Abstract
Although numerous bioaerosol samplers for counting and identifying airborne microorganisms are available, the considerably high purchase and maintenance costs for the sampler often prevent broad monitoring campaigns for occupational or environmental surveillance of bioaerosols. We present here a newly developed simple adapter and filter system (TOP filter system) designed to collect bioaerosol particles from a defined air volume using conventional vacuum cleaners as air pumps. We characterized the physical properties of the system using air flow measurements and validated the biological performance. The culture-based detection capacities for airborne fungal species were compared to a standard impaction sampler (MAS-100 NT) under controlled conditions in a bioaerosol chamber (using Trichoderma spores as the test organism) as well as in the field. In the chamber, an overall equivalent detection capacity between all tested filters was recorded, although a significant underrepresentation of the TOP filter system for Trichoderma spores were seen in comparison to the MAS-100 NT. In a comparative field study (n = 345), the system showed similar biological sampling efficiencies compared to the MAS-100 NT impactor, only the diversity of identified fungal communities was slightly lower on the filters. Thus, the system is suitable for large-scale environmental sampling operations where many samples have to be taken in parallel at a given time at distant locations. This system would allow endeavors such as antibiotics resistance monitoring or hygiene surveys in agricultural or occupational settings.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
1 Introduction
Modern life in industrialized countries dictate that people usually spend more than 80% of their daily time indoors (Bernstein et al. 2008; Klepeis et al. 2001; Normand et al. 2015). Hence, healthy indoor air and monitoring indoor air quality is of great importance and public concern. Exposure to microbial air pollutants present as bioaerosols has been related to a wide range of diseases such as allergy, asthma or respiratory disorder (Dannemiller et al. 2016; Madsen et al. 2016; Nevalainen et al. 2015; Osborne et al. 2015). Increased awareness of such potential adverse health effects caused by airborne microorganisms raised the interest in reliable methods for the characterization and collection of bioaerosols. In addition, fungi and bacteria are prominent pathogens and monitoring of these microbes in aerosols and testing their antibiotic or antimycotic resistance profiles is an important task in public health management (Jones and Brosseau 2015).
Besides air filtration-based sampling, where airborne particles are drawn through a filter holder and captured on filters, impaction and impingement are among the most frequently used and well-established air sampling techniques (Ghosh et al. 2015). Mostly, microbial analysis of bioaerosol loads relies on cultivation-based techniques that provide an estimation of culturable microorganisms expressed as “colony forming units” (CFUs) (Chang et al. 1994; Ghosh et al. 2015). Although culturable and non-culturable microorganisms are usually present in bioaerosols international standards for the detection and enumeration of these organisms usually rely on CFU counting and species determination based on morphological traits. If using a culture-based analysis, it has to be considered that an accurate enumeration of the collected airborne microorganisms strongly depends on parameters such as cultivability and viability of the microbial species and the selection of an appropriate cultivation medium and growth conditions, sampling method and analytical strategy. Furthermore, environmental factors such as relative humidity, UV radiation, and temperature show a significant impact on the survival rate of the microorganisms and, therefore, influence a cultivation-based overall assessment (Juozaitis et al. 1994; Tseng and Li 2005; Zhen et al. 2018). For air filtration-based sampling, desiccation effects are likely to impair the survival of collected microorganisms and might lead to an underestimation of the microbial particles present, particularly of the sensitive ones, such as vegetative cells. (Ghosh et al. 2015; Haig et al. 2016; Wang et al. 2001). Spore forming bacteria and fungi, which are highly resistant to environmental stresses, are more amenable to monitoring with filtration sampling. Only recently, molecular methods such as DNA-based quantification of microbes or quantitative biochemical assays (Kespohl et al. 2013; Zahradnik and Raulf 2019) and fungal-specific enzyme assays (Reeslev et al. 2003) are starting to be commonly used for routine measurements and will be implemented in ISO Standards worldwide (Rylander et al. 2010; Unterwurzacher et al. 2018).
Air filtration is usually easy to accomplish and depending on the reason for sampling, a broad spectrum of filter materials featuring different characteristics and pore sizes are available (Deacon et al. 2009). Commonly used filter types include membrane filters composed of gelatin (Wu et al. 2010), polymer layers and fibrous filters consisting of cellulose, glass or variant copolymers (Burton et al. 2007; Haig et al. 2016; Miaskiewicz-Peska and Lebkowska 2012; Raynor et al. 2011). Compared to impactors, filtration-based samplers cover a wider range of particle sizes with high collection efficiency depending on the pore size of the membrane (Aizenberg et al. 2000). One advantage of air filtration is the compatibility with various analytical techniques including cultivation, DNA-based, biochemical, immunochemical, and gravimetrical analyses. Generally, high volume sampling is desired but particularly for a culture-based analysis, long-term air filtration may also lead to overloading and extended sampling stress (Eduard and Heederik 1998; Wang et al. 2001).
Besides validation of the physical collection efficiency according to directives such as DIN ISO 13205-1 (2014), air samplers are often compared in field studies to characterize their overall performance (An et al. 2004; Frankel et al. 2012b; Wang et al. 2015). Discrepancies between such comparative measurements occur due to the generally low microbial particle concentrations in the environment, particle agglomerations, and the unequal distribution of bioaerosols in the air (Fierer et al. 2008; Ghosh et al. 2015; Nazaroff 2004, 2016). Specific technical limitations of each sampling device might amplify these differences. To compensate for such uncertainties, various bioaerosol chambers have been implemented to enable stabilized sampling condition (Dybwad et al. 2014; Estill et al. 2008; Feather and Chen 2003; Kesavan and Sagripanti 2015; Pogner et al. 2019; Simon and Duquenne 2013; Wang et al. 2001).
In the present study, we validated a novel filter-based sampling system that uses conventional vacuum cleaners as air pumps. The initiative to develop such a system comes from the necessity of parallel air samplings in large sample numbers at many different locations in a short period of time. Typically, such sampling campaigns are performed in hygiene surveys or when antibiotic resistance profiles need to be determined over a wide geographic area at a given time point. For this, many samplers need to be employed at each of the distant locations and it is financially not feasible to equip each of the operators with one of the costly commercial bioaerosol samplers. The system described here is low-cost and easy to handle and would thus be well suited for such large sampling campaigns.
2 Materials and Methods
2.1 Description of the Adaptor and Filter System
The system consists of a cylindrical adaptor with a conical inner wall (see Fig. 1A) that is designed to place the adaptor on the tube of any conventional or industrial vacuum cleaner with varying diameters from 2.600 to 3.873 cm. On top of the adaptor the filter unit is placed. The ring-shaped filter unit carries a grid at the bottom where the filter material is placed and then fixed by a hold-down bracket ring. The filter unit is closed by a cap (not shown) that is placed on top of the filter unit after the sampling has been completed to avoid additional microbial contamination of the filter or loss of samples from the filter surface during handling or transportation. The whole system can be packed after complete mounting in a plastic bag and sterilized by gamma irradiation directly inside the packaging. It thus represents a ready-to-use sterile filter unit that is unpacked at the sampling site and placed at the adaptor for each individual aerosol sampling procedure. Before the first sampling, one filter unit is used to calibrate the vacuum cleaner-filter system combination for air sampling volume per time interval. For this, an anemometer is placed on top of the filter system mounted already with the adaptor and the vacuum cleaner is started with highest power settings. The settings of the vacuum cleaner (if available) should be adjusted to yield a total sampled air volume between 100 and 150 L per minute and sampling times between 1 and 5 min would hence be required, depending on the expected bioaerosol load. After this calibration step, the first and all subsequent samplings can be performed using the same settings. After each individual sampling is completed, the cap is placed and fixed on top of the filter unit which is then put back into the plastic bag being ready for shipment to the laboratory for analysis. For all sampling operations the same adaptor can be used as only the filter units are exchanged after each round of collection.
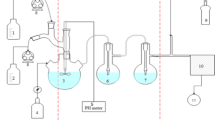
A. Photograph of the adaptor with support grid and ring holding the filters to be inserted. B Schematic representation of the experimental setup for the validation of the volume streams. \(~v_{{{\text{FA}}}}\) velocity at filter inlet, \({\dot{V}}_{FA}\) volume flow, M1 hydrodynamic vane, M2 pressure measurement, M3 diaphragm gas meter, \({\dot{V}}_{FL}\) volume flow due to leakage, \({\dot{V}}_{BGZ}\) total volume flow. For the dimensions of the unit see Supplemental Figure S2
For analysis of the collected airborne material in the laboratory the filter unit is demounted in a sterile hood. For this, the cap is carefully removed from the filter unit and rinsed with buffer to collect the material that has fallen off the filter during transportation and handling (details see below). For CFU-based analysis of filter-captured microbes, the filter is removed from the unit using sterile forceps and directly placed onto suitable growth medium exposing the collection side of the filter to the air. To collect the particles from the cap, the inner side is rinsed thoroughly with 500 µL sterile 1 × phosphate buffered saline (PBS, pH 7.4), 0.01% Tween 20 (Sigma–Aldrich, USA) solution. After gentle shaking for half a minute, this solution is placed directly on top of the filter and distributed over the whole surface of the medium in the petri dish using a Drigalski spatula. This procedure was found to be optimal not only to collect particles from the cap but also to evenly distribute microbial cells from the filter over the whole area of the medium thus preventing growth restriction due to exceedingly high colony densities on the filter area. The adaptor and the filter units with the cap have been fabricated by low density polyethylene SPB608 (Braskem Europe GmbH, Germany) injection molding and can be reused after rinsing in 70% ethanol and washing in a laboratory dish washer. The system has received intellectual property rights protection and is registered as EP2730912A2.
2.2 Experimental Setup and Determination of Airstream Correction Factors
The newly developed system was validated for air stream parameters and possible leakage problems that would allow the air stream passing through the adaptor without passing through the filter. This situation would result in a total air volume measurement by the anemometer that is actually higher than the real air volume passing through the filter. To know the portion of air passing through the system but not through the filter, a leakage analysis was performed and an air volume correction factor was determined. The experimental setup and the main components for these tests are described below and graphically shown in Fig. 1B. The gas velocity at the filter inlet (vFA) was measured with a hydrodynamic vane (M1) model PCE-TA 30 (PCE Instruments, Germany). At the drain of the filter holder, a diaphragm gas meter (M3) model BK-G6 (Elster GmbH, Germany) measured the total volume flow (\({\dot{V}}_{\mathrm{B}\mathrm{G}\mathrm{Z}}\)) including the inlet air by leakage before and after the filter holder. To provide a wide range of different volume flows for each filter, a side channel blower model SKV ND-150-3-935 (SKV-tec GmbH, Germany) was used in vacuum mode. At last, for the determination of an alternative measurement method, a pressure measurement device was placed at the pipe connecter (M2). The determination of the volume flow by means of speed measurement by a hydrometric vane took place via the equation of continuity (1).
\(v_{{FA}}\) denotes the gas velocity measured by hydrometric vane. In Fig. 1B this point is marked by the circle M1. It can be assumed that the volume flow, which is determined by means of the speed of the vane anemometer and the filter surface (A), is higher than the volume flow actually flowing through the filter:
To stabilize the filter during sampling, a grid was inserted into the filter carrier. The diameter of the grid and the inner diameter of the ring are the same size (40 mm). However, the grid reduces the effective filter area and this fact is considered by modification of the equation used to determine the active filter area. The filter surface is thus composed of an active and a passive part. Equation (1) calculates the entire filter area (A) but the actual volume flow \(\dot{V}_{{{\text{FA.k}}}} ~\) through the filter needs to be corrected applying Eq. (2).
This correction factor \(k_{{{\text{ges}}}}\) is calculated according to Eq. (3) and refers to the total volume measured by the anemometer but not passing through the filter.
The change of condition between in- and outlet of the sample, can be modelled as isothermal. Therefore, at atmospheric condition, a pressure drop of 40 mbar corresponds to a maximum error of 4% (according to ideal gas equation) of the measured volume flow with the vane anemometer. As mentioned, many other factors (air leakage, grid, turbulences etc.) influence the vane anemometer signal. All factors are summarized by the correction factor kges which was empirically determined (Fig. 2). This factor is only valuable for the used filters and the tested range of volume flow. By correcting the filter surface and the air stream not passing through the filter but through the system, the determination of the real volume flow \(\dot{V}_{{{\text{FA.k}}}} ~\) is now possible by means of gas velocity measurement.
Diagram showing the relationship between air stream velocity and the correction factor kges with different filter materials, n = 28. Experimental setup of establishing the correction factors of the filter holder was according to Fig. 1 and for details on the used filters see descriptions in Materials and Methods
Distinct types of membrane filter discs commonly used for air-based sampling in occupational hygiene assessment (Soo et al. 2016) were used for the physical characterization of the newly developed filter holders. All filters had a diameter of 47 mm to fit into the filter holder and included the following filter types: acrylic copolymer membrane filters Versapor 1200 (pore size 1.2 µm, Pall Inc. product #66394) and Versapor 3000 (pore size 3 µm, Pall Inc. product #66387), mixed cellulose ester/cellulose nitrate membrane filters GN-4 (pore size 0.8 µm, Pall Inc. product # 64679) and GN-6 (pore size 0.45 µm, Pall Inc. product # 66191), hydrophilic polycarbonate membrane filters Isopore (pore size 0.4 µm, Merck-Millipore product # HTTP04700), polytetrafluoroethylene (PTFE) membrane filters type TF1000 (pore size 1 µm, Pall Inc. product # 66155), type Zefluor 0.45 (pore size 0.45 µm, Pall Inc. product # P5PQ047), and type Zefluor 2 (pore size 2 µm, Pall Inc. product #P5PJ047).
2.3 Air Sampling
The newly developed filter system was tested under controlled conditions in a bioaerosol test chamber (Pogner et al. 2019) and during a field study with different indoor mold situations. In both cases the MAS-100 NT impactor was used as a reference device for testing the suitability of the novel filter holders for bioaerosol sampling.
2.3.1 Bioaerosol Chamber
The sampling conditions in the bioaerosols chamber and the according experimental setup used for this study are based on Pogner et al. 2019. In short, the test system that was used for the biological validation of the sampling device, comprised of the CCB 3000 bioaerosol chamber (Calibration Chamber for Bioaerosols 3000, Palas, Germany) and an LSA nebulizer (Liquid Sparging Aerosolizer, CH Technologies, United States). The bioaerosol chamber features a laminar airflow, even particle distribution and a testing area of 0.57 m3. The MAS-100 NT and the filter holder of the air filtration-based system were placed in the chamber. The casing of the vacuum cleaner was placed outside and was connected to the filter holder through a tube connection in the chamber wall. For the spore suspensions, fungal strains were cultivated on malt extract agar (Merck, United States) for 2–3 weeks at room temperature (22 °C). Trichoderma longibrachiatum was used as test organism, since it was used to validate the CCB 3000 bioaerosol chamber as described in Pogner et al. 2019 and its conidia can be aerosolized without detergent. As described previously, the use of detergents in aerosolization creates a high load of unspecific particle background and thus disqualifies parallel control measurements with the particle counter installed inside the chamber. Harvesting of the spores was performed by taking mycelia and spores from the plate with a spatula, adding 0.01% Tween 20 to the plates, vortexing and filtering the suspensions through sterilized glass wool to remove mycelial cells. The spore suspension’s concentrations (given in spores/mL) were determined by counting a 10 µL aliquot of a proper dilution in a C-Chip Haemocytometer (Neubauer improved; Incyto, South Korea) disposable counting chambers. All spores in the nine big squares were counted using a microscope and a 400-fold magnification, the mean and final concentration were calculated. Homogenous spore suspensions (2.0 × 105 spores/mL) of T. longibrachiatum in 0.01% Tween 20 were aerosolized and subsequently sampled with the test devices.
2.3.2 Cultivation
For sampling in the bioaerosol chamber modified HYCON-YM medium (RCS-T; peptone 6 g/L; D (+)-glucose monohydrate 15 g/L; malt extract 5 g/L; yeast extract 1 g/L; agar–agar 16 g/L; pH 7.3 ± 0.2; 0.1% Triton™ X-100) was used. For the field study, microbial particles were cultivated on malt extract agar (MEA; Merck, United States); MEA containing Rose Bengal (MEA-BR; 25 mg/L, respectively, 50 mg/L Rose Bengal); dichloran glycerol agar (DG18; Merck, United States); and RCS-T medium. No significant differences were detected for the tested Rose Bengal concentrations (data not shown). Colony forming units (CFUs) were counted after 6–7 days of incubation at room temperature (22 °C). CFUs were converted to CFU per m3 sampled air (CFU/m3).
2.3.3 Impaction Sampling
Air sampling with the MAS-100 NT (MAS) (MBV, Switzerland) was performed with a constant airflow of 100 L/min for 2 min for the bioaerosol chamber and field study sampling. 18 replicates were conducted for the bioaerosol chamber experiment. For the field study 4–6 replicates were taken per sampling campaign, site, and medium. A statistical correction of the impactor was not needed, since CFUs were well within the countable range (Ghosh et al. 2015; Mandal and Brandl 2011).
2.3.4 Filtration-Based Microbial Sampling
The filter system combined with a vacuum cleaner (VAC) was used for air filtration sampling with different membrane filter types commonly employed in bioaerosol sampling: Versapor 1200 acrylic copolymer membrane disc filters (V 1200; 47 mm, pore size 1.2 µm, Pall Inc., United States), cellulose nitrate/mixed cellulose ester filters (GN-6, 47 mm, pore size 0,45 µm, both from Pall Inc., United States), and polycarbonate filters (PC; Isopore membrane, 47 mm, pore size 0.4 µm, Merck-Millipore, Germany). Air filtration-based sampling for the field study was performed with Versapor 1200 filters (V 1200). A series of different air velocities was measured with an impeller anemometer (airflow meter PCE-TA 30, PCE Instruments, Germany) for all filters used in this study and according air sampling (L/min) were established (Table ST1). By measuring the air velocity (m/s) of each vacuum cleaner prior to air sampling, the corresponding sampling volume can be calculated according to Table S1 independent from the performance and type of the vacuum cleaner. For the bioaerosol chamber measurements a “Siemens Super XS dino e, 2200 W” vacuum cleaner (Siemens, Germany) was used. Based on the same air velocity of the vacuum cleaner (0.89 m/s), sampling was performed for 3 min for each of the tested filters (Versapor 1200 filter, Polycarbonate filter, and Cellulose Nitrate filter). During the field study sampling was conducted with Versapor 1200 filters (V 1200) and a “Simpex 17730, 800 W” (Simpex, United States) vacuum cleaner for 3 min 20 s at an air velocity of 0.80 m/s. Five to 6 replicates on 2 independent sampling days were sampled during for the bioaerosol chamber experiments. Two replicates on 4 independent sampling campaigns were taken per sampling site and medium during the field study. Previously determined correction factors for each filter material were considered in the calculations (see Sect. 2.2).
2.3.5 Field Study Sampling Sites
The field study was conducted at an industrial unit of a sparkling wine factory. Further details on the sampling site are described in Unterwurzacher et al. (2018). The sampling sites covered a wine cellar with obvious mold growth, an office that is primarily used as storage room without indications of mold infestation and an inner courtyard that separated the indoor sampling sites and served as outdoor reference. During sampling, a minimum distance of 1 m was kept between each sampling device and any other object (including other sampling devices, walls, and/or personnel). Relative humidity and temperature were monitored. The observed relative humidity and temperature ranges are listed in Table ST 2. Four independent sampling campaigns were performed between June 2015 and January 2016. In detail, sampling was conducted at 23 June 2015 (sampling 1), 5 October 2015 (sampling 2), 23 November 2015 (sampling 3) and 27 January 2016 (sampling 4).
Besides the wine cellar with obvious mold growth, an indoor sampling site (office) without indications of mold infestation was chosen to represent both, a highly contaminated site as well as a sampling site with low levels of airborne mold particles. Outdoor samples were used as reference to determine background levels at the location and to check for distinct differences between the fungal communities indoors and outdoors. The MAS-100 NT impactor (MAS) was used for validation of the results obtained by the newly designed filter holders. Furthermore, 4 different cultivation media were compared, to investigate if certain fungal communities might be underrepresented, either dependent on the media, the sampling device or a combination of these factors.
2.3.6 Field Study Fungal Community Spectra
For the field study, CFUs were counted and classified according to the groups Alternaria, Cladosporium, Aspergillus/Penicillium, Aureobasidium, Fusarium, Neurospora, Rhizopus/Mucor, Trichoderma, Chaetomium, Epicoccum, Botrytis, and yeast. The selection of these taxa was based on literature indicating frequently occurring outdoor and indoor airborne fungal genera (Bowers et al. 2013; Fierer et al. 2008; Hyvärinen et al. 2001; Lymperopoulou et al. 2016; Yamamoto et al. 2012). To enable a fast and efficient analytical evaluation, the taxa Aspergillus/Penicillium and Rhizopus/Mucor were combined in 1 classification due to their high morphological similarities, as suggested by other studies (Li and Kendrick 1995). Unidentified fungal colonies were classified in the categories "other sporulating fungi" and sterile mycelia. Rankings were determined corresponding to the relative abundance within the sample (1 = low abundance; 2 = medium abundance; 3 = dominant). A taxon was considered as dominant if CFUs for that community fraction exceeded 75% of the total cell counts. Subsequently, low and medium abundances were defined accordingly. Data were illustrated in a heat map showing relative abundances within one sample. Technical replicates of each sampling day, sampling location, and cultivation medium were combined.
2.4 Statistical Analysis and Linear Regression Models
Statistical analyses were conducted using R version 3.2.4 (R Core Team 2015). Pearson's correlation coefficients between sampling devices and environmental factors (temperature and relative humidity) were calculated using cor.test (x, y). Pearson’s coefficients were used in respect to creating first order linear models and avoiding overfitting of these models for comparison between different sampling devices. Normal distribution of the residuals was tested by the Shapiro–Wilk test and p values less than 0.05 were considered significant.
For the comparison of different sampling devices, multi-dimensional linear models were developed. These models consider besides the different sampling methods (CFUs) also the factors temperature and relative humidity. The respective coefficients for each factor were estimated by minimizing the sum of squared residuals and the assumption that y-intercept is zero. For estimating the regression coefficients of linear models, the means of 5 technical repeats were considered. For linear least squares regression computation, the R function “lm” was used.
Differences between mean CFU values of different media or sampling devices were calculated using the R function t test (x,y). p values less than 0.01 were considered significant.
Standard deviations of measurements conducted during the field study are shown as boxplot. In the boxplot the calculated upper and lower quartiles (75th and 25th percentile), the maximum and minimum values (upper and lower whiskers), and the median (excluding outliers) are indicated. If data exceeded 1.5 times, the interquartile range of these measurements were considered as outliers (circles).
3 Results
3.1 Filter System Characterization
To determine the correction factor kges, the parameters kRest and kFL were measured individually. For \(k_{{{\text{Rest}}}}\), determination, the filter attachment site with the ring fixing the filter and the adaptor were glued airtight, so that it can be assumed that the air volume flow due to leakage \(\dot{V}_{{{\text{FL}}}}\) is equal to zero. For this experimental setup, \(k_{{{\text{Rest}}}}\) can be calculated using Eq. (4), where vFA is the gas velocity at the filter inlet, \(\dot{V}_{{{\text{BGZ}}}}\) is the total volume flow, \(v_{{{\text{FA}}}}\) denotes the velocity measured by hydrometric vane and A is the filter surface area.
The air passing through the system but not through the filter is termed “false air”. In our setup this results from the difference between the volume flow of the diaphragm gas meter \(\dot{V}_{{{\text{BGZ}}}} ~\) and the corrected volume flow of the hydrometric vane \(\dot{V}_{{{\text{FA.k}}}}\) according to Eqs. (5) and (6).
and,
The test series for determining the correction factor \(k_{{{\text{Rest}}}}\), was carried out with airtight glued filter components. As a result, the entry of false air \(\dot{V}_{{{\text{FL}}}}\) due to leaks in the filter attachment could be excluded and the volume flow of the diaphragm gas meter \(\dot{V}_{{{\text{BGZ}}}}\) can be considered as the actual flow rate through the filter. To determine the false air portion of the filter system, the experiments for the determination of \(k_{{{\text{Rest}}}}\) were repeated with non-bonded components. Identical to the test series of the bonded components, the volume flow of each filter was plotted over the velocities at four different partial loads and a linear regression model was formed. The difference in the volume flows at the same speeds results in the false air entry \(\dot{V}_{{{\text{FL}}}}\). The numerical results are listed in Fig. S1 and represent the mean of three measurements, each of it was determined at four different partial loads. The filters GN-6 and Isopore (PC) have a comparatively high correction factor \(k_{{{\text{FL}}}}\) due to lower porosity and, therefore, a higher pressure drop of the filter. Results are shown and summarized in Fig. 2. In summary, it was established that even with a simple vacuum cleaner the adaptor system can be used and correct measurements can be performed as air leakage is not critical factor for the novel bioaerosol collection system.
3.2 Sampling Device Comparison Under Controlled Conditions
A bioaerosol chamber was used to establish stable sampling conditions for the reference sampler (MAS) and the air filtration-based system to be validated (VAC) as described before (Pogner et al. 2019). Three different filter materials were tested with the air filtration-based sampler in this comparative study under controlled conditions. Similar sampling concentrations between the distinct filter materials were detected (162–206 CFU/m3), with the exception of the comparison of the Cellulose Nitrate and the Polycarbonate filter, where mean CFU/m3 values were in a comparable range, but differences were significant (p value < 0.01). Independently from the tested filter materials, the air filtration-based system showed highly significant impairment in sampling Trichoderma longibrachiatum spores compared to the MAS-100 NT impactor (MAS), with on average 66.9% lower CFU counts detected by the air filtration system (Fig. 3). Considering the broad stable range of different air velocities (roughly 0.7–2.0 m/s) tested during the establishment of the filters’ correction factors (Fig. S1) and the comparable sampling efficiency with the 2 other tested filter materials in the bioaerosol chamber, in further consequence the Versapor 1200 filter was used for the field study.
Results of the bioaerosol chamber experiments. Comparison of CFU detection of Trichoderma longibrachiatum spores under controlled conditions in a bioaerosol chamber using the filter-adaptor system. The MAS-100 NT (MAS) was used as a reference device (n = 18). Different membrane filters were tested for the air filtration-based sampling using the vacuum cleaner-adaptor system (VAC): VAC Versapor: Versapor 1200 filter (n = 5), VAC Cell.Nitr.: cellulose nitrate GN-6 filter (n = 6), and VAC Polycarb: Isopore polycarbonate filter (n = 6). Asterisk indicates significant difference to MAS reference sampler
3.3 Comparative Field Study
For further validation of the air filtration-based sampler, a comparative field study was conducted at the production site of a sparkling wine factory during 4 independent sampling campaigns.
3.3.1 Detected Colony Forming Units
Independent from the sampling site and sampling device, there was a clear seasonal shift in airborne fungal particles (Fig. 4). Overall highest CFU values (up to 2054 CFU/m3 outdoors; 3196 CFU/m3 wine cellar; 724 CFU/m3 office) were measured at sampling campaigns 1 and 2.
Results of the field studies. CFU counts detected at a comparative field sampling study at four different sampling days (1–4) conducted with the MAS-100 NT (MAS) and air filtration (VAC; on Versapor 1200 filters) for the tested cultivation media DG18, MEA, MEA-BR, and RCS-T. Results are given for each sampling site. Asterisk indicates significant difference to MAS reference sampler
While cultivation medium had little influence on CFU counts from filtration-based sampling, significant differences (p value < 0.01) were found for 41.7% of measurements from the MAS-100 NT (MAS) (Table ST3).
Comparing differences in sampling devices, particularly outdoors the MAS-100 NT impactor (MAS) and the air filtration-based sampler detected fungal CFUs in a highly similar range (MAS: 2054–230 CFU/m3; VAC: 1618–241 CFU/m3). In the wine cellar, the air filtration-based sampler measured significantly higher CFU values compared to the MAS-100 NT impactor (MAS: 1731–463 CFU/m3; VAC: 3196 ± 365 CFU/m3). The CFU levels detected in the office were similar between the two sampling systems and ranged between MAS: 703 ± 48 CFU/m3 and VAC: 724 ± 98 CFU/m3.
3.3.2 Measurement Stability Based on Relative Standard Deviations
Overall, sampling with the MAS-100 NT (MAS) showed a high reproducibility with slightly lower relative standard deviations (rsd) of the measurement standard deviations compared to the air filtration-based sampler (median of rsd: MAS: 12.8%; VAC: 19.4%) (Fig. 5). Particularly outdoor measurements conducted with the air filtration-based sampler varied strongly, while MAS-100 NT data were more robust (median of rsd: MAS: 12.1%; VAC: 24.1%). Highest similarities in measurement reproducibility were found in the office with little expected spore counts in the aerosols, where both sampling devices showed relatively high data fluctuations (median of rsd: MAS: 18.4%; VAC: 22.4%). Contrary to that, both samplers displayed the lowest relative standard deviations in the wine cellar, which generally exhibited high CFU levels due to the obvious indoor mold infestation (median of rsd: MAS: 9.3%; VAC: 14.6%).
Measurement reproducibility of the air filtration system. Relative standard deviations [rsd (%)] of field samplings were calculated from the technical replicates to the means of the sampling campaign for the MAS-100 NT (MAS) and the air filtration-based sampler (VAC; on Versapor 1200 filters). Boxplots include data from four independent sampling campaigns and four different media, resulting in 16 data points for each sampling location and sampler. In the boxplot the calculated upper and lower quartiles (75th and 25th percentile), the maximum and minimum values (upper and lower whiskers), and the median (excluding outliers) are indicated. If a given SD value exceeded 1.5 times the interquartile range, this value was considered as outlier (circle). Values used for statistical analyses shown in the boxplots are listed in Supplementary Table ST 4
3.3.3 Sampler Correlations and Linear Regression Models
A linear regression model for comparison of sampling devices and methodologies based solely on CFU counts showed already high comparability between the sampling devices (MAS-100 NT and air filtration) with an R2 value of 0.90 (Fig. 6). Furthermore, Pearson’s correlations (overall data Shapiro–Wilk test of normality, p value > 0.05) at each sampling site substantiated these strong correlations between the compared air samplers (outdoors: cor = 0.96, p value < 0.01; cellar: cor = 0.89, p value < 0.01; and office: cor = 0.77, p value < 0.01).
Correlations of detected CFU counts between the reference sampler MAS-100 NT (MAS) and the air filtration-based sampler (VAC; on Versapor 1200 filters). All sampling campaigns and cultivation media are shown. Data are indicated dependent on the sampling locations of the comparative field study (outdoor reference; office; wine cellar) and the continuous line represents the overall linear correlation
Furthermore, we developed multi-dimensional linear models that implement also the factors of temperature and relative humidity (Supplementary Table ST2). Considering these additional parameters, the adjusted R2 value reached nearly 0.95 (Table 1). Equation 7 allows calculating CFU counts from a specified device in our study given the CFU counts from another device in this study, considering environmental temperature and relative humidity:
Besides CFU counts, only the influence of humidity was statistically significant. Temperature was of minor importance. The herein presented correlations are valid for this study. Further comparative measurement campaigns would be needed for more general correlations. Also, polynomial functions were investigated but could not bring a significant improvement to our models (data not shown). Measurements taken outdoors could not sensibly be fitted into these models as there may be influencing factors that were not measured in our experiments (e.g., wind velocity and air turbulence).
3.3.4 Culture-Based Fungal Community Spectra
Besides CFU values, the influence of cultivation media and sampling devices on the detected fungal community spectra was investigated during the field study. To cover all possible underrepresented fungal genera, 4 different sampling media (DG18; MEA; MEA-BR; RCS-T) with distinct cultivation characteristics were chosen. Community composition was highly influenced by both, sampling method and growth medium. While outdoors especially Cladosporium and Alternaria species were detected, in the wine cellar Aspergillus/Penicillium species were dominant, indicating an indoor mold infestation. In the vacant office, besides typical outdoor taxa Cladosporium and Alternaria also Aspergillus/Penicillium species were detected more frequently, which could be explained by whirled up dust during sampling rather than by indoor mold growth.
Generally, fungal community compositions sampled with the MAS-100 NT (MAS) impactor showed slightly higher diversity at all sampling locations. For example, colonies belonging to the fungal taxa Aureobasidium, Trichoderma, and Epicoccum were almost exclusively detected with the MAS-100 NT (MAS). For the most dominant genera such as Alternaria, Cladosporium, and Aspergillus/Penicillium similar abundances were found with both sampling devices (Fig. 7).
Community spectra of cultivated airborne microbial particles sampled with the MAS-100 NT (MAS) and the air filtration-based system (VAC) during the field study. All sampling days (1–4), sampling locations (outdoor reference; office; wine cellar) and different cultivation media (DG18, MEA, MEA-BR; RCS-T) are illustrated. The heat map displays relative abundances of the fungal communities detected within the sample. The color code is indicated. Yellow: highest abundance; dark blue: lowest abundance; black: below detection limit (bdl). Mean values of replicates are represented (color figure online)
4 Discussion
Here, we present the development and evaluation of a new filter-based bioaerosol sampling system that can be operated with commercially available vacuum cleaners as air pumps. It opens the possibility to perform multiple sampling campaigns in parallel in a short period of time at many different, geographically distant locations. Such samplings are done for instance for epidemiologic studies to screen pathogens or monitor antibiotic resistance genes by molecular biology analysis. Until now, large-scale comparative samplings are not routinely performed because at each of these distant sampling locations, a standard bioaerosol sampler is needed and the associated costs usually exceed the budget available for such studies. The adaptor and sterile filter unit composite serve as a feasible option for routine analytics as the system is highly cost-effective compared to other sampling devices, since beside a vacuum cleaner that is usually available and an anemometer, no further equipment is required. The physical tests of the sampling system with different air filters resulted in the formulation of a correction factor for each filter to be applied. This factor in combination with a simple anemometer gave consistent sampling volumes in each of the tests run. Therefore, the combination of the filter-adaptor and vacuum cleaner can thus be regarded as reliable bioaerosol sampling system. An uncertainty of the system that has so not been tested in the long run may be a reduction of vacuum generation by the vacuum cleaner. The performance of the system might suffer from reduced electric engine performance after a long sampling campaign. However, field data from more than 300 individual test runs as presented here gave quite consistent CFU counts in comparison to the reference sampler and thus we do not foresee large problems due to reduction of performance of the vacuum cleaner used for sampling.
Although the biological sampling efficiency was comparable to the reference device in our field studies, under controlled conditions in a bioaerosol chamber, the air filtration-based sampler showed significantly lower CFU-based collection efficiency of aerosolized Trichoderma longibrachiatum spores. The susceptibility of the sampled microorganism due to diverging sampling stress pursued by the different sampling principles might explain this discrepancy (Wang et al. 1999). Although a variety of different species could have been tested with the new filter system in the chamber, we decided to perform this first step in validation only under the most suitable conditions for our MAS-100 NT control device as defined in our previous study (Pogner et al., 2019). As outlined in the Sect. 2Trichoderma was used because it is one of the few species whose spores can be aerosolized without detergent and thus allows parallel control measurements with a particle counter installed inside the chamber. Moreover, in the subsequent field sampling comparative studies were broad in terms of conditions, locations, and expectable fungal biodiversity.
Generally, the testing in the bioaerosol chamber demonstrated measurement stabilities of viable T. longibrachiatum spores similar to values reached by the MAS-100 NT, suggesting high detection reproducibility of the novel air filtration system under conditions without wind effects. In future, more microbial species will be compared between the filter-based sampler and a reference sampler in the bioaerosol chamber to gain better insights into possible bias for certain types of microbes.
As reported by Elmashae and coworkers (Elmashae et al. 2017), which tested different filter materials accordingly to identify airborne bacterial cells, our results showed similar detection rates with all tested filter materials. It has to be mentioned, that pore sizes of the different membrane filter discs tested in the bioaerosol chamber were not identical, but a similar size range (Versapor 1.2 µm pore size; Cellulose Nitrate and Polycarbonate 0.8 µm pore sizes). Nevertheless, considering that typical fungal spores feature sizes from approx. 1–30 µm (Yoo et al. 2017), the effect of the different pore sizes is expected to be rather small. This would be in agreement with the findings of Soo et al. (2016), who detected similar particle collection efficiencies in the range of 1 µm pore sizes and strong deviations only compared to higher pore sizes among different commonly used air filter materials.
The new filter system was also functional in displaying expected seasonal fluctuations in cultivatable fungal cells. It is well known that the composition and biodiversity of airborne microbial cells as well as their concentrations change strongly with seasons (Bowers et al. 2013; Frankel et al. 2012a; Heo et al. 2014; Matthias-Maser et al. 2000). As reported before by Bowers and colleagues (Bowers et al. 2013), also our results showed highest fungal quantities between summer and fall and lowest in winter. The diverse sampling time points covering different seasons were chosen for this study to determine if not only number but also biodiversity of monitored microbes correlate between our device and the reference sampler. Strong overall correlations between the tested and the reference devices indicate high measurement reliability of the new filter holder system for environmental samples (Ghosh et al. 2015; Kalogerakis et al. 2005; Yao and Mainelis 2006).
To investigate if certain fungal species are underrepresented in the newly developed sampling system, four different cultivation media were compared during the field study. The tested cultivation media covered highly divergent growth properties including different carbon and nitrogen sources, water activities and inhibitory compounds. The air filters detected comparable CFU counts on all tested cultivation media, which was not the case for the MAS-100 NT. A reduction in biodiversity due to higher sampling stress (Zhen et al. 2018) during filtration (see below) could explain the minor influence of cultivation media on enumeration of fungal propagules. Fungi resistant to the stress imposed by filtration probably grow equally well on different growth media.
There was also no significant bias towards indoor or outdoor locations. Particle counts detected outdoors and in the office were in a highly similar range for both sampling devices. In the wine cellar, particularly at sampling campaigns 1 and 2, the filter-based system reached higher CFU counts compared to the MAS-100 NT. Since the community spectra of the office and the outdoor reference are more alike as shown in another study conducted at this site (Unterwurzacher et al. 2018), discrepancies in particle detections in the wine cellar might be again a result of species related underrepresentation.
However, one weakness of the newly developed system might be the higher variability of detected fungal counts. Overall, relative standard deviations were higher for the air filters compared to the impactor at the chosen conditions and experimental design, i.e., two technical replicates per cultivation media, per sampling site, and per sampling date. Although this sampling strategy still concurs with official directives and guidelines, e.g., DIN ISO 16000-16 (2008) the comparatively low number of repeats might explain higher relative standard deviations. Furthermore, filters need additional manipulation steps after sampling, while plates from the MAS-100 NT impactor are directly incubated after sampling. Handling of filters can potentially increase variation in enumeration of fungal propagules.
Similar to other comparative studies of bioaerosol samplers, we also found differences in community composition depending on the sampling device (Lukaszuk et al. 2017; Unterwurzacher 2016). Nevertheless, it cannot be excluded that particle dispersal not only occurred through the overall sampling activity (Meadow et al. 2014) but especially during the use of the vacuum cleaner (Corsi et al. 2008; Knibbs et al. 2012).
Sampling with the MAS-100 NT impactor showed slightly higher diversity at all investigated sites. Although most dominant taxa including Alternaria, Cladosporium, and Aspergillus/Penicillium were detected in a similar range, other taxa such as Aureobasidium, Trichoderma, and Epicoccum were almost exclusively detected with the MAS-100 NT impactor. This is consistent with the findings in the bioaerosol chamber experiment, where Trichoderma longibrachiatum spores were strongly underrepresented by the air filtration system. Higher susceptibility to desiccation stress during air filtration might explain the underrepresentation of more desiccation-sensitive species (Ghosh et al. 2015; Wang et al. 2001).
Overall, the field study indicated a strong and highly significant correlation between the newly designed filter holders and the well-established impactor system. The implementation of multi-dimensional linear models for the indoor measurements considered the environmental factors temperature and relative humidity, which further strengthened the correlation. Given the results from these models, it can be deduced that especially relative humidity has a significant impact on the comparability between the different sampling methods used in this study. The temperature did not show a significant influence on the measurement correlations. Both bioaerosol samplers were subjected to some species related restrictions in detecting viable fungal cells, but overall the CFU-based results were comparable and the CFU regression coefficient of 1.026 (p value < 0.01) indicates the reproducibility of the compared sampling devices during environmental conditions.
Obviously, presented results are valid for the sampling settings chosen in this study, different sampling durations or airflow rates might lead to diverging correlations, since sampling settings are crucial for the overall performance of bioaerosol samplers (Grinshpun et al. 2007; Han et al. 2015; Uhrbrand et al. 2017). Although a broad range of environmental conditions, including different seasons, indoor sampling sites with high, respectively, low fungal particle levels and outdoor reference measurements were covered within this experimental setup, further studies at different sampling sites are needed to verify these finding. One limitation of the herein presented field study is the rather small number of technical replicates, which possibly explains the higher relative standard deviations compared to the MAS-100 NT reference device.
5 Conclusion
The findings underline the applicability of the novel vacuum-cleaner-based filter sampling system to detect viable fungal cells and spores in bioaerosols. The system is largely equivalent to other commonly used samplers as it gave similar results as the reference MAS-100 NT impaction sampler. The advantage of this system is that (1) no extra air pump or collector needs to be purchased as vacuum cleaners are usually available, (2) materials used for collection (sterile filters) endure harsh temperature conditions during storage and thus have a long shelf life, and (3) the simplicity of the system allows large number sampling campaigns to be run in parallel at very distant locations (e.g., broad antibiotic resistance sampling campaigns). A weakness of the system may be the need of an anemometer to calibrate the air volume sampled per time unit, a possible underrepresentation of some genera depending on the chosen filters and a slightly higher variability of the obtained results due to changing power capacities of the vacuum cleaners. However, these slight disadvantages may be compensated by opening the possibility to perform large parallel sampling campaigns at distant locations. Such scientifically valuable endeavors would not be economically feasible if at each of these locations a costly commercial sampler needs to be purchased. Other bioaerosol compounds such as spore-forming bacteria or pollen can equally be sampled with the described system once validation studies with these components have been performed similar to the work presented here.
Code availability
Not applicable.
Abbreviations
- bdl:
-
Below detection limit
- CFU:
-
Colony forming unit
- MCE:
-
Mixed cellulose ester
- DG18:
-
Dichloran glycerol agar
- MAS:
-
MAS-100 NT (Microbial Air Sampler)
- MCE:
-
Mixed cellulose esters
- MEA:
-
Malt extract agar
- MEA-BR:
-
Malt extract agar containing Rose Bengal
- PTFE:
-
Polytetrafluoroethylene (TF)
- RCS-T:
-
Modified HYCON-YM medium
- sd:
-
Standard deviation
- VAC:
-
Air filtration by filter holder and vacuum cleaner
References
Aizenberg V, Reponen T, Grinshpun SA, Willeke K (2000) Performance of Air-O-Cell, Burkard, and Button Samplers for total enumeration of airborne spores. Am Ind Hyg Assoc J 61(6):855–864. https://doi.org/10.1202/0002-8894(2000)061%3c0855:POAOCB%3e2.0.CO;2
An HR, Mainelis G, Yao M (2004) Evaluation of a high-volume portable bioaerosol sampler in laboratory and field environments. Indoor Air 14(6):385–393. https://doi.org/10.1111/j.1600-0668.2004.00257.x
Bernstein JA, Alexis N, Bacchus H, Bernstein IL, Fritz P, Horner E, Li N, Mason S, Nel A, Oullette J, Reijula K, Reponen T, Seltzer J, Smith A, Tarlo SM (2008) The health effects of nonindustrial indoor air pollution. J Allergy Clin Immunol 121(3):585–591. https://doi.org/10.1016/j.jaci.2007.10.045
Bowers RM, Clements N, Emerson JB, Wiedinmyer C, Hannigan MP, Fierer N (2013) Seasonal variability in bacterial and fungal diversity of the near-surface atmosphere. Environ Sci Technol 47(21):12097–12106. https://doi.org/10.1021/es402970s
Burton NC, Grinshpun SA, Reponen T (2007) Physical collection efficiency of filter materials for bacteria and viruses. Ann Occup Hyg 51(2):143–151. https://doi.org/10.1093/annhyg/mel073
Chang CW, Hwang YH, Grinshpun SA, Macher JM, Willeke K (1994) Evaluation of counting error due to colony masking in bioaerosol sampling. Appl Environ Microbiol 60:3732–3738
Corsi RL, Siegel JA, Chiang C (2008) Particle resuspension during the use of vacuum cleaners on residential carpet. J Occup Environ Hyg 5(4):232–238. https://doi.org/10.1080/15459620801901165
Dannemiller KC, Gent JF, Leaderer BP, Peccia J (2016) Influence of housing characteristics on bacterial and fungal communities in homes of asthmatic children. Indoor Air 26(2):179–192. https://doi.org/10.1111/ina.12205
Deacon LJ, Pankhurst LJ, Drew GH, Hayes ET, Jackson S, Longhurst PJ, Longhurst JWS, Liu J, Pollard SJT, Tyrrel SF (2009) Particle size distribution of airborne Aspergillus fumigatus spores emitted from compost using membrane filtration. Atmos Environ 43:5698–5701. https://doi.org/10.1016/j.atmosenv.2009.07.042
DIN-ISO16000-16 (2008) Indoor air. Part 16: detection and enumeration of moulds—sampling by filtration, vol 16000-16. International Standards Organization
DIN-ISO13205-1:2014 (2014) Workplace atmospheres-Assessment of performance of instruments for measurement of airborne particle concentrations German Version. International Standards
Dybwad M, Skogan G, Blatny JM (2014) Comparative testing and evaluation of nine different air samplers: end-to-end sampling efficiencies as specific performance measurements for bioaerosol applications. Aerosol Sci Technol 48(3):282–295. https://doi.org/10.1080/02786826.2013.871501
Eduard W, Heederik D (1998) Methods for quantitative assessment of airborne levels of noninfectious microorganisms in highly contaminated work environments. Am Ind Hyg Assoc J 59(2):113–127. https://doi.org/10.1080/15428119891010370
Elmashae Y, Grinshpun SA, Reponen T, Yermakov M, Riddle R (2017) Performance of two respiratory protective devices used by home-attending health care workers (a pilot study). J Occup Environ Hyg. https://doi.org/10.1080/15459624.2017.1319571
Estill CF, Baron PA, Beard JK, Hein MJ, Larsen LD, Deye GJ, Rose L, Hodges L (2008) Comparison of air sampling methods for aerosolized spores of B. anthracis sterne. J Occup Environ Hyg 8(3):179–186. https://doi.org/10.1080/15459624.2011.556981
Feather GA, Chen BT (2003) Design and use of a settling chamber for sampler evaluation under calm-air conditions. Aerosol Sci Technol 37(3):261–270. https://doi.org/10.1080/02786820390125115
Fierer N, Liu Z, Rodrguez-Hernndez M, Knight R, Henn M, Hernandez MT (2008) Short-term temporal variability in airborne bacterial and fungal populations. Appl Environ Microbiol 74(1):200–207. https://doi.org/10.1128/AEM.01467-07
Frankel M, Bekö G, Timm M, Gustavsen S, Hansen EW, Madsen AM (2012a) Seasonal variations of indoor microbial exposures and their relation to temperature, relative humidity, and air exchange rate. Appl Environ Microbiol 78:8289–8297. https://doi.org/10.1128/AEM.02069-12
Frankel M, Timm M, Hansen EW, Madsen AM (2012b) Comparison of sampling methods for the assessment of indoor microbial exposure. Indoor Air 22(5):405–414. https://doi.org/10.1111/j.1600-0668.2012.00770.x
Ghosh B, Lal H, Srivastava A (2015) Review of bioaerosols in indoor environment with special reference to sampling, analysis and control mechanisms. Environ Int 85:254–272. https://doi.org/10.1016/j.envint.2015.09.018
Grinshpun S, Adhikari A, Cho S-H, Kim K-Y, Lee T, Reponen T (2007) A small change in the design of a slit bioaerosol impactor significantly improves its collection characteristics. JEM 9(8):855–861. https://doi.org/10.1039/b702743e
Haig CW, Mackay WG, Walker JT, Williams C (2016) Bioaerosol sampling: sampling mechanisms, bioefficiency and field studies. J Hosp Infect 93(3):242–255. https://doi.org/10.1016/j.jhin.2016.03.017
Han T, Wren M, Dubois K, Therkorn J, Mainelis G (2015) Application of ATP-based bioluminescence for bioaerosol quantification: effect of sampling method. J Aerosol Sci 90:114–123. https://doi.org/10.1016/j.jaerosci.2015.08.003.Application
Heo KJ, Kim HB, Lee BU (2014) Concentration of environmental fungal and bacterial bioaerosols during the monsoon season. J Aerosol Sci 77:31–37. https://doi.org/10.1016/j.jaerosci.2014.07.001
Hyvärinen A, Vahteristo M, Meklin T, Jantunen M, Nevalainen A, Moschandreas D (2001) Temporal and spatial variation of fungal concentrations in indoor air. Aerosol Sci Technol 35:688–695. https://doi.org/10.1080/02786820117763
Jones RM, Brosseau LM (2015) Aerosol transmission of infectious disease. J Occup Environ Med 57(5):501–508. https://doi.org/10.1097/JOM.0000000000000448
Juozaitis A, Willeke K, Grinshpun SA, Donnelly J (1994) Impaction onto a glass slide or agar versus impingement into a liquid for the collection and recovery of airborne microorganisms. Appl Environ Microbiol 60(3):861–870
Kalogerakis N, Paschali D, Lekaditis V, Pantidou A, Eleftheriadis K, Lazaridis M (2005) Indoor air quality—bioaerosol measurements in domestic and office premises. J Aerosol Sci 36:751–761. https://doi.org/10.1016/j.jaerosci.2005.02.004
Kesavan J, Sagripanti JL (2015) Evaluation criteria for bioaerosol samplers. Environ Sci Process Impacts 17(3):638–645. https://doi.org/10.1039/c4em00510d
Kespohl S, Maryska S, Zahradnik E, Sander I, Bruning T, Raulf-Heimsoth M (2013) Biochemical and immunological analysis of mould skin prick test solution: current status of standardization. Clin Exp Allergy 43(11):1286–1296. https://doi.org/10.1111/cea.12186
Klepeis NE, Nelson WC, Ott WR, Robinson JP, Tsang AM, Switzer P, Behar JV, Hern SC, Engelmann WH (2001) The National Human Activity Pattern Survey (NHAPS): a resource for assessing exposure to environmental pollutants. J Expos Anal Environ Epidemiol 11:231–252. https://doi.org/10.1038/sj.jea.7500165
Knibbs LD, He C, Duchaine C, Morawska L (2012) Vacuum cleaner emissions as a source of indoor exposure to airborne particles and bacteria. Environ Sci Technol 46(1):534–542. https://doi.org/10.1021/es202946w
Li DW, Kendrick B (1995) A year-round comparison of fungal spores in indoor and outdoor air. Mycologia 87:190–195
Lukaszuk C, Krajewska-Kulak E, Guzowski A, Kulak W, Kraszyska B (2017) Comparison of the results of studies of air pollution fungi using the SAS super 100, MAS 100, and air IDEAL. Int J Environ Res Public Health. https://doi.org/10.3390/ijerph14070815
Lymperopoulou DS, Adams RI, Lindow SE (2016) Contribution of vegetation to the microbial composition of nearby outdoor air. Appl Environ Microbiol 82:3822–3833. https://doi.org/10.1128/AEM.00610-16
Madsen AM, Larsen ST, Koponen IK, Kling KI, Barooni A, Karottki DG, Tendal K, Wolkoff P (2016) Generation and characterization of indoor fungal aerosols for inhalation studies. Appl Environ Microbiol 82:2479–2493. https://doi.org/10.1128/AEM.04063-15
Mandal J, Brandl H (2011) Bioaerosols in indoor environment—a review with special reference to residential and occupational locations. Open Environ Biol Monit J 4(1):83–96. https://doi.org/10.2174/1875040001104010083
Matthias-Maser S, Obolkin V, Khodzer T, Jaenicke R (2000) Seasonal variation of primary biological aerosol particles in the remote continental region of Lake Baikal/Siberia. Atmos Environ 34(22):3805–3811. https://doi.org/10.1016/S1352-2310(00)00139-4
Meadow JF, Altrichter AE, Kembel SW, Kline J, Mhuireach G, Moriyama M, Northcutt D, O’Connor TK, Womack AM, Brown GZ, Green JL, Bohannan BJM (2014) Indoor airborne bacterial communities are influenced by ventilation, occupancy, and outdoor air source. Indoor Air 24:41–48. https://doi.org/10.1111/ina.12047
Miaskiewicz-Peska E, Lebkowska M (2012) Comparison of aerosol and bioaerosol collection on air filters. Aerobiologia 28(2):185–193. https://doi.org/10.1007/s10453-011-9223-1
Nazaroff WW (2004) Indoor particle dynamics. Indoor Air 14:175–183. https://doi.org/10.1111/j.1600-0668.2004.00286.x
Nazaroff WW (2016) Indoor bioaerosol dynamics. Indoor Air. https://doi.org/10.1111/ina.12174
Nevalainen A, Täubel M, Hyvärinen A (2015) Indoor fungi: companions and contaminants. Indoor Air 25:125–156. https://doi.org/10.1111/ina.12182
Normand A-C, Ranque S, Cassagne C, Gaudart J, Sallah K, Charpin D-A, Piarroux R (2015) Comparison of air impaction and electrostatic dust collector sampling methods to assess airborne fungal contamination in public buildings. Ann Occup Hyg 60:161–175. https://doi.org/10.1093/annhyg/mev075
Osborne NJ, Thornton CR, Sharpe RA (2015) Indoor fungal exposure and allergic respiratory disease. Curr Allergy Asthma Rep 15:1–9. https://doi.org/10.1007/s11882-015-0572-7
Pogner C, Konlechner A, Unterwurzacher V, Kolk A, Hinker M, Mölter L, Strauss J, Gorfer M, Strauss-Goller S (2019) A novel laminar-flow-based bioaerosol test system to determine biological sampling efficiencies of bioaerosol samplers. Aerosol Sci Technol 53(4):355–370. https://doi.org/10.1080/02786826.2018.1562151
Raynor PC, Leith D, Lee DK, Mukund R (2011) Sampling and analysis using filters. Wiley, Hoboken, pp 107–128
Reeslev M, Miller M, Nielsen KF (2003) Quantifying mold biomass on gypsum board: comparison of ergosterol and beta-N-acetylhexosaminidase as mold biomass parameters. Appl Environ Microbiol 69(7):3996–3998. https://doi.org/10.1128/aem.69.7.3996-3998.2003
Rylander R, Reeslev M, Hulander T (2010) Airborne enzyme measurements to detect indoor mould exposure. J Environ Monit 12(11):2161. https://doi.org/10.1039/c0em00336k
Simon X, Duquenne P (2013) Feasibility of generating peaks of bioaerosols for laboratory experiments. Aerosol Air Qual Res. https://doi.org/10.4209/aaqr.2012.12.0340
Soo J-C, Monaghan K, Lee T, Kashon M, Harpera M (2016) Air sampling filtration media: Collection efficiency for respirable size-selective sampling. Aerosol Sci Technol 50(1):76–87
Tseng CC, Li CS (2005) Collection efficiencies of aerosol samplers for virus-containing aerosols. J Aerosol Sci 36:593–607. https://doi.org/10.1016/j.jaerosci.2004.12.004
Uhrbrand K, Koponen IK, Schultz AC, Madsen AM (2017) Evaluation of air samplers and filter materials for collection and recovery of airborne norovirus. J Appl Microbiol. https://doi.org/10.1111/jam.13588
Unterwurzacher V (2016) Sampling methods of airborne mold—limits and strengths in qualitative and quantitative analysis
Unterwurzacher V, Pogner C, Berger H, Strauss J, Strauss-Goller S, Gorfer M (2018) Validation of a quantitative PCR based detection system for indoor mold exposure assessment in bioaerosols. Environ Sci Process Impacts 20(10):1454–1468. https://doi.org/10.1039/c8em00253c
Wang Z, Reponen T, Willeke K, Grinshpun SA (1999) Survival of bacteria on respirator filters. Aerosol Sci Technol 30(3):300–308. https://doi.org/10.1080/027868299304651
Wang RT, Grinshpun SA, Grny RL, Willeke K (2001) Effect of sampling time and air humidity on the bioefficiency of filter samplers for bioaerosol collection. J Aerosol Sci 32(5):661–674. https://doi.org/10.1016/S0021-8502(00)00108-7
Wang CH, Chen BT, Han BC, Liu ACY, Hung PC, Chen CY, Chao HJ (2015) Field evaluation of personal sampling methods for multiple bioaerosols. PLoS ONE 10:1–19. https://doi.org/10.1371/journal.pone.0120308
Wu Y, Shen F, Yao M (2010) Use of gelatin filter and BioSampler in detecting airborne H5N1 nucleotides, bacteria and allergens. J Aerosol Sci 41(9):869–879. https://doi.org/10.1016/j.jaerosci.2010.05.006
Yamamoto N, Bibby K, Qian J, Hospodsky D, Rismani-Yazdi H, Nazaroff WW, Peccia J (2012) Particle-size distributions and seasonal diversity of allergenic and pathogenic fungi in outdoor air. ISME J 6:1801–1811. https://doi.org/10.1038/ismej.2012.30
Yao M, Mainelis G (2006) Investigation of cut-off sizes and collection efficiencies of portable microbial samplers. Aerosol Sci Technol 40:595–606. https://doi.org/10.1080/02786820600729146
Yoo K, Lee TK, Choi EJ, Yang J, Shukla SK, Hwang SI, Park J (2017) Molecular approaches for the detection and monitoring of microbial communities in bioaerosols: a review. J Environ Sci China 51:234–247. https://doi.org/10.1016/j.jes.2016.07.002
Zahradnik E, Raulf M (2019) Development of mold antigen-specific enzyme-linked immunosorbent assays (ELISA) to quantify airborne antigen exposure. Methods Mol Biol 2020:115–130. https://doi.org/10.1007/978-1-4939-9591-2_8
Zhen H, Krumins V, Fennell DE, Mainelis G (2018) Analysis of airborne microbial communities using 16S ribosomal RNA: potential bias due to air sampling stress. Sci Total Environ 621:939–947. https://doi.org/10.1016/j.scitotenv.2017.10.154
Acknowledgements
The authors wish to express their gratitude to Dragana Bandian from Austrian Institute of Technology GmbH (AIT) and the AQA team for technical support and sampling assistance. This work was supported by the Lower Austria Corporation for Research and Education (NFB; Grant LS12-011) and by the DGUV together with the AUVA (Grant FF-FP-0337). Special thanks to the occupants of the site, where sampling was conducted.
Funding
Open access funding provided by University of Natural Resources and Life Sciences Vienna (BOKU). This work was supported by the Lower Austria Corporation for Research and Education (NFB; Grant LS12-011) and by the DGUV together with the AUVA (Grant FF-FP-0337).
Author information
Authors and Affiliations
Contributions
Study design: SSG, MG, SB, CP, GT, and JS. Performed research: VU, MB, CP, AK, and HB. Analysis of data: SB, GT, HB, SSG, CP, JS, and MG. Writing of manuscript: VU, SSG, CP, GT, HB, JS, and MG. All authors read and approved the manuscript.
Corresponding author
Ethics declarations
Conflict of interest
The authors declare a potential conflict of interest as a member of the company that may in future bring the sampling system to market is a coauthor of this manuscript. All authors declare that this fact, however, has not influenced the design or interpretation of the results and that the research was conducted in the absence of any commercial or financial interest.
Ethics approval
Not applicable.
Consent to participate
Not applicable.
Consent for publication
Not applicable.
Research involving human and animal participants
This article does not contain any studies with human participants or animals performed by any of the authors.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Unterwurzacher, V., Bruck, S., Biedermann, M. et al. Development and Validation of a Simple Bioaerosol Collection Filter System Using a Conventional Vacuum Cleaner for Sampling. Aerosol Sci Eng 5, 404–418 (2021). https://doi.org/10.1007/s41810-021-00110-9
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s41810-021-00110-9