Abstract
Purpose of Review
Fomites are inanimate objects that become colonized with microbes and serve as potential intermediaries for transmission to/from humans. This review summarizes recent literature on fomite contamination and microbial survival in the built environment, transmission between fomites and humans, and implications for human health.
Recent Findings
Applications of molecular sequencing techniques to analyze microbial samples have increased our understanding of the microbial diversity that exists in the built environment. This growing body of research has established that microbial communities on surfaces include substantial diversity, with considerable dynamics. While many microbial taxa likely die or lay dormant, some organisms survive, including those that are potentially beneficial, benign, or pathogenic. Surface characteristics also influence microbial survival and rates of transfer to and from humans. Recent research has combined experimental data, mechanistic modeling, and epidemiological approaches to shed light on the likely contributors to microbial exchange between fomites and humans and their contributions to adverse (and even potentially beneficial) human health outcomes.
Summary
In addition to concerns for fomite transmission of potential pathogens, new analytical tools have uncovered other microbial matters that can be transmitted indirectly via fomites, including entire microbial communities and antibiotic-resistant bacteria. Mathematical models and epidemiological approaches can provide insight on human health implications. However, both are subject to limitations associated with study design, and there is a need to better understand appropriate input model parameters. Fomites remain an important mechanism of transmission of many microbes, along with direct contact and short- and long-range aerosols.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
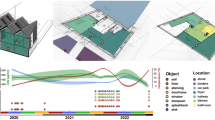
Conceptualized as early as the 1500s, fomites (or fomes) were first thought of as “seeds of disease,” found in the clothing of infected individuals that spread contagion long distances by indirect human contact [1]. Today, fomites are generally considered any inanimate object that, when contaminated with infectious organisms, can serve as a means of transferring disease-causing agents to a new human host. Because people in industrialized countries spend approximately 90% of their time indoors [2], the most important fomites for contimination and transmission tend to be those found in the built environment and those that humans frequently come into direct contact with, such as doorknobs, countertops, medical equipment, handrails, clothing, and mobile phones. As our understanding of microbes in the built environment has greatly expanded in the last decade, so has our understanding of fomites and their role in the transmission of infectious agents and other microbial matter to and from humans. Here, we review the recent body of literature on fomite contamination and microbial survival in the built environment, factors that affect transmission of microbes between fomites and humans, and the implications for human health. Table 1 summarizes what the authors consider to be some of the most important or influential recent research outcomes within the major categories of articles reviewed herein. These highlighted articles were determined to stand out in the field as being particularly novel, comprehensive, and/or addressing a fundamental question that had not been thoroughly addressed before.
Indoor Microbiomes
We live in a microbial world. Viruses, bacteria, protists, fungi, and archaea exist in all of our inhabited environments [25,26,27]. In buildings, we shed microbes directly to the indoor air and onto building surfaces [28,29,30], microbes are transported indoors from outdoors [31, 32], and we also acquire microbes from our surroundings [3••, 33, 34]. Human occupancy and activity, the outdoor environment, and building design and operation each influence the abundance and diversity of microbes in buildings—or what is collectively referred to as the indoor microbiome [23••, 35,36,37,38,39,40,41,42]. Many molecular analyses have identified considerable microbial diversity on built surfaces. Most microbes found in indoor environments appear to be dormant, inactive, or dead [43], and either has no known impact on human health or are possibly even beneficial to human health [24•, 44,45,46]. For example, early life exposures to particular microbes or assemblages of microbes have been shown to shape our innate immune responses to protect against allergy and asthma [47,48,49]. However, potentially pathogenic organisms can also reside within the microbial milieu of our built worlds, which can have a variety of negative health consequences.
Microbes on Surfaces
Inanimate objects in the built environment can serve as reservoirs of microbial matter. Each of these objects is host to an entire community composed of a wide variety of bacterial, viral, archaeal, protistan, and fungal organisms, including potential pathogens and microbial metabolic products harmful to humans.
Microbial Community Ecology on Fomite Surfaces
On indoor surfaces that lack abundant moisture and nutrient availability, most microorganisms that arrive from other environments (such as from human occupants) are generally considered unlikely to survive, and those viable microbes that do survive are generally considered to be inactive or dormant until transferred to other host locations or until they experience an influx of moisture and nutrients that help them proliferate [4••, 50,51,52]. Surveys of fungal communities in indoor environments, conducted using high-throughput molecular sequencing, have shown that they tend to be driven primarily by transport from the local outdoor environment [31]. However, similar surveys of bacterial communities in the built environment have revealed high abundances of skin-associated bacteria (e.g., Propionibacterium acnes, Corynebacterium, and Streptococcus), particularly in buildings and on surfaces with high human occupancy and frequency of interactions [35]. Source-tracking efforts have also provided insight into the origin of the bacteria that reside on various indoor surfaces. For example, urine- and feces-associated bacteria have been shown to be more common on toilet seats and toilet handles than on other surfaces [53]; bacteria associated with fresh produce have been shown to be more common on kitchen countertops and inside refrigerators [54]; and bacteria associated with leaves and soil have been shown to be more common on the interior and exterior trim of doors that open to the outside than other (more interior) home surface locations [55]. Conversely, on surfaces that frequently have high moisture levels, such as those in bathrooms and kitchens (e.g., shower curtains, sinks, and countertops), rich microbial biofilms can form community assemblages that closely resemble those found in plumbing systems and water reservoirs [56,57,58]. Investigating differences both within and between homes, Lax et al. (2014) demonstrated that bacterial communities on different surfaces in an individual home showed strong similarities for some surfaces (e.g., kitchen floors were similar to bedroom floors and both were similar to human feet; and kitchen light switches were similar to the front doorknob, which were also similar to occupants’ hands) but not for others (e.g., kitchen countertops and human noses were distinct from doorknobs) [3••]. Moreover, when a family moved homes, the bacterial community composition on surfaces in the new home rapidly converged toward the composition of bacteria from surfaces in the previous home, suggesting that the new occupants quickly deposited their own unique signatures of human-associated bacteria to the new space.
While much has been revealed about bacterial and fungal communities in indoor environments in recent years, much less is known about viral communities and total viral abundance on surfaces in buildings [59]. However, much has been learned about the presence, abundance, and survival of specific viruses and other potential pathogens that cause concern for infectious disease transmission and other emerging microbial hazards.
Viral Pathogens on Fomite Surfaces
Presence and Abundance
The study of fomites has traditionally involved determining whether the presence of specific potentially pathogenic organisms—primarily those of viruses or bacteria—resided on environmental surfaces. For example, in an early influential survey, Boone and Gerba (1982) sampled over 300 fomites from daycare centers and homes to determine the presence of influenza A virus on each surface [60]. During flu seasons, approximately half of all common building surfaces from both types of indoor environments had measurable levels of influenza virus, suggesting that contaminated fomite surfaces could play a role in influenza transmission.
Since then, numerous similar studies targeting influenza and other viruses have also discovered that, for example:
Norovirus and influenza A virus were found on frequently used fomites (e.g., desktops, faucet handles, and paper towel dispensers) in elementary school classrooms [61].
Widespread norovirus contamination was found on fomite surfaces on houseboats on which an outbreak of norovirus gastroenteritis was suspected [62].
Picornavirus (including rhinovirus and/or enterovirus) was detected on approximately 20% of toys in pediatric office waiting rooms [63].
Human rhinovirus (hRV) was detected on 5% of clothing samples from teachers working in childcare centers [64].
Rotavirus was detected on about 20% of fomite samples in daycare centers, including on telephone receivers, drinking fountains, water-play tables, and toilet handles [65]; and on nearly half of surfaces sampled in a pediatric unit, with higher prevalence on surfaces that are commonly in direct contact with children (e.g., thermometers and play mats) than on other environmental surfaces (e.g., door handles and wash basins) [66].
Severe acute respiratory syndrome (SARS) coronavirus RNA was found on 30% of surface swab samples in hospitals, including in patient rooms, on computer mice at nurse stations, and on the handrail of a public elevator [67].
Human parainfluenza virus 1 (HPIV1) was detected on 37% of a total of 328 fomites from 12 different office buildings, most frequently isolated on desktops [68].
HPIV3, HPIV1, and norovirus GII RNA were detected on 16 (12%), 7 (5%), and 4 (3%) of a total of 130 surfaces sampled in offices, with computer keyboards, computer mice, telephones, and desktops having significantly higher abundances than other fomite surfaces such as door handles, light switches, or ventilation ducts [5•].
Human adenoviruses (HAdV) were detected from 63 of 141 (45%) fomite samples in an adult intensive care unit (ICU) in a hospital in Rio de Janeiro, Brazil, with viral loads ranging from 2.48 × 101 to 2.1 × 103 genomic copies per milliliter [69].
Middle East respiratory syndrome coronavirus (MERS-CoV) was detected on 2 of 51 (4%) high-touch surfaces in patient rooms with laboratory-confirmed MERS-CoV patients [70].
These studies and many others confirm that viruses that are known to cause communicable diseases in humans are commonly found on surfaces, but it then must be determined whether they are viable and potentially infectious to humans.
Viability and Survival
Weber and Stilianakis (2008) reviewed numerous studies that investigated the environmental inactivation of influenza A viruses, finding that daily inactivation rate constants differ by several orders of magnitude depending on the nature of surface characteristics and that influenza virus can survive in aerosols for several hours, but only for a few minutes on human hands [71]. As an example from this body of literature, Bean et al. (1982) tracked the survival of laboratory-grown influenza A and B viruses on various surfaces, finding that both viruses survived up to 48 h on hard, non-porous surfaces, such as stainless steel and plastic and up to 12 h on porous surfaces, such as cloth, paper, and tissues [72]. Moreover, fomite transmission of influenza viruses was considered possible because influenza virus could be transferred from stainless steel surfaces to hands for up to 24 h after deposition (and from tissues to hands for up to 15 min after deposition). The viruses then subsequently survived on hands for an additional 5 min after transfer from the tested fomites.
More recently, Greatorex et al. (2011) combined the two main types of approaches commonly used in the literature to evaluate the survival of influenza A and pandemic H1N1 viruses inoculated onto a wide range of surfaces common to work and home environments [73]: (i) molecular (genomic) detection by reverse transcription polymerase chain reaction (RT-PCR), which provides a quantitative measure of presence/abundance of genetic material, and (ii) virus viability by plaque assay (for influenza A) or fluorescent focus assay (for H1N1), which provides a measure of virus survivability. The genome of both viruses was detected on most surfaces up to 24 h after inoculation with minimal decrease in gene copy number (except for unsealed wood surfaces), while virus viability decreased more rapidly to a level below detection on all surfaces at 24 h. However, viruses did survive up to 4 h on most surfaces and up to 9 h on non-porous surfaces. The authors concluded that influenza A transmission via fomites is possible, but it is unlikely if contact occurs after long periods following surface contamination, unless re-inoculation occurs during that time. Similarly, Mukherjee et al. (2012) investigated the viability of H1N1 virus on naturally contaminated hands and household surfaces of 20 individuals with laboratory-confirmed infection, finding that H1N1 has a short period of survival on naturally contaminated skin and fomites, and secretions deposited on hands by coughing or sneezing have a concentration of approximately 20–30 TCID50/mL [74].
Others have found that influenza virus can survive (i.e., remain viable and/or potentially infectious) much longer on fomite surfaces, using a variety of approaches. Thomas et al. (2008) tested the survival of influenza A viruses on banknotes after intentional contamination, finding viruses could survive up to 3 days after inoculation at high concentrations [75]. Additionally, when the virus was encapsulated in respiratory mucus (which may more realistically reflect human contributes to fomite surfaces), survival was as high as 17 days. And when nasopharyngeal secretions from naturally infected children were used to inoculate banknote surfaces, influenza virus survived at least 2 days in one-third of the test cases. Similarly, Oxford et al. (2014) found that influenza A H1N1sw virus particles survived and remained infectious for up to 48 h on a wooden surface, for 24 h on stainless steel and plastic surfaces, and for 8 h on a cloth surface [76]. Perry et al. (2016) found that two influenza A (H1N1) virus strains deposited on stainless steel surfaces remained infectious over a weeklong period, with a 2-log10 loss (99%) in infectivity over 7 days [77]. Moreover, infectivity decreased more rapidly over time at higher absolute humidity, which is consistent with other similar studies [78, 79]. Thompson et al. (2017) tested the viability and RNA abundance (via qt-RT-PCR signal) of five influenza strains seeded on three surfaces (cotton, microfiber, and stainless steel) over time, finding that viable virus was detected for up to 2 weeks on stainless steel and up to 1 week on cotton and microfiber samples [6•]. Times to achieve 99% reductions in viability were ~ 18 h for cotton, ~ 34 h for microfiber, and ~ 175 h for stainless steel. Specific to materials used in personal protective equipment (PPE), Sakaguchi et al. (2010) found that the infectivity of influenza A virus was maintained for ~ 8 h on the surface of an N95 particulate respirator, a non-woven fabric surgical mask, a Tyvek gown, a coated wooden desk, and stainless steel, and for ~ 24 h on a rubber glove [80], suggesting that frequent replacement of PPE and clothing worn by healthcare professionals is warranted to minimize cross-infection. While there is high variability among these studies in influenza inactivation rates and survival on fomite surfaces over time (influenced by a combination of material type, environmental conditions, virus strain, inoculation methods, and viral presence/abundance/viability detection methods), there is general consistency in the literature that influenza viruses can survive for up to several days after being deposited on some surface types and in some conditions.
The survival of other viruses on fomites has also been investigated in recent years. For example, in the aforementioned study of HAdV in an ICU unit in Brazil, a subset of 10 samples that were positive for HAdV were selected for viability assessment, and exactly half of those samples were indeed still viable [69]. Boone and Gerba (2007) reviewed prior studies of the viability of numerous respiratory and enteric viruses on surfaces, reporting virus inactivation rates ranging from ~ 0.01–0.1 log10 per hour for avian influenza and influenza A and B to ~ 0.2–0.6 log10 per hour for rhinovirus 14, PIV2, and respiratory syncytial virus [81]. Inactivation rates for enteric viruses were lower, from ~ 0.002–0.003 log10 per hour for astrovirus (serotype 4) and rotavirus p13 to ~ 0.01 log10 per hour for adenovirus 40. van Doremalen et al. (2013) reported that MERS-CoV viability was more stable at low temperature and low humidity conditions and could still be recovered after 48 h, suggesting fomite transmission of MERS-CoV is possible [82].
Bacterial Hazards on Fomite Surfaces
In addition to viruses, bacterial hazards have also been found on fomite surfaces, including potentially pathogenic and antibiotic-resistant bacteria, which are often not mutually exclusive.
Potentially Pathogenic Bacteria
Marks et al. (2014) detected viable Streptococcus pyogenes and Streptococcus pneumoniae in samples from a daycare and then verified in laboratory tests that isolates of both organisms remained viable over extended periods of time and remained infectious in a mouse model when present as a biofilm (rather than as desiccated cells on surfaces) [83]. These findings suggest that fomite transmission in the environment could be an important pathway if fomites are contaminated with oropharyngeal secretions containing biofilm streptococci. Jones and Lutz (2014) measured the mean survival time of Pseudomonas aeruginosa on laminate, glass, and stainless steel surfaces to be 3.75, 5.75, and 6.75 h, respectively [84]. Malcolm et al. (2017) evaluated the growth and survival of the non-tuberculous mycobacterium (NTM) Mycobacterium abscessus in the presence of mineral particles, kaolin, halloysite, silicon dioxide, and house dust. Mycobacterium abscessus interacted with the particulates, with increased survival rates in the presence of house dust, surviving desiccation for as long as 2 weeks [7•]. These studies and others confirm that potentially pathogenic bacteria are present in the built environment and that they can survive on fomites for long periods of time.
Antibiotic-Resistant Bacteria
Antibiotic-resistant bacteria have been studied in even more detail than potentially pathogenic bacteria. In 2013, the US Centers for Disease Control and Prevention (CDC) published an analysis of the major antibiotic-resistant threats in the USA [85]. Davis et al. (2012) reviewed published works about the household transmission of Staphylococcus aureus and other staphylococci, and suggested that household microbial communities might have a role in the transfer of antimicrobial resistance genes and could be reservoirs for recolonization of humans [86]. Public transit environments can also play an important role, as handrails of public buses [87], as well as the hands of bus riders [88], in two cities in Portugal were tested positive for contamination by methicillin-resistant Staphylococcus aureus (MRSA).
Some of the greatest concerns for antibiotic-resistant bacteria transmission occur in healthcare environments where contamination and transmission are possible through numerous fomites, ranging from mobile phones [89] to medical devices [90] to surgical tape [91] to doctors’ handbags [92]. While it has been hypothesized that many of these fomites have been important sources, closer investigation often reveals a more nuanced understanding. For example, Julian et al. (2011) sampled the surfaces of cellular phones carried by personnel at a veterinary hospital for both MRSA and methicillin-resistant Staphylococcus pseudintermedius (MRSP). MRSP was isolated from only 2 of 123 phones, and MRSA was isolated from only 1 of 123 phones [93]. Similarly, Missri et al. (2018) assessed bacterial colonization on healthcare workers’ mobile phones in a hospital that were sampled immediately before and 5 min after sanitization with bactericidal wipes [8•]. All phones were colonized with bacteria, and healthcare workers had higher bacterial colonization than administrative staff. However, potential pathogens were detected on approximately one-third of phones (most commonly by Staphylococcus aureus), while only one phone was colonized with MRSA. No multi-drug resistant bacteria were detected. Smibert et al. (2018) swabbed medical staff personal mobile phones, departmental phones, and ICU keyboards and cultured for 94 multi-drug resistant organisms (MRDOs) that had been previously cultured from ICU patients, including 11 MRSA, 2 VRE, and 81 Gram-negative bacteria [9••]. MRSA was isolated from only two phones, and whole-genome sequencing of mobile phone isolates demonstrated the isolates on mobile phones had different single nucleotide polymorphism (SNPs) compared with the clinical isolates, which suggests that these fomites are unlikely to contribute to hospital-acquired MRDOs. Given the ubiquitous nature of bacteria in the built environment, studies that have characterized bacterial colonization alone tend to be less useful for yielding mechanistic or health-relevant insights than those that have targeted specific pathogens and other microbial hazards.
In addition to MRSA, other major microbial hazards in healthcare environments include Clostridioides difficile (C. diff), carbapenem-resistant Enterobacteriaceae, vancomycin-resistant Enterococcus (VRE), and a number of single- and multi-drug-resistant organisms [85]. Haun et al. (2016) reviewed 72 studies that assessed contamination of fomites in healthcare settings and found high variability in contamination rates by fomite type, by microbial agent (including MRSA, Gram-negative rods, enterococci, and C. diff), and by microbiological sampling and analysis technique [10•]. Grimmond et al. (2018) sampled for C. diff on 50 disposable and 50 reusable sharps containers in seven hospitals, finding that 8% and 16% of containers had detectable, albeit sub-infective, levels of C. diff, suggesting that sharps containers are not likely to pose a risk of C. diff transmission [94]. Jackson et al. (2019) sampled the bacterial burden on body sites of ICU patients who were colonized with vancomycin-resistant Enterococcus (VRE) and the healthcare workers (HCWs) who tended to those patients [11••]. HCW contamination on gloves and gowns (i.e., personal protective equipment or PPE) was associated with the VRE burden on body sites of patients with VRE, including perianal, stool, and skin swab samples, suggesting that ICU patients with a higher bacterial burden were more likely to transmit VREs to HCWs via their PPE.
A number of methods for controlling antibiotic-resistant bacteria and other microbial hazards on fomite surfaces have been investigated, including UV light, disinfectant cleaners, material coatings, and others. For example, Mitchell et al. (2019) quantified the doses of UV light that are required to inactivate MRSA, VRE, C. diff, and murine norovirus on stainless steel and Formica laminate fomite surfaces [95]. Reitzel et al. (2014) characterized the ability of a novel chlorhexidine and gentian violet antiseptic coating to kill bacterial and fungal pathogens on the surface of disposable medical gloves, finding that the coating eradicated MRSA, VRE, and multi-drug-resistant Pseudomonas aeruginosa, among others [96]. Despite the effectiveness of antimicrobial cleaners, other studies suggest that caution should be practiced in their use. For example, Hartmann et al. (2016) identified antibiotic resistance genes (ARGs) in settled dust from athletic and educational facilities, and found that ARG abundance was positively correlated with the concentration of antimicrobial chemicals found in the same dust samples [12••]. Similarly, Mahnert et al. (2019) compared the microbial communities and their resistomes (the total antibiotic resistance gene profile of a community) on surfaces of clinical settings using metagenomic genome and plasmid reconstruction, where they found that the microbiome of highly maintained built environments has a different resistome compared with other built environments, as well as a greater diversity of resistance genes [13••]. How these results are best applied is still an active area of research, as ARGs are also natural components of environments rich with bacteria (e.g., soils), and their role in shaping bacteria in indoor environments is not yet well understood.
One promising area of research that has emerged in recent years may offer an alternative to traditional cleaning methods. Unlike antimicrobials that kill microbes, probiotic cleaners that contain spores from Bacillus species (i.e., B. subtilis, B. pumilus, and B. megaterium) are thought to work primarily through biological competition to prevent the survival and proliferation of pathogenic bacteria [97, 98]. Probiotic cleaners have been found to be more effective than traditional cleaning methods, with several studies demonstrating that their use decreased pathogen load on surfaces by an average of 90% more than conventional chemical cleaners (ranging from 70 to 99%; [99, 100]). Furthermore, Caselli et al. (2019) showed that in hospitals where probiotic cleaners were used, the abundance of antibiotic resistance genes on treated surfaces was reduced by up to 99% [101]. Importantly, Caselli et al. (2016) confirmed their safety for use in healthcare facilities by measuring the infection rate from over 30,000 patients across seven facilities and found no evidence of infection by Bacillus spp., regardless of whether patients were at high risk for infection by opportunistic pathogens [102]. Safe sterilization or eradication of antibiotic-resistant bacteria on fomite surfaces remains an active area of study.
The aforementioned studies confirm that not only do pathogenic viruses, non-pathogenic viruses, and bacteria deposit and exist on fomites in the built environment they can also remain viable for hours, or even days, dependent upon the fomite material, microorganism type, and indoor environmental characteristics. From there, are they transmitted to humans and, if so, what are the implications of fomites for human health?
Transmission Between Fomites and Humans
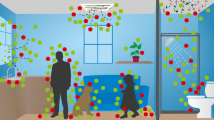
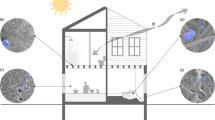
In addition to detecting genetic material and viable microbes on numerous fomite surfaces, it is also crucial to understand the factors that affect the likelihood of transmission between fomites and humans. Three key approaches have been used to provide insight on the importance of fomites and other potential modes of transmission for various microbial hazards and their impacts on human health: (i) experimental measurements of the transfer of microbes to/from fomites and humans, (ii) mathematical modeling of microbial exchange between fomites and humans and subsequent health risks in the context of other exposure pathways (e.g., direct contact and aerosol exposure), and (iii) epidemiological studies designed to elucidate the importance of different modes of transmission in causing disease. Figure 1 illustrates routes of microbial transfer to and from fomites, air, and humans in a typical indoor environment.
Measurements of Microbial Transfer to/from Fomites
Numerous studies have experimentally characterized the transfer of microbes to and from fomites and humans in the built environment, including those that have focused on dynamics at the surface-scale and room-scale.
Surface-Scale
In one surface-level transfer dynamics study, Tuladhar et al. (2013) measured the transfer of human norovirus (NoV) between fingers and fomites, as well as between fingers and food products. They artificially contaminated human finger pads and pressed them on laminate surfaces, stainless steel surfaces, whole tomatoes, and cucumber slices. In addition, they contaminated the surfaces themselves and pressed clean human finger pads against those same surfaces [103]. Initial transfer efficiencies on the first pressing averaged ~ 13%, decreasing over time and after drying of the contaminated finger pads. The transfer efficiency for a viable, infectious virus from surfaces to finger pads was between 2 and 4%, on average, even after 40 min of drying the contaminated surfaces. A number of other surface-scale dynamic studies have focused on the transfer efficiency between different types of organisms and different types of fomites that are common in healthcare settings, such as medical gloves. For example, Moore et al. (2013) evaluated MRSA transmission between different types of gloves worn by HCWs and fomite surfaces, finding that bacterial transfer ranged from ~ 0 to ~ 20% and varied depending on glove material and material hydrophobicity, while the adsorption of simulated body fluids increased bacterial transfer and also made transfer more uniform across glove types [104]. Greene et al. (2015) estimated the transfer efficiency of Acinetobacter baumannii—a drug-resistant healthcare-associated pathogen—with and without latex glove use from the finger pad to a fomite and from a fomite to the finger pad, testing six materials (i.e., glass, stainless steel, porcelain, polypropylene, polycarbonate, and rubber) [14•]. Without gloves, the fomite-to-finger-pad transfer efficiency was ~ 24% and the finger-pad-to-fomite transfer efficiency was ~ 6%. Latex gloves reduced both of these transfer efficiencies by about half, and material type was not a major determining factor. Koenig et al. (2016) measured the transfer efficiency of Staphylococcus aureus between nitrile exam gloves and non-porous fomites via handshaking with another person with gloved hands, touching a plastic cellular phone back, and touching a stainless steel rod [105]. The highest transfer efficiency was with the steel rod, followed by the cellular phone back. Glove-to-glove transfer occurred but had the lowest transfer efficiency among the three scenarios studied. Lopez et al. (2014) quantified fomite-to-finger microbial transfer of Escherichia coli, Staphylococcus aureus, Bacillus thuringiensis spores, and poliovirus 1 seeded on ceramic tile, laminate, and granite after treatment with a disinfectant wipe; microbial transfer was much lower on treated fomites than non-treated fomites (i.e., up to < 0.1% vs. up to 36%, on average) [106]. These studies and others provide helpful quantitative information about the physical microbial exchange to/from fomites and humans, which provides helpful context for fomite transmission and also informs mathematical models of microbial exchange in the built environment.
Room-Scale
Room-level dynamics studies have used a number of experimental approaches to elucidate the importance of fomite transmission to and from humans. For example, Winther et al. (2007) assessed rhinovirus contamination of environmental surfaces by housing 15 adults with naturally acquired rhinovirus colds in a hotel overnight and having them conduct a combination of natural and scripted activities [107]. Thirty-five percent (35%) of 150 environmental sites sampled in the rooms were contaminated with rhinovirus. Moreover, rhinovirus was successfully transferred from surfaces to fingertips in 60% of samples taken 1 h after scripted activities to intentionally transfer the virus from humans to surfaces, as well as in 33% of samples taken 18 h after the scripted activities. Killingley et al. (2016) quantified viral loads recovered from the nostrils of subjects infected with influenza A(H1N1)pdm09 and correlated those amounts with viral loads recovered from their immediate environment in community and hospital settings [15•]. The mean duration of virus shedding was ~ 6 days by PCR (molecular detection) and ~ 4 days by culture (viability detection). Only ~ 5% of surface swabs were PCR positive for influenza, and only 0.3% yielded viable virus; however, room air near a subset of the subjects was also sampled and was found PCR positive for influenza virus in ~ 40% of the samples, suggesting that the importance of aerosol influenza transmission is likely greater than indirect transmission via fomites. Suwantarat et al. (2017) combined microbial sampling with observations of hospitalized patients and reported that patients frequently had direct or indirect interactions with medical equipment and other fomites that are shared among patients, and that those items were often contaminated with health care–associated pathogens [108]. The surfaces that patients interacted with most frequently included medication carts, wheelchairs, food trays, and cleaning carts, resulting in between ~ 0.2 and ~ 1.4 interactions per hour.
Microbial tracers have also been used to investigate fomite transmission at the room-scale with some success. For example, Kunkel et al. (2017) used a human respiratory activity simulator to aerosolize two model organisms—Escherichia coli K12 and bacteriophage T4—in an unoccupied apartment unit operating with four different particle filters installed in the recirculating central forced air heating, ventilation, and air-conditioning (HVAC) system [16•]. Size-resolved aerosol sampling and settle plate swabbing were conducted in multiple locations, and samples were analyzed by DNA extraction and qPCR. DNA from both organisms was detected under all test conditions in all air samples up to 7 m away from the bioaerosol source, with concentrations decreasing at greater distances. A greater fraction of T4 DNA was recovered from the aerosol size fractions smaller than 1 μm than E. coli K12 at all air sampling locations, suggesting that smaller virus-like organisms can transport longer distances than the larger bacterial organisms. Moreover, higher efficiency particle filters in the HVAC system reduced the amount of DNA recovered in air samples, as well as on settle plates located 3–7 m from the source. In another microbial tracer study, Sassi et al. (2018) assessed the amount of surface contamination that occurs in restrooms during toilet flushing using coliphage MS2 added to the toilet bowl [109]. The toilet bowl rim, toilet seat top, and toilet seat underside were contaminated in all tests where no disinfectant was added to the bowl water before flushing, while the addition of disinfectant to the toilet bowl prior to flushing reduced concentrations on fomites after flushing. Similarly, Booth and Frost (2019) used a vomit simulator to investigate the distribution and survival of Feline calicivirus (FCV) as a surrogate for norovirus, demonstrating that viable virus was recovered from almost all samples taken from the floor up to 3 m away from the source, while no air samples contained viable virus [110]. In a highly novel tracer experiment, Reynolds et al. (2019) evaluated microbial transmission in an outpatient clinic and the impact of an ethanol-based disinfectant by placing a viral tracer (bacteriophage MS2) on two fomites at the beginning of the day: a patient room door handle and a front desk pen [17••]. Fomites and the hands of patients and staff were sampled after 2, 3.5, and 6 h. For the disinfectant intervention trials, high-touch surfaces were cleaned 4 h after seeding and sampled 2 h later. The viral tracer was detected on all surfaces and all hands sampled at all three time points, with examination room door handles and nurse station chair arms yielding the greatest concentrations. MS2 concentrations were greatest 2 h after inoculation, and virus concentrations decreased by ~ 94% after application of the disinfectant spray. If one can assume that microbial transfer efficiencies for MS2 are similar to those for other health-relevant organisms (which may not be the case [111]), then tracer studies like this can provide meaningful experimental evidence for the potential for fomite transmission in the built environment.
Combined, both surface-scale and room-scale studies clearly demonstrate that the role of fomites in the transmission of microbes to humans can be an important exposure pathway.
Mathematical Modeling of Fomite Transmission
The combination of quantitative measures of (i) abundance of specific microorganisms on fomite surfaces, (ii) human contact frequency and interactions with fomites, and (iii) microbial transfer efficiency between humans and fomites also makes it possible to quantify the likelihood of microbial exchange and potential health risks using more detailed mathematical models of entire built spaces. These approaches are commonly referred to as quantitative microbial risk assessments (QMRA) [112].
Mechanistic Models
Detailed mechanistic models of disease transmission in the built environment combine (i) probabilistic fate and transport models to estimate the dose of potential pathogens delivered to infection sites of susceptible individuals and (ii) dose-response models to estimate the probability of infection based on estimates of the quantities of pathogens delivered to infection sites [113]. The underlying fate and transport models in these studies commonly use Markov chain models combined with single-zone or multi-zone mass balance models or computational fluid dynamics (CFD) models to simulate physical transport mechanisms, such as bioaerosol emissions, removal by ventilation, and deposition to fomite surfaces. A Markov chain is a random process that undergoes transitions from one state to another on a state space. Physical elements (e.g., room air, skin, and mucus membranes) and pathogen removal mechanisms (e.g., loss of viability, ventilation, and filtration) in the source environment-receptor pathways are represented as states in a discrete-time Markov chain model. Pathogens can be transferred and exchanged between states due to physical mechanisms. Markov chain models have been widely used for estimating doses of influenza virus in several environments, including healthcare facilities and airplanes [114,115,116,117,118,119].
As an early example of a Markov chain model application for estimating transmission routes of a respiratory pathogen, Nicas and Sun (2006) illustrated a hypothetical scenario in which a viral pathogen was emitted by a patient via coughing and transmitted to an attending HCW [115]. The model required detailed inputs on pathogen loads in coughs; deposition rates to and survival on surfaces; airborne inactivation and ventilation rates; rates of pathogen transfer to hands, mucous membranes, and respiratory tract; and dose-response model parameters. Their case study demonstrated the importance of the hand-to-mucous-membrane contact and droplet spray exposure routes for the case study and demonstrated model sensitivity to assumptions for each input parameter.
Other models have been used to simulate the transmission of specific pathogens. For example, Kraay et al. (2018) developed a compartmental model that accounts for fomite transmission of viral pathogens, including contacting fomites after shedding onto those surfaces and shedding onto hands. This model then predicted influenza, rhinovirus, and norovirus transmission in a daycare, subway, office, and school [120]. They predicted that fomite transmission for rhinovirus and norovirus can sustain transmission in all locations, while fomite transmission is likely not sustained for influenza. Xiao et al. (2017) used a multi-agent model to predict the distributions of infection risk during the well-known Ward 8A SARS outbreak in 2003 in the Prince of Wales Hospital in Hong Kong, concluding that the SARS coronavirus was most likely spread via a combination of long-range airborne and fomite routes [18•]. Sze-To et al. (2013) modeled the impacts of surface material, ventilation rates, and human behavior (e.g., close contact rates between individuals) on the transmission of influenza A, respiratory syncytial virus (RSV), and rhinovirus in a hospital ward and aircraft cabin. They predicted that a reduction in close contact rates is more effective than an increase in ventilation rates at decreasing infection risk, and fabric surfaces present a much lower risk of transmission than non-fabric surfaces [121]. Similar results have been obtained for SARS and MERS in other studies as well [122]. For example, Lei et al. (2018) modeled in-flight outbreaks of influenza A H1N1, SARS, and norovirus in an air cabin, and predicted that the dominant route of transmission was a close contact for influenza and contact with fomites for SARS and norovirus [123]. Many others have also developed and applied models for investigating the transmission dynamics of generic [124] and specific pathogens, including influenza [125,126,127,128], norovirus [129], rotavirus [130], and cholera [127].
Improving Model Inputs
A number of recent studies have also focused on improving the accuracy of model inputs to increase the overall accuracy and relevance of mechanistic transmission models. For example, since models have shown that human behaviors (e.g., close human-to-human and human-to-fomite contact rates) are major drivers of variability in infection risk models, several studies have used videography and visual observations to characterize and quantify these inputs. Nicas and Best (2008) videotaped 10 subjects for 3 h while performing office-type work in isolation from others, recording the number of contacts to the eyes, nostrils, and lips [131]. Julian and Pickering (2015) combined videography with a microbial sampling of fomite surfaces to develop fecal indicator bacterial exposure profiles at very high time resolution (i.e., 1-s intervals) in Tanzania [132]. Zhang and Li (2018) characterized more than 3500 person-to-person contacts and 127,000 surface touches in a student office space obtained by video camera. They then used those data as model inputs to predict the intranasal dose of influenza A viruses to students and surrounding fomites and to evaluate the effectiveness of various control strategies [19•]. Hertzberg et al. (2018) chronicled the behaviors and movements of individuals in the economy cabin on single-aisle aircraft during 10 transcontinental US flights to provide information on the movements of passengers and crew that may facilitate disease transmission [133]. They simulated droplet-mediated transmission, predicting that there is a low probability of direct transmission to passengers not seated in close proximity to an infectious passenger. They also collected 229 air and surface samples during flights, and although eight flights were during the flu season, all qPCR assays for 18 common respiratory viruses were negative. Smieszek et al. (2019) used wireless sensors to measure the location and close proximity contacts among individuals at a high school in the USA, which allowed for modeling droplet and aerosol transmission of influenza both in isolation and in combination [134]. Importantly, Greene et al. (2018) used a model of Acinetobacter baumannii to demonstrate how incorrect assumptions for pathogen transfer efficiency between fomites and fingers (and between fingers and fomites) can adversely impact model predicted results [20•].
Another key parameter that affects not only model results but also the underlying mechanistic studies of surface interactions is sample recovery efficiency (SRE). Herzog et al. (2012) demonstrated that the SRE of bacteriophage P22 applied to a number of different fomites under a variety of conditions was most influenced by sampling time, fomite surface area, wetting agent use, and relative humidity [135]. Ganime et al. (2015) evaluated swab sampling as a method to recover murine norovirus 1 (MNV-1) and bacteriophage PP7 from porous, non-porous, and rubberized fomite surfaces, finding a highly variable recovery efficiency ranging from < 1 to 77% [136]. Weir et al. (2016) demonstrated the impact that variability in fomite recovery from surfaces using different sampling methods can have on QMRA model results [21•]. They also reported measurements of the recovery efficiency for enterobacteria phage P22 using combinations of different sampling tools (i.e., swabs and wipes) and eluents (i.e., polysorbate 80, trypticase soy broth, and beef extract) on different non-porous fomites (i.e., aluminum, ceramic, glass, plastic, steel, and wood laminate), finding that polysorbate 80 wipes with a surface area of 10–100 cm2 had the highest sample recovery efficiency. As models continue to be developed, refined, and applied, the importance of having accurate input parameters continues to grow, as it has become increasingly clear that they are essential in the development of realistic and useful models.
Epidemiology of Fomite Transmission
Epidemiological investigations offer the benefit of increasing our understanding of overall disease transmission and attack rates in exposed populations, but they are often limited in their ability to disentangle infectious disease transmission through various exposure routes. In large observational cohort studies, conducting exposure assessments that parse out the various modes of transmission is impractical. As such, researchers often rely on so-called natural experiments, like an outbreak in transportation environments, to make inferences about exposure pathways. For example, in a norovirus outbreak on an airplane, flight crews across several subsequent shifts became ill up to 5 days after an infectious passenger vomited in the airplane, confirming research on fomite transmission of norovirus [137]. For other viruses, like influenza, where fomite transmission is less well understood, natural experiments offer less clarity. For example, in another study of an airplane disease outbreak, 54 passengers aboard an airplane that experienced a 3-h ground delay with no ventilation and one infectious person, 72% of passengers developed symptoms [138].
As an example of an indirect human epidemiological study, Knox et al. (2012) investigated intra-household S. aureus transmission using a sample of multiple member households in New York City, NY [139]. Household members and standardized household environmental fomites were swabbed and cultured for S. aureus. Staphylococcus aureus colonized individuals in 62% of households and contaminated the environment in 54% of households. Environmental contamination was associated with transmission among households. Somewhat similarly, Cowling et al. (2013) applied a mathematical model to empirical data on influenza A transmission from randomized controlled trials of hand hygiene and surgical face masks in Hong Kong and Bangkok households [140]. They used inferences on the importance of close-range and long-range transmission modes, including information on the timing of secondary infections and apparent differences in the clinical presentation of secondary infections resulting from aerosol transmission. They estimated that long-range aerosol transmission via small particles (i.e., droplet nuclei) accounted for approximately half of all influenza transmission events. Lee and Wong (2015) conducted an epidemiological analysis of MERS-CoV transmission in South Korea and concluded that fomite transmission might have explained a significant proportion of the infections that occurred in the absence of direct contact with infected cases [141]. Despite some of these useful approaches, Kutter et al. (2018) recently summarized the state of knowledge of dominant transmission routes for a number of human respiratory viruses and noted that many studies on inter-human transmission routes remain inconclusive [22•].
Intervention studies, while rare and difficult to conduct, provide the best opportunity to empirically isolate transmission pathways. In an early study on this subject, Dick et al. (1987) experimentally investigated the plausible routes of transmission of rhinovirus colds by infecting healthy male adults with rhinovirus type 16, waiting for the onset of symptoms, and then having the infected individuals play cards with susceptible male adults in a room for 12 h [142]. Some of the recipients were allowed to function normally, retaining the ability to touch the cards and their faces, meaning that infection could plausibly occur via a combination of aerosol, direct contact, and/or indirect fomite contact. Meanwhile, other recipients were restrained with a harness, such that they could not touch their faces, thereby eliminating the possibility of fomite transmission. Strikingly, there was no statistically significant difference in infection rates between the two groups, suggesting that aerosol transmission was the dominant mode of transmission among individuals. Moreover, another experiment reported in the same study attempted to spread rhinovirus via heavily contaminated fomites alone, and no infections occurred among the recipients. This highly novel study, with an approach that has not been repeated for other organisms or other settings to date, provided unique epidemiological evidence of the importance of aerosol transmission—and the lack of importance of fomite transmission—for rhinovirus colds in humans.
Conclusions
This review summarizes our understanding of fomite contamination, microbial survival, microbial exchange, and associated human health risks in the built environment. Past efforts have relied on a combination of empirical measurements, mathematical models, and epidemiological approaches to yield novel insight into the magnitude of health risks to humans, the important modes of transmission, and therefore, effective strategies to use for controlling exposures. However, despite numerous recent advances, significant knowledge gaps remain regarding the dynamics of the transmission of infectious disease and other microbial hazards, the relative importance of long-range aerosol, short-range aerosol, direct contact, and fomites, and effective means for controlling exposures and reducing human health risks. Interdisciplinary research that integrates public health, microbiology, engineering, and architectural design is needed to address these knowledge gaps and to inform our current design standards and guidelines.
Change history
19 October 2019
The article Microbial Exchange via Fomites and Implications for Human Health, written by Brent Stephens, Parham Azimi, Megan S. Thoemmes, Mohammad Heidarinejad, Joseph G. Allen, and Jack A. Gilbert, was originally published electronically on the publisher’s internet portal
References
Papers of particular interest, published recently, have been highlighted as: • Of importance •• Of major importance
Nutton V. The seeds of disease: an explanation of contagion and infection from the Greeks to the renaissance. Med Hist. 1983;27:1–34.
Klepeis NE, Nelson WC, Ott WR, Robinson JP, Tsang AM, Switzer P, et al. The National Human Activity Pattern Survey (NHAPS): a resource for assessing exposure to environmental pollutants. J Expo Anal Environ Epidemiol. 2001;11:231–52.
•• Lax S, Smith DP, Hampton-Marcell J, Owens SM, Handley KM, Scott NM, et al. Longitudinal analysis of microbial interaction between humans and the indoor environment. Science. 2014;345:1048–52. This study characterized the longitudinal succession of bacterial communities on indoor surfaces in 7 homes, finding that microbial communities were largely sourced from humans and were identifiable by family.
•• Chase J, Fouquier J, Zare M, Sonderegger DL, Knight R, Kelley ST, et al. Geography and location are the primary drivers of office microbiome composition. mSystems. 2016;1:e00022-16. This study investigated the impacts of a wide variety of parameters including geography, material type, human interaction, location in a room, seasonal variation, and indoor and microenvironmental parameters on bacterial communities in offices.
• Stobnicka A, Gołofit-Szymczak M, Wójcik-Fatla A, Zając V, Korczyńska-Smolec J, Górny RL. Prevalence of human parainfluenza viruses and noroviruses genomes on office fomites. Food Environ Virol. 2018;10:133–40. This study evaluated the potential role of office fomites in respiratory and enteric virus transmission by assessing the occurrence of viruses on 130 surfaces from both open-space and non-open-space rooms in office buildings during a 9-month period.
• Thompson K-A, Bennett AM. Persistence of influenza on surfaces. J Hosp Infect. 2017;95:194–9. This study assessed the viability of five influenza strains seeded on three surfaces over the course of several weeks, finding that viable influenza was recovered from surfaces for up to 2 weeks while influenza genetic material could be detected by PCR for more than 7 weeks.
• Malcolm KC, Caceres SM, Honda JR, Davidson RM, Epperson LE, Strong M, et al. Mycobacterium abscessus displays fitness for fomite transmission. Appl Environ Microbiol. 2017;83. https://doi.org/10.1128/AEM.00562-17. This article demonstratedM. abscessusgrowth on surfaces is enhanced in the presence of house dust, surviving desiccation for up to 2 weeks.
• Missri L, Smiljkovski D, Prigent G, Lesenne A, Obadia T, Joumaa M, et al. Bacterial colonization of healthcare workers’ mobile phones in the ICU and effectiveness of sanitization. J Occup Environ Hyg. 2018:1–4. This article assessed the prevalence of bacterial colonization of 56 healthcare workers’ mobile phones in an intensive care unit both immediately before and 5 min after sanitization of the phones with bactericidal wipes, finding that colonization with pathogens was frequent but colonization with multi-drug resistant bacteria was rare.
•• Smibert OC, Aung AK, Woolnough E, Carter GP, Schultz MB, Howden BP, et al. Mobile phones and computer keyboards: unlikely reservoirs of multidrug-resistant organisms in the tertiary intensive care unit. J Hosp Infect. 2018;99:295–8. This article sampled for a large number of multi-drug resistant organisms (MRDOs) on medical staff personal mobile phones, departmental phones, and ICU keyboards, finding that MRSA was isolated from only two phones and that these fomites appear unlikely to contribute much to hospital-acquired MRDOs.
• Haun N, Hooper-Lane C, Safdar N. Healthcare personnel attire and devices as fomites: a systematic review. Infect Control Hosp Epidemiol. 2016;37:1367–73. This article reviewed a large number of studies that assessed contamination of fomites in healthcare settings and found high variability in contamination rates by fomite type, microbial agent, and sampling and analysis technique.
•• Jackson SS, Harris AD, Magder LS, Stafford KA, Johnson JK, Miller LG, et al. Bacterial burden is associated with increased transmission to health care workers from patients colonized with vancomycin-resistant Enterococcus. Am J Infect Control. 2019;47:13–7. This article found that the bacterial contamination of HCW gloves and gowns was associated with the vancomycin-resistantEnterococcus(VRE) on body sites of patients with VRE, suggesting that ICU patients with a higher bacterial burden were more likely to transmit VREs to HCWs.
•• Hartmann EM, Hickey R, Hsu T, Betancourt Román CM, Chen J, Schwager R, et al. Antimicrobial chemicals are associated with elevated antibiotic resistance genes in the indoor dust microbiome. Environ Sci Technol. 2016;50:9807–15. This article was the first to find an association between antibiotic resistance genes and antimicrobial chemicals in dust samples from indoor environments.
•• Mahnert A, Moissl-Eichinger C, Zojer M, Bogumil D, Mizrahi I, Rattei T, et al. Man-made microbial resistances in built environments. Nat Commun. 2019;10:968. This article demonstrated that the loss of microbial diversity on surfaces is correlated with an increase in antibiotic resistance, suggesting there is a need for implementing strategies to restore bacterial diversity in certain built environments.
• Greene C, Vadlamudi G, Eisenberg M, Foxman B, Koopman J, Xi C. Fomite-fingerpad transfer efficiency (pick-up and deposit) of Acinetobacter baumannii—with and without a latex glove. Am J Infect Control. 2015;43:928–34. This article estimated the transfer efficiency ofAcinetobacter baumanniiwith and without latex glove use from the finger pad to a fomite and from a fomite to the finger pad.
• Killingley B, Greatorex J, Digard P, Wise H, Garcia F, Varsani H, et al. The environmental deposition of influenza virus from patients infected with influenza A(H1N1)pdm09: implications for infection prevention and control. J Infect Public Health. 2016;9:278–88. This article used a novel approach to quantify and correlate the amount of virus recovered from the nares of infected subjects with that recovered from their immediate environment in the community and hospital settings.
• Kunkel SA, Azimi P, Zhao H, Stark BC, Stephens B. Quantifying the size-resolved dynamics of indoor bioaerosol transport and control. Indoor Air. 2017;27:977–87. This article used a human respiratory activity simulator to aerosolize two model organisms and measured the abundance of microbes on surfaces and in bioaerosols in multiple locations in an apartment unit operating with different HVAC particle filters, finding that DNA from both organisms was detected under all test conditions in all air samples up to 7 m away from the source, with concentrations decreasing at greater distances from the bioaerosol source and with higher efficiency filters.
•• Reynolds KA, Sexton JD, Pivo T, Humphrey K, Leslie RA, Gerba CP. Microbial transmission in an outpatient clinic and impact of an intervention with an ethanol-based disinfectant. Am J Infect Control. 2019;47:128–32. This article used a novel method to evaluate microbial transmission in an outpatient clinic and the impact of a disinfectant by placing a viral tracer on two fomites at the beginning of the day and tracking its presence throughout the remainder of the day.
• Xiao S, Li Y, Wong T, Hui DSC. Role of fomites in SARS transmission during the largest hospital outbreak in Hong Kong. PLoS One. 2017;12:e0181558. This article used a multi-agent model to predict the distributions of infection risk during the well-known Ward 8A SARS outbreak in 2003 in the Prince of Wales Hospital in Hong Kong, concluding that the SARS coronavirus was most likely spread via a combination of long-range airborne and fomite routes.
• Zhang N, Li Y. Transmission of influenza A in a student office based on realistic person-to-person contact and surface touch behaviour. Int J Environ Res Public Health. 2018;15:1699. This study simulated the transmission of influenza A virus in a graduate student office via three transmission routes, informed by novel data on more than 3500 person-to-person contacts and 127,000 surface touches obtained by video camera recording.
• Greene C, Ceron NH, Eisenberg MC, Koopman J, Miller JD, Xi C, et al. Asymmetric transfer efficiencies between fomites and fingers: impact on model parameterization. Am J Infect Control. 2018;46:620–6. This article used a model ofAcinetobacter baumanniito demonstrate the effects that incorrect assumptions for pathogen transfer efficiency between fomites and fingers (and between fingers and fomites) can have on QMRA model predicted results.
• Weir MH, Shibata T, Masago Y, Cologgi DL, Rose JB. Effect of surface sampling and recovery of viruses and non-spore-forming Bacteria on a quantitative microbial risk assessment model for fomites. Environ Sci Technol. 2016;50:5945–52. This article demonstrated the impact that QMRA model inputs can have on model results, including recovery efficiency from several non-porous fomites, fomite material, surface area, recovery tool, and initial fomite concentrations.
• Kutter JS, Spronken MI, Fraaij PL, Fouchier RA, Herfst S. Transmission routes of respiratory viruses among humans. Curr Opin Virol. 2018;28:142–51. This article summarized the state of knowledge of dominant transmission routes for a number of human respiratory viruses and noted that many studies on inter-human transmission routes remain inconclusive.
•• Dannemiller KC, Gent JF, Leaderer BP, Peccia J. Influence of housing characteristics on bacterial and fungal communities in homes of asthmatic children. Indoor Air. 2016;26:179–92. This article found associations between housing characteristics and bacterial and fungal communities in homes of asthmatic children.
• O’Connor GT, Lynch SV, Bloomberg GR, et al. Early-life home environment and risk of asthma among inner-city children. J Allergy Clin Immunol. 2018;141:1468–75. This article found that higher indoor levels of pet or pest allergens in infancy, as well as the abundance of some bacterial taxa, were associated with a lower risk of asthma in a birth cohort of high-risk inner-city children.
Kelley ST, Gilbert JA. Studying the microbiology of the indoor environment. Genome Biol. 2013;14:202.
Konya T, Scott JA. Recent advances in the microbiology of the built environment. Curr Sustain Renew Energy Rep. 2014;1:35–42.
Gilbert JA, Stephens B. Microbiology of the built environment. Nat Rev Microbiol. 2018;16:661–70.
Adams RI, Bhangar S, Pasut W, Arens EA, Taylor JW, Lindow SE, et al. Chamber bioaerosol study: outdoor air and human occupants as sources of indoor airborne microbes. PLoS One. 2015;10:e0128022.
Hospodsky D, Qian J, Nazaroff WW, Yamamoto N, Bibby K, Rismani-Yazdi H, et al. Human occupancy as a source of indoor airborne bacteria. PLoS One. 2012;7:e34867.
Qian J, Hospodsky D, Yamamoto N, Nazaroff WW, Peccia J. Size-resolved emission rates of airborne bacteria and fungi in an occupied classroom. Indoor Air. 2012;22:339–51.
Adams RI, Miletto M, Taylor JW, Bruns TD. Dispersal in microbes: fungi in indoor air are dominated by outdoor air and show dispersal limitation at short distances. ISME J. 2013;7:1262–73.
Hospodsky D, Yamamoto N, Nazaroff WW, Miller D, Gorthala S, Peccia J. Characterizing airborne fungal and bacterial concentrations and emission rates in six occupied children’s classrooms. Indoor Air. 2015;25:641–52.
Lax S, Sangwan N, Smith D, et al. Bacterial colonization and succession in a newly opened hospital. Sci Transl Med. 2017;9:eaah6500.
Lai PS, Allen JG, Hutchinson DS, Ajami NJ, Petrosino JF, Winters T, et al. Impact of environmental microbiota on human microbiota of workers in academic mouse research facilities: an observational study. PLoS One. 2017;12:e0180969.
Adams RI, Bateman AC, Bik HM, Meadow JF. Microbiota of the indoor environment: a meta-analysis. Microbiome. 2015;3. https://doi.org/10.1186/s40168-015-0108-3.
Kembel SW, Jones E, Kline J, Northcutt D, Stenson J, Womack AM, et al. Architectural design influences the diversity and structure of the built environment microbiome. ISME J. 2012;6:1469–79.
Kembel SW, Meadow JF, O’Connor TK, Mhuireach G, Northcutt D, Kline J, et al. Architectural design drives the biogeography of indoor bacterial communities. PLoS One. 2014;9:e87093.
Meadow JF, Altrichter AE, Kembel SW, Moriyama M, O’Connor TK, Womack AM, et al. Bacterial communities on classroom surfaces vary with human contact. Microbiome. 2014;2:7.
Meadow JF, Altrichter AE, Kembel SW, Kline J, Mhuireach G, Moriyama M, et al. Indoor airborne bacterial communities are influenced by ventilation, occupancy, and outdoor air source. Indoor Air. 2014;24:41–8.
Meadow JF, Altrichter AE, Bateman AC, Stenson J, Brown GZ, Green JL, et al. Humans differ in their personal microbial cloud. PeerJ. 2015;3:e1258.
Stephens B. What have we learned about the microbiomes of indoor environments? mSystems. 2016;1:e00083-16.
Barberán A, Dunn RR, Reich BJ, Pacifici K, Laber EB, Menninger HL, et al. The ecology of microscopic life in household dust. Proc R Soc B Biol Sci. 2015;282:20151139.
Gibbons SM. The built environment is a microbial wasteland. mSystems. 2016;1:e00033-16.
Dannemiller KC, Mendell MJ, Macher JM, Kumagai K, Bradman A, Holland N, et al. Next-generation DNA sequencing reveals that low fungal diversity in house dust is associated with childhood asthma development. Indoor Air. 2014;24:236–47.
Karvonen AM, Hyvärinen A, Rintala H, Korppi M, Täubel M, Doekes G, et al. Quantity and diversity of environmental microbial exposure and development of asthma: a birth cohort study. Allergy. 2014;69:1092–101.
Lynch SV, Wood RA, Boushey H, et al. Effects of early-life exposure to allergens and bacteria on recurrent wheeze and atopy in urban children. J Allergy Clin Immunol. 2014;134:593–601.e12.
Ege MJ, Mayer M, Normand A-C, Genuneit J, Cookson WOCM, Braun-Fahrländer C, et al. Exposure to environmental microorganisms and childhood asthma. N Engl J Med. 2011;364:701–9.
Stein MM, Hrusch CL, Gozdz J, Igartua C, Pivniouk V, Murray SE, et al. Innate immunity and asthma risk in Amish and Hutterite farm children. N Engl J Med. 2016;375:411–21.
Schuijs MJ, Willart MA, Vergote K, Gras D, Deswarte K, Ege MJ, et al. Farm dust and endotoxin protect against allergy through A20 induction in lung epithelial cells. Science. 2015;349:1106–10.
Gibbons SM, Schwartz T, Fouquier J, Mitchell M, Sangwan N, Gilbert JA, et al. Ecological succession and viability of human-associated microbiota on restroom surfaces. Appl Environ Microbiol. 2015;81:765–73.
Hegarty B, Dannemiller KC, Peccia J. Gene expression of indoor fungal communities under damp building conditions: implications for human health. Indoor Air. 2018;28:548–58.
Hu J, Ben Maamar S, Glawe AJ, Gottel N, Gilbert JA, Hartmann EM. Impacts of indoor surface finishes on bacterial viability. Indoor Air. 2019. https://doi.org/10.1111/ina.12558.
Flores GE, Bates ST, Knights D, Lauber CL, Stombaugh J, Knight R, et al. Microbial biogeography of public restroom surfaces. PLoS One. 2011;6:e28132.
Flores GE, Bates ST, Caporaso JG, Lauber CL, Leff JW, Knight R, et al. Diversity, distribution and sources of bacteria in residential kitchens. Environ Microbiol. 2013;15:588–96.
Dunn RR, Fierer N, Henley JB, Leff JW, Menninger HL. Home life: factors structuring the bacterial diversity found within and between homes. PLoS One. 2013;8:e64133.
Kelley ST, Theisen U, Angenent LT, St. Amand A, Pace NR. Molecular analysis of shower curtain biofilm microbes. Appl Environ Microbiol. 2004;70:4187–92.
Adams RI, Lymperopoulou DS, Misztal PK, de Cassia Pessotti R, Behie SW, Tian Y, et al. Microbes and associated soluble and volatile chemicals on periodically wet household surfaces. Microbiome. 2017;5. https://doi.org/10.1186/s40168-017-0347-6.
Yano T, Kubota H, Hanai J, Hitomi J, Tokuda H. Stress tolerance of methylobacterium biofilms in bathrooms. Microbes Environ. 2012. https://doi.org/10.1264/jsme2.ME12146.
Prussin AJ, Garcia EB, Marr LC. Total concentrations of virus and bacteria in indoor and outdoor air. Environ Sci Technol Lett. 2015;150310105417006.
Boone S, Gerba C. The occurrence of influenza A virus on household and day care center fomites. J Infect. 2005;51:103–9.
Bright KR, Boone SA, Gerba CP. Occurrence of bacteria and viruses on elementary classroom surfaces and the potential role of classroom hygiene in the spread of infectious diseases. J Sch Nurs. 2010;26:33–41.
Jones EL, Kramer A, Gaither M, Gerba CP. Role of fomite contamination during an outbreak of norovirus on houseboats. Int J Environ Health Res. 2007;17:123–31.
Pappas DE, Hendley JO, Schwartz RH. Respiratory viral RNA on toys in pediatric office waiting rooms. Pediatr Infect Dis J. 2010;29:102–4.
Gralton J, McLaws M-L, Rawlinson WD. Personal clothing as a potential vector of respiratory virus transmission in childcare settings: personal clothing as transmission vector. J Med Virol. 2015;87:925–30.
Butz AM, Fosarelli P, Dick J, Cusack T, Yolken R. Prevalence of rotavirus on high-risk fomites in day-care facilities. Pediatrics. 1993;92:202–5.
Soule H, Genoulaz O, Gratacap-Cavallier B, Mallaret MR, Morand P, François P, et al. Monitoring rotavirus environmental contamination in a pediatric unit using polymerase chain reaction. Infect Control Hosp Epidemiol. 1999;20:432–4.
Dowell SF, Simmerman JM, Erdman DD, Wu J-SJ, Chaovavanich A, Javadi M, et al. Severe acute respiratory syndrome coronavirus on hospital surfaces. Clin Infect Dis. 2004;39:652–7.
Boone SA, Gerba CP. The prevalence of human parainfluenza virus 1 on indoor office fomites. Food Environ Virol. 2010;2:41–6.
Ganime AC, Carvalho-Costa FA, Santos M, Costa Filho R, Leite JPG, Miagostovich MP. Viability of human adenovirus from hospital fomites: viability of human adenovirus from fomites. J Med Virol. 2014;86:2065–9.
Khan RM, Al-Dorzi HM, Al Johani S, Balkhy HH, Alenazi TH, Baharoon S, et al. Middle East respiratory syndrome coronavirus on inanimate surfaces: a risk for health care transmission. Am J Infect Control. 2016;44:1387–9.
Weber TP, Stilianakis NI. Inactivation of influenza A viruses in the environment and modes of transmission: a critical review. J Infect. 2008;57:361–73.
Bean B, Moore BM, Sterner B, Peterson LR, Gerding DN, Balfour HH. Survival of influenza viruses on environmental surfaces. J Infect Dis. 1982;146:47–51.
Greatorex JS, Digard P, Curran MD, Moynihan R, Wensley H, Wreghitt T, et al. Survival of influenza A(H1N1) on materials found in households: implications for infection control. PLoS One. 2011;6:e27932.
Mukherjee DV, Cohen B, Bovino ME, Desai S, Whittier S, Larson EL. Survival of influenza virus on hands and fomites in community and laboratory settings. Am J Infect Control. 2012;40:590–4.
Thomas Y, Vogel G, Wunderli W, Suter P, Witschi M, Koch D, et al. Survival of influenza virus on banknotes. Appl Environ Microbiol. 2008;74:3002–7.
Oxford J, Berezin EN, Courvalin P, Dwyer DE, Exner M, Jana LA, et al. The survival of influenza A(H1N1)pdm09 virus on 4 household surfaces. Am J Infect Control. 2014;42:423–5.
Perry KA, Coulliette AD, Rose LJ, Shams AM, Edwards JR, Noble-Wang JA. Persistence of influenza A (H1N1) virus on stainless steel surfaces. Appl Environ Microbiol. 2016;82:3239–45.
McDevitt J, Rudnick S, First M, Spengler J. Role of absolute humidity in the inactivation of influenza viruses on stainless steel surfaces at elevated temperatures. Appl Environ Microbiol. 2010;76:3943–7.
Shaman J, Kohn M. Absolute humidity modulates influenza survival, transmission, and seasonality. Proc Natl Acad Sci. 2009;106:3243–8.
Sakaguchi H, Wada K, Kajioka J, Watanabe M, Nakano R, Hirose T, et al. Maintenance of influenza virus infectivity on the surfaces of personal protective equipment and clothing used in healthcare settings. Environ Health Prev Med. 2010;15:344–9.
Boone SA, Gerba CP. Significance of fomites in the spread of respiratory and enteric viral disease. Appl Environ Microbiol. 2007;73:1687–96.
van Doremalen N, Bushmaker T, Munster VJ. Stability of Middle East respiratory syndrome coronavirus (MERS-CoV) under different environmental conditions. Euro Surveill. 2013;18.
Marks LR, Reddinger RM, Hakansson AP. Biofilm formation enhances fomite survival of Streptococcus pneumoniae and Streptococcus pyogenes. Infect Immun. 2014;82:1141–6.
Jones TM, Lutz EA. Environmental survivability and surface sampling efficiencies for Pseudomonas aeruginosa on various fomites. J Environ Health. 2014;76:16–20.
CDC. Antibiotic resistance threats in the United States: U.S. Department of Health and Human Services Centers for Disease Control and Prevention; 2013.
Davis MF, Iverson SA, Baron P, Vasse A, Silbergeld EK, Lautenbach E, et al. Household transmission of meticillin-resistant Staphylococcus aureus and other staphylococci. Lancet Infect Dis. 2012;12:703–16.
Simões RR, Aires-de-Sousa M, Conceição T, Antunes F, da Costa PM, de Lencastre H. High prevalence of EMRSA-15 in Portuguese public buses: a worrisome finding. PLoS One. 2011;6:e17630.
Conceição T, Diamantino F, Coelho C, de Lencastre H, Aires-de-Sousa M. Contamination of public buses with MRSA in Lisbon, Portugal: a possible transmission route of major MRSA clones within the community. PLoS One. 2013;8:e77812.
Bhoonderowa A, Gookool S, Biranjia-Hurdoyal SD. The importance of mobile phones in the possible transmission of bacterial infections in the community. J Community Health. 2014;39:965–7.
Kanamori H, Rutala WA, Weber DJ. The role of patient care items as a fomite in healthcare-associated outbreaks and infection prevention. Clin Infect Dis. 2017;65:1412–9.
Harris PNA, Ashhurst-Smith C, Berenger SJ, Shoobert A, Ferguson JK. Adhesive tape in the health care setting: another high-risk fomite? Med J Aust. 2011;196:34.
Feldman J, Feldman J, Feldman M. Women doctors’ purses as an unrecognized fomite. Del Med J. 2012;84:277–80.
Julian T, Singh A, Rousseau J, Weese J. Methicillin-resistant staphylococcal contamination of cellular phones of personnel in a veterinary teaching hospital. BMC Res Notes. 2012;5:193.
Grimmond T, Neelakanta A, Miller B, Saiyed A, Gill P, Cadnum J, et al. A microbiological study to investigate the carriage and transmission-potential of Clostridium difficile spores on single-use and reusable sharps containers. Am J Infect Control. 2018;46:1154–9.
Mitchell JB, Sifuentes LY, Wissler A, Abd-Elmaksoud S, Lopez GU, Gerba CP. Modelling of ultraviolet light inactivation kinetics of methicillin-resistant Staphylococcus aureus , vancomycin-resistant Enterococcus , Clostridium difficile spores and murine norovirus on fomite surfaces. J Appl Microbiol. 2019;126:58–67.
Reitzel R, Rosenblatt J, Jiang Y, Hachem R, Raad I. Disposable gendine antimicrobial gloves for preventing transmission of pathogens in health care settings. Am J Infect Control. 2014;42:55–9.
Falagas ME, Makris GC. Probiotic bacteria and biosurfactants for nosocomial infection control: a hypothesis. J Hosp Infect. 2009;71:301–6.
Vandini A, Temmerman R, Frabetti A, Caselli E, Antonioli P, Balboni PG, et al. Hard surface biocontrol in hospitals using microbial-based cleaning products. PLoS One. 2014;9:e108598.
Caselli E, D’Accolti M, Vandini A, Lanzoni L, Camerada MT, Coccagna M, et al. Impact of a probiotic-based cleaning intervention on the microbiota ecosystem of the hospital surfaces: focus on the Resistome Remodulation. PLoS One. 2016;11:e0148857.
Caselli E, Brusaferro S, Coccagna M, Arnoldo L, Berloco F, Antonioli P, et al. Reducing healthcare-associated infections incidence by a probiotic-based sanitation system: a multicentre, prospective, intervention study. PLoS One. 2018;13:e0199616.
Caselli E, Arnoldo L, Rognoni C, D’Accolti M, Soffritti I, Lanzoni L, et al. Impact of a probiotic-based hospital sanitation on antimicrobial resistance and HAI-associated antimicrobial consumption and costs: a multicenter study. Infect Drug Resist. 2019;12:501–10.
Caselli E, Antonioli P, Mazzacane S. Safety of probiotics used for hospital environmental sanitation. J Hosp Infect. 2016;94:193–4.
Tuladhar E, Hazeleger WC, Koopmans M, Zwietering MH, Duizer E, Beumer RR. Transfer of noroviruses between fingers and fomites and food products. Int J Food Microbiol. 2013;167:346–52.
Moore G, Dunnill CW, Wilson APR. The effect of glove material upon the transfer of methicillin-resistant Staphylococcus aureus to and from a gloved hand. Am J Infect Control. 2013;41:19–23.
Koenig DW, Korir-Morrison C, Hoffman DR. Transfer efficiency of Staphylococcus aureus between nitrile exam gloves and nonporous fomites. Am J Infect Control. 2016;44:245–6.
Lopez GU, Kitajima M, Havas A, Gerba CP, Reynolds KA. Evaluation of a disinfectant wipe intervention on fomite-to-finger microbial transfer. Appl Environ Microbiol. 2014;80:3113–8.
Winther B, McCue K, Ashe K, Rubino JR, Hendley JO. Environmental contamination with rhinovirus and transfer to fingers of healthy individuals by daily life activity. J Med Virol. 2007;79:1606–10.
Suwantarat N, Supple LA, Cadnum JL, Sankar T, Donskey CJ. Quantitative assessment of interactions between hospitalized patients and portable medical equipment and other fomites. Am J Infect Control. 2017;45:1276–8.
Sassi HP, Reynolds KA, Pepper IL, Gerba CP. Evaluation of hospital-grade disinfectants on viral deposition on surfaces after toilet flushing. Am J Infect Control. 2018;46:507–11.
Booth CM, Frost G. Potential distribution of viable norovirus post simulated vomiting. J Hosp Infect. 2019;102:304–10. https://doi.org/10.1016/j.jhin.2019.02.010.
Pitol AK, Bischel HN, Boehm AB, Kohn T, Julian TR. Transfer of enteric viruses adenovirus and Coxsackie virus and bacteriophage MS2 from liquid to human skin. Appl Environ Microbiol. 2018;84. https://doi.org/10.1128/AEM.01809-18.
Haas CN, Rose JB, Gerba CP. Quantitative microbial risk assessment: Haas/quantitative microbial risk assessment; 2014. https://doi.org/10.1002/9781118910030.
Sze To GN, Chao CYH. Review and comparison between the Wells-Riley and dose-response approaches to risk assessment of infectious respiratory diseases. Indoor Air. 2010;20:2–16.
Nicas M, Jones RM. Relative contributions of four exposure pathways to influenza infection risk. Risk Anal. 2009;29:1292–303.
Nicas M, Sun G. An integrated model of infection risk in a health-care environment. Risk Anal. 2006;26:1085–96.
Jones RM, Masago Y, Bartrand T, Haas CN, Nicas M, Rose JB. Characterizing the risk of infection from Mycobacterium tuberculosis in commercial passenger aircraft using quantitative microbial risk assessment. Risk Anal. 2009;29:355–65.
Jones RM, Adida E. Influenza infection risk and predominate exposure route: uncertainty analysis. Risk Anal. 2011;31:1622–31.
Jones RM, Nicas M. Benchmarking of a Markov multizone model of contaminant transport. Ann Occup Hyg. 2014;58:1018–31.
Jones RM, Nicas M. Experimental evaluation of a Markov multizone model of particulate contaminant transport. Ann Occup Hyg. 2014;58:1032–45.
Kraay ANM, Hayashi MAL, Hernandez-Ceron N, Spicknall IH, Eisenberg MC, Meza R, et al. Fomite-mediated transmission as a sufficient pathway: a comparative analysis across three viral pathogens. BMC Infect Dis. 2018;18:540. https://doi.org/10.1186/s12879-018-3425-x.
Sze-To GN, Yang Y, Kwan JKC, Yu SCT, Chao CYH. Effects of surface material, ventilation, and human behavior on indirect contact transmission risk of respiratory infection: effects of different factors on indirect contact risk. Risk Anal. 2014;34:818–30.
Otter JA, Donskey C, Yezli S, Douthwaite S, Goldenberg SD, Weber DJ. Transmission of SARS and MERS coronaviruses and influenza virus in healthcare settings: the possible role of dry surface contamination. J Hosp Infect. 2016;92:235–50.
Lei H, Li Y, Xiao S, Lin C-H, Norris SL, Wei D, et al. Routes of transmission of influenza A H1N1, SARS CoV, and norovirus in air cabin: comparative analyses. Indoor Air. 2018;28:394–403.
Li S, Eisenberg JNS, Spicknall IH, Koopman JS. Dynamics and control of infections transmitted from person to person through the environment. Am J Epidemiol. 2009;170:257–65.
Atkinson MP, Wein LM. Quantifying the routes of transmission for pandemic influenza. Bull Math Biol. 2008;70:820–67.
Xiao S, Tang JW, Hui DS, Lei H, Yu H, Li Y. Probable transmission routes of the influenza virus in a nosocomial outbreak. Epidemiol Infect. 2018;146:1114–22.
Breban R. Role of environmental persistence in pathogen transmission: a mathematical modeling approach. J Math Biol. 2013;66:535–46.
Zhao J, Eisenberg JE, Spicknall IH, Li S, Koopman JS. Model analysis of fomite mediated influenza transmission. PLoS One. 2012;7:e51984.
Canales RA, Reynolds KA, Wilson AM, Fankem SLM, Weir MH, Rose JB, et al. Modeling the role of fomites in a norovirus outbreak. J Occup Environ Hyg. 2019;16:16–26.
Julian TR, Canales RA, Leckie JO, Boehm AB. A model of exposure to rotavirus from nondietary ingestion iterated by simulated intermittent contacts. Risk Anal. 2009;29:617–32.
Nicas M, Best D. A study quantifying the hand-to-face contact rate and its potential application to predicting respiratory tract infection. J Occup Environ Hyg. 2008;5:347–52.
Julian TR, Pickering AJ. A pilot study on integrating videography and environmental microbial sampling to model fecal bacterial exposures in peri-urban Tanzania. PLoS One. 2015;10:e0136158.
Hertzberg VS, Weiss H, Elon L, Si W, Norris SL, The FlyHealthy Research Team. Behaviors, movements, and transmission of droplet-mediated respiratory diseases during transcontinental airline flights. Proc Natl Acad Sci. 2018;115:3623–7.
Smieszek T, Lazzari G, Salathé M. Assessing the dynamics and control of droplet- and aerosol-transmitted influenza using an indoor positioning system. Sci Rep. 2019;9. https://doi.org/10.1038/s41598-019-38825-y.
Herzog AB, Pandey AK, Reyes-Gastelum D, Gerba CP, Rose JB, Hashsham SA. Evaluation of sample recovery efficiency for bacteriophage P22 on fomites. Appl Environ Microbiol. 2012;78:7915–22.
Ganime AC, Leite JPG, de Abreu Corrêa A, Melgaço FG, Carvalho-Costa FA, Miagostovich MP. Evaluation of the swab sampling method to recover viruses from fomites. J Virol Methods. 2015;217:24–7.
Thornley CN, Emslie NA, Sprott TW, Greening GE, Rapana JP. Recurring norovirus transmission on an airplane. Clin Infect Dis. 2011;53:515–20.
Moser MR, Bender TR, Margolis HS, Noble GR, Kendal AP, Ritter DG. An outbreak of influenza aboard a commercial airliner. Am J Epidemiol. 1979;110:1–6.
Knox J, Uhlemann A-C, Miller M, Hafer C, Vasquez G, Vavagiakis P, et al. Environmental contamination as a risk factor for intra-household Staphylococcus aureus transmission. PLoS One. 2012;7:e49900.
Cowling BJ, Ip DKM, Fang VJ, Suntarattiwong P, Olsen SJ, Levy J, et al. Aerosol transmission is an important mode of influenza A virus spread. Nat Commun. 2013;4. https://doi.org/10.1038/ncomms2922.
Lee SS, Wong NS. Probable transmission chains of Middle East respiratory syndrome coronavirus and the multiple generations of secondary infection in South Korea. Int J Infect Dis. 2015;38:65–7.
Dick EC, Jennings LC, Mink KA, Wartgow CD, Inborn SL. Aerosol transmission of rhinovirus colds. J Infect Dis. 1987;156:442–8.
Acknowledgements
BS and JAG were supported by the Alfred P. Sloan Foundation’s program on the Microbiology of the Built Environment (MoBE); BS was supported in part by an ASHRAE New Investigator Award.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of Interest
The authors declare that they have no conflicts of interest.
Human and Animal Rights and Informed Consent
This article does not contain any studies with human or animal subjects performed by any of the authors.
Additional information
Publisher’s Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
This article is part of the Topical Collection on Biology and Pollution
The original version of this article was revised due to a retrospective Open Access order.
Rights and permissions
Open Access This article is distributed under the terms of the Creative Commons Attribution 4.0 International License (http://creativecommons.org/licenses/by/4.0/), which permits unrestricted use, distribution, and reproduction in any medium, provided you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made.
About this article
Cite this article
Stephens, B., Azimi, P., Thoemmes, M.S. et al. Microbial Exchange via Fomites and Implications for Human Health. Curr Pollution Rep 5, 198–213 (2019). https://doi.org/10.1007/s40726-019-00123-6
Published:
Issue Date:
DOI: https://doi.org/10.1007/s40726-019-00123-6