Abstract
Geo-referenced and temporal data are becoming more and more ubiquitous in a wide range of fields such as medicine and economics. Particularly in the realm of medical research, spatio-temporal data play a pivotal role in tracking and understanding the spread and dynamics of diseases, enabling researchers to predict outbreaks, identify hot spots, and formulate effective intervention strategies. To forecast these types of data we propose a Probabilistic Spatio-Temporal Neural Network that (1) estimates, with computational efficiency, models with spatial and temporal components; and (2) combines the flexibility of a Neural Network—which is free from distributional assumptions—with the uncertainty quantification of probabilistic models. Our architecture is compared with the established INLA method, as well as with other baseline models, on COVID-19 data from Italian regions. Our empirical analysis demonstrates the superior predictive effectiveness of our method across multiple temporal ranges and offers insights for shaping targeted health interventions and strategies.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
In recent years, the surge in spatial and spatio-temporal data availability has been remarkable, largely related to technological advancements in computational tools. These tools enable the real-time acquisition of data from sources like GPS satellites, cellular network triangulation, Wi-Fi location tracking, etc. Consequently, researchers across diverse domains, from epidemiology, ecology, and climatology all the way to social sciences, often find themselves dealing with geo-referenced and time-stamped data that encapsulate spatial information as well as temporal aspects.
Machine learning and deep learning have attracted tremendous attention from researchers in various fields, e.g., AI, computer vision, and language processing, but also from more traditional sciences, e.g., physics, biology, and manufacturing. The added value provided by these algorithms is the few or no assumptions to be met. They are far more flexible than traditional statistical models, as they have weaker requirements in terms of collinearity, Gaussianity of residuals, and similar. Thus, they have high model uncertainty tolerance. In spite of the many pros, neural networks are often blamed for lack of interpretability (black-box models) and of uncertainty quantifications, and for high computational costs.
Image processing components such as convolutional neural networks, sequence processing models, such as recurrent neural networks, and regularization layers, such as dropouts [1], are used extensively, and they contribute to the lack of interpretability.
Yet, in sectors like physics, biology, business, and manufacturing, the representation of model uncertainty remains paramount. As these sectors increasingly lean toward embracing uncertainty, deep learning presents novel opportunities. Coherently, the goal of the current work is twofold: to create a model that is able to infer both the spatial and temporal components and to combine the advantages of both approaches, namely the flexibility of a neural network and the quantification of uncertainty offered by a traditional probabilistic regression model.
To implement this model different approaches were used:
-
Embeddings, a relatively low-dimensional space into which we translate high-dimensional vectors, to model spatio-temporal components and feed them to neural networks;
-
A neural network architecture able to handle sequences of data and quantify the uncertainty of each prediction.
We conduct a comprehensive analysis on the historical series of COVID-19 deaths. We evaluate the accuracy of our forecasts in comparison with other state-of-the-art models in the machine learning and spatio-temporal statistical literature, over various forecasting ranges.
1 Related works
In this section we reference, to the best of our knowledge, some studies related to the use of statistical models and machine learning for the analysis of COVID-19 data.
In [2] and [3] authors use machine learning models to forecast the number of upcoming patients affected by COVID-19. In particular, in [2] four standard forecasting models, such as linear regression (LR), least absolute shrinkage and selection operator (LASSO), support vector machine (SVM), and exponential smoothing (ES), have been used, while [3] uses different ML algorithms for predicting the chance of being infected and leverages an autoregressive integrated moving average time series for forecasting confirmed cases for various states of India.
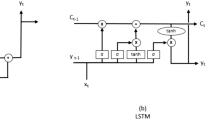
Authors in [4] proposed comparative forecasting results using machine learning methods. The classical SIR model was used to fit COVID-19 data using different techniques and tools for forecasting, including machine learning with fitting functions. In [5] a multilayer perceptron for predicting the spread of COVID-19 is proposed, while in [6] simple Recurrent Neural Network (RNN), Long short-term memory (LSTM), Bidirectional L-STM (BiLSTM), Gated recurrent units (GRUs), and Variational AutoEncoder (VAE) algorithms have been applied for global forecasting of COVID-19 cases based on a small volume of data.
Authors of [7] compared the performance of several machine learning methods to predict the COVID-19 spread in different countries. In [8] a multimodel machine learning technique for forecasting COVID-19-related parameters in the long term both within India and on a global scale has been proposed.
In [9] the author proposes a semi-parametric approach to estimate the evolution of COVID-19 (SARS-CoV-2) in Spain using a combination of both a Deep learning model and a Poisson-Gamma Bayesian regression model to take into account uncertainty quantification. The goal was to elicit the expected number of counts and their reliability. In [10] they use a INLA spatio-temporal stochastic model to explain the temporal and spatial variations in the daily number of new confirmed cases in Spain, Italy, and Germany. In [11] authors present a Poisson autoregressive model to monitor the temporal evolution of COVID-19 contagion and associated reproduction rate, dynamically adapting parameters to explain the epidemic propagation in terms of short- and long-term case count dependencies, demonstrating how health policies can impact contagion trends. In [12], authors use a Poisson autoregressive model to analyze daily new observed cases, revealing whether the contagion exhibits a trend and determining the position of each country on that trend, while in [13] an endemic–epidemic model is proposed in order to track COVID-19 contagion dynamics both temporally and spatially, exemplified through an empirical analysis of Northern Italy’s provinces affected by the pandemic. Authors of [14] use a discrete latent variable model with spatial and time dependences, for the analysis of SARS-CoV-2 infections. Finally, [15] reviews different spatial and spatio-temporal approaches to identify spatial clusters and associated risk factors.
2 Methods
2.1 Modeling spatial and temporal components with embeddings
Embeddings are numerical representations of categorical variables, commonly used in machine learning [16]. They capture the semantic meaning of a variable by mapping it to a dense vector of real numbers. Models based on embeddings can thus learn the relationships among the variables in a continuous space rather than a discrete one.
The idea is to embed spatial and temporal components by synthesizing “context” information (i.e., locations with similar behavior have similar latent representation). Entity Embedding [17] serves this purpose: the idea is to map categorical variables into Euclidean spaces using a function approximation problem where categories are turned into “Entity” (a.k.a. category) Embeddings. It is expected that similar categories are close in the embedding space.
In this work, we take into account categorical information related to regions, week-day, month, season, and year. To give an intuition on how embeddings work we can observe Fig. 1. In this case, data come from the COVID-19 daily deaths time series for each of the 20 Italian regions (as discussed extensively in Sect. 4) and the goal is to embed each region in a latent space using the entity embedding approach. Figure 1 represents the embeddings in a two-dimensional-reduced space for visualization purposes, after applying the t-SNE [18] to the (n-dimensional) latent embeddings vectors. It can be observed that regions that have had a similar incidence of deaths from COVID-19 lie close together in the two-dimensional embedding representation of Fig. 1. The resulting clusters among regions can be intuitively justified.
2.2 Model architecture
We aim to construct a neural network (NN) architecture capable of discerning patterns within spatio-temporal count data. To achieve this, we will model the outcome using a probability distribution [19, 20] suitable for count data, such as the Poisson distribution. Additionally, our NN will incorporate uncertainty in its predictions, akin to conventional statistical models.
The proposed architecture is shown in Fig. 2, and it is based on a multi-head CNN-LSTM [21] structure. We will now delve into the various components of the architecture, emphasizing their novel aspects.Footnote 1
Consider a scenario where we wish to study the temporal progression of a specific phenomenon across \(N\) distinct locations. Formally, for each location \(j \in \{1, \dots , N\}\) an event is observed at \(T\) time intervals, and our objective is to forecast for subsequent intervals up to a horizon of \(T+h\).
The first layer consists of N inputs, where N is the number of sites under consideration, ensuring that the T temporal data of each site are individually accounted for.
The temporal data are processed by a 1D convolutional layer, or Conv1D. This layer serves to provide temporal smoothing, ensuring that fluctuations over time are harmonized. Moreover, it is instrumental in identifying pertinent patterns within the time series. Following this process, the output from the convolutional layer is flattened [22] and reshaped, making it compatible with subsequent layers. The next step in the architecture involves employing two stacked LSTMs [23]. These are useful to extract insights from sequential data. In parallel with the spatio-temporal data, the network is also fed with additional information about the region under prediction, the day of the week, month, and year. These details are processed through different embedding layers.
These different processing flows are then joined together: the additional information embeddings (focusing on the region under prediction, ..., month, and year), which encompass both temporal and spatial information, are merged with the LSTMs outputs. This ensures that the model has a comprehensive view of the data, priming it to make accurate predictions.
For each time instance, identical input data are supplied to the network \(N\) times, each paired with the spatial data pertinent to the specific location for which a prediction is being generated. The rationale behind supplying both data and spatial embeddings as network inputs is that the embeddings will provide invaluable insights to compute the output for a specific site, while considering input data from multiple sites. Furthermore, replicating the same information N times acts as a data augmentation strategy: more complex models demand larger datasets for effective training. For instance, if we were to rely solely on a year’s worth of observations, we would be limited to 365 input data points to train our architecture. However, by iterating this process N times, we effectively amplify the number of input observations available to the model. This not only enhances the robustness of our model, but also aids in parameter estimation, ultimately leading to more accurate predictions.
After concatenating the embeddings and the LSTM layer outputs, a dense layer is added, culminating in the output layer. The latter is a dense layer with as many neurons as the range of forecast, representing the rate parameters \( \lambda \) of a Poisson distribution, which fully identifies the conditional probability distribution (CPD) of the outcome \( y \) given the input \( x \). The whole NN input scheme is summarized in Algorithm 1.
Conventionally, a NN updates its parameters based on the minimization of a loss function. In our context, the approach is centered around maximizing the likelihood, ensuring that the resulting model can predict observed values with high probability. The likelihood of an arbitrary CPD can be maximized within a neural network framework by interpreting the probabilistic neural network’s output as a unique distribution parameter. The neural network “learns” to predict the \( \lambda \) value that maximizes the likelihood—or minimizes the negative log likelihood (NLL)—of the observed data
where
-
\(\lambda _i\) is the value predicted by the neural network for the \(i^{th}\) data point.
-
\(x_i\) is the \(i^{th}\) observed data point.
-
n is the total number of data points.
2.3 Alternative models
To evaluate the performance of our model, the results are compared with a pool of alternatives that are commonly used in the literature for spatio-temporal data forecasting: two ensemble models: Random Forest [24] and XGBoost [25]; and a Bayesian statistical model: INLA [26].
In particular, regarding the ensemble models, the embeddings obtained from the embedding layers of a neural network are used as input. These embeddings are tasked to learn meaningful data representations along with the lagged time series. Regarding INLA, we have used a Poisson distribution for modeling the outcomes. The choice of the Poisson distribution was driven by the estimated dispersion parameter in our data being close to one, indicating that the Poisson distribution adequately captures the data variability. Additionally, we incorporated a spatial component using the Besag-York-Mollié (BYM) model [27] and an autoregressive temporal component, in order to capture spatio-temporal dynamics in our data.
3 Data
Our primary data source for the analysis is derived from the GitHub repository maintained by the Italian Civil Protection, accessible via this link: https://github.com/pcm-dpc/COVID-19. This repository provides daily updates, offering a comprehensive overview of the pandemic’s progression. It includes different time series such as the number of new positive cases, ICU occupancy, swabs made, and deceased, both from a national and regional perspective.
Our analysis primarily centers on the historical series of daily deaths for several reasons:
-
This time series exhibits significant variability and is prone to abrupt fluctuations. The daily death count often undergoes revisions in the days following its initial publication. Consequently, discerning the genuine signal amidst this noise requires the deployment of complex modeling techniques.
-
It is used as an indicator of the pandemic’s severity. Unlike time series like New Daily Positive cases (which are influenced by the number of swabs conducted) and ICU occupancy (that exhibit a degree of temporal persistence), the count of new deaths provides a more direct and unfiltered reflection of the pandemic’s impact.
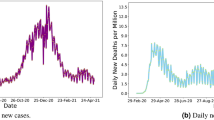
Figure 3 showcases the historical series of COVID-19-related deaths across five Italian regions: LOMBARDY, VENETO, BASILICATA, CALABRIA, and VALLE D’AOSTA. A cursory examination reveals notable disparities in death counts across regions. Furthermore, on certain days, there are significant spikes, indicating abrupt surges in fatalities.
4 Experiments and results
Table 1 presents two metrics to evaluate the performance of the proposed model in comparison with benchmarks: the mean absolute error (MAE), which measures the average absolute prediction error, and the mean squared error (MSE), which more heavily penalizes large prediction errors. Comparisons were made using different forecasting ranges (7, 14, 21, and 28 days) for the period considered in December 2023. This approach aimed to assess the accuracy and reliability of forecasts over varying time spans within the specific month. Table 2 shows the two error metrics across different regions, specifically for each model, in the context of the 28-day forecast. It is observed that the regions with the largest discrepancies between the observed and predicted cases are also those with higher incidence and greater variability (Lombardy, Piedmont, and Veneto).
The proposed model, which we name probabilistic neural network (PNN), outperforms the others in terms of both metrics in all scenarios. Additionally, Fig. 4 shows that PNN returns highly accurate predictions at national level (obtained summing the daily regional forecasts): the 0.025 and 0.975 quantiles of the conditional probability distribution define a 95% prediction interval that quantifies the uncertainty of each prediction, and in most cases it includes the true number of COVID-19 deaths. To more accurately assess the accuracy of prediction intervals, in Table 3 we compared the intervals generated by the PNN model with those derived from the INLA approach. This analysis reveals that the coverage of the PNN’s prediction intervals is close to the theoretical level, but not perfectly in line. The regional coverages in both cases do not reach the nominal 95%, possibly because of anomalies in regional data, such as recounts of previous days’ deaths. These events introduce significant and sudden variations in the observed regional time series, making predictions more uncertain and affecting the coverage of the prediction intervals.
5 Strength and weaknesses
The model we propose offers several advantages, compared to competing models:
-
It is designed to handle input data characterized by spatial and temporal changes, and it delivers accurate results, offering a comprehensive and detailed view of trends and patterns. Moreover, this is a lightweight approach, which can be easily run on standard laptops, making it accessible with no advanced hardware resources.
-
Compared to a traditional neural approach, it adopts a probabilistic approach, thus providing an estimate of the probability of a particular outcome. This makes it flexible and particularly suitable for the analysis of count data. Specifically, the Poisson distribution that we adopt returns outputs that are integer values, capturing the inherent nature of count data and ensuring more meaningful predictions.
-
Embedding-related representations can be extracted to provide insights on specific features, such as locations. This means that the model can identify and place similar entities close in the embedding space, facilitating the interpretation and the understanding of relationships between different entities.
However, alongside the numerous advantages, it is also appropriate to analyze the potential weaknesses of our model. First, like all deep learning models, a substantial amount of data is needed to train the models, while a limited amount of data can, in fact, compromise parameters estimation accuracy and reliability. Furthermore, an incorrect representation of the embeddings can lead to a model that cannot correctly discriminate between different locations. This means that if the embeddings are not properly calibrated or correctly interpreted, the model might fail to distinguish between different positions or categories, leading to inaccurate or misleading results.
6 Conclusion
We introduce a neural network architecture capable of delivering forecasts for spatio-temporal data, together with a measure of uncertainty. Our model is evaluated across various range of forecasting intervals, outperforming benchmarks.
The use of a neural network, particularly a probabilistic one, offers a level of flexibility that traditional statistical models often lack. This flexibility is especially crucial when dealing with complex datasets, such as the spatio-temporal one we focus on. Neural networks can adapt to intricate patterns and complex relationships in the data, which might be challenging to capture by conventional statistical models. Moreover, by employing a probabilistic neural network, we not only benefit from the adaptability of neural architectures but also retain the advantages of statistical models in estimating uncertainty. This combination ensures that our predictions are both accurate and endowed with a reliable measure of confidence.
Moreover, embeddings have proven to be a valuable tool in guiding the network’s learning process, especially when forecasting COVID-19-related deaths across different Italian regions. These embeddings allow the model to understand and represent the similarities and differences between regions, enhancing predictive capabilities.
The proposed model serves as an efficient foundational framework and, in light of the results we discuss, is versatile enough to be extended for other related series and geographies.
Notes
The codebase is available to facilitate reproducibility at the following link: https://github.com/Fede-stack/Probabilistic-COVID19.
References
Srivastava, N., Hinton, G., Krizhevsky, A., Sutskever, I., Salakhutdinov, R.: Dropout: a simple way to prevent neural networks from overfitting. J. Mach. Learn. Res. 15(1), 1929–1958 (2014)
Rustam, F., Reshi, A.A., Mehmood, A., Ullah, S., On, B.-W., Aslam, W., Choi, G.S.: Covid-19 future forecasting using supervised machine learning models. IEEE Access 8, 101489–101499 (2020)
Painuli, D., Mishra, D., Bhardwaj, S., Aggarwal, M.: Forecast and prediction of Covid-19 using machine learning. In: Kose, U., Gupta, D., de Albuquerque, V.H.C., Khanna, A. (eds.) Data Science for COVID-19, pp. 381–397. Academic Press, Cambridge (2021)
Baldé, M.A.: Fitting sir model to Covid-19 pandemic data and comparative forecasting with machine learning. medRxiv, 2020–04 (2020)
Car, Z., Baressi Šegota, S., Anelić, N., Lorencin, I., Mrzljak, V., et al. (2020) Modeling the spread of Covid-19 infection using a multilayer perceptron. Comput. Math. Methods Med
Zeroual, A., Harrou, F., Dairi, A., Sun, Y.: Deep learning methods for forecasting Covid-19 time-series data: a comparative study. Chaos Solitons Fractals 140, 110121 (2020)
Ghafouri-Fard, S., Mohammad-Rahimi, H., Motie, P., Minabi, M.A., Taheri, M., Nateghinia, S.: Application of machine learning in the prediction of Covid-19 daily new cases: a scoping review. Heliyon (2021). https://doi.org/10.1016/j.heliyon.2021.e08143
Mohan, S., Abugabah, A., Kumar Singh, S., Kashif Bashir, A., Sanzogni, L.: An approach to forecast impact of Covid-19 using supervised machine learning model. Softw. Pract. Exp. 52(4), 824–840 (2022)
Cabras, S.: A Bayesian-deep learning model for estimating Covid-19 evolution in Spain. Mathematics 9(22), 2921 (2021)
Jalilian, A., Mateu, J.: A hierarchical spatio-temporal model to analyze relative risk variations of Covid-19: a focus on Spain, Italy and Germany. Stoch. Env. Res. Risk Assess. 35, 797–812 (2021)
Agosto, A., Campmas, A., Giudici, P., Renda, A.: Monitoring COVID-19 contagion growth. Stat. Med. 40, 4150–4160 (2021)
Agosto, A., Giudici, P.: A poisson autoregressive model to understand COVID-19 contagion dynamics. Risks 8, 77–77 (2020)
Celani, A., Giudici, P.: Endemic-epidemic models to understand COVID-19 spatio-temporal evolution. Spat. Stat. 49, 100528 (2022)
Bartolucci, F., Farcomeni, A.: A spatio-temporal model based on discrete latent variables for the analysis of covid-19 incidence. Spat. Stat. 49, 100504 (2022)
Nazia, N., Butt, Z.A., Bedard, M.L., Tang, W.-C., Sehar, H., Law, J.: Methods used in the spatial and spatiotemporal analysis of covid-19 epidemiology: a systematic review. Int. J. Environ. Res. Public Health 19(14), 8267 (2022)
Hancock, J.T., Khoshgoftaar, T.M.: Survey on categorical data for neural networks. J. Big Data 7(1), 1–41 (2020)
Guo, C., Berkhahn, F.: Entity embeddings of categorical variables. arXiv preprint arXiv:1604.06737 (2016)
Maaten, L., Hinton, G.: Visualizing data using t-sne. J. Mach. Learn. Res. 9(11), 2579–2605 (2008)
Dürr, O., Sick, B., Murina, E.: Probabilistic deep learning: with python. Keras and Tensorflow Probability, Manning Publications, New York (2020)
Dillon, J.V., Langmore, I., Tran, D., Brevdo, E., Vasudevan, S., Moore, D., Patton, B., Alemi, A., Hoffman, M., Saurous, R.A.: Tensorflow distributions. arXiv preprint arXiv:1711.10604 (2017)
Canizo, M., Triguero, I., Conde, A., Onieva, E.: Multi-head CNN-RNN for multi-time series anomaly detection: an industrial case study. Neurocomputing 363, 246–260 (2019)
Jin, J., Dundar, A., Culurciello, E.: Flattened convolutional neural networks for feedforward acceleration. arXiv preprint arXiv:1412.5474 (2014)
Hochreiter, S., Schmidhuber, J.: Long short-term memory. Neural Comput. 9(8), 1735–1780 (1997)
Breiman, L.: Random forests. Mach. Learn. 45, 5–32 (2001)
Chen, T., He, T., Benesty, M., Khotilovich, V., Tang, Y., Cho, H., Chen, K., Mitchell, R., Cano, I., Zhou, T. et al.: Xgboost: extreme gradient boosting. R package version 0.4-2 1(4), 1–4 (2015)
Rue, H., Martino, S., Chopin, N.: Approximate Bayesian inference for latent gaussian models by using integrated nested laplace approximations. J. R. Stat. Soc. Ser. B Stat Methodol. 71(2), 319–392 (2009)
Besag, J., York, J., Mollié, A.: Bayesian image restoration, with two applications in spatial statistics. Ann. Inst. Stat. Math. 43, 1–20 (1991)
Acknowledgements
Antonietta Mira has been supported by SNSF project 200021_208249 “Feature Learning for Bayesian Inference.”
Funding
Open access funding provided by Universitá della Svizzera italiana.
Author information
Authors and Affiliations
Contributions
Federico Ravenda wrote the main manuscript text. All authors reviewed the manuscript.
Corresponding author
Ethics declarations
Conflict of interest
The authors declare no competing interests.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Ravenda, F., Cesarini, M., Peluso, S. et al. A probabilistic spatio-temporal neural network to forecast COVID-19 counts. Int J Data Sci Anal (2024). https://doi.org/10.1007/s41060-024-00525-w
Received:
Accepted:
Published:
DOI: https://doi.org/10.1007/s41060-024-00525-w