated with 1 μM cytochalasin D for 1 h before fixation.
Jurkat T cells climbing across the micro-fences
The coverslip manufactured with micro-fences was packed into a chamber and immersed in 75% alcohol for 5 min, and then cleaned with O2 plasma cleaner for 2 min. Jurkat T cells were added into the chamber and let most of the cells drop down into the micro-fences. After no more floating cells were seen, cell imaging was performed every 15 s for a total of 30 min. For experiments with ICAM-1, 10 μg/mL of ICAM-1-Fc was incubated on the coverslip before cell addition for 1 h at 37 °C. For the cytochalasin D inhibition experiments, cells were incubated with 1 μM cytochalasin D during imaging. The imaging buffer contained DMEM medium, 10% FBS, and 1% PS. The ICAM-1 experimental buffer contained additional 1.5 mM MgCl2 and 1.5 mM CaCl2. To measure the time for a Jurkat T cell to migrate across the micro-fences, the start time point was the moment when the cell started to touch the micro-fence boundary, and the end time point was the moment when the whole body of cell migrated across the micro-fence boundary. The migrating percentage was calculated based on the number of migrating cells and total cells in the micro-fences within 30 min.
Results and discussion
Advantages and versatility of TPP-manufactured micro-fences
The main principle and chemical information of TPP-manufactured micro-fences are schematically illustrated in Fig. 1a. The coverslip was first functionalized with the reagent “MPS” (3-(methacryloyloxy)propyltrimethoxysilane) as a basal layer. Free radicals were generated from the photoinitiator in the focal spot of the femtosecond laser, and these free radicals attacked the carbon–carbon double bonds (C = C) of the acrylate groups of the monomer in the negative photoresist IP-L 780 (main component: 2-(Hydroxymethyl)-2-[[(1-oxoallyl)oxy]methyl]-1,3-propanediyl diacrylate, > 95%), which formed carbon–carbon single bonds (C–C) to propagate the radical reaction by linking the polymer chain. With the galvo scanning in the X–Y plane and piezoelectric displacement in the Z direction, the multiple polymerized voxel along the scanning path can produce a continuous structure with a coalescence effect. To extend the working range up to 1000 μm, a large translation platform was used to shift the fabrication area and assemble the fabrication blocks. As shown in Fig. 1b, the generated micro-fence array had a height and width of 1 μm, a side length of 15 μm, and a whole range of 450 μm, providing promising applications for the investigation of multiscale restricted cellular dynamics, as described in Fig. 1e. The photoresists used in the study were autofluorescent, with an emission spectrum from 450 to 600 nm [37], which is useful to acquire structures under microscopy.
To demonstrate the advantages (i.e., manufacturing efficiency and costs) of using TPP to manufacture micro-fences, we compared it to the most popular EBL technique (Fig. 1c). We utilized a scanning electronic microscope (SEM), an optical microscope, and a step profiler to verify that both methods could manufacture the desired micro-fences (Fig. S1). Considering the required procedure steps, material types, instruments, financial cost, and time cost, the TPP method performed much better than the EBL method (Fig. 1d and Table S1). Specifically, the TPP method to manufacture one sample of micro-fences required 72 min and cost 18.97 dollars, which is only 1/3 of the time and 1/9 of the financial requirement of the EBL method (Table S1 and Fig. 1d). In massive production, the manufacturing time and financial costs of a sample fabricated by the TPP method would be further sharply reduced to 15 min and 12 dollars. However, EBL also possesses the ability to process multiple samples at the same time, which could potentially reduce the time cost; however, considering the process of vacuum pumping (Table S1), the EBL technique needs more time for each coverslip with micro-fences.
Furthermore, due to the long manufacturing time and financial burden of the EBL method, the height of the micro-fences was limited to 10 nm, which is only slightly higher than the thickness of the cell plasma membrane (5 nm). Therefore, with these EBL-manufactured micro-fences, we were unable to investigate the micron-level questions, such as organelle deformation and cell climbing. In contrast, TPP possessed a micron-level manufacturing capacity for both 2D and 3D investigation (Fig. 1e). Taken together, the TPP method has obvious advantages over the EBL method concerning the number of required procedure steps, material types, instruments, financial and time cost, as well as potential biological applications. More importantly, the TPP method also presents a much lower technical barrier to most laboratories and beginners without micro-fabrication access or experience.
Quantification of SLB fluidity and investigation of restricted Lck diffusion on SLB
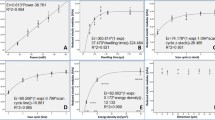
We utilized TPP-manufactured micro-fences to quantify the lipid fluidity, and compared their performance with those produced by the EBL method. We first reconstituted liposomes in vitro [34, 35], and spread the liposome solution into micro-fences to form SLB (Fig. 2a) [27]. As expected, micro-fences successfully separated SLB into dozens of units (Fig. 2b). Next, we simultaneously bleached two regions of interest (ROIs), which, respectively, covered a total micro-fence unit (ROI-2) and an area out of micro-fences (ROI-1) (Fig. 2b). After 90 s, the bleached fluorescence intensity in ROI-1 recovered to almost 100%. However, the fluorescence intensity in ROI-2 remained unrecovered even after 120 s (Figs. 2b and 2c). The diffusion coefficient of lipid in ROI-1 was determined to be 0.56 μm2/s (Fig. 2c), which was comparable with previously reported ones [38, 39]. The diffusion coefficient was 0 μm2/s in ROI-2 (Fig. 2c), as every single micro-fence unit separated SLB well and thus completely blocked lipid exchange between the neighboring micro-fences. To experimentally further support this blocking effect and test whether fence boundaries would impair lipid diffusion or not, we bleached the SLB covering a quarter of the area of a micro-fence unit (Fig. 2d), and the lipid diffusion coefficient in micro-fences was found to be comparable with that out of micro-fences (0.50 vs. 0.56 μm2/s) (Fig. 2e). Moreover, SLB could be well separated by different micro-fences, and be photo-bleached with patterns in a long duration (Fig. S2), suggesting a much more widespread application of micro-fences for probing different lipid-based biophysical phenomena. Thus far, the precise and low-cost micro-fences have ultra-stable performance in separating SLB into parallel units without damaging lipid fluidity, which performed better than the EBL method, as in some reported cases, the boundaries of EBL-manufactured micro-fences were occasionally broken and failed to prevent lipid exchange over a long period [40].
SLB preparation and quantification of its fluidity. a Schematic for preparing SLB and spreading SLB into micro-fences. b, c Photo-bleaching of SLB out of (ROI-1, blue) and inside of (ROI-2, green) a micro-fence unit (b) and their quantifications of fluidity (c). d, e Photo-bleaching of SLB covering a quarter of a micro-fence unit (ROI, black) (d) and the quantification of its fluidity (e). The boundary of micro-fences is marked in b and d. The calculated diffusion coefficients of SLB are indicated in c and e. f Schematic of micro-fences blocking lipid diffusion
Since micro-fences can restrict the lipids within each unit, we next applied them to investigate the restricted Lck diffusion on SLB. First, we purified human wild-type Lck with a 10 × His tag on its N-terminal, labeled it with maleimide-Alexa Fluor 488 (named Lck-488 thereafter) (Fig. 3a and Fig. S3), and verified its specific attachment to NAT(Ni) (the binding affinity between His tag and NTA(Ni) is high, KD = 10−4 nM) [41], but not to PC lipid, through flow cytometry (Fig. 3b). Then, we incubated Lck-488 separately either onto NTA(Ni) or PC SLB in micro-fences (Fig. 3c). The statistically analysis proved that the attachment of Lck-488 to NTA(Ni) was significantly different (****P < 0.0001) from that to PC SLB (Figs. 3d and 3e), which was consistent with the flow cytometry result. By this method, we could observe that Lck-488 formed micro-clusters and tracked their diffusion on SLB within a micro-fence unit (Figs. 3f and 3g). Lck contains multiple domains interacting with itself (Fig. 3a) and neighboring Lck molecules, which may account for the forming of micro-clusters by cis interactions. We observed that Lck-488 diffusion in the area away from the micro-fence boundary (distance > 0.5 μm) was much faster than that near the micro-fence boundary (distance < 0.5 μm), suggesting that the biophysical restrictions of micro-fences largely restrict the diffusion of Lck. This may be because biophysical restrictions change the direction of molecular movement and reduce its kinetic energy (Fig. 3h). In cells, these restrictions can constitute the polymerized F-actin networks, superclustered molecules, and so on [5, 13]. The mechanism of how these restrictions regulate the function of Lck will be further investigated. With this newly developed method, apart from Lck, we can also quantitatively determine the restricted dynamics of many other membrane proteins to understand their biophysical functions on the lipid membrane (e.g., TCR triggering [8] and phase separation [42]).
Real-time tracking of the restricted localization of Lck on SLB. a Schematic of wild-type Lck protein and the labeling. b Verification of the attachment of Lck-488 onto NTA(Ni) lipid but not onto PC lipid. c-e Schematic of incubating Lck-488 onto PC or NTA(Ni) SLB in micro-fences (c), the related representative fluorescence images in micro-fences (d), and the corresponding time-dependent mean fluorescence intensity inside of micro-fences (e). f Real-time tracking of the localization of a Lck-488 micro-cluster within a micro-fence unit. g Diffusion pathways and velocities of the Lck-488 on NTA(Ni) lipid, where the related non-restricted and restricted diffusion coefficients are indicated. h Schematic of the diffusion of Lck on SLB restricted by micro-fences. Error bars from 40 or 60 independent fence units represent Mean ± SEM, ****P < 0.0001 by unpaired Student’s t test in e
Investigation of cell nucleus mechanical bending
Beyond 2D patterning, TPP possesses a 3D structure manufacturing capacity; therefore, we extended the application of micro-fences to probe the nucleus mechanical bending in response to biophysical restrictions during cell adhesion. First, we cultured U2OS cells onto micro-fences and stained the cell nucleus (Fig. 4a), and found that the cell nucleus slightly bent onto the micro-fences of 1 μm height (green arrows, sectional view). However, for 5 μm height, the nucleus bent much more significantly (0.28 vs. 0.1 μm−1, ****P < 0.0001), clearly demonstrating an arch structure over the micro-fences (Figs. 4b and 4c). It has been reported that F-actin is vital for the cell’s mechanical sensing and its maintenance [43, 44], but whether it regulates the nucleus mechanical bending or not is still unclear. Here, we treated cells with cytochalasin D (Cyto. D) to inhibit F-actin polymerization, and revealed that the nucleus mechanical bending was largely abolished (0.06 vs. 0.28 μm−1, ****P < 0.0001) (Figs. 4b and 4c, and Fig. S4), indicating that the intact F-actin network regulates the nucleus mechanical bending when cells adhere on and adapt to 3D biophysical restrictions (Fig. 4d). Therefore, we speculate that DNA damage might be inevitable inside the cell nucleus, and gene transcription and translation might also be severely altered. What is more, concerning the TPP method, some previously published papers have reported that different 3D scaffolds regulate the nucleus shapes in neuron-like cells [45], bone-like cells [46], and mesenchymal stem cells [47], which not only support our observed results, but also elicit that nucleus mechanical deformation in cells is universal when they respond to physical micro-environments. Nonetheless, the molecular mechanism by which cells mechanically deform the nucleus needs further elucidation. Besides the cell nucleus, other organelles (e.g., mitochondria) can be mechanically deformed or repositioned in response to 3D biophysical restrictions [16]. It will be meaningful to reveal the unexplored organelle mechanobiology using multiple methods.
U2OS cell nucleus mechanical bending. a Schematic of the experiment. b Confocal Z-stack images of U2OS cells on micro-fences with different heights. The green arrows indicate the nucleus mechanical bending, and section views indicate the overall perspectives of nucleus bending. c Bending curvature’s definition and quantification. d Schematic of nucleus mechanical bending on micro-fences. The error bars represent Mean ± SEM from at least 15 cells in three replicates, ****P < 0.0001 by unpaired Student’s t test in c
Investigation of the T cell climbing ability across the restrictions
Other than impacting the cell nucleus mechanical bending process, the mechanism of how 3D biophysical restrictions affect the climbing ability of a cell is a hardly touched subject. To investigate this question, we further extended the application of micro-fences to the single-cell level. Although Jurkat T cells can freely migrate on a flat surface [48], we observed that 3D biophysical restrictions (micro-fences) largely suppressed Jurkat T cells’ climbing ability by increasing their migration time (Figs. 5a and 5c). Specifically, with the increasing height of micro-fences (from 0.5 μm to 1, 2, 3 and 5 μm), the time for a Jurkat T cell to climb across the micro-fences significantly increased (Figs. 5a and 5c, **P < 0.01 for 2 μm, ****P < 0.0001 for 3 and 5 μm) in the absence of cell-adhesion molecule (ICAM-1) on the substrate. However, when ICAM-1 (Fig. S3) was coated on the substrate, Jurkat T cells climbed across higher micro-fences faster. Thus, the time for a Jurkat T cell to climb across the higher micro-fences did not increase (Figs. 5b and 5d), showing no significance among all heights (n.s., P > 0.05), which was totally different from the group without ICAM-1 (Figs. 5a and 5c). On the other hand, the percentage of Jurkat T cells that were able to climb across the micro-fences was still similar to those without ICAM-1, that is, higher micro-fences block more climbing cells (Fig. 5e). In our opinion, when ICAM-1 is coated onto the coverslip surface, Jurkat T cells are more adhesive and thus have more visible filopodia (Fig. S5a), and low micro-fences cannot prevent Jurkat T cells’ filopodia from touching the microenvironment behind micro-fences, hence most Jurkat T cells can easily migrate across the barriers in a short period of time. However, the filopodia of only few Jurkat T cells can touch the high micro-fences. Once touched, ICAM-1 binding to Jurkat T cells’ integrin provides the biophysical linkage to sustain the mechanical force for Jurkat T cells, helping them quickly climb across micro-fences, thus the climbing time was short, but the climbing percentage was still low (Fig. S5b). Moreover, the climbing ability of Jurkat T cells was totally abolished when F-actin polymerization was inhibited by cyto. D treatment (Fig. 5e and Fig. S5c). Since the de-polymerization of F-actin largely disrupts the adhesion and the mechanical force generation by Jurkat T cells, this completely impairs the Jurkat T cells’ climbing ability. Overall, the results demonstrate that both 3D biophysical restrictions and integrin/F-actin dynamic signaling couple together to regulate Jurkat T cells’ climbing ability (Fig. 5f). As T cells frequently migrate to find the targeted cancer cells, the ability of climbing across biophysical restrictions should be crucial for T cell function. The detailed regulating mechanism will be investigated in the near future.
Jurkat T cells’ climbing ability. a, b Bright-field images of Jurkat T cells climbing across the micro-fences with or without ICAM-1, respectively. c, d Time for climbing with or without ICAM-1. e Percentage of climbing cells under different conditions. f Summary of micro-fences, F-actin polymerization and integrin signal regulating Jurkat T cells’ climbing ability. The error bars represent Mean ± SEM, n.s. represents no significance, and **P < 0.01, ****P < 0.0001 by unpaired Student’s t test in c, d. Climbing cells from three to five replicates are used to calculate the percentage in e
Conclusions
In this paper, we utilized TPP to efficiently manufacture multiscale and low-cost 3D micro-fences with different aspect ratios, which were shown to possess obvious advantages over previous methods in efficiency, cost, and applications. Therefore, the micro-fences enabled us to investigate the dynamics within a cell from molecular to subcellular and cellular levels. With these versatile micro-fences, we quantified the SLB fluidity and tracked the restricted diffusion of clustered Lck on SLB in real time. We further demonstrated the coupling of F-actin polymerization and 3D biophysical restrictions in regulating nucleus mechanical bending during cell adhesion. Lastly, we revealed that biophysical restrictions coupled with integrin/F-actin signaling to regulate Jurkat T cells to climb across 3D restrictions, which might be crucial for the immunological function of T cells. Since the TPP method can effectively manufacture 3D microstructures (i.e., not limited to micro-fences), we believe that it can potentially become a preferred method to investigate many other biophysical and biochemical phenomena.
References
Vogel V, Sheetz M (2006) Local force and geometry sensing regulate cell functions. Nat Rev Mol Cell Biol 7:265–275. https://doi.org/10.1038/nrm1890
Hanson L, Zhao WT, Lou HY et al (2015) Vertical nanopillars for in situ probing of nuclear mechanics in adherent cells. Nat Nanotechnol 10:554–592. https://doi.org/10.1038/Nnano.2015.88
Reversat A, Gaertner F, Merrin J et al (2020) Cellular locomotion using environmental topography. Nature 582:582–585. https://doi.org/10.1038/s41586-020-2283-z
Yu H, Sheetz M (2018) Life at low Reynolds number and the mesoscale leads to stochastic phenomena. The cell as a machine. Cambridge University Press, Cambridge, pp 95–116
Ostrowski PP, Grinstein S, Freeman SA (2016) Diffusion barriers, mechanical forces, and the biophysics of phagocytosis. Dev Cell 38:135–146. https://doi.org/10.1016/j.devcel.2016.06.023
van der Merwe PA, Dushek O (2011) Mechanisms for T cell receptor triggering. Nat Rev Immunol 11:47–55. https://doi.org/10.1038/nri2887
Razvag Y, Neve-Oz Y, Sajman J et al (2018) Nanoscale kinetic segregation of TCR and CD45 in engaged microvilli facilitates early T cell activation. Nat Commun 9:732. https://doi.org/10.1038/s41467-018-03127-w
Courtney AH, Lo WL, Weiss A (2018) TCR signaling: mechanisms of initiation and propagation. Trends Biochem Sci 43:108–123. https://doi.org/10.1016/j.tibs.2017.11.008
Nika K, Soldani C, Salek M et al (2010) Constitutively active Lck kinase in T cells drives antigen receptor signal transduction. Immunity 32:766–777. https://doi.org/10.1016/j.immuni.2010.05.011
Rossy J, Williamson DJ, Gaus K (2012) How does the kinase Lck phosphorylate the T cell receptor? Spatial organization as a regulatory mechanism. Front Immunol 3:167. https://doi.org/10.3389/fimmu.2012.00167
Yurchak LK, Sefton BM (1995) Palmitoylation of either Cys-3 or Cys-5 is required for the biological activity of the Lck tyrosine protein kinase. Mol Cell Biol 15:6914–6922. https://doi.org/10.1128/mcb.15.12.6914
Kim PW, Sun ZY, Blacklow SC et al (2003) A zinc clasp structure tethers Lck to T cell coreceptors CD4 and CD8. Science 301:1725–1728. https://doi.org/10.1126/science.1085643
Freeman SA, Goyette J, Furuya W et al (2016) Integrins form an expanding diffusional barrier that coordinates phagocytosis. Cell 164:128–140. https://doi.org/10.1016/j.cell.2015.11.048
Lo WL, Shah NH, Ahsan N et al (2018) Lck promotes Zap70-dependent LAT phosphorylation by bridging Zap70 to LAT. Nat Immunol 19:733–741. https://doi.org/10.1038/s41590-018-0131-1
Douglass AD, Vale RD (2005) Single-molecule microscopy reveals plasma membrane microdomains created by protein-protein networks that exclude or trap signaling molecules in T cells. Cell 121:937–950. https://doi.org/10.1016/j.cell.2005.04.009
Helle SCJ, Feng Q, Aebersold MJ et al (2017) Mechanical force induces mitochondrial fission. eLife. https://doi.org/10.7554/eLife.30292
Liu XN, Liu RL, Gu YX et al (2017) Nonmonotonic self-deformation of cell nuclei on topological surfaces with micropillar array. ACS Appl Mater Interf 9:18521–18530. https://doi.org/10.1021/acsami.7b04027
Denais C, Lammerding J (2014) Nuclear mechanics in cancer. Adv Exp Med Biol 773:435–470. https://doi.org/10.1007/978-1-4899-8032-8_20
Shivashankar GV (2011) Mechanosignaling to the cell nucleus and gene regulation. Ann Rev Biophys 40:361–378. https://doi.org/10.1146/annurev-biophys-042910-155319
Pfeifer CR, Vashisth M, Xia YT et al (2019) Nuclear failure, DNA damage, and cell cycle disruption after migration through small pores: a brief review. Mech Cell Migration 63:569–577. https://doi.org/10.1042/Ebc20190007
Pfeifer CR, Xia Y, Zhu K et al (2018) Constricted migration increases DNA damage and independently represses cell cycle. Mol Biol Cell 29:1948–1962. https://doi.org/10.1091/mbc.E18-02-0079
Renkawitz J, Kopf A, Stopp J et al (2019) Nuclear positioning facilitates amoeboid migration along the path of least resistance. Nature 568:546–550. https://doi.org/10.1038/s41586-019-1087-5
Hou Y, Carrim N, Wang Y et al (2015) Platelets in hemostasis and thrombosis: novel mechanisms of fibrinogen-independent platelet aggregation and fibronectin-mediated protein wave of hemostasis. J Biomed Res 29:546–574. https://doi.org/10.7555/JBR.29.20150121
Krummel MF, Bartumeus F, Gerard A (2016) T cell migration, search strategies and mechanisms. Nat Rev Immunol 16:193–201. https://doi.org/10.1038/nri.2015.16
Mossman KD, Campi G, Groves JT et al (2005) Altered TCR signaling from geometrically repatterned immunological synapses. Science 310:1191–1193. https://doi.org/10.1126/science.1119238
Salaita K, Nair PM, Petit RS et al (2010) Restriction of receptor movement alters cellular response: physical force sensing by EphA2. Science 327:1380–1385. https://doi.org/10.1126/science.1181729
Nair PM, Salaita K, Petit RS et al (2011) Using patterned supported lipid membranes to investigate the role of receptor organization in intercellular signaling. Nat Protoc 6:523–539. https://doi.org/10.1038/nprot.2011.302
Huang WYC, Alvarez S, Kondo Y et al (2019) A molecular assembly phase transition and kinetic proofreading modulate Ras activation by SOS. Science 363:1098–1103. https://doi.org/10.1126/science.aau5721
Hippler M, Lemma ED, Bertels S et al (2019) 3D scaffolds to study basic cell biology. Adv Mater 31:e1808110. https://doi.org/10.1002/adma.201808110
Ding H, Zhang Q, Gu H et al (2019) Controlled microstructural architectures based on smart fabrication strategies. Adv Funct Mater 30:e1901760. https://doi.org/10.1002/adfm.201901760
Ciuciu AI, Cywinski PJ (2014) Two-photon polymerization of hydrogels-versatile solutions to fabricate well-defined 3D structures. RSC Adv 4:45504–45516. https://doi.org/10.1039/c4ra06892k
Su X, Ditlev JA, Hui E et al (2016) Phase separation of signaling molecules promotes T cell receptor signal transduction. Science 352:595–599. https://doi.org/10.1126/science.aad9964
Sou K, Naito Y, Endo T et al (2003) Effective encapsulation of proteins into size-controlled phospholipid vesicles using freeze-thawing and extrusion. Biotechnol Prog 19:1547–1552. https://doi.org/10.1021/bp0201004
Hui E, Vale RD (2014) In vitro membrane reconstitution of the T-cell receptor proximal signaling network. Nat Struct Mol Biol 21:133–142. https://doi.org/10.1038/nsmb.2762
Ma L, Cai Y, Li Y et al (2017) Single-molecule force spectroscopy of protein-membrane interactions. eLife. https://doi.org/10.7554/eLife.30493
Perez MA, Moriones OH, Bastus NG et al (2019) Mechanomodulation of lipid membranes by weakly aggregating silver nanoparticles. Biochemistry 58:4761–4773. https://doi.org/10.1021/acs.biochem.9b00390
Wollhofen R, Buchegger B, Eder C et al (2017) Functional photoresists for sub-diffraction stimulated emission depletion lithography. Opt Mater Express 7:2538–2559. https://doi.org/10.1364/Ome.7.002538
Almeida PF, Vaz WL, Thompson TE (2005) Lipid diffusion, free area, and molecular dynamics simulations. Biophys J 88:4434–4438. https://doi.org/10.1529/biophysj.105.059766
Machan R, Hof M (2010) Lipid diffusion in planar membranes investigated by fluorescence correlation spectroscopy. BBA-Biomembranes 1798:1377–1391. https://doi.org/10.1016/j.bbamem.2010.02.014
Moulick RG, Panaitov G, Choi SE et al (2018) Patterning artificial lipid bilayer on nanostructured surfaces. Int J Nanomed 13:55–58. https://doi.org/10.2147/IJN.S125005
Bornhorst JA, Falke JJ (2000) Purification of proteins using polyhistidine affinity tags. Method Enzymol 326:245–254. https://doi.org/10.1016/s0076-6879(00)26058-8
Li P, Banjade S, Cheng HC et al (2012) Phase transitions in the assembly of multivalent signalling proteins. Nature 483:336–340. https://doi.org/10.1038/nature10879
Garzon-Coral C, Fantana HA, Howard J (2016) A force-generating machinery maintains the spindle at the cell center during mitosis. Science 352:1124–1127. https://doi.org/10.1126/science.aad9745
Blumenthal D, Burkhardt JK (2020) Multiple actin networks coordinate mechanotransduction at the immunological synapse. J Cell Biol 219(2):e201911058
Marino A, Ciofani G, Filippeschi C et al (2013) Two-photon polymerization of sub-micrometric patterned surfaces: investigation of cell-substrate interactions and improved differentiation of neuron-like cells. ACS Appl Mater Interf 5:13012–13021. https://doi.org/10.1021/am403895k
Marino A, Filippeschi C, Genchi GG et al (2014) The Osteoprint: a bioinspired two-photon polymerized 3-D structure for the enhancement of bone-like cell differentiation. Acta Biomater 10:4304–4313. https://doi.org/10.1016/j.actbio.2014.05.032
Raimondi MT, Eaton SM, Lagana M et al (2013) Three-dimensional structural niches engineered via two-photon laser polymerization promote stem cell homing. Acta Biomater 9:4579–4584. https://doi.org/10.1016/j.actbio.2012.08.022
Kwon KW, Park H, Song KH et al (2012) Nanotopography-guided migration of T cells. J Immunol 189:2266–2273. https://doi.org/10.4049/jimmunol.1102273
Acknowledgements
This work was financially supported by the National Basic Research Program of China (Nos. 2015CB910800 and 2017ZX10203205), the National Natural Science Foundation of China (Nos. 31522021, 31971237, and 61671308), the National Key Research and Development Program of China (No. 2017YFA0700500), and the China Postdoctoral Science Foundation (No. 2021M690617). Authors also appreciated the Medicine-Engineering Cross Fund program in Zhejiang University.
Author information
Authors and Affiliations
Contributions
PYF, HBD, ZCP, ZZG, and WC conceived the project. PYF performed the experiments and analyzed data. HBD, YD, and MXW performed micro-fences manufacturing. XYW performed cell culture and staining. WH and PW purified proteins. PYF, HBD, and WC wrote the manuscript. All authors have given approval to the final version of the manuscript. All authors have read and agreed to the published version of the manuscript.
Corresponding authors
Ethics declarations
Conflict of interest
The authors declare that there is no conflict of interest.
Ethical approval
This article does not contain any studies with human or animal subjects performed by any of the authors.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Fei, P., Ding, H., Duan, Y. et al. Utility of TPP-manufactured biophysical restrictions to probe multiscale cellular dynamics. Bio-des. Manuf. 4, 776–789 (2021). https://doi.org/10.1007/s42242-021-00163-2
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s42242-021-00163-2