Abstract
Vegetation monitoring is important for many applications, e.g., agriculture, food security, or forestry. Optical data from space-borne sensors and spectral indices derived from their data like the normalised difference vegetation index (NDVI) are frequently used in this context because of their simple derivation and interpretation. However, optical sensors have one major drawback: cloud coverage hinders data acquisition, which is especially troublesome for moderate and tropical regions. One solution to this problem is the use of cloud-penetrating synthetic aperture radar (SAR) sensors. Yet, with very different image characteristics of optical and SAR data, an optical sensor cannot be easily replaced by SAR sensors. This paper presents a globally applicable model for the estimation of NDVI values from Sentinel-1 C-band SAR backscatter data. First, the newly created dataset SEN12TP consisting of Sentinel-1 and -2 images is introduced. Its main features are the sophisticated global sampling strategy and that the images of the two sensors are time-paired. Using this dataset, a deep learning model is trained to regress SAR backscatter data to NDVI values. The benefit of auxiliary input information, e.g., digital elevation models, or land-cover maps is evaluated experimentally. After selection of the best model configuration, another experimental evaluation on a carefully selected hold-out test set confirms that high performance, low error, and good level of spatial detail are achieved. Finally, the potential of our approach to create dense NDVI time series of frequently clouded areas is shown. One limit of our approach is the neglect of the temporal characteristics of the SAR and NDVI data, since only data from a single date are used for prediction.
Similar content being viewed by others
Explore related subjects
Discover the latest articles, news and stories from top researchers in related subjects.Avoid common mistakes on your manuscript.
1 Introduction
1.1 Motivation
Vegetation monitoring is an important task and used, for example, in agriculture (Weiss et al. 2020), mangrove forest applications (Wang et al. 2019), and carbon sink estimation (He et al. 2022). For large-scale or global monitoring, space-borne remote sensing is typically utilised, with a frequent preference for multi-spectral optical sensors because of their easy interpretability and well-established analysis techniques.
A particularly common approach to vegetation monitoring is the derivation of the normalised difference vegetation index (NDVI) (Rouse et al. 1974) from the red and infrared spectral bands
NDVI values range from \(-\) 1 to 1. Values below zero indicate water surfaces, and values between 0 and 0.25 typically indicate barren land and above 0.25 vegetation (Filgueiras et al. 2019). The main advantage of the NDVI is the high sensitivity to vegetation photosynthesis and its wide usage in research. Another benefit is its low sensitivity to changes in illumination due to changing sun position (Reed et al. 1994); however, cloud shadows seem to influence the NDVI to some extent (Jiang et al. 2018; Park 2012).
Despite the common use of optical space-borne sensors, they also have a major drawback: they need an unobstructed view of the area of interest, and they rely on an external energy source for illumination. This limits their usefulness as images can only be obtained during the day and from cloud-free areas. This is especially a problem in heavily clouded areas like the tropics where the cloud coverage is above 75% (King Michael et al. 2013). Also, subtropic regions are affected by cloud cover in the rainy season which is often also the single growing season. For example, almost all images of an area in Burkina Faso are at least partly clouded in the rainy season between June and October (Belesova et al. 2017), as shown in Fig. 1.
One solution to this problem is the use of cloud-penetrating radar sensors. Synthetic aperture radar (SAR) sensors are unaffected by clouds or mist and can capture images day or night and provide a spatial resolution similar to or slightly worse than optical sensors (Moreira et al. 2013). These sensors are used for vegetation monitoring or the retrieval of crop parameters (Mandal et al. 2021, pp. 177–228) and especially useful for biomass estimation (Sinha et al. 2015). Furthermore, similar to the analysis of multi-spectral data, different indices are modelled to enable easier analysis of the data, such as the radar vegetation index (RVI) (Kim and van Zyl 2000), the radar forest degradation index (Mitchard et al. 2012), the polarimetric radar vegetation index (Chang et al. 2018), or the dual polarimetric SAR vegetation index (Periasamy 2018), allowing the use of radar data for different applications.
Sentinel-2 RGB images of an area next to the city Gaoua in southern Burkina Faso. Clouds prevent the monitoring of the area between June and October, the rainy and single growing season in that region (Belesova et al. 2017). Completely white images are fully covered with clouds
Even with the advantages of radar data, optical data still have benefits. First of all, literature on vegetation monitoring using optical data is more abundant than with radar data (McNairn and Shang 2016; Xie et al. 2008). There also exists a plethora of established approaches to use NDVI time series for different applications, like cropland mapping (Estel et al. 2016), forest phenological parameter retrieval (Prabakaran et al. 2013), phenological change detection (Verbesselt et al. 2010), or land degradation and regeneration detection (Eckert et al. 2015). This wealth of publications might also be increased be the easy use of optical data. Additionally, longer time series exist of optical data than of radar data. For example, the Landsat programme provides multi-spectral optical data since 1972 (Madry 2013). In contrast, radar data suitable for monitoring over land have only been available since 1991 with the data of the ERS-1 satellite (Alpers et al. 2019). This is further compounded by the fact that radar data of different sensors are harder to combine than optical data of different sensors, where long time series are created from different sensors (Swinnen and Veroustraete 2008).
Due to these issues, combining the advantages of both sensor types by translating radar data to optical vegetation indices would be of great interest. Existing applications based on NDVI images and time series could be improved by making them independent of the cloud coverage and sun illumination. No adjustments or modifications would be needed to have the downstream tasks profit from the SAR data. All existing approaches could be enhanced easily and their performance increased by having more frequent NDVI images and denser, gap-free time series. Examples that would profit from this approach include land-cover classification (Gómez et al. 2016), or biomass estimation (Ali et al. 2017) amongst others.
1.2 Related Work
The NDVI and the SAR backscatter cannot be directly regressed to each other. Whereas the NDVI is influenced by the spectral characteristics, chemical properties, and especially the chlorophyll content of a medium (Gitelson and Merzlyak 1997), the backscattered radar waves are influenced by physical and electrical properties, e.g., roughness, geometry, and the water content of a medium (Moreira et al. 2013). However, the NDVI and radar backscatter are related to each other through a multitude of plant parameters (McNairn and Shang 2016). For example, the leaf area index (LAI) can be derived using NDVI values (for mangroves (Tian et al. 2017), for wheat (Duchemin et al. 2006)) or radar backscatter (for mangroves (Kovacs John et al. 2013), for wheat (Harfenmeister et al. 2019), for corn and wheat (Vreugdenhil et al. 2018)). But also the amount of water in the vegetation can be correlated to both modalities: using NDVI values Castro et al. (2014) showed a correlation between the fuel water content (FWC) of a meadow and Jiang et al. (2009) showed a relationship with the equivalent water thickness (EWT). For radar backscatter, this is shown for the vegetation water content (VWC) for English Rye Grass (Khabbazan et al. 2019), for wheat (Vreugdenhil et al. 2018), and for oilseed-rape, corn, and wheat (Vreugdenhil et al. 2018). A third parameter influencing both radar backscatter and NDVI values is the biomass as shown for radar backscatter for different crops (Harfenmeister et al. 2019; Khabbazan et al. 2019; Vreugdenhil et al. 2018) and for NDVI values (Hansen and Schjoerring 2003). Finally, the land cover is also affecting both modalities as shown for NDVI values (Hansen et al. 2000; Huete 1997; Pettorelli et al. 2005) and radar backscatter (Balzter et al. 2015; Mahdianpari et al. 2017). This clearly shows that NDVI values and the radar backscatter are related to each other through a multitude of different mechanisms.
Some studies also show the relationship between radar backscatter and NDVI values directly. A correlation of NDVI values for croplands with C-Band radar was shown using Sentinel-1 data (Alvarez-Mozos et al. 2021; Holtgrave et al. 2020) and Radarsat-2 data (Jiao et al. 2021; Moreno et al. 2012). For X-Band radar, this correlation was shown using TerraSAR-X data by Baup et al. (2016) and using COSMO-SkyMed data by Capodici et al. (2013) and by Segalini et al. (2014). This relationship was also found for forested areas: Frison et al. (2018) shows a correlation between the NDVI from Landsat 8 and the radar cross ratio of Sentinel-1 data for a temperate mixed forest.
Going beyond a mere correlation, three studies already demonstrated the pixel-wise estimation of NDVI values from radar data. Two studies use random forest (RF) regression for areas in Brazil: Filgueiras et al. (2019) investigate irrigated soybean and maize crops, whereas Santos et al. (2022) study a mixed area with forests, mangroves, savanna, croplands, pastures, and man-made structures. The third study by Scarpa et al. (2018) is focussed on an agricultural area in Burkina Faso and utilises a small convolutional neural network (CNN). All three studies estimate NDVI values derived from optical sensors (Sentinel-2 in Filgueiras et al. 2019; Scarpa et al. 2018, Landsat 8 in Santos et al. 2022). Model input for all studies is the Sentinel-1 SAR backscatter values together with auxiliary data, for example the elevation, land use, land cover, or the month. All models achieved a good performance for their investigated area. However, the investigated areas are small and relatively homogeneous. Since all three studies trained their models with data covering only single regions and lack experiments regarding the transferability of these models to other areas on the Earth, one can safely assume that existing machine learning models are not applicable on a global scale—even though Sentinel-1, -2, and Landsat 8 data would be available worldwide.
Overall this research shows that the estimation of NDVI values using radar backscatter data is feasible, even though the exact mechanisms and principles that allow this estimation are not properly researched yet.
1.3 Contribution
This paper has two main contributions. First, the dataset SEN12TP (Sentinel-1 and -2 images, time paired) is presented. It contains paired Sentinel-1 and -2 imagery with small time separation together with auxiliary data for all regions and land conditions worldwide. This dataset is created using a sophisticated sampling strategy to ensure a balanced and global distribution of the images and enables an accurate translation from radar to optical data regardless of geolocation. SEN12TP can not only be used for the NDVI prediction task outlined in this paper, but also for single-time cloud removal, SAR-to-optical conversion, or optical-to-SAR image-to-image translation.
Second, a globally applicable model to estimate NDVI values from SAR backscatter data is presented and experimentally validated. The importance of the radar backscatter data and auxiliary data on model performance is determined. The evaluation on hold-out test data shows the high performance and good level of spatial detail of the predicted NDVI maps. Additionally, we demonstrate the successful densification of NDVI time series using radar data. Overall, the final model is able to estimate NDVI values despite cloud coverage and to monitor vegetation continuously, even in frequently clouded areas like the tropics. As a universal model, it can be applied to specific areas without the need for adaptation or fine-tuning, e.g., training it further with data of the area of interest.
2 The SEN12TP Dataset
The SEN12TP dataset is created for the training of deep learning models that are supposed to estimate NDVI values from SAR backscatter. The dataset consists of paired imagery from radar and optical satellites. The images have to be of the same area and with only a small difference in acquisition time (on the order of one day), because a large temporal difference weakens the correlation of radar to optical data. This is due to changes in the observed area, for example growing or harvested vegetation. A similar dataset is SEN12MS (Schmitt et al. 2019), but as the image pairs in that dataset can have an acquisition time gap of up to 3 months, it is not suitable to precisely relate SAR backscatter and NDVI values. In the following subsections, first, the selection of the images is described, then the used remote-sensing data characterised, and finally, the resulting dataset analysed. All three steps were done using Google Earth Engine (GEE) (Gorelick et al. 2017).
2.1 Selection of Image Regions
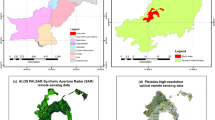
The regions of interest (ROI) for the dataset are selected in a sophisticated manner to create a balanced and geospatially uniformly distributed dataset. The main steps are the selection of candidate points, subsequent filtering, and finally the creation of ROIs from these points. These steps are also depicted in Fig. 2.
To ensure capturing all vegetation conditions globally all land-cover types, climates and seasons have to be taken into account and balanced to ensure an unskewed dataset. Candidate points are selected using the climate zone and land-cover class. For the climate zones, the Köppen–Geiger classification is used (Peel Murray et al. 2007), as it is based on the vegetation found on the ground. To ensure including all land-cover types, the global land-cover product Copernicus Global Land Service (CGLS) 100 m version 3 is chosen (Buchhorn et al. 2021). Both climate zones and land-cover classes are simplified as described in Tables 1 and 2, respectively. Additionally, the CGLS land-cover classes unknown and water bodies and open sea were excluded as well as the polar areas north of \({78}^{\circ }\) N and south of \({60}^{\circ }\) S as they are not vegetated. The land-cover product is used in a downscaled resolution of 2 km to ensure that the surroundings of a sampled point contain pixels of the same class. If the original 100 m resolution of the CGLS classes would be used to sample candidate points for ROIs, there is a chance that only one pixel of the this land-cover class would be in the ROI. For example, a small park in an urban area: it could be selected as candidate for a forest, but in fact, most of the ROI is an urban area. However, choosing a too large of a scale results in missed land-cover types with smaller spatial extents like small villages or forest stands. Therefore, the chosen resolution for sampling is a trade-off between including smaller objects and a perfectly balanced pixel distribution.
From the combination of simplified and downsampled land-cover classes and climate zones, a stratified sample of point is drawn. For each of the 112 combined classes (14 climate zones times 8 land-cover classes), 25 candidate points are selected. As some combinations of climate and land cover like Tropical rainforest with Snow and Ice are not present in the data, only 6754 points could be sampled. From these, all neighbouring points which would lead to overlapping ROIs are excluded, resulting in 4082 points. For each of these filtered points, Sentinel-1 and -2 sensing times between 28 March 2017 and 31 December 2020 which are free of clouds and cover the entire ROI were retrieved. Using the retrieved sensing times, pairs of radar and optical images are selected which were taken within 12 h from each other. This avoids temporal decorrelation between radar and optical values. To incorporate different seasons, the acquisition month of each image pair is used. The distribution of the combination of climate, land cover, and month is highly imbalanced; therefore, samples were randomly selected to ensure a balanced distribution.
With this balanced set of sample points, ROIs are created. For this, a square with an edge length of 20 km is determined around each sample point. This leads to an imbalanced land-cover distribution, as only the centre of an ROI is required to be of the specified land cover. However, choosing ROIs with a smaller size would require a higher number of ROIs when aiming for the same dataset size. As this would become computationally expensive and prohibitively time-consuming using GEE, we refrained from doing so.
All ROIs contain at most 5% clouds and 5% no data areas. For this the cloud probabilities were calculated using the s2cloudless package (Zupanc 2017), which are contained in the GEE data collection COPERNICUS/S2_CLOUD_PROBABILITY. From the cloud probability, the cloud masks are generated with a cloud probability threshold of 20%. Stricter filtering would exclude too many ROIs. Overall 2319 image pairs are selected that cover 484,400 km\(^{2}\) and 1236 distinct areas, given that from some areas, images from several months were selected. Their spatial distribution is shown in Fig. 3.
Global distribution of the selected ROIs, split into training (blue squares) and test regions (red squares). Underlayed are the simplified Köppen–Geiger climate classes. For the colour legend, refer to Table 1
2.2 Remote-sensing Data
Multiple data modalities are selected to be included for SEN12TP with SAR and optical imagery being the main ones. Next to that, a digital elevation model (DEM) and a land-cover map is included, because for both modalities could be shown, that they improve the estimation of NDVI values from SAR backscatter (Santos et al. 2022; Scarpa et al. 2018). All data were downloaded from GEE. Images and bands with a lower resolution than 10 m were upsampled to 10 m using nearest-neighbour upsampling.
The GRD Sentinel-1 product COPERNICUS/S1_GRD provided in the GEE data catalogue consists of \(\sigma ^\circ\) backscatter values in dB. The following preprocessing steps are applied as described in the documentationFootnote 1: apply orbit file, GRD border noise removal, thermal noise removal, and radiometric calibration. The final terrain correction and orthorectification step normalises the radar response as if the response is received from a flat surface on the earth reference ellipsoid. Since the earth is not flat, the terrain, however, influences the radar backscatter (Small 2011). This could negatively impact the relation to the NDVI, due to different radar backscatter values being sensed for the same ground conditions depending on the terrain and radar geometry. To counter this effect, radiometric terrain correction (RTC) can be applied resulting in \(\gamma ^\circ\) backscatter (Small 2011). This RTC is done on GEE using the model optimised for volume scattering on the ground as described in Vollrath et al. (2020). Including uncorrected \(\sigma ^\circ\) as well as the corrected \(\gamma ^\circ\) backscatter allows a comparison of both backscatter values.
As optical data, the Sentinel-2 Level-2A data collection COPERNICUS/S2_SR is used. It contains 12 spectral bands of the Bottom Of Atmosphere (BOA) reflectance where atmospheric effects are corrected using sen2cor (Main-Knorn et al. 2017). Additionally, the cloud probability from COPERNICUS/S2_CLOUD_PROBABILITY is added as a thirteenth band.
The 30 m resolution ALOS World 3D DEM (version 3.2) product (Takaku et al. 2020) is included because of its global coverage and high accuracy (Uuemaa et al. 2020) from the GEE collection JAXA/ALOS/AW3D30/V3_2. As land-cover product, the 10 m ESA WorldCover v100 product from ESA/WorldCover/v100 is selected, because of its high accuracy and resolution as well as its global coverage (Zanaga et al. 2021).
2.3 Final Dataset
Downloading the SAR, optical, DEM, and land-cover data for all ROIs results in dataset size of 221.6GB. One example of the dataset is shown in Fig. 4 to visualise the different data modalities.
Despite the balanced sampling, the distribution is unbalanced: for example, only 3% of all pixels belong to Snow and Ice, but 26.5% belong to Grassland and 25% belong to Trees. The imbalance stems from the selection of points: the sampling points have a balanced land-cover distribution with the surrounding 2 km square area considered. However, the surrounding area of the ROI can be different and in theory only 1% (\(2\,\hbox {km} \times 2\,\hbox {km}\) of \(20 \,\hbox {km} \times 20\,\hbox {km}\)) of each ROI is required to be of this land-cover class.
However, because Grassland, Trees, and Cropland are the most important land covers for vegetation monitoring and also the most common classes in the dataset, this does not pose a problem for our application. If this dataset is used for other applications, this imbalance has to be taken into consideration.
The distribution of sensing times of all optical and radar images is shown in Fig. 5. All sensor platforms have a sun-synchronous orbit, and therefore, the sensing time does not differ much for each platform. Most images from each platform are taken in a 2–4 h range, mostly due to differences between the local time zone and local solar time. Optical images are only acquired during the ascending orbit by the Sentinel-2 platforms, as the best illumination is around noon and images cannot be acquired at night during the descending orbit. Both orbits can be used for sensing radar data: the descending orbit takes data in the morning around 7AM local time, whereas the ascending orbit occurs in the evening around 7PM local time.
This affects the dataset in two ways: First, pairs of optical and radar data cannot have the same sensing time. The minimum time difference of image pairs is around 6 h. Second, diurnal weather cycles affect data differently for ascending and descending radar orbit, mainly due to water on the vegetation. This water changes the radar response without changing the vegetation condition and thereby complicates the retrieval of vegetation parameters from radar data. In the tropics, the water stems from precipitation which is more frequent in the afternoon and evening (Kazuyoshi and Wang 2008) and leads to water throughfall. Dew affects images of areas with moderate climate in the morning (Khabbazan et al. 2019). Therefore, the model has to compensate the effect of water on the vegetation which might cause a significant bias and an imperfect pairing relation of optical and radar data.
The dataset is available on ZenodoFootnote 2. To load the images a PyTorch dataloader is available on GithubFootnote 3.
3 NDVI Estimation from SAR Backscatter
With the SEN12TP dataset created as described in the previous section, a deep learning model to estimate NDVI values from radar backscatter can be trained. The convolutional neural network architecture used for this task is described in Sect. 3.1. After that, the training regime and data preprocessing steps are reported in Sect. 3.2.
3.1 Deep Learning Model
To estimate the NDVI from SAR backscatter data, a slightly adapted U-Net architecture is used (Ronneberger et al. 2015). This is one of the most established architectures for semantic segmentation and pixel-wise regression purposes, and is also commonly used in remote sensing (Yuan et al. 2021). U-Net models consist of two parts: an encoder branch and a decoder branch. The encoder extracts meaningful features, and to do so, reduces the spatial resolution of the input, in our case SAR backscatter values. For this, the spatial neighbourhoods of each pixel is considered using convolution kernels. The decoder calculates the output NDVI values from the extracted and compressed features. To avoid losing spatial detail in the output, skip connections are added between the encoder and decoder which transfer high-frequency details.
The original U-Net used two output channels with no activation function to create probability maps of the background and foreground class. In contrast, we use only one output channel which is passed into a sigmoid activation function. This ensures that the model output \(\hat{y}\) is limited to [0, 1]. To get the predicted NDVI values \(y_{\text {NDVI}}\), this output has to be transformed to \([-1, 1]\) using the equation \(y_{\text {NDVI}}=2(\hat{y} - 0.5)\). Additionally, instead of up-convolution layers, bilinear upsampling with standard convolution layers was used in the decoder. This avoids checkerboard artefacts in the output image (Odena et al. 2016). The model architecture is depicted in Fig. 6. To train the model, the \(\mathcal {L}_1\) loss is utilised, because the amount of spatial detail is better than with the MSE loss (Zhao et al. 2017).
U-Net architecture based on Ronneberger et al. (2015). Features are extracted from the model inputs in the Encoder and transformed by the Decoder to the model output. To extract abstract features, the data size is reduced in the encoder. Spatial details are added in the decoder by the skip connections to end up with a spatially accurate prediction. Unlike the original paper, the feature maps are enlarged by a bilinear upsampling. Conv, BN, and ReLU denote convolutional layer, batch normalisation, and ReLU activation function, respectively. The number of filters a convolutional layer has is shown below each layer, next to it are the width and height wxh
To evaluate the model and compare the prediction with the optical derived values, multiple metrics are used. In the following, \(\hat{y}\) denotes the Sentinel-1-based NDVI prediction, y the Sentinel-2 derived NDVI, and N the number of pixel values which are compared. The mean absolute error
(also \(\mathcal {L}_1\) loss) and the mean squared error
are commonly used for regression tasks in deep learning. The main difference is the effect of outliers in the data to which the MSE is more sensitive at the expense of the distinction of small value differences. The root-mean-squared error (RMSE)
is used to convert the MSE to the same unit as the MAE.
To allow a comparison with Scarpa et al. (2018), the peak signal-to-noise ratio (PSNR), the Pearson correlation coefficient \(\rho\), and the structural similarity index measure (SSIM) (Wang et al. 2004) are also reported. The PSNR is calculated using
with the maximum interval of the data \(MAX_I\), which equals two in our case as the NDVI is inside \([-1, 1]\). The Pearson correlation coefficient \(\rho\) is calculated using the mean of the target values \(m_y\) and of the predictions \(m_{\hat{y}}\)
The SSIM incorporates the luminance, the contrast, and the structure of many windows of two images. For each window x, y of the two images, it is calculated as
where \(\mu _x\), \(\mu _y\) are the pixel sample means, \(\sigma _x^2\), \(\sigma _y^2\) the variances, and \(\sigma _{xy}\) the covariance of the image windows x, y. \(c_1\) and \(c_2\) are factors to ensure the numerical stability. They are calculated using L, the dynamic range of the pixel values and \(k_1\), \(k_2\) as small values as \(c_n=(k_n L)^2\) with \(k_1=0.01\) and \(k_2=0.03\) as default values. To get the final SSIM, the SSIM over all windows is averaged using the mean.
3.2 Training Procedure and Data Preprocessing
To input the data into the model, it has to be preprocessed and normalised which is done in multiple steps.
First, to ensure a fair evaluation, the \(20\,\hbox {km} \times 20\,\hbox {km}\), ROI images are split up spatially into a training, validation, and test set. This spatial split ensures that the model is not trained with images which are also used for evaluation. 80% of the ROI images are used for training, and both validation and test set consist of 10% of the scenes.
For training, the large ROIs are split up into smaller patches with a size of \(256\,\hbox {px}\times 256\,\hbox {px}\). A stride of \({249}\,{\hbox {px}}\) is chosen to split each \({2000}\,\hbox {px}\) scene into \({8\times 8}\) tiles. Then, patches containing clouds or areas with missing or invalid data are removed on the fly, since some scenes contain small clouds or missing data areas. Filtering bad data areas is done in this two step fashion, because creating a perfectly clean dataset with large ROIs is infeasible. Only a small number of tiles are excluded in this step, which shows the high data quality of the dataset: for the train set, only 2172 of the 134 080 (1.6%) tiles of the train set are excluded. After filtering the data, it is clipped to remove outliers and anomalous sensor values. Values exceeding the clipping threshold are set to the threshold value. The multi-spectral channels of the Sentinel-2 data, the VV and VH polarisations of the Sentinel-1 data, and the DEM are clipped to [0, 10000], \([{-25}\,\hbox {dB}, {0}\,\hbox {dB}]\), \([{-32.5}\,\hbox {dB}, {0}\,\hbox {dB}]\), and \([{-450}\,{\hbox {m}}, {9000}\,{\hbox {m}}]\), respectively. The SAR clipping values are the same as in (Meraner et al. 2020), the DEM clipping values are the approximate lowest and highest possible elevation of the earth. The Sentinel-2 values are clipped at 10 000, because this equals a reflectance of 100%. Finally, all values, including the land-cover classification, are normalised to be in the range [0, 1] using min–max-normalisation.
The NDVI is calculated on the fly from the Sentinel-2 images with the red band B4 and the infrared band B8 as in Eq. (1). As the model output is bounded to [0, 1], the NDVI is transformed to be in that interval for loss calculation.
For model training, the Adam optimiser (Kingma and Ba 2014) with an initial learning rate of 0.001 is used. Learning rate scheduling is used and the learning rate multiplied by 0.5 every seven epochs resulting in a learning rate of 0.0005 after seven epochs, 0.000 25 after 14 epochs and so on. Early stopping is used to avoid overfitting by ending training when the performance does not improve anymore. For each epoch, the MAE of the validation set was calculated. When after a patience period of ten epochs no improvement of the MAE was achieved, training was stopped. The batch size is set to 32 and not data augmentation was used.
All models were implemented using Python 3.7, PyTorch 1.10, and PyTorch Lightning 1.3.5. Training was conducted on two GPUs and took approximately 27 min and 38 min per epoch using an NVIDIA RTX A6000 or an NVIDIA Quadro RTX 8000, respectively.
4 Experiments
The model is trained with the SEN12TP dataset and the results are evaluated. The usefulness of the radar backscatter and auxiliary data for NDVI estimation are investigated and quantitative results presented in Sect. 4.1. Then, in Sect. 4.2, the radar-derived NDVI is qualitatively evaluated for different example regions. To compare this model with existing research, its performance is compared with another study in Sect. 4.3. Finally, the model is applied to a time series of radar images and the optical and radar-derived NDVI time series compared in Sect. 4.4.
4.1 Backscatter and Auxiliary Input Performance
To compare the importance of the radar backscatter and auxiliary input data for NDVI estimation, models trained with different inputs are compared. For this, the predictions of the SEN12TP test set are calculated and compared to the actual optical NDVI using different metrics (cf. Sect. 3.1). This allows a fair evaluation of the model performance, as these scenes are not used during training or validation and contain completely new and unseen data for the model. For this evaluation, the scenes were not split up into smaller patches like in during training to avoid border artefacts. A batch size of 1 had to be used, as only one \({2000\times 2000}\,{\hbox {px}}\) large image fit into the GPU memory. Each model was trained five times with different splits of the dataset to get the average model performance.
For the first set of experiments, the performance is evaluated using radar backscatter data without (\(\sigma ^\circ _\text {VV}\), \(\sigma ^\circ _\text {VH}\)) and with radiometric terrain correction (\(\gamma ^\circ _\text {VV}\), \(\gamma ^\circ _\text {VH}\)), and the elevation retrieved from the DEM. The elevation data are included here, as it is a requirement for the terrain correction and is therefore available at least for all experiments using \(\gamma ^\circ _\text {VV}\) and \(\gamma ^\circ _\text {VH}\).
All models using both radar polarisations VV and VH achieve a good performance very similar to each other with a low error \(\text {MAE}=0.12\). There are only insignificant performance differences between the different models with the exception of the model trained with (\(\gamma ^\circ _\text {VV}\), \(\gamma ^\circ _\text {VH}\)), which is slightly better, as shown in Table 4. Using only one radar polarisation increases the error, in the case of \(\sigma ^\circ _\text {VH}\) to \(\text {MAE}=0.14\), in case of \(\sigma ^\circ _\text {VV}\) to \(\text {MAE}=0.15\).
For the next set of experiments, auxiliary data in form of the ESA WorldCover landcover map are included. This decreases the error to \(\text {MAE}=0.10\). Similar to to experiments without the land-cover map, adding the DEM or applying radiometric terrain flattening does not improve the performance. Results are given in Table 5.
In the last set of experiments, no backscatter data are used, only the auxiliary data from the ESA WorldCover and DEM. This is meant to provide a lower end baseline. The performance using the WorldCover is \(\text {MAE}=0.15\) and worse than using only backscatter data or using backscatter and land-cover data together. Using only the DEM the performance is even worse with a \(\text {MAE}=0.26\) (see Table 6).
To have an understanding of the performance of our model for different landcovers, we evaluate the performance for each land cover class separately. The model shows a similar performance for almost all of the land-cover classes, with an MAE between 0.10 and 0.11. Only for Open Water, a slightly better performance is achieved (\(\text {MAE}=0.099\)), and for Barren/sparse vegetation, the performance is slightly worse (\(\text {MAE}=0.115\)). The full results are shown in Table 7.
4.2 NDVI Image Prediction
Using the trained model, NDVI images can be predicted for all areas of the world from SAR backscatter. When evaluating the model on the test set of SEN12TP, a low error with a good level of spatial details is achieved, as shown in Fig. 7. Especially for forested, grassland, and cropland areas, the NDVI is predicted with high accuracy. Even some fine details like roads or urban areas can be predicted properly. Despite the generally very good results, some remaining shortages of the predicted NDVI images become apparent. One problem is the loss of some spatial details making the predicted NDVI images seem slightly less sharp than their original counterparts. One example for this phenomenon is the delineation between field plots. All delineations between land-cover types, however, are captured well due to the inclusion of the WorldCover landcover map. Water bodies and wetlands also present a problem, as their NDVI values are predicted with quite large errors.
Comparison of the NDVI calculated from Sentinel-2 data with the Sentinel-1 radar backscatter derived NDVI using the proposed model on images of the hold-out test set. The model uses as input the two Sentinel-1 polarisations \(\sigma ^\circ _\text {VV}\) and \(\sigma ^\circ _\text {VH}\), the ESA WorldCover landcover map, and the ALOS World 3D DEM. Also shown are the RGB representation of the area and the error between optical and radar-derived NDVI. The Sentinel-1 data is displayed as false-colour image (R: \(\sigma ^\circ _\text {VV}\), G: \(\sigma ^\circ _\text {VH}\), B: \(\sigma ^\circ _\text {VV} / \sigma ^\circ _\text {VH}\)). The location of each image is shown on the map above; the number next to each location denotes the row of each prediction image below
4.3 Comparison to a Regional Model
To assess the performance of our approach in comparison to another, previously published approach, we compare it using the same area of Scarpa et al. (2018). This study uses data from a small region in the southwest of Burkina Faso to train their model. Part of this area they designate as test region to evaluate their model performance. Because the training and test data are next to each other, their model is very likely only applicable to that region or similar ones and will likely achieve a suboptimal performance for other regions.
In contrast, our model trained using the globally distributed SEN12TP dataset is globally applicable and does not need fine-tuning for specific regions. The closest image our model has seen during training, validation, or testing has a distance of more than 150 km from the region used by Scarpa et al. (2018).
To compare both approaches, we downloaded the SAR, DEM, and WorldCover data from Google Earth Engine of the area. We did not use the originally provided data out of two reasons: first, we wanted to ensure the same data preprocessing as used for the data to train our models. Second, we noticed a slight shift between SAR data and NDVI label pixels in the provided dataset by approximately half a pixel. We avoided this using the georeferencing of the NDVI images of Scarpa et al. (2018) for the data downloaded by us. The original NDVI images were also used by us to calculate the metric values to ensure a fair evaluation.
A quantitative comparison of our results with the results of Scarpa et al. (2018) is shown in Table 8. Our trained model achieves a higher SSIM for all input modalities. This higher level of spatial detail can also be seen in the predicted NDVI images, together with a lower amount of noise, as shown in Fig. 8. In contrast to our high spatial performance, their model achieves a higher Pearson correlation. For the PSNR, our model is better when using only radar backscatter, but their model is better using backscatter and elevation data. The best-performing model of all possible inputs is our model using backscatter, elevation, and land-cover data. However, this comparison is not fully fair, as Scarpa et al. (2018) did not evaluate the effectiveness of using land-cover information. The results still demonstrate that our model is able to achieve a similar or even better performance than a area-specific model trained for a small region.
Visual comparison of the optical and radar-derived NDVI for the data of Scarpa et al. (2018), located in the Houet province, Burkina Faso, Africa. The optical NDVI is from the 4 June 2016. Both models use the Sentinel-1 radar backscatter from 30 May 2016 together with the elevation data for NDVI estimation. Figures 8b and 8c are taken from Scarpa et al. (2018). The test site mainly consists of three land-cover types:  Cropland,
Cropland,  Grassland, and
Grassland, and  Shrubland. For a full legend of the WorldCover classes, see Fig. 7
Shrubland. For a full legend of the WorldCover classes, see Fig. 7
4.4 Densification of NDVI Time Series
For many applications, time series of vegetated surfaces are more interesting than an image of a single date. Therefore, we predicted the NDVI for all Sentinel-1 images of the area used in Fig. 1 for the year 2019. As model input, we used the \(\sigma ^\circ\) SAR backscatter, the DEM, and the WorldCover data. We predicted the NDVI using the same five models as used for the evaluation which enables us to calculate the mean and standard deviation of the predicted NDVI values. Together with the NDVI retrieved from all Sentinel-2 images, we created multiple time series for several example points with different land covers. The time series show a high correspondence between the mean predicted NDVI values and the optical retrieved values from cloud-free images as can be seen in Fig. 9. The increase of the NDVI during the rainy season between June and October is captured by the radar-derived NDVI predictions. In contrast to the optical values, a higher temporal resolution is achieved, because all images can be used and not only cloud-free ones.
The standard deviation is small for most of the predicted pixels, showing a high agreement and certainty between the models. One lower outlier of the mean predicted NDVI at the beginning of August is especially notable. This outlier is observable for all four examples and is most pronounced for Shurbland where it coincides with a large standard deviation.
NDVI time series for different points of the area in Fig. 1. Shown are NDVI time series derived with our model and calculated from Sentinel-2 optical data of four image pixel with different land-cover classes. Five model predictions were created using the SAR data \(\sigma ^\circ\), the DEM, and the WorldCover as input. The mean of the predicted values is denoted with a diamond, and the bar shows the standard deviation of the models. The land-cover information is taken from the ESA WorldCover v100 map. Cloud covered values are de-emphasised
5 Discussion
5.1 General Validity of the Model
The results summarised in Tables 4 and 5 and illustrated in Fig. 7 show the general high performance of our approach, unbiased by the land-cover type (cf. Table 7). The successful prediction of the NDVI from radar data indicates that there is indeed some relationship between NDVI and radar data despite the different sensing principle and wavelength used. This is in accordance to the research literature which could also relate radar backscatter with biophysical parameters and NDVI values (cf. Sect. 1.2). A relation in the data of both sensors could be exploited to translate the values.
One problem of the predicted NDVI maps is the spatial resolution: the predicted NDVI is blurrier compared to the Sentinel-2 derived NDVI as visible in Fig. 7. This is caused by the lower resolution of the SAR input data. The used Sentinel-1 data have a spatial resolution of \({20\times 22}\,{\hbox {m}}\) (Level-1 product, IW mode, after multi-looking) (Collecte Localisation Satellites 2016), whereas the Sentinel-2 data have a spatial resolution of 10 m (Drusch et al. 2012), even though both products are provided with a 10 m pixel spacing in the GEE data catalogue. Another factor deteriorating the spatial detail is the speckle noise of the radar which makes it hard to predict small objects. Combining these two factors results in a blurrier NDVI of the prediction compared to the optical retrieved NDVI and a loss of fine spatial detail.
Another problem in the predicted NDVI maps are wetlands and water surfaces. In these areas, the predicted NDVI is very imprecise (cf. Fig. 7). One possible reason for this might be the low amount of water pixels in the dataset with 4.9% (cf. Table 3). The interaction of radar waves with water might also be another cause. First, radar waves do not penetrate the water but are reflected at the water surface. Second, the signal energy is reflected away from the sensor, so that only very little energy is received back at the sensor and the weak received signal does not contain any information of what is below the water surface. In contrast, optical light penetrates into the water to some extent and is also reflected back to the sensor by the plants, algae, bacteria, and other particles on the water surface and in the water. Therefore, optical sensors can be used to retrieve meaningful information about the water composition, and the amount of chlorophyll containing species. This drawback of our approach does not impose many restrictions on monitoring vegetation on land which are mainly agricultural or forested areas. To avoid an erroneous retrieval of NDVI values, all water bodies can be masked using the existing water body layers such as the MODIS/Terra Land Water Mask (Carroll et al. 2017). The only problem could be paddy fields, which are similar to wetland areas, as rice is an important crop grown on paddy fields. However, other studies showed the feasibility to monitor paddy rice fields using SAR data (Inoue et al. 2014). Therefore, our model and dataset could be adapted and improved for this special case to allow the retrieval of NDVI values from these areas as well.
5.2 Influence of Auxiliary Model Inputs
The choice of the model inputs changes how accurate the NDVI can be derived, as shown in Tables 4, 5, and 6. The two polarisations of the radar backscatter are the most important input data as they achieve the highest performance when using a single modality. This good performance can be increased when the land-cover map is also fed to the model as auxiliary input. Using the two SAR polarisations separately shows a higher relative importance of the VH polarisation compared to the VV polarisation. The performance of using solely \(\sigma ^\circ _{\text {VV}}\) data is on par with using only the WorldCover as input; however, both achieve only a substandard performance. The low performance when using the land-cover information without the radar backscatter shows that it is only supporting the model, but not being the main information source of the model.
The improved model performance when using the ESA WorldCover can have multiple reasons. One reason could be the influence of a prior knowledge of the underlying land cover. It is likely that different mathematical relationships between SAR backscatter and NDVI exist for different land-cover types, as it is an important feature for NDVI estimation (Santos et al. 2022). For example, high intensities in urban environments will certainly propagate into lower NDVI values than high intensities in croplands. If the model does not have to learn the distinction between the different land-cover classes itself, but receives prior knowledge about them, this leaves more model capacity for the actual implicit regression models. Another reason could be the high resolution and sharp edges included in the land-cover map, which help to add spatial detail, at least at the edges of different land covers.
Adding the DEM as model input does not improve model performance. The radar backscatter is influenced by the elevation and the terrain due to the side looking nature of the sensor. Two identical areas with the same vegetation and land cover will have a different backscatter if the terrain is different (e.g., slope and aspect) (Small 2011). This weakens the relationship between radar backscatter and NDVI values, and should reduce the model performance. However, a similar performance is achieved independent whether these terrain effects are corrected for or not and adding the DEM as model input does not have an effect on model performance either. The reason for this might be that the model is able to learn the influence of the terrain on the backscattered signal and correct for it. This suggests that terrain flattening as preprocessing step or including the DEM as model input might be unnecessary when deep neural networks are used. Additionally, the elevation correlates with the NDVI only very little, as shown in Table 6 by the poor performance when using only the DEM as model input.
5.3 Creation of Dense NDVI Time Series
As shown in Fig. 9, the model prediction seems to be useful for filling gaps due to cloud coverage in NDVI time series of a location. Thereby, at each date, the SAR image is used for NDVI estimation and the NDVI of the prediction is extracted neglecting any information contained in the temporal characteristics of a surface. Another approach to acquire dense NDVI time series would be to first extract a time series of SAR values and then estimate the NDVI of all dates of the series as done for example by Zhao et al. (2020). When comparing these two approaches, ours is more flexible, since only one SAR image is needed and not a whole time series. Therefore, our method enables a real-time application where the NDVI is predicted for the most recent image and is not only applicable in retrospect when the whole time series is acquired and NDVI prediction is carried out in a sequence-to-sequence manner.
The standard deviation of the NDVI predictions of the different models could be an indicator of the certainty of the estimations. It could be used to detect and exclude outliers and predictions with a low confidence which would increase the overall quality of the time series and avoid wrongly predicted NDVI values.
6 Summary and Conclusion
This study demonstrates the feasibility to translate SAR backscatter values to NDVI values with one globally applicable model. This allows gap-free vegetation monitoring even in heavily clouded areas worldwide. We also showcased the densification of NDVI time series using the predicted values as an outlook experiment. To achieve this, we created a global and balanced dataset of radar and optical imagery with a maximum time difference of 12 h together with auxiliary data. The dataset consists of images of all climates, land-cover types, and months of the year. It is used to train a modified U-Net architecture to translate radar backscatter values to NDVI values. Next to the Sentinel-1 C-Band SAR data, the ALOS World 3D DEM and the ESA WorldCover v100 land-cover product are used to improve the accuracy and spatial detail. The model reaches a high performance and low error (\(\text {MAE}=0.10\)) with a good amount of spatial detail (\(\text {SSIM}=0.61\)). The comparison with a model trained with local data of a small region (Scarpa et al. 2018) shows a similar performance of our model for that region even without any fine-tuning or region-specific adaptions. We also show that NDVI time series can be retrieved from the model prediction which are similar to time series from the optical data but with a higher temporal resolution. Whilst our approach neglects information contained in the temporal behaviour of a surface, its application only needs an SAR image of a single date allowing an easy application.
Availability of data and Materials
The dataset SEN12TP is available at https://zenodo.org/record/7342059, code for a PyTorch dataloader is available at https://github.com/oceanites/sen12tp.
References
Ali I, Cawkwell F, Dwyer E et al (2017) Modeling managed grassland biomass estimation by using multitemporal remote sensing data—a machine learning approach. IEEE J Sel Top Appl Earth Obs Remote Sens 10(7):3254–3264
Alpers W, Liu AK, Wu SY (2019) Satellite remote sensing SAR. In: Kirk Cochran J, Bokuniewicz HJ, Yager PL (eds) Encyclopedia of Ocean sciences. Elsevier, pp 429–442
Alvarez-Mozos Jesús, Villanueva J, Arias Maria et al (2021) Correlation between NDVI and sentinel-1 derived features for maize. In: Proc. of IEEE international geoscience and remote sensing symposium IGARSS. IEEE, pp 6773–6776
Balzter H, Cole B, Thiel C et al (2015) Mapping CORINE land cover from sentinel-1A SAR and SRTM digital elevation model data using random forests. Remote Sens 7(11):14876–14898
Baup F, Villa L, Fieuzal R et al (2016) Sensitivity of X-Band (\(^{0}, \gamma\)) and optical (NDVI) satellite data to corn biophysical parameters. Adv Remote Sens 05(02):103–117
Belesova K, Gasparrini A, Sié A et al (2017) Household cereal crop harvest and children’s nutritional status in rural Burkina Faso. Environ Health 16(1):65
Buchhorn M, Smets B, Bertels L et al (2021) Copernicus global land service: land cover 100 m: version 3 Globe 2015-2019: Product User Manual. Version Dataset v3.0, doc issue 3.4
Capodici F, D’Urso G, Maltese A (2013) Investigating the relationship between X-Band SAR data from COSMO-SkyMed satellite and NDVI for LAI detection. Remote Sens 5(3):1389–1404
Carroll M, DiMiceli C, Wooten M et al (2017) MOD44W MODIS/terra land water mask derived from MODIS and SRTM L3 global 250 m SIN grid V006
Castro M, Parra JC, Morales LJ et al (2014) Establishment of empirical relations between fuel moisture content and the normalised difference vegetation index. J Soil Sci Plant Nutr 14(3):670–675
Chang JG, Shoshany M, Oh Y (2018) Polarimetric radar vegetation index for biomass estimation in desert fringe ecosystems. IEEE Trans Geosci Remote Sens 56(12):7102–7108
Collecte Localisation Satellites (2016) Sentinel-1 product definition
dos Santos EP, da Silva DD, do Amaral CH et al (2022) A machine learning approach to reconstruct cloudy affected vegetation indices imagery via data fusion from Sentinel-1 and Landsat 8. Comput Electron Agric 194:106753
Drusch M, Del Bello U, Carlier S et al (2012) Sentinel-2: ESA’s optical high-resolution mission for GMES operational services. Remote Sens Environ 120:25–36
Duchemin B, Hadria R, Erraki S et al (2006) Monitoring wheat phenology and irrigation in Central Morocco: on the use of relationships between evapotranspiration, crops coefficients, leaf area index and remotely-sensed vegetation indices. Agric Water Manag 79(1):1–27
Eckert S, Hüsler F, Liniger H et al (2015) Trend analysis of MODIS NDVI time series for detecting land degradation and regeneration in Mongolia. J Arid Environ 113:16–28
Estel S, Kuemmerle T, Levers C et al (2016) Mapping cropland-use intensity across Europe using MODIS NDVI time series. Environ Res Lett 11(2):024015
Filgueiras R, Mantovani EC, Daniel A et al (2019) Crop NDVI monitoring based on sentinel 1. Remote Sens 11(12):1441
Frison P-L, Fruneau B, Kmiha S et al (2018) Potential of sentinel-1 data for monitoring temperate mixed forest phenology. Remote Sens 10(12):2049
Gitelson AA, Merzlyak MN (1997) Remote estimation of chlorophyll content in higher plant leaves. Int J Remote Sens 18(12):2691–2697
Gómez C, White JC, Wulder Michael A (2016) Optical remotely sensed time series data for land cover classification: a review. ISPRS J Photogramm Remote Sens 116:55–72
Gorelick N, Hancher M, Mike D et al (2017) Google Earth engine: planetary-scale geospatial analysis for everyone. Remote Sens Environ 202:18–27
Hansen PM, Schjoerring JK (2003) Reflectance measurement of canopy biomass and nitrogen status in wheat crops using normalized difference vegetation indices and partial least squares regression. Remote Sens Environ 86(4):542–553
Hansen MC, Defries RS, Townshend JRG et al (2000) Global land cover classification at 1 km spatial resolution using a classification tree approach. Int J Remote Sens 21(6–7):1331–1364
Harfenmeister K, Spengler D, Weltzien C (2019) Analyzing temporal and spatial characteristics of crop parameters using sentinel-1 backscatter data. Remote Sens 11(13):1569
He W, Wu J, Fei M et al (2022) China’s terrestrial carbon sink over 2010–2015 constrained by satellite observations of atmospheric CO\(_2\) and land surface variables. J Geophys Res Biogeosci 127(2):e2021JG006644
Holtgrave A-K, Röder N, Ackermann A et al (2020) Comparing sntinel-1 and-2 data and indices for agricultural land use monitoring. Remote Sens 12(18):2919
Huete A (1997) A comparison of vegetation indices over a global set of TM images for EOS-MODIS. Remote Sens Environ 59(3):440–451
Inoue Y, Sakaiya E, Wang C (2014) Capability of C-band backscattering coefficients from high-resolution satellite SAR sensors to assess biophysical variables in paddy rice. Remote Sens Environ 140:257–266
Jiang Z, Li L, Ustin SL (2009) Estimation of canopy water content with MODIS spectral index. In: SPIE proceedings, vol 7454. International Society for Optics and Photonics. SPIE, Art no. 74540V
Jiang H, Wang S, Cao X et al (2018) A shadow-eliminated vegetation index (SEVI) for removal of self and cast shadow effects on vegetation in rugged terrains. Int J Digit Earth 12(9):1013–1029
Jiao X, McNairn H, Robertson LD (2021) Monitoring crop growth using a canopy structure dynamic model and time series of synthetic aperture radar (SAR) data. Int J Remote Sens 42(17):6433–6460
Khabbazan S, Vermunt P, Steele-Dunne S et al (2019) Crop monitoring using sentinel-1 data: a case study from The Netherlands. Remote Sens 11(16):1887
Kikuchi K, Wang B (2008) Diurnal precipitation regimes in the global tropics. J Clim 21(11):2680–2696
Kim Y, van Zyl J (2000) On the relationship between polarimetric parameters. In: Proc. of IEEE international geoscience and remote sensing symposium IGARSS, vol 3. IEEE, pp 1298–1300
King MD, Platnick S, Menzel WP et al (2013) Spatial and temporal distribution of clouds observed by MODIS onboard the terra and aqua satellites. IEEE Trans Geosci Remote Sens 51(7):3826–3852
Kingma DP, Ba J (2014) Adam: a method for stochastic optimization. Preprint at https://arxiv.org/abs/1412.6980
Kovacs JM, Jiao X, Flores-de-Santiago F et al (2013) Assessing relationships between Radarsat-2 C-band and structural parameters of a degraded mangrove forest. Int J Remote Sens 34(20):7002–7019
Madry S (2013) Introduction and history of space remote sensing. Handbook of satellite applications. Springer, New York, pp 657–666
Mahdianpari M, Salehi B, Mohammadimanesh F et al (2017) Random forest wetland classification using ALOS-2 L-band, RADARSAT- 2 C-band, and TerraSAR-X imagery. ISPRS J Photogramm Remote Sens 130:13–31
Main-Knorn M, Pflug B, Louis et al (2017) Sen2Cor for sentinel-2. In: Image and Signal Processing for Remote Sensing XXIII. SPIE
Mandal D, Bhattacharya A, Rao YS (2021) Radar remote sensing for crop biophysical parameter estimation. Springer, Singapore
McNairn H, Shang J (2016) A review of multitemporal synthetic aperture radar (SAR) for crop monitoring. In: Ban Y (ed) Multitemporal remote sensing. Methods and applications, vol 20. Springer International Publishing, pp 317–340
Meraner A, Ebel P, Xiao XZ et al (2020) Cloud removal in sentinel-2 imagery using a deep residual neural network and SAR-optical data fusion. ISPRS J Photogramm Remote Sens 166:333–346
Mitchard ETA, Saatchi SS, White LJT et al (2012) Mapping tropical forest biomass with radar and spaceborne LiDAR in Lopé National Park, Gabon: overcoming problems of high biomass and persistent cloud. Biogeosciences 9(1):179–191
Moran MS, Alonso L, Moreno JF et al (2012) A RADARSAT-2 quad-polarized time series for monitoring crop and soil conditions in Barrax, Spain. IEEE Trans Geosci Remote Sens 50(4):1057–1070
Moreira A, Prats-Iraola P, Younis M et al (2013) A tutorial on synthetic aperture radar. IEEE Geosci Remote Sens Mag 1(1):6–43
Odena A, Dumoulin V, Olah C (2016) Deconvolution and Checkerboard Artifacts. In: Distill
Park S (2012) Cloud and cloud shadow effects on the MODIS vegetation index composites of the Korean Peninsula. Int J Remote Sens 34(4):1234–1247
Peel Murray C, Finlayson BL, McMahon TA (2007) Updated world map of the Köppen–Geiger climate classification. Hydrol Earth Syst Sci 11(5):1633–1644
Periasamy S (2018) Significance of dual polarimetric synthetic aperture radar in biomass retrieval: an attempt on Sentinel-1. Remote Sens Environ 217:537–549
Pettorelli N, Vik JO, Mysterud A et al (2005) Using the satellite-derived NDVI to assess ecological responses to environmental change. Trends Ecol Evol 20(9):503–510
Prabakaran C, Singh CP, Panigrahy S et al (2013) Retrieval of forest phenological parameters from remote sensing-based NDVI time-series data. Curr Sci 105(6):795–802
Reed BC, Brown JF, Darrel VZ et al (1994) Measuring phenological variability from satellite imagery. J Veg Sci 5(5):703–714
Ronneberger O, Fischer P, Brox T (2015) U-Net: convolutional networks for biomedical image segmentation. In: Medical image computing and computer-assisted intervention (MICCAI), vol 9351. Springer International Publishing, pp 234–241
Rouse JW, Haas RH, Schell JA et al (1974) Monitoring vegetation systems in the great plains with ERTS. In: Proceedings of the third earth resources technology satellite-1 symposium, vol 1, pp 309–317
Scarpa G, Gargiulo M, Mazza A et al (2018) A CNN-based fusion method for feature extraction from Sentinel data. Remote Sens 10(2):236
Schmitt M, Hughes LH, Qiu C et al (2019) SEN12MS—a curated dataset of georeferenced multi-spectral Sentinel-1/2 imagery for deep learning and data fusion. In: ISPRS annals of photogrammetry, remote sensing and spatial information sciences IV-2/W7, pp 153–160
Segalini F, Guarini R, Notarnicola C et al (2014) COSMO-SkyMed\({\circledR}\) imagery for crops characterization. In: EARSeL 34th symposium proceedings, pp 7.1–7.11
Sinha S, Jeganathan C, Sharma LK et al (2015) A review of radar remote sensing for biomass estimation. Int J Environ Sci Technol 12(5):1779–1792
Small D (2011) Flattening gamma: radiometric terrain correction for SAR imagery. IEEE Trans Geosci Remote Sens 49(8):3081–3093
Swinnen E, Veroustraete F (2008) Extending the SPOT-vegetation NDVI time series (1998–2006) back in time with NOAA-AVHRR data (1985–1998) for Southern Africa. IEEE Trans Geosci Remote Sens 46(2):558–572
Takaku J, Tadono T, Doutsu M et al (2020) Updates of ‘AW3D30’ ALOS global digital surface model with other open access datasets. In: The international archives of the photogrammetry, remote sensing and spatial information sciences XLIIIB4-2020, pp 183–189
Tian J, Wang L, Li X et al (2017) Comparison of UAV and WorldView-2 imagery for mapping leaf area index of mangrove forest. Int J Appl Earth Obs Geoinf 61:22–31
Uuemaa E, Ahi S, Montibeller B et al (2020) Vertical accuracy of freely available global digital elevation models (ASTER, AW3D30, MERIT, TanDEM-X, SRTM, and NASADEM). Remote Sens 12(21):3482
Verbesselt J, Hyndman R, Zeileis A et al (2010) Phenological change detection while accounting for abrupt and gradual trends in satellite image time series. Remote Sens Environ 114(12):2970–2980
Vollrath A, Mullissa A, Reiche J (2020) Angular-based radiometric slope correction for sentinel-1 on google earth engine. Remote Sens 12(11):1867
Vreugdenhil M, Wagner W, Bauer-Marschallinger B et al (2018) Sensitivity of sentinel- 1 backscatter to vegetation dynamics: an Austrian case study. Remote Sens 10(9):1396
Wang Z, Alan CB, Hamid RS et al (2004) Image quality assessment: from error visibility to structural similarity. IEEE Trans Image Process 13(4):600–612
Wang L, Jia M, Yin D et al (2019) A review of remote sensing for mangrove forests: 1956–2018. Remote Sens Environ 231:111223
Weiss M, Jacob F, Duveiller G (2020) Remote sensing for agricultural applications: a meta-review. Remote Sens Environ 236:111402
Xie Y, Sha Z, Yu M (2008) Remote sensing imagery in vegetation mapping: a review. J Plant Ecol 1(1):9–23
Yuan X, Shi G, Jianfang L (2021) A review of deep learning methods for semantic segmentation of remote sensing imagery. Expert Syst Appl 169:114417
Zanaga D, Van De Kerchove R, De Keersmaecker W et al (2021) ESA WorldCover 10 m 2020 v100
Zhao H, Gallo O, Frosio I et al (2017) Loss functions for image restoration with neural networks. IEEE Trans Comput Imaging 3(1):47–57
Zhao W, Qu Y, Chen J et al (2020) Deeply synergistic optical and SAR time series for crop dynamic monitoring. Remote Sens Environ 247:111952
Zupanc A (2017) Improving cloud detection with machine learning. https://medium.com/sentinel-hub/improving-cloud-detection-with-machine-learning-c09dc5d7cf13. Accessed 02 July 2023
Acknowledgements
We thank Andrew Madden for his help revising figures and proof-reading the manuscript. The authors gratefully acknowledge the computation time granted by the Institute for Distributed Intelligent Systems and provided on the GPU cluster Monacum One at the University of the Bundeswehr Munich.
Funding
Open Access funding enabled and organized by Projekt DEAL. This work was supported by the German Federal Ministry for Economic Affairs and Energy in the project “DESTSAM—Dense Satellite Time Series for Agricultural Monitoring” (FKZ 50EE2018A).
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of Interest
Not applicable.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Roßberg, T., Schmitt, M. A Globally Applicable Method for NDVI Estimation from Sentinel-1 SAR Backscatter Using a Deep Neural Network and the SEN12TP Dataset. PFG 91, 171–188 (2023). https://doi.org/10.1007/s41064-023-00238-y
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s41064-023-00238-y