Abstract
Blackleg (stem canker) of crucifers is a globally important disease caused by multiple genetic subclades of the fungi Plenodomus lingam (syn.: Leptosphaeria maculans) and Plenodomus biglobosus (syn.: Leptosphaeria biglobosa). In our study, we monitored the geographical distribution of these two pathogen species from rapeseed growing areas in Hungary. Multiplex PCR identified 48.7% of the isolates as Plenodomus biglobosus, which indicates the non-recent introduction of the pathogen into Hungary. In addition, multi-locus analysis revealed low genetic diversity within the species, as all isolates were clustered to the Plenodomus lingam ‘brassicae’ and Plenodomus biglobosus ‘brassicae’ subclades. The low genetic diversity of a population generally means reduced adaptation potential, which is essential information in breeding and in developing more effective management strategies.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Oilseed rape (Brassica napus L.) is one of the world’s most important oilseed crops. Stem canker also known as ‘blackleg’ of Brassica crops is the most damaging disease causing yield loss worldwide (Fitt et al. 2008). Stem canker is caused by two closely related fungal species sharing a similar life pattern: Plenodomus lingam (Tode) Desm. and Plenodomus biglobosus (Shoemaker & H. Brun) Gruyter, Aveskamp & Verkley (Dilmaghani et al. 2009). Their virulence differs significantly, with the greater yield loss being attributed to P. lingam (Williams and Fitt 1999). In the early 2000s, the species were distinguished by morphological differences of pseudothecia and classified in the L. genus (Shoemaker and Brun 2001), but later studies suggested that they rather belong to the P. genus (de Gruyter et al. 2012; Wijayawardene et al. 2014).
In Hungary, only P. lingam was previously reported as the causal agent of blackleg, but the presence of P. biglobosus was also confirmed a few years ago (Bagi et al. 2020). Despite some divergence, the pathogens cannot always be distinguished by morphological characteristics (Williams and Fitt 1999). DNA-based identification is required for reliable identification (Rouxel et al. 2004) and subclade differentiation. Plenodomus lingam can be divided into two subclades, whereas P. biglobosus includes seven distinct subclades (Mendes-Pereira et al. 2003; Vincenot et al. 2008; Zou et al. 2019). The subclade classification is based on geographical distribution, natural host range and phylogenetic analysis (Zou et al. 2019). The Plenodomus isolates previously reported from oilseed rape belong to P. lingam ‘brassicae’ (Mendes-Pereira et al. 2003), P. biglobosus ‘brassicae’ (Liu et al. 2014), ‘canadensis’ (Van de Wouw et al. 2008; Dilmaghani et al. 2009), ‘australensis’ (Plummer et al. 1994; Voigt et al. 2005) and ‘occiaustralensis’ (Vincenot et al. 2008; Dilmaghani et al. 2009) subclades. As pointed out by King and West (2022), the distribution patterns of P. biglobosus subclades need to be mapped because P. biglobosus is becoming an increasingly important pathogen of oilseed rape. The rDNA sequences, partial small subunit nrDNA (18S, SSU) and partial large subunit nrDNA (28S, LSU) are highly conserved and can discriminate at the levels of orders and kingdoms (Balesdent et al. 1998; de Gruyter et al. 2009). Molecular analysis of the ITS regions and the 5.8S rRNA gene has been used for many years to estimate diversity and classify Ascomycota fungi to species level (White et al. 1990; Capote et al. 2012). In the subclade related studies, ITS rDNA has been used for the phylogenetic analysis and reliable subclade identification of both species (Zou et al. 2019). Fragments of β-tubulin 2 gene sequences were also involved in the cluster analysis (Mendes-Pereira et al. 2003), but these sequences have not been published in international databases (NCBI, ENA). In order to distinguish closely related Plenodomus species, the RNA polymerase II second largest subunit (rpb2) region, which is more informative and variable than the ITS region, was also analyzed (Chen et al. 2015). This region can indicate the differences at species level (Drehmel et al. 2008).
We aimed to monitor and determine the distribution of the P. lingam and the newly reported P. biglobosus in Hungary, in order to estimate the arising importance of the fungi in Central Europe. To elucidate the genetic diversity within the Plenodomus population infecting oilseed rape in Hungary, our goal was to observe the phylogenetic relationship among the Plenodomus isolates by multi-locus sequence analysis of ITS1-5.8S-ITS2 region, partial sequences of the 28S nrDNA (LSU), the β-tubulin 2 gene (tub2) and the RNA polymerase second largest subunit (rpb2 region) and provide available sequence data in the NCBI.
Materials and methods
Collection of Plenodomus isolates
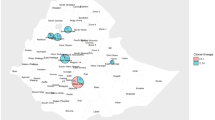
Leaf and stem tissues of oilseed rape with blackleg symptoms were collected for the survey in major producing counties across Hungary in commercial fields in 2017–2021. The fungus was induced to sporulate by incubating unsterilized, diseased tissues in humidity chambers during 2–3 days to allow cirri exudation from pycnidia. After formation of pycnidiospores, the cirrus was transferred with a sterile glass needle (Goh 1999) and suspended in sterile distilled water. Suspensions of conidia were transferred to PDA (potato dextrose agar, BioLab Zrt., Hungary) plates and incubated at 24 ± 1 °C temperature for 3–4 days. The cultures were purified by single-spore colony isolation. Growing hyphal tips from germinating conidia were transferred onto fresh PDA plates with the aid of a sterile dissection needle. Identification was initially based on morphological criteria, and cultures identified as Plenodomus spp. (Mitrovic et al. 2016) were grown on PDA plates and maintained at 4 °C.
Identification and selection of isolates for analysis
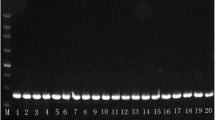
In five years, 308 Hungarian Plenodomus isolates (data not shown) were obtained and identified to species level by multiplex PCR with specific primers: LmacR, LmacF and LbigF (Liu et al. 2006). The rapid and simultaneous detection of species is performed by the length of specific bands of the target sequence: 331 bp for P. lingam isolates, and 444 bp for P. biglobosus isolates.
For the detailed phylogenetic analysis and comparison, 26 P. lingam and 17 P. biglobosus isolates were selected from different locations (Table 1). We aimed to include at least one P. lingam and one P. biglobosus isolate from each studied county. For two counties, this was not possible because we could not identify P. biglobosus at all. From these counties, two P. lingam isolates were chosen for the analysis.
DNA isolation, amplification and sequencing
Hyphae were collected from a PDA plate of each isolate. Then, genomic DNA was extracted using the cetyl-trimethyl-ammonium-bromide (CTAB) method (Maniatis et al. 1983), followed by chloroform/isoamyl alcohol (24:1, v/v) extraction and isopropanol precipitation. The concentration and the purity of the DNA were evaluated by NanoDrop™ Spectrophotometer.
The PCR-based assay was conducted by amplifying the 18S-28S rRNA region (a partial sequence of the 18S ribosomal gene, the ITS1 region, the 5.8S ribosomal gene, the ITS2 region and a partial sequence of the 28S ribosomal gene), other section of 28S nrDNA (LSU), part of the tub2 gene and partial rpb2 region. Amplifications of fragments were carried out in a total reaction volume of 50 μL containing 15 ng of genomic DNA, 0.2–0.2 µM forward and reverse primers, DreamTaq Green PCR Master Mix (2X) (Thermo Scientific™).
To amplify the 18S-28S region, PN3 and PN10 primers and PCR conditions were performed according Mitrovic et al. 2016. The length of the target sequence without primers was 507 bp for P. lingam and 535 bp at P. biglobosus. The LSU region was amplified with the primers LR0R and LR7 (Rehner and Samuels 1994) according to the protocol of Chen et al. 2015. The target sequence length without the primers was 1328 bp for P. lingam and 1327 bp for P. biglobosus. The part of the tub2 gene was amplified with the primer pair Btub2Fd and Btub4Rd (Woudenberg et al. 2009) by amplification conditions of Chen et al. 2015. The target sequence length without primers was 345 bp for P. lingam and 337 bp for P. biglobosus. To amplify partial region of rpb2, RPB2F (5′-AGGCTTGTGGTTTGGTCAAGA-3′) and the RPB2R (5′-ATCATAGCRGTCTCTTCCTCCT-3′), we designed primers based on sequences from the P. lingam rpb2 region (Accession Nos. DQ470894; KT389669; KY064047; XM_003841144) and P. biglobosus rpb2 region (Accession Nos. KY064037; MT683512). For rpb2 region, thermal cycling conditions consisted of denaturation at 95 °C for 3 min, followed by 35 cycles of the following steps: denaturation at 95 °C for 30 s, annealing at 58 °C for 30 s and extension at 72 °C for 45 s, with a final extension step at 72 °C for 10 min. The target sequence length for both species was 492 bp without primers.
Amplification products were analyzed first by gel electrophoresis on 1% agarose gel (Sigma) stained with ECO Safe Nucleic Acid Staining Solution (Biocenter), then visualized under UV light. Amplicons were purified by High Pure PCR Product Purification Kit (Roche) according to the manufacturer’s protocol. Fragments of the 18S-28S were sequenced by PN10 primer, fragment of the LSU, fragment of the tub2 gene, fragment of the rpb2 region were sequenced in both directions using the primers for PCR, in an ABI Prism automatic sequencer (BaseClear B.V.). Sequences were manually checked, edited and compared to reference sequences deposited in the NCBI BLAST database (Altschul et al. 1999).
Phylogenetic analyses of the P. lingam and P. biglobosus isolates
Further phylogenetic analyses were carried out with the MEGA11 program package (Tamura et al. 2021). The trees were obtained by applying neighbor-joining algorithms to matrixes of pairwise distances estimated using the maximum composite likelihood (MCL) approach (Tamura et al. 2004). The tree is drawn to scale, with branch length measured by the number of substitutions per site. The clade stability was assessed with 1000 replicates of bootstrap values, and Leptosphaeria doliolum (CBS 505.75) was designated as the outgroup in all rooted trees.
Sequences of the ITS regions including the 5.8S gene of rDNA used for subclade identification (Mendes-Pereira et al. 2003) were compared to P. lingam strain CBS 275.63, P. biglobosus ‘brassicae’ UBIP01000001 genome, ITS1-5.8S-ITS2 sequences of the P. lingam subclades ‘brassicae’ (AJ550885; AJ550887), ‘lepidii’ (AJ550890) and the P. biglobosus subclades ‘thlaspi’ (AJ550891), ‘australensis’ (AJ550869; AJ550870), ‘erysimii’ (AJ550872), ‘canadensis’ (AJ550868; FJ172238; AJ550867), ‘occiaustralensis’ (AM410082), ‘americensis’ (MG321243) and ‘brassicae’ (DQ133890). For the multi-locus analysis (ITS1-5.8S-ITS2, partial LSU, partial tub2 gene, partial rpb2 region), the whole-genome isolates (P. lingam ‘brassicae’ CBS 275.63 and P. biglobosus ‘brassicae’ UBIP01000001) were included, because only these two isolates had published sequences from all these regions.
Results
Across all sites and surveillance years, 308 Hungarian Plenodomus isolates were identified: 158 isolates were P. lingam (51.3%), while P. biglobosus was detected in case of 150 isolates (48.7%) (Table 2). The newly reported Plenodomus biglobosus was identified from six new counties, so it can be concluded that the pathogen is widespread and common in Hungary.
Phylogenetic analysis
For phylogenetic analyses, ITS1-5.8S-ITS2 sequences (468 bp for P. lingam and 496 bp for P. biglobosus), partial LSU sequences (877–881 bp for P. lingam and 874–881 bp for P. biglobosus), partial tub2 gene sequences (343–345 bp for P. lingam and 336–337 bp for P. biglobosus) and partial rpb2 region sequences (492–493 bp for P. lingam and 492–493 bp for P. biglobosus) of the examined 43 isolates were generated and deposited in GenBank.
Phylogenetic tree based on ITS1-5.8S-ITS2 region
The ITS1-5.8S-ITS2 sequences of Hungarian isolates were 100% identical in all 26 P. lingam isolates with the reference isolate from UK (JF740234 from CBS275.63 complete genome). Similarly, the ITS1-5.8S-ITS2 sequences of the 17 P. biglobosus isolates were also 100% identical with the reference sequence (UBIP01000001) (Fig. 1). Based on these results, it can be stated that all Plenodomus isolates investigated in the present study could be classified into P. lingam ‘brassicae’ and P. biglobosus ‘brassicae’ subclades. This region is highly conserved, as it has been determined by Mendes-Pereira et al. (2003).
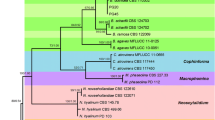
Phylogenetic neighbor-joining tree based on ITS1-5.8S-ITS2 sequences of 26 isolates of Plenodomus lingam and 17 isolates of Plenodomus biglobosus from Hungary and isolates from NCBI database. The numbers are the percent bootstrap support for 1000 resampling and evolutionary analyses conducted in MEGA11. Leptosphaeria doliolum (CBS 505.75) was used as the outgroup
Phylogenetic tree based on multi-locus phylogenetic analysis
Sequences of the P. lingam isolates: the partial tub2 gene sequences were 99.42–100%, the partial LSU sequences 99.43–100% and the partial rpb2 region sequences 98.37–100% identical with the reference strain sequences (CBS 275.63). Sequences of the P. biglobosus isolates: the partial tub2 gene sequences were 99.70–100%, the partial LSU sequences 99.09–100% and the partial rpb2 region sequences 100% identical with the reference strain sequences (UBIP01000001) (Fig. 2). Noteworthy, these are the first sequence data in relation to the genes and regions of interest of the P. lingam and P. biglobosus in Hungary.
Phylogenetic tree obtained from the combined partial tub2 gene, ITS1-5.8S-ITS2, partial LSU, partial rpb2 gene sequence alignment of 26 Plenodomus lingam and 17 Plenodomus biglobosus isolates under survey and reference strains. The tree was rooted using Leptosphaeria doliolum (CBS 505.75) as outgroup taxon. Bootstrap support values > 95% are indicated near the nodes
The combined four‐locus data set consisted of 46 isolates with the whole-genome isolates of P. lingam ‘brassicae’ and P. biglobosus ‘brassicae’ and with Leptosphaeria doliolum as the outgroup taxon. In the tree (Fig. 2), the surveyed P. lingam and P. biglobosus isolates were clustered separately to their reference strains with 96% bootstrap support, respectively. Within species, we observed only parsimony-uninformative variability. The low genetic diversity among isolates in this investigation is not surprising, as the isolates belong to the same subclade (Fig. 1).
Discussion
Outbreak of a new pathogen or changes in genetic composition of a population could compromise the efficiency of established plant protection strategies. Plenodomus biglobosus was at first described in 2020 in Hungary (Bagi et al. 2020). The monitoring of new pathogens is of high importance as it can help to optimize breeding strategies and crop protection technologies (Huang et al. 2014). There is a lack of information about the distribution of Plenodomus species causing blackleg of brassicas in Central Europe; therefore, our goal was to identify and describe the local pathogens in oilseed rape cultivation. Based on the occurrence and frequency of P. lingam and P. biglobosus, it can be stated that P. biglobosus is more common and widespread in Hungary than previously thought.
Furthermore, we also tried to investigate the sources of variation in genetic diversity observed in the Hungarian P. lingam and P. biglobosus population. According to some views, in the UK additional genetic subclades may be responsible for the growing importance of P. biglobosus (King and West 2022). Molecular identification and characterization clearly identified the Hungarian P. lingam isolates as members of the P. lingam ‘brassicae’ subclade. This subclade is distributed worldwide and can infect several Brassica species (Mendes-Pereira et al. 2003), while P. lingam ‘lepidii’ subclade has been only isolated from Lepidium sp. from Canada (Mendes-Pereira et al. 2003). Similarly, the analysis of Hungarian P. biglobosus isolates showed that all isolates belong to the P. biglobosus ‘brassicae’ subclade, admittedly. The subclade ‘brassicae’ that infects Brassica species (Mendes-Pereira et al. 2003) is the most widely distributed subclade of P. biglobosus (Liu et al. 2014). Plenodomus biglobosus ‘canadensis’ is the most closely related subclade to P. biglobosus ‘brassicae’ and has been isolated from oilseed rape and Chinese mustard (Van de Wouw et al. 2008; Dilmaghani et al. 2009). Plenodomus biglobosus ‘australensis’ (Voigt et al. 2005), ‘occiaustralensis’ (Vincenot et al. 2008) and ‘americensis’ (Zou et al. 2019) subclades can also infect Brassica species, incl. oilseed rape, while other subclades (Mendes-Pereira et al. 2003) ‘thlaspi’ (obtained from Thlaspi arvense) and ‘erysimii’ (isolated from Erysimum sp.) have not been reported from brassicas yet.
Molecular methods have been used in taxonomic studies of Plenodomus to reveal phylogenetic relationship among the species and subclades (Zou et al. 2019). Combined DNA phylogenetic analysis based on ITS, 28S nrDNA (LSU) and β-tubulin, sequences are often used to reconstruct these relationships.
Hungarian P. lingam and P. biglobosus isolates, based on four DNA regions, resulted consistent phylogenetic trees and the sequences were extremely similar to each other. The low molecular diversity among the P. biglobosus isolates also suggests that the pathogen was introduced to Hungary much earlier than it was first identified. Most likely its emergence has remained hidden due to very similar symptoms. These fungi coexist in hosts and cause leaf lesions, in addition P. lingam is associated with basal stem canker, while P. biglobosus rather causes upper stem lesions (Eckert et al. 2010; Sprague et al. 2017). Low genetic diversity in a population is more likely to mean reduced adaptation potential, which may also provide important information on the risk of fungicide resistance development.
Worldwide large-scale monitoring and surveys are required to prevent the extreme economic losses. Our results indicate that for both pathogens only the ‘brassicae’ subclades are present in the Hungarian populations at the moment. As a result of globalization, there is a risk that additional subclades will emerge in Hungary on oilseed rape, but in the meantime, it can be concluded that the importance of blackleg pathogens will not change in the near future.
References
Altschul SF, Gish W, Miller W, Myers EW, Lipman DJ (1999) Basic local alignment search tool. J Mol Biol 215:403–410. https://doi.org/10.1016/S0022-2836(05)80360-2
Bagi B, Csaba N, Tóth A, Palkovics L, Petróczy M (2020) Plenodomus biglobosus on oilseed rape in Hungary. Phytopathol Mediterr 59(2):345–351. https://doi.org/10.14601/Phyto-11099
Balesdent MH, Jedryczka M, Jain L, Mendes-Pereira E, Bertrandy J, Rouxel T (1998) Conidia as substrate for internal transcribed spacer-based PCR identification of components of the Leptosphaeria maculans species complex. Phytopathology 88:1210–1217. https://doi.org/10.1094/PHYTO.1998.88.11.1210
Capote N, Pastrana AM, Aguado A, Sánchez-Torres P (2012) Molecular tools for detection of plant pathogenic fungi and fungicide resistance. Plant Pathol. https://doi.org/10.5772/38011
Chen Q, Jiang JR, Zhang GZ, Cai L, Crous PW (2015) Resolving the Phoma enigma. Stud Mycol 82:137–217. https://doi.org/10.1016/j.simyco.2015.10.003
de Gruyter J, Aveskamp MM, Woudenberg JHC, Verkley GJM, Groenewald JZ, Crous PW (2009) Molecular phylogeny of Phoma and allied anamorph genera: towards a reclassification of the Phoma complex. Mycol Res 113(4):508–519. https://doi.org/10.1016/j.mycres.2009.01.002
de Gruyter J, Woundenberg JHC, Aveskamp MM, Verkley GJM, Groenewald JZ, Crous PW (2012) Redisposition of Phoma-like anamorphs in Pleosporales. Stud Mycol 75:1–36. https://doi.org/10.3114/sim0004
Dilmaghani A, Balesdent MH, Didier JP, Wu C, Davey J, Barbetti MJ, Li H, Moreno-Rico O, Phillips D, Despeghel P, Vincenot L, Gout L, Rouxel T (2009) The Leptosphaeria maculans –Leptosphaeria biglobosa species complex in the American continent. Plant Pathol 58:1044–1058. https://doi.org/10.1111/j.1365-3059.2009.02149.x
Drehmel D, James T, Vilgalys R (2008) Molecular phylogeny and biodiversity of the boletes. Fungi 1(4):17–23
Eckert EM, Rossall S, Selley A, Fitt BDL (2010) Effects of fungicides on in vitro spore germination and mycelial growth of the phytopathogens Leptosphaeria maculans and L. biglobosa (Phoma stem canker of oilseed rape). Pest Manag Sci 66(4):396–405. https://doi.org/10.1002/ps.1890
Fitt BDL, Hu BC, Li ZQ, Liu SY, Lange RM, Kharbanda PD, Butterworth MH, White RP (2008) Strategies to prevent spread of Leptosphaeria maculans (phoma stem canker) onto oilseed rape crops in China; costs and benefits. Plant Pathol 57:652–664. https://doi.org/10.1111/j.1365-3059.2008.01841.x
Goh TK (1999) Single-spore isolation using a hand-made glass needle. Fungal Divers 2:47–63
Huang YJ, Karandeni-Dewage CS, Fitt BDL (2014) Importance of Leptosphaeria biglobosa as a cause of phoma stem canker on winter oilseed rape in the UK. Asp Appl Biol 127:117–122
King KM, West JS (2022) Detection of the Phoma pathogens Plenodomus biglobosus subclades ‘brassicae’ and ‘canadensis’ on wasabi, and ‘canadensis’ in Europe. Eur J Plant Pathol 162:751–756. https://doi.org/10.1007/s10658-021-02428-z
Liu SY, Liu Z, Fitt BDL, Evans N, Foster SJ, Huang YJ, Latunde-Dada AO, Lucas JA (2006) Resistance to Leptosphaeria maculans (phoma stem canker) in Brassica napus (oilseed rape) induced by L. biglobosa and chemical defence activators in field and controlled environments. Plant Pathol 55:401–412. https://doi.org/10.1111/j.1365-3059.2006.01354.x
Liu Z, Latunde-Dada AO, Hall AM, Fitt BDL (2014) Phoma stem canker disease on oilseed rape (Brassica napus) in China is caused by Leptosphaeria biglobosa ‘brassicae.’ Eur J Plant Pathol 140:841–857. https://doi.org/10.1007/s10658-014-0513-7
Maniatis T, Sambrook J, Fritsch EF (1983) Molecular cloning: a laboratory manual. Cold Spring Harbor Laboratory, Cold Spring Harbor, New York
Mendes-Pereira E, Balesdent MH, Hortense B, Rouxel T (2003) Molecular phylogeny of the Leptosphaeria maculans –L. biglobosa species complex. Mycol Res 107:1287–1304. https://doi.org/10.1017/S0953756203008554
Mitrovic P, Jeromela AM, Trkulja V, Milovac Z, Terzic S (2016) The First Occurrence of Stem Canker on Oilseed Rape Caused by Leptosphaeria biglobosa in Serbia. Ratarstvo I Povrtarstvo 53(2):53–60. https://doi.org/10.5937/ratpov53-8997
Plummer KM, Dunse K, Howlett BJ (1994) Non-aggressive strains of the blackleg fungus, Leptosphaeria maculans, are present in Australia and can be distinguished from aggressive strains by molecular analysis. Aust J Bot 42:1–8. https://doi.org/10.1071/BT9940001
Rehner SA, Samuels GJ (1994) Taxonomy and phylogeny of Gliocladium analysed from nuclear large subunit ribosomal DNA sequences. Mycol Res 98:625–634. https://doi.org/10.1016/S0953-7562(09)80409-7
Rouxel T, Mendes-Pereira E, Brun H, Balesdent MH (2004) Species complex of fungal phytopathogens: the Leptosphaeria maculans–L. biglobosa case study. In: Sharma AK, Sharma A (eds) Plant genome: biodiversity and evolution, vol 2. Science Publishers, Inc., Enfield, pp 33–75
Shoemaker RA, Brun H (2001) The teleomorph of the weakly aggressive segregate of Leptosphaeria maculans. Can J Bot 79:412–419. https://doi.org/10.1139/b01-019
Sprague SJ, Marcroft SJ, Lindbeck KD, Ware AH, Khangura RK, Van de Wouw AP (2017) Detection, prevalence and severity of upper canopy infection on mature Brassica napus plants caused by Leptosphaeria maculans in Australia. Crop Pasture Sci 69(1):65–78. https://doi.org/10.1071/CP17140
Tamura K, Nei M, Kumar S (2004) Prospects for inferring very large phylogenies by using the neighbor-joining method. Proc Natl Acad Sci USA 101(30):11030–11035. https://doi.org/10.1073/pnas.0404206101
Tamura K, Stecher G, Kumar S (2021) MEGA11: molecular evolutionary genetics analysis version 11. Mol Biol Evol 38:3022–3027. https://doi.org/10.1093/molbev/msab120
Van de Wouw AP, Thomas VL, Cozijnsen AJ, Marcroft SJ, Salisbury PA, Howlett BJ (2008) Identification of Leptosphaeria biglobosa ‘canadensis’ on Brassica juncea stubble from northern New South Wales, Australia. Australas Plant Dis Notes 3:124–128. https://doi.org/10.1007/BF03211265
Vincenot L, Balesdent MH, Li H, Barbetti MJ, Sivasithamparam K, Gout L, Rouxel T (2008) Occurrence of a new subclade of Leptosphaeria biglobosa in Western Australia. Phytopathology 98:321–329. https://doi.org/10.1094/PHYTO-98-3-0321
Voigt K, Cozijnsen AJ, Kroymann J, Pöggeler S, Howlett BJ (2005) Phylogenetic relationships between members of the crucifer pathogenic Leptosphaeria maculans species complex as shown by mating type (MAT1-2), actin, and β-tubulin sequences. Mol Phylogenet Evol 37:541–557. https://doi.org/10.1016/j.ympev.2005.07.006
White TJ, Bruns T, Lee S, Taylor J (1990) Amplification and direct sequencing of fungal ribosomal RNA genes for phylogenetics. In: Innis MA, Gelfand DH, Sninsky JJ, White TJ (eds) PCR protocols: a guide to methods and applications, 1989. Academic Press, Inc., San Diego, pp 315–322
Wijayawardene NN, Crous PW, Kirk PM, Hawksworth DL, Boonmee S, Braun U, Dai D-Q et al (2014) Naming and outlike of Dothideomycetes—2014 including proposals for the protection or suppression of generic names. Fungal Divers 69:1–55. https://doi.org/10.1007/s13225-014-0309-2
Williams RH, Fitt BDL (1999) Differentiating A and B groups of Leptosphaeria maculans, causal agent of stem canker (blackleg) of oilseed rape. Plant Pathol 48:161–175. https://doi.org/10.1046/j.1365-3059.1999.00333.x
Woudenberg JHC, Aveskamp MM, de Gruyter J, Spiers AG, Crous PW (2009) Multiple Didymella teleomorphs are linked to the Phoma clematidina morphotype. Persoonia 22:56–62. https://doi.org/10.3767/003158509X427808
Zou Z, Zhang X, Parks P, du Toit LJ, Van de Wouw AP, Dilantha Fernando WG (2019) A new subclade of Leptosphaeria biglobosa identified from Brassica rapa. Int J Mol Sci 20(7):1668. https://doi.org/10.3390/ijms20071668
Acknowledgements
The study was supported by the Hungarian Ministry for Innovation and Technology within the framework of the Thematic Excellence Program 2020. (TKP2020-IKA-12).
Author information
Authors and Affiliations
Corresponding author
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Bagi, B., Palkovics, L. & Petróczy, M. Phylogenetic analysis of Plenodomus lingam and Plenodomus biglobosus isolates in Hungary. J Plant Dis Prot 130, 875–882 (2023). https://doi.org/10.1007/s41348-023-00720-0
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s41348-023-00720-0