Abstract
Purpose of Review
Since the late 1990s, researchers have been increasingly utilising digital methodologies to assess the branch structure of trees. The emergence of commercial terrestrial laser scanners during this period catalysed an entirely new domain focused on point cloud-based research. Over the years, this field has transformed from a complex computational discipline into a practical tool that effectively supports research endeavours. Through the combined use of non-destructive remote sensing techniques and advanced analytical methods, branch characterisation can now be carried out at an unprecedented level.
Recent Findings
While terrestrial laser scanning has traditionally been the dominant methodology for this research domain, the increased use of mobile laser scanners and unmanned aerial vehicles indicates a transition towards more mobile platforms. Quantitative structural modelling (QSM) has been pivotal in advancing this field, enhancing branch characterisation capabilities across diverse fields. The past five years have seen increased uptake of 2D and 3D deep learning techniques as alternatives.
Summary
This article presents a comprehensive synthesis of approximately 25 years of research in the field of digital branch characterisation, reviewing the data capture technologies and analytical methods, along with the forest types and tree species to which these technologies have been applied. It explores the current trends in this dynamic field of research, research gaps and some of the key challenges that remain within this field. In this review, we placed particular emphasis on the potential resolution of the significant challenge associated with occlusion through the utilisation of mobile technologies, such as mobile laser scanners and unmanned aerial vehicles. We highlight the need for a more cohesive method for assessing point cloud quality and derived structural model accuracy, and benchmarking data sets that can be used to test new and existing algorithms.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
The characterisation of branch structure is a fundamental practice for a variety of purposes. Understanding the way that trees grow empowers urban planners to plant the right trees in the right place for the right purpose. It also enables arborists and utilities managers to implement appropriate pruning regimes for managed trees. A comprehensive understanding of branch structure facilitates the development of high-fidelity models for modelling the carbon storage capacity of forests, supporting carbon sequestration initiatives. From an academic perspective, the ability to characterise branch structure enables a greater understanding of tree function and physiology, such as tree mechanics [1], tree respiration [2] or seed dispersal techniques [3]. Branch characteristics such as branch angle and branch frequency have been demonstrated to influence the overlap and aggregation of foliage, which are important in determining light-harvesting capacity [4, 5]. For foresters and tree breeders, branch characteristics are highly significant as branching is considered to be one of the most important phenotypic traits.
The value of timber is influenced by various factors, including the size, angle, frequency, and dispersion of branches, and internodal length [6], because branching is directly linked to knottiness in timber products [7]. Consequently, the phenotypic traits associated with branching are closely tied to the value of a specific genotype [8]. Branch form is influenced by multiple factors [6, 9] including climate [10], terrain (exposure, elevation; [11]), nutrition [12], site and genetics [13], silvicultural practices such as pruning and thinning [14], and site and environment parameters [15].
Traditionally, branch characteristics are measured using manual tools such as callipers and protractors [16, 17]. Manual branch measurement provides a means of deriving highly accurate measurements that can be used for developing or testing allometric equations [18] and have been conducted intensively on some of the largest trees in the world [19]. This process is labour intensive and often involves working at height, reducing throughput, and increasing cost and safety concerns [20]. From an ecological perspective, the acquired knowledge is often worth the additional cost and time. Conversely, these methods are too costly for production forestry where large numbers of trees are regularly required to be assessed for inventory purposes. Consequently, visual assessments are often preferred for phenotyping and forest inventory due to their cost-effectiveness and rapid execution. However, visual assessments are inherently subjective and assessor variability can influence precision [21]. In addition, the assessment of tree structure and form, from a physiological perspective, are impractical with these methods. Testing the validity of scaling laws, such as the metabolic scaling theory [22, 23] or pipe model theory [24] are crucial for understanding biomass, carbon and water dynamics; however, scaling these measurements to large trees with traditional methods are unfeasible due to time, skill and cost requirements [25].
Laser scanning technologies offer a rapid and non-destructive means for tree structural characterisation, encompassing individual tree level as well as forest stand level captures [26]. Laser scanning is an active sensing technology utilising laser ranging to measure distances between the sensor and target objects and creating a three dimensional (3D) model or point cloud which can be scrutinised to study targets in great detail. Depending on the size and power of the laser, laser scanning offers capabilities for characterisation of individual branches with handheld scanners, to the global scale characterisation of forest resources using satellite-borne scanners [27]. Airborne laser scanning (ALS) enables tree height and productivity metric extraction at the landscape scale [28], surpassing the spatial limitations of traditional plot-based methods. Laser scanning technologies also hold potential for phenotyping breeding trials. While ALS is well-suited to characterising canopy structure [29] and height measurement [30], inherent low resolution and occlusion issues render this technology unsuitable for the characterisation of branching structure. Consequently, laser scanning from below the forest canopy provides a more suitable approach for fine-detailed tree characterisation [31•]. Proximal sensors, such as terrestrial laser scanners (TLS), mobile laser scanners (MLS) or unmanned aerial vehicle (UAV) laser scanners (ULS), enable individual tree characterisation at the branch scale. The majority of studies on branch structure have primarily focused on larger branches, with limited attention given to the finer-level branch architecture [32, 33]. This research gap represents a significant limitation in the field of phenotyping, particularly considering that the breeding process requires information at a young age when branch sizes are smaller.
The measurement of branch structure across different age classes is critical in production forestry and forest research for different purposes. Branch characterisation across age classes can help to develop and refine accurate allometric equations for calculating tree age or biomass [18]. In production forestry, branch characterisation of mature trees facilitates the inventory of timber grades prior to harvest, whereas evaluating branch-related phenotypes in young trees accelerates tree improvement programmes, streamlining the breeding process. To expedite the breeding process and supply optimal genetic material to growers, phenotypic assessments are typically conducted at a young age [34]. This early assessment is particularly important in forestry where crop rotation cycles are notably longer than, for example, cereal crops; and enables characterisation of important traits prior to the trees being affected by competition, masking the impact of genetics on branch form [35]. Studies on Pinus radiata have shown strong phenotype-genotype correlations as early as age one to two [36].
The primary objective of this review is to explore the existing technologies and analytical methods employed in characterising branch structure from point cloud-producing technologies. Through a comprehensive review of approximately 25 years of research, this review aims to assess the current state of branch characterisation, identify research gaps, and provide recommendations for future studies.
Methods
The research reviewed in this study was collated through a literature search focused on branch characterisation using point cloud-generating technologies. To achieve this, an initial series of nested queries was performed searching the abstracts, titles, and key words from articles within the Scopus database. The Scopus database was selected as it is one of the premier databases for remote sensing, covering a larger number of journals than other leading databases [37]. The first part of the initial query string included terms related to branch and tree architecture and modelling, while the second part was focused on terms associated with point clouds and their production, such as laser scanning, photogrammetry and lidar. The Scopus database was searched for all available publications up to 1st August 2023 using the following key words:
-
(TITLE-ABS-KEY (branch* AND ("detect*" OR "characteri*" OR "model*" OR "Skeleton*" OR "structur*" OR "pattern*" OR "angle*" OR "point*" OR "length*" OR "scar*") OR "Tree" AND (model* OR "Individual tree model*" OR "architecture") OR "QSM" OR "Quantitative Structure Model" OR "Tree components" OR "Tree attributes" OR "Tree structure")) AND (TITLE-ABS-KEY ("TLS" OR "Terrestrial laser scanning" OR "Terrestrial lidar" OR "T-lidar" OR "ULS" OR "UAV laser scanning" OR "UAV laser scanner" OR "UAV lidar" OR "Drone laser scanning" OR "Drone lidar" OR "MLS" OR "Mobile laser scanner" OR "Mobile laser scanning" OR "Mobile lidar" OR "Laser scan" OR "laser scanning" OR "Lidar" OR "Lidar scan" OR "Lidar scanning" OR "Light detection and ranging" OR "Structure from motion" OR "Sfm" OR "Photogrammetry" OR "Point cloud")).
To ensure the research was limited to relevant sources, only peer-reviewed articles and review papers were considered as they are recognised as the highest standard for trusted scientific communication [38]. Furthermore, the search was restricted to papers written in English and within the fields of computer science, agriculture and biological sciences, earth and planetary sciences and environmental science. It is, therefore, acknowledged that the scope of this review is limited to the database used for the search, along with the key words used in the string, the language and field parameters used to limit the search. The identified literature was then reviewed, and a process of forward and reverse chaining was implemented in Google Scholar to identify any additional literature that was crucial to the field of branch characterisation using point cloud technologies.
The Scopus database search produced 771 results. An initial cursory review of the results revealed that there were a number studies that shared key words with the search, which were either in the relevant field but focused on other aspects of tree characterisation, such as stem form or crown morphology, or were from different disciplines, such as coral reef ecology, astronomy, or computer architecture, which share common terminology or tools. Removing these brought the total number of applicable studies to 142. Forward and reverse chaining found an additional 70 pertinent studies, bringing the total to 212. More focused review of these studies found an additional 46 of these to be irrelevant, resulting in 166 publications being included in this review.
Literature Review
Traditional Methods
Forests are grown for a number of reasons including for recreation, timber production, biodiversity conservation and ecosystem services. In the context of timber production, the value placed on trees is often associated with their large, straight stems that are free from deformities [36]. From a tree improvement perspective, breeding programmes aim to produce high-quality trees for timber by combining desirable stem form characteristics such as straight stems and small branches with rapid stem volume increment [39, 40]. To support these programs, it is essential to measure a substantial number of trees accurately, using methods that are both simple and cost-effective [35]. Traditional manual measurement of stem form parameters such as diameter at breast height (DBH) and height are relatively fast using a diameter tape or callipers for stem and branch diameters, and a height pole or hypsometer for height. Branch structure, however, is generally too labour-intensive and is often limited to small sample sizes, if carried out at all [41,42,43,44].
A variety of branch characteristics, including diameter, angle, length, number and their dispersion across the stem are of interest in the study, growth and breeding of trees. Branch angle is assessed using methods such as direct measurement with a protractor [45], a digital goniometer [46], or by climbing and placing a clinometer at the base of the branch [47]. Kint et al. [48] used a Field-Map scope (IFER Ltd., Strašice, Czech Republic) system to calculate branch angle automatically. This involved positioning a reflector on an extendable pole at the branch insertion point and at a point 5 cm along the branch and measuring their position. This enabled elevated branch measurement from the ground; however, this method was noted to produce relatively large measurement errors [48]. Alternatively, trees can be felled and branch parameters measured on the ground [46]; however, destructive sampling is not practical for ongoing genetic trials or most types of inventory. Many traditional measurement methods of branch angle are time-consuming and are better suited for measuring small subsamples or individual trees. To expedite measurement, branch angles are commonly estimated visually. These scale-based assessments, such as the six-point scale introduced by Raymond, Cotterill [49], enable genotype comparisons for breeding programmes; however, they are inherently subjective.
More comprehensive methods for tree form measurement include crown mapping (Fig. 1a). This objective methodology was developed for the thorough measurement of large gymnosperms [19]. Data are systematically recorded in spreadsheets and subsequently analysed and modelled using the Sitchensis package (Fig. 1b; Kramer et al. [19]). The 3D visualisation of the data within Sitchensis enhances data integrity through rapid error checking in the field.
a Research crew measuring branch structure of a mature P. radiata, demonstrating the technical climbing and mensuration skill requirements; b Example of a model generated in the Sitchensis software using the sample data provided with the package. Stem and branches are colour-coded to represent different types of appendages: main stem (black), limbs (red), live branches (green) and dead branches (grey)
Technologies Available
A number of additional technologies have been trialled in the literature for assessing the structure of trees and plants including machine vision techniques like time of flight [50], structured light [51] or shape from silhouette [52]. X-ray computerised tomography (CT) has also been used to derive the structure of live plants [53] and to assess the branch structure of trees after harvest [54]. A common issue with these technologies is that they are often complex, require controlled laboratory environments, or large and expensive equipment that make them impractical for tree measurement which often takes place in remote locations like forests [50, 55]. Proximal and remote sensing techniques offer alternative methods that can increase the measurement efficiency whilst retaining objectivity and accuracy. A study found time savings of 50–76% per tree by using TLS for tree structure measurements in contrast to destructive sampling methods [56].
In the literature, three main techniques have been discussed for capturing tree-branching structures and representing them as point clouds [57]: three-dimensional electromagnetic digitising (3D EMD; Sinoquet et al. [58], Sellier, Fourcaud [59]), terrestrial photogrammetry (T-SfM; Morgenroth, Gomez [60], Dong et al. [61], Miller et al. [62]) and laser scanning, which includes airborne, UAV, terrestrial, and mobile methods [63,64,65•]. A summary of the advantages and disadvantages of the different point cloud-generating technologies can be found in Table 1.
Three-dimensional Electromagnetic Digitising
3D EMD involves measuring the architecture of a tree using a 3D electromagnetic digitising device (Polhemus, USA, 3space Fastrak; Fig. 2a) to record the spatial coordinates, diameter, and order of branch structure [59]. This method generates highly accurate and comprehensive models of tree architecture; however, involves labour-intensive data collection [26]. Additionally, it is limited by the 3 m range of the magnetic field [209], making it logistically challenging for larger trees (Fig. 2a).
a Application of the 3D digitising method to macadamia trees in an orchard (reprinted from White, Hanan [210] with permission from the Queensland Government Department of Agriculture and Fisheries); b A T-SfM model of a radiata seedling, showing the multiple image points (green dots) required to capture data using this methodology
Terrestrial Photogrammetry
Terrestrial photogrammetry (T-SfM) is arguably the most cost-effective method as cameras are more economical than laser scanners or 3D digitisers [62]. However, T-SfM data capture routines are more complex than laser scanning, requiring imagery from numerous viewpoints to reconstruct a single tree (Fig. 2b; Bournez et al. [57]) and are sensitive to light conditions [211]. Due to the passive nature of cameras, the accuracy of the T-SfM model reconstruction is highly dependent on the quality of the input imagery [212]. Achieving optimal camera settings, including low light sensitivity (ISO) and fast shutter speed, is essential to acquire the high quality imagery required for creating quality point clouds [213]. These parameters can be challenging to achieve simultaneously in a forest setting due to limited sunlight [213]. In addition, feature matching algorithms used in photogrammetric workflows may not be efficient for complex plant structures with self-occlusion, leading to reconstruction errors [66]. Thus, reproducing the branch structure of densely foliated crowns using T-SfM is unlikely due to canopy penetration limitations and associated reconstruction challenges. Another form of photogrammetry commonly used in forestry for characterising tree structure is UAV photogrammetry (UAV-SfM). Due to the aerial perspective and the passive nature of the sensors, UAV-SfM is generally unsuitable for characterising branch structure in forest environments as demonstrated in Fig. 3. A recent study observed that fine-level branch detail in an orchard environment in leaf-off conditions was difficult to detect with UAV-SfM, proving that the technology is not suitable for branch detection [214].
Figure showing point clouds from a single P. radiata tree captured with five different technologies. From left: ALS (20 points per m2 (ppm2); synthesised from ULS), UAV-SfM (637 ppm2), ULS (19,265 ppm2), TLS (203,424 ppm2) and personal laser scanning PLS (196,047 ppm2). Points are coloured by height and ALS and UAV-SfM points are displayed in larger size to increase visibility of their sparse points
Laser Scanning
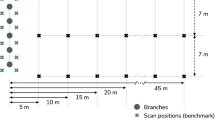
Laser scanning offers a more rapid and efficient capture process along with a high level of reconstruction accuracy [26, 57, 78]. In contrast to photogrammetry, laser scanning is an active sensing technology and is therefore not dependant on lighting conditions. It also enables the efficient collection of hundreds or thousands of data points across the structural surface of a tree, without the need for physical contact that is required for 3D EMD. Laser scanning can be conducted using a variety of airborne and terrestrial platforms (Fig. 4). For the purposes of this review, airborne platforms are being divided into two categories: airborne laser scanning (ALS) from a manned aircraft and UAV laser scanning (ULS) from unmanned aircraft. Terrestrial platforms will be divided into terrestrial laser scanning (TLS) and mobile laser scanning (MLS). Being an active sensing technology, laser scanning in all its forms is less prone to environmental factors such as lighting conditions or sun angle that affect passive remote sensing techiques. However, laser scanning is not immune to the adverse effects of wind, which can introduce “ghost artefacts” to point clouds, where a tree or part of a tree is recorded in different locations at different times [213]. The impact of wind is greatest at the tree top and lowest towards the base of the stem, therefore, a greater or lower wind speed can be tolerated depending on the data capture objective. It is generally recommended to capture data in windless conditions [213], especially when characterising smaller structures such as branches, which restricts possible capture windows. Rain, fog and high humidity can also introduce noise into laser scanned point clouds by refracting, scattering, reflecting and absorbing the infrared light wavelengths that are used in laser scanners, thus affecting the laser beam [215,216,217,218].
Airborne Laser Scanning
ALS enables rapid capture of forest structure and vegetation information across vast areas, spanning hundreds or even thousands of square kilometres [219]. ALS has been successfully used to derive tree heights [30] and estimate DBH indirectly [220] with a high level of precision and accuracy. Branch characterisation, however, may face challenges when occlusion of the stem and branches by other tree structures leads to a reduction in point cloud density, resulting in gaps in the point cloud [199]. Guidelines for the forestry sector state low pulse density for ALS to be 0.5–1 pulse per m2, and high as being > 3 pulses per m2 [221]. Reviewing the literature, [222] found ALS used in studies ranged between 1 point per m2 (ppm2۔) and > 25 ppm2, with 4–12 ppm2 being the most common. Given these low point cloud densities, ALS is typically not well-suited for branch characterisation (Fig. 3).
ALS data has been utilised to derive branch structure of mature coniferous trees for the purpose of species prediction [67•]. This involved computing internal crown parameters like branch length, width, slope, and density by fitting an internal spline from the base to the top of the tree, modelling the branch peaks and applying region growing and Euclidean clustering algorithms to model the branch skeleton. It is important to note that this promising method for branch characterisation was exclusively tested on open-grown trees with full canopies. Its suitability for forest-grown gymnosperms, which often exhibit substantial inter-tree occlusion and have dead or broken lower branches, has not been examined. Additionally, the study did not offer ground truth data to assess the accuracy of the reproduced branch structures.
Tree canopies have naturally evolved to effectively intercept light from above, making branch characterisation using laser scanning more effective when performed from beneath the canopy. One notable example of laser scanning applied to phenotyping is the study by du Toit et al. (2021), which focused on using ALS to derive the internal crown structure of Pseudotsuga menziesii [68••]. Their approach was similar to that of Harikumar et al. [67•], using a clustering algorithm to segment branches and subsequently growing clusters based on the similarity of the best-fit planes. Once clusters were identified, various metrics, including branch angle, width, length, and volume density, were calculated. This study also lacked field-measurements for validation, with branch length validated against manual measurements taken from the point clouds and branch angle validated against an independent dataset providing average measurements for the species in a given growing condition.
One final study used high-resolution TLS data to calibrate ALS data and scale assessments of forest canopy structure across large areas [29]. This method, however, would not be suitable for branch characterisation in a forest setting, particularly for phenotyping, which requires an accurate description of each individual tree. Overall, although ALS offers significant advantages for assessing forest structure, its suitability for detailed branch characterisations may vary between different forest conditions and the intended applications.
UAV Laser Scanning
UAV laser scanning (ULS; Fig. 4a) enables branch-level tree characterisation at both plot and stand levels [26] and serves as an effective tool for capturing vegetation metrics at an intermediate scale, bridging the gap between ALS and TLS [31•, 223]. The lower altitude and slower speeds typically result in high point densities for ULS data, which are orders of magnitude greater than ALS data, enabling characterisation of tree and branch structure that is more typical of TLS [224]. A previous study comparing five different ULS sensors found a range of point densities from 17 to 1942 ppm2 [225]. Much higher point densities are attainable with ULS, as is demonstrated in Fig. 3 (19,265 ppm2). Despite these higher resolutions, ULS remains constrained by the occlusion and canopy penetration challenges, which are commonly associated with ALS.
Adapting the methodology developed for ALS in a previous paper, du Toit et al. [73], successfully determined the heritability of branching traits such as branch angle (h2 = 0.277). Their study also found that branch length and angle could be derived with R2 values of 0.81 and 0.37 respectively, when validated against manual measurements from the point cloud [73]. Nevertheless, despite demonstrating that branch metrics could be derived from ULS data, this study primarily focused on upper canopy branching and did not address the measurement of suppressed branching on the lower stem, which is a key indicator of timber quality.
ULS point clouds have been used to accurately estimate volumes of stems and large branches using quantitative structural models (QSMs) of multiple tree species [31•]. Their study found that when assessing the volume of all cylinders from the QSMs exceeding 30 cm in diameter, there was a maximum concordance correlation coefficient (CCC) of 0.85 and an RMSE of 1.12 m3 between ULS and TLS volume measurements. This, however, significantly reduced to a CCC of 0.51 and an RMSE of 6.59 m3 when all branches were included, indicating that fine branch structure could not be accurately reconstructed [31•]. Their results suggest that scanning from below the forest canopy was more favourable for accurate modelling of the trunk and branches.
While ULS studies represent an improvement upon ALS for branch characterisation, some key limitations remain. First, regulatory restrictions currently limit the application of ULS technology in forest environments, as UAVs must typically operate within visual line of sight [226]. Second, flying above the canopy results in occlusion limiting the potential to describe branches lower in tree crowns.
Terrestrial Laser Scanning
Research into the characterisation of branching metrics from point clouds has predominantly focused on TLS (Fig. 4b). The first commercial TLS was released in 1998 [227]. The technology was trialled for forestry applications in the early 2000s [228,229,230,231], primarily focusing on assessing metrics used for volume estimation, such as height and diameter. The first studies applying TLS to branch characterisation were published shortly afterward [79, 232]. TLS has been used to accurately measure branch diameters, for example a study of Magnolia denudata Desr. attained a high level of precision (R2 of 0.84 and an RMSE of 8.4 mm) for branches with a minimum diameter of approximately 1 cm [80]. It was also used in a mature stand of P. sylvestris L. to detect whorls and estimate branch diameters and insertion angles with relatively low errors, yielding an RMSE of 1.31 cm (40.4%) for branch diameter and 15.4° (23.4%) for insertion angle, respectively [81•]. While these results demonstrate that TLS can effectively resolve smaller branching structures, it’s worth noting that some studies have highlighted that methods like QSM tend to overestimate the girth of branches with diameters less than 4 cm [82].
The literature shows that TLS offers a rapid and less labour-intensive approach to collecting and analysing data for branch characterisation, enabling detailed, tree-level forest structure characterisation at larger spatial scales than field measurements [26, 83]. It also provides more objective measurements compared to traditional ground-based observations by foresters [26]. TLS is considered an optimal technology for branch characterisation due to its capability for below-canopy, proximal sensing with a small beam divergence and high resolution, facilitating the measurement of small branches [84]. Indeed, the mm-level accuracy and ability to resolve fine scale branching has been hailed a revolution in the study of tree architecture, enabling researchers to explore the ‘missing link’ between large diameter branches and leaves [3, 55]. Point densities are characteristically an order of magnitude greater than ULS data (Fig. 3); however, the lack of reporting on point density in TLS studies makes it difficult to determine a typical point density. A recent study tested a range of point densities from 4800 to 79,700 ppm2 for estimating forest structural parameters [233]. Another study found an average point density of 1500 ppm2 for a data set with areas as high as 300,000 ppm2 [234].
While TLS shows great promise, occlusion remains a significant limitation [26, 85]. The inner branch structure of densely branched or foliated trees, especially in dense coniferous species, is often not visible using TLS [199]. Various methods have been proposed to address occlusion issues [235, 236], however, the static nature of TLS requires precise co-registration of point clouds collected from multiple locations to attain adequate coverage [31•]. Increasing the number of scan positions around a tree can help reduce branch occlusion [26]; however, the “observation angles” of TLS that enable penetration of gaps in the canopy can be limited in comparison to mobile systems like mobile laser scanning (MLS). Depending on the TLS system utilised, the increase in scan positions can also lead to reduced efficiency, which can make the operational use of TLS impractical for characterising a large number of trees, such as in a genetic trial. For example, in a recent unpublished study, the authors captured TLS and personal laser scanner (PLS) data of a 3 ha breeding trial. The PLS capture was conducted within a single day, whereas the TLS took eight days to complete (an example of the data can be found in Fig. 3). Recent advances in high-end TLS units that offer real-time, on-board registration of individual scans, negating the need for the placement of control points in the field, represent a significant advancement in the practicality of TLS deployment. For a recent, comprehensive review of TLS technology, please refer to Calders et al. [32].
Mobile Laser Scanning
MLS has emerged as a practical and scalable alternative to TLS (Fig. 4c; [8]). MLS involves using laser scanners on mobile platforms such as cars or all-terrain vehicles. As with TLS, the range of point density that can be expected is much larger than its airborne counterparts. Also similar to TLS, the studies seldom report on the point density. Of the studies compiled within this literature review, the point densities ranged from 10,000 to ~ 30,000 ppm2 [8, 74, 203]. The PLS data set used in Fig. 3 had a point density of 196,047 ppm2. Recent innovations in lightweight laser scanning have led to the adaptation of scanners for handheld and backpack-mounted platforms, enabling the operator to navigate through environments that are inaccessible to vehicles [237]. This increased mobility offers a greater flexibility in data capture within complex environments, significantly reducing object occlusion and maximising data coverage [213]. These backpack or handheld scanning systems are often referred to as “personal laser scanning” (PLS) in recent literature [213]. PLS generally uses simultaneous localisation and mapping (SLAM) algorithms to match scan scenes concurrently during the capture, ensuring accurate data collection while in motion. SLAM systems do not rely on global-navigation-satellite system (GNSS) locations, making them particularly well-suited for forest surveys, where GNSS accuracy is inherently poor under the canopy [8]. For the purposes of this study, we distinguish PLS from MLS as being smaller, SLAM-based systems that can be carried on a backpack, or handheld, for example the Li-Backpack (Green Valley, Beijing, China). MLS is defined here as being a larger, more accurate laser scanning system that is mounted to a vehicle, for example the SSW-IV mobile mapping system [204, 238]. A detailed appraisal of MLS technologies can be found in [213]. An additional advantage of the SLAM method is that it automatically co-registers the scans, an aspect which generally requires a significant amount of time when utilising TLS for deriving structural attributes. Conversely, PLS point clouds are inherently noisy, caused by a combination of the motion of the capture, high beam divergence, and co-registration issues within the SLAM process due to the complexity of the forest environment [239, 240].
To date, the limited number of studies employing MLS and PLS have primarily focused on deriving tree position, DBH and stem volume, with little emphasis on tree branch characterisation [205]. PLS has demonstrated significant efficiency gains, with a fourfold increase in capture efficiency for DBH and height compared to direct measurements [241], and six to sevenfold increases for DBH measurement alone compared to direct measurement and TLS measurement, respectively [242]. However, it’s important to note that in the latter study, understorey vegetation was cleared prior to scanning to facilitate the process, highlighting the need for testing PLS in more operational environments. Furthermore, PLS data, along with other laser scanning technologies and SfM, has the potential to yield additional metrics, such as stem curve and canopy metrics, without incurring additional fieldwork costs. For example, using QSM methods, PLS was employed to derive stem-to-branch volume ratios for sparsely-stocked plots of Pinus contorta with an RMSE of 21% [75]. In contrast, stem-to-branch volume ratios derived from ULS data overestimated volume by more than 300%, underscoring the argument that branch structure is more accurately characterised from beneath the canopy [75]. Additionally, PLS has been used to assess whorl location by modelling diametric profiles of the stems and employing an algorithm to detect stem swellings, which were assumed to be whorls [8]. A strong correlation was observed between field-measured and detected whorl heights; however, whorl detection was hindered by high numbers of false positives, mainly attributed to the prevalent noise in PLS point clouds [8]. To the authors’ knowledge, this is the only phenotyping study to date using MLS and one of the few laser scanning studies with a specific focus on phenotyping.
A recent study tested the efficacy of measuring branch structure of Picea abies L. using PLS, comparing two different methods (QSM and DBSCAN clustering) for extracting branching parameters [203••]. Derived branching metrics, such as branch location, diameter and angle were used to predict timber-related quality metrics, which were assessed against knottiness metrics derived from x-ray scans of the logs recovered from these stems after harvest. This study found QSM-derived branch features were superior for training machine learning models, which predicted features such as whorl distance, whorl count, knot index and knot volume with R2 values ranging 0.47–0.54 [203••]. Among all the reviewed methods, PLS stands out as combining optimal efficiency, scalability and level of detail, with minimal occlusion issues. Conversely, the lower quality of the data produced by PLS systems impacts the quality of the data analysis, inflating branch sizes and reducing the efficacy of TLS-based procedures, such as QSM, to delineate and measure individual branches [86]. The practicality of these systems is currently offset by these issues; however, recent additions of more powerful laser scanners to PLS system will hopefully improve upon this. Therefore, it remains crucial to continue to test these systems in structurally complex forests to further their development [32].
Summary of Available Technologies
TLS is the most extensively studied technology, with approximately four times the number of studies dedicated to this technology than to all of the other technologies combined (Table 1, Fig. 5a). PLS is the second most studied method and has arguably the largest and fastest-growing literature currently. When combining MLS with PLS, the overall field of MLS exhibits two distinct publication periods (Fig. 5b). The first MLS study was published in 2010, with an additional four studies published by 2014. The second period of publication began in 2020, with 12 studies published by 2023. The earlier studies primarily focused on vehicle-mounted MLS, whereas, with the exception of Li et al. [77], all studies since 2020 have employed PLS. This recent increase in publications coincides with the emergence of commercial PLS systems that combine automotive laser scanners with SLAM algorithms, such as the Hovermap (Emesent, Milton, QLD, Australia) and the Zeb Horizon (GeoSLAM Ltd., Nottingham, UK). Similarly, the use of ULS point clouds for branch characterisation is a relatively recent development, with the first papers published in 2017 and 2019, and an additional four studies published between 2022 and 2023 (Fig. 5b). This recent trend for ULS and PLS research reflects both a desire to explore methods that enhance data capture efficiencies and the increasing availability of these new technologies.
Branch Characterisation Methods
Two major approaches have emerged in the field of branching. The first approach, known as skeletonisation, involves fitting a central spline to the point cloud data representing the tree stem and major branches [26]. This is the older of the two approaches, with initial methods introduced shortly after the inception of forest TLS studies [79, 124]. Skeletonisation generates a more streamlined representation of the complex tree architecture, making it computationally less intensive compared to working with the original point cloud data [108]. This method enables the assessment of tree structure, and branch number, order, length and angle [93].
The second approach involves reconstructing the surface geometry of the stem and its appendages using an approach known as quantitative structure modelling (QSM). QSM was first introduced by Raumonen et al. [174••], with later methods introduced by Hackenberg et al. [65•], Du et al. [117] and Fan et al. [121]. The main difference between the two approaches is in the outputs, where skeletonisation aims to simplify the representation of tree structure to a skeleton based on the central axes of the trunk and branches, whereas QSM aims to facilitate the extraction of additional metrics like stem and branch diameter [26], volume [82] and biomass [146]. QSM relies on highly accurate TLS scans to generate reliable representations of tree structures. Consequently, studies often utilise individual trees within idealised conditions to maximise the quality of the input data [111]. It’s important to note that QSM cannot reconstruct missing branch structures from heavily occluded scans [243]. Generally, QSM methodologies are designed for individual tree-level analysis, necessitating an initial process of individual tree segmentation [206, 243]. Similar to QSM, other methodologies have opted for a simplified process of fitting geometric primitives, such as cones or cylinders, to the point cloud without the prior step of skeletonisation [81, 169, 172].
Several studies have combined skeletonisation with complementary modelling approaches to address occlusion issues. Gymnosperms are especially prone to occlusion and Côté et al. [109] combined skeletonisation with allometric relationships to account for data gaps. Other studies have used topological rules or crown shape to determine a best path for extending branch skeletons through noise or data voids within point clouds [76, 197, 202]. These hybrid skeletonisation approaches enhance the creation of more complete tree models that are more robust against occlusion and noise in the input data [101].
There are additional branch characterisation methods worth noting, particularly those focused on assessing timber quality from the bark surface of TLS scans. Kretschmer et al. [152•] proposed the “bark surface modelling” (BSM) approach, which involves flattening the 3D stem point cloud and then rasterising the height of the bark surface, in an approach somewhat similar to canopy height modelling. Another method relies on machine learning to train a defect classification algorithm, enabling the detection and classification of abnormalities on the stem point cloud, such as branches, branch scars, burls and epicormic shoots [166]. These methods require an accurate laser scanner with a clear field of view of the tree stem to provide a smooth and detailed representation of the bark surface within the point cloud.
The authors are not aware of any study that has attempted to create such a scan from young gymnosperms with heavy stem occlusion and apply either BSM, skeletonisation or QSM techniques. In addition, there is a noticeable gap in the research concerning the application of BSM techniques to assess the surface quality of the stems scanned with MLS. For a comprehensive list of studies focused on branch characterisation and the methods they employed, please refer to Table 2.
QSM methodologies are widely used in the field, with approximately half of the studies relying on them (Fig. 6). Among these studies, a significant majority (60) utilised only two software tools, namely SimpleTree [65•] and TreeQSM [174••]. By comparison, of the 40 studies employing skeletonisation, 29 are methodological, proposing new skeletonisation methods. This suggests that QSM is more commonly applied in research, while skeletonisation sees more methodological developments.
An emerging trend in the tree structural modelling is the application of 3D deep learning (DL). Since 2015, when the first DL models were trained on 3D point clouds [32], there have been several publications using DL to characterise stem form [251], segment forest structures [252] and assess branch structure, which has been conducted using 2D DL techniques, in which the individual tree point clouds are first converted to 2D image projections [171••], and 3D DL techniques [87, 191, 244, 245]. DL offers advantages over traditional heuristics because these models can adapt and improve over time through transfer-learning methods [171••]. There is potential for DL algorithms to be trained to recognise branch patterns within noisy point clouds, such as MLS point clouds. However, DL methods require substantial amounts of labelled training data to achieve accurate results, and so far, they have not been widely applied in diverse and complex environments [206].
Heuristic algorithms, such as Dijkstra’s shortest path algorithm [253], depend on a well-defined trunk to establish a root-position for path calculation [206]. This can be challenging when noise and stem returns cannot be distinguished. To address this, using biologically accurate training datasets generated from synthetic point clouds, augmented with additional noise, could aid in training DL models to differentiate noise from the tree structure.
Applied Branch Characterisation
When reviewing the location of the studies, it was interesting to see that there are several hotspots for branch characterisation research with China, Germany and Canada producing the most studies, according to the lead author’s location (Fig. 7). Other notable clusters of research include the contributions from Finland, France, the Netherlands and the United States (USA). The pattern for the countries in which the study sites were located was similar, with hotspots once again located in China, Germany and Canada. There are some notable additions from countries such as Australia, the UK, Cameroon, Guyana, Indonesia and Peru (Fig. 7). This could be due to a combination of various factors including the research interest in some of these locations being of international importance (for example tropical forest research for carbon studies), or a potential lack of funding or lower resource availability to some of these nations to conduct the research solely within their national research agencies. Additionally, several of the datasets from these studies were made available publicly, such as [43, 114, 163], expanding the number of studies that can be conducted utilising the same data.
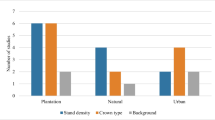
The multitude of methods developed for characterising branch structures have not been uniformly applied to a standardised set of tree species and environment conditions. The lack of benchmark makes it challenging to evaluate the suitability of different algorithms for deriving branch-level data across various species. These branch characterisation methods have been tested on a range of forests, including tropical, broadleaf and coniferous forests, urban and simulated trees. Table 3 provides a comprehensive list of studies and the types of forests they focus on, while Fig. 8 illustrates the cumulative number of studies per forest type.
Out of 143 studies listed in Table 3, that reported tree heights, only 24 were conducted on trees with an average height of less than 10 m. Among these 24 studies, six focused on deciduous trees [97, 123, 134, 153, 158, 178], seven on urban trees [57, 99, 167, 168, 182, 188, 200], and six on fruit trees [63, 103, 108, 150, 170, 196], and only one partially focused on coniferous trees [242]. These results may have limited applicability to researchers seeking to employ branch characterisation methods on young gymnosperms, as the structural characteristics of these trees differ significantly from those studied in these particular works.
Previous studies have focused on a wide range of tree sizes, ranging from young deciduous saplings [134] to emergent rainforest trees [161]. Compounding this issue is the variety of different tree species or vegetation types that have been studied. This variability of tree size, age class and species across different branch characterisation studies, along with the diverse range of metrics derived from point cloud data, indeed presents a challenge for comparing and transferring these techniques to various scenarios. Further complicating the issues of transferability and comparability, various studies have extracted a wide array of metrics from the point cloud data. The studies have investigated diverse methods for assessing branch parameters, including diameter [67•, 81•, 82, 122, 134, 139, 172], angle [81•, 122, 172, 173, 189, 254], and length [33, 80, 132, 146, 158]. Other metrics that have been explored include whorl height [8, 81•, 133, 171••], branch count [33, 83, 125, 129, 134, 158], branch volume [146, 169] and crown dimensions [135, 139, 173].
Early studies that developed branching algorithms focused on well-branched angiosperms, including tropical trees [43], orchard species [108] and natural deciduous forests [118, 152•, 173]. Several studies have examined the branching structure of mature gymnosperms in a range of boreal [130] and coniferous forests [81•], as well as plantations [205]. However, there is a lack of studies focusing on juvenile gymnosperms aged ten or less. This is a significant gap as phenotyping juvenile trees can shorten the breeding cycle [34] and enable the assessment of branching traits before canopy closure and competition begin to impact tree form [35]. The dense foliage on juvenile trees also causes greater occlusion, which has less of an impact in mature plantations due to natural branch senescence, branch shedding and management practices such as pruning.
Coniferous trees are particularly prone to occlusion of their inner branch structure and stem by the canopy and outer branches [199]. Data voids resulting from occlusion are a widely reported challenge and a significant cause of failure in branch characterisation algorithms, leading to inaccuracies in tree models [29, 85, 100]. It is worth noting that the majority of recent studies in this field have predominantly focused on deciduous trees during leaf-off conditions [83]. This trend implies that the issues related to occlusion and the need for further research in this area, as highlighted by Bayer et al. [93], have not yet been adequately addressed.
Branch characterisation and tree structural assessment have found various applications, including means of non-destructive measurement of biomass in a range of forest environments [78, 82, 125, 130, 146, 147]. Other research applications include assessing the aerodynamics of forest plantations [139], the effect of ice accretion on tree canopies [168], the impact of neighbouring tree species on crown structure [173], the effect of slope on tree shape and volume [118] and the effect of stocking on tree structure [145]. Some of these applications have used TLS and QSM techniques and revealed significant variations in the branch structure of four different oak species, highlighting the potential of these technologies for phenotyping [185]. In another study, a methodology was introduced for characterising branch volume, angle, length and width using ALS point clouds [68••]. This method was then applied to ULS data, achieving a high correlation of 0.93 for branch length [73]. Furthermore, when comparing the phenotypic traits derived from ULS data with genetic information using narrow-sense heritability, the study found that branch angle exhibited the highest heritability value, measuring at h2 = 0.277. QSM techniques have been employed with TLS data to phenotype fruit trees, allowing the characterisation of branching structures down to the fourth order while maintaining a low estimation error for the number of branches [150]. Significantly, the ability to characterise higher branch order levels through the combination of TLS and QSM holds the potential to uncover previously undiscovered branch architectures [150]. The authors are unaware of any research specifically focused on branch-level tree phenotyping using MLS or PLS point clouds.
Discussion
Emerging Trends
The application of point cloud technologies, such as T-SfM, ALS, TLS, and PLS, has revolutionised forestry research by providing efficient and non-destructive data capture methods that offer insights into overall tree structure, including branching. These technologies have enabled measurements that were previously labour-intensive or unattainable through traditional approaches, greatly enhancing forest assessment capabilities. This review highlights emerging trends in the utilisation of point cloud technologies for branch characterisation across diverse forest environments.
Since their introduction in 1997 and 2004, 3D EMD and TLS were among the first technologies to explore branch characterisation, with TLS becoming the most extensively studied tool in this field. Recent research has shifted towards mobile technologies like PLS and ULS, which offer potential solutions to issues related to occlusion and data capture inefficiencies associated with TLS. However, due to fundamental differences in noise levels and point density between these technologies, algorithms developed for TLS may not be fully optimised for mobile technologies. Hence, refining algorithms tailored to these emerging technologies is essential for their continued development and effective utilisation.
QSM is the primary method employed in branch characterisation studies, using various QSM software tools such as TreeQSM, SimpleForest and its predecessor SimpleTree. There is increased accessibility of these methods to experts in diverse fields such as forest ecology, genetics, and fire research. Research efforts also continue to refine tree skeletonisation methods, indicating their ongoing significance.
Alternative approaches to branch characterisation include BSM, initially explored for branch scars, and later used predominantly for timber quality studies since 2013. BSM has evolved with the integration of machine learning techniques. Furthermore, over the last five years, there has been a significant increase in the uptake of both 2D and 3D DL techniques. This is particularly promising, as these models can be tailored to enhance their performance, indicating substantial potential for continued growth within this research domain.
Research Gaps
A moderately-sized body of research has developed around the use of laser scanning, particularly TLS, for branch characterisation, while MLS has seen limited use in this context. Various methods have been proposed for characterising branch structure; however, only a small subset of studies has focused on juvenile trees, with gymnosperms receiving minimal attention. This research gap is notable as phenotyping juvenile trees can accelerate the breeding process, and young gymnosperms face specific challenges related to occlusion due to their dense foliage. Notably, branch characterisation from point clouds for phenotyping has been largely restricted to mature trees, with only a handful of studies focusing on this aspect, revealing significant research gaps in this area for tree breeders and foresters.
The field of branch characterisation from point cloud data is rapidly evolving with various methodologies, algorithms, and software being published regularly. However, comparisons across different methods and species are challenging due to limited validation with a narrow range of species and age classes. Whilst a framework for comparing the structural models created from different methods has been introduced [98•], such standardised tests have not been widely adopted within this field. To address this, there is a growing need for an international open-access library that encompasses a diverse range of trees, including various developmental stages and environmental conditions. Additionally, despite the creation of realistic tree models, many studies lack comprehensive evaluation to assess their fidelity to actual tree structures. While a framework for quantifying structural similarity exists, it has yet to be widely adopted in the field, highlighting the importance of standardised testing in future research.
Branch characterisation studies using technologies like TLS, ALS, and PLS often lack ground truth measurements, making it difficult to determine the precision of these methods accurately. While some studies introduce virtual ground truth, the absence of field measurements hinders the assessment of measurement accuracy and potential oversights. To advance tree phenotyping and modelling, it's crucial to understand technology limitations, and thus future research should incorporate field measurements for benchmarking. Achieving cohesion and synergy in the field, standardising data quality, and leveraging open-access resources, which are starting to be created [114, 255, 256], will further enhance research quality and avoid duplicative efforts.
Remaining Challenges and Potential Solutions
The field of branch characterisation has witnessed significant growth, driven by accessible methodologies like QSM and emerging point cloud technologies such as ULS and PLS. However, a number of challenges remain unresolved. Laser scanners, while enabling detailed measurements, face issues like occlusion caused by leaves and branches in forest environments. Novel approaches, such as high-density data capture from above the canopy with ULS, and sub-canopy assessment using PLS, aim to address these challenges by minimising occlusion through increased mobility and enhancing data coverage through co-registration of multiple scan scenes. Sub-canopy UAVs, which rely on SLAM technology and operate independently of GNSS, hold great promise by combining the advantages of both PLS and ULS. While initial studies have explored these technologies in forestry applications [257,258,259], their application to branch characterisation remains unexplored.
Challenges in branch characterisation often arise from understory vegetation, which can occlude individual trees and hinder accurate segmentation. Therefore, standardising understory removal would greatly benefit branch characterisation workflows, as they typically require individual tree point clouds. Various methodologies, including heuristic [260] and DL [252] approaches, have been proposed to remove understorey from forest point clouds. The overlapping tree crowns pose challenges in segmenting individual trees, leading to potential errors such as multiple trees being represented as a single entity or incorrect branch measurements being incorporated into neighbouring tree models. These challenges highlight the need for robust solutions that can address understorey-related issues.
A possible solution to the segmentation issues between understory and target tree, as well as for intra-crown branch segmentation is the use of multispectral laser scanning. This approach combines lasers with multiple wavelengths, so that both spatial and spectral data can be captured simultaneously [213]. Previous research has noted promising results with the combination of dual-wavelength terrestrial laser scanners to aid in the segmentation of foliage from the tree skeleton through the use of indices such as NDVI [261, 262]. Hyperspectral TLS systems have also been developed in laboratories and portable versions have been trialled for field deployment [213, 263,264,265]. Multispectral systems are being developed commercially for ALS and have been deployed for forestry research [266, 267], however, at the time of writing there are no commercially available multispectral or hyperspectral TLS systems. There is also a notable lack of development for multispectral and hyperspectral systems for other platforms, such as MLS, PLS or ULS.
Most methods for individual tree modelling start with a seed point at the base of the tree and use processes like region growing to cluster points into stem segments. An approach originally developed for assessing branch structure from ALS point clouds used topology to improve individual tree reconstruction [67•]. This was achieved by reducing uncertainty in branch paths through the utilisation of seed (segmented stem) and end points (branch tips). Recent studies have adapted similar approaches to both TLS and PLS point clouds [206, 268]. Allometry has also been used to fill gaps in point clouds caused by occlusion [109]. Current research focuses on data capture methods that reduce voids in point clouds by increasing vantage points (e.g., in MLS), exploring new perspectives (e.g., in ULS), or combining both. Despite these advances, uncertainties persist regarding branch reconstruction accuracy, partly due to the lack of studies with rigorous field measurements for parameters like branch angle or length.
Considerable uncertainties exist throughout the tree characterisation process when using technologies like TLS, and these uncertainties likely extend to other technologies such as ALS, PLS, and T-SfM. The results of some studies are based on idealised conditions, which may not reflect real-world challenges. The quality of output in methods like QSM is closely tied to the quality of the input point cloud, and QSM may not perform optimally with lower-grade sensors found in PLS or ULS units, which introduce more noise compared to higher-grade TLS sensors. Further research is needed to understand the limitations of these emerging technologies and develop noise mitigation procedures. Although initial efforts have been made to quantify uncertainty in PLS [269], additional work is required to calibrate these devices, address uncertainty, and improve precision in tree characterisation.
Though the advent of QSM has semi-democratised branch characterisation algorithms, there is still a reasonable amount of coding knowledge required to operate them. Some packages, such as TreeQSM and aRchi [270], have training material available online; however, require a software license and/or some basic knowledge of the respective programming language to use them. SimpleForest is arguably the most accessible and open-source option available as it resides within the CompuTree platform [271], presenting a graphical user interface (GUI)-based system for users along with very detailed instructions. As for methods outside of QSM, various skeletonisation packages such as Skeletor [272] and Plantscan3D [98, 273] are available online, largely in open source languages such as Python [274], requiring programming skills to operate them. Other methods such as BSM methodologies and the branch segmentation methodologies of Harikumar et al. [67•] and du Toit et al. [68••] do not have readily compiled packages available and would require considerable programming ability to reproduce. Likewise, deep learning methodologies, though available as Python packages online [87, 171, 244], would require coding ability and considerable computing resources to implement. A recent package developed using Python to derive basic inventory metrics from TLS and PLS data (3DFIN; [275]) was released with a variety of plugin options for commonly used geospatial GUIs including QGIS [276] and CloudCompare [277]. The authors applaud this multi-platform approach and recommend this as a model template to make software available to people with all levels of programming ability.
Aside from the platform and the algorithms, the size of the data is also something that remains a significant challenge for some of these technologies. With point densities in the tens or hundreds of thousands, or even millions of ppm2, data sets can quickly use up available computing power and become unwieldy. That is not to mention storage, and for practitioners whose current methods might involve storage of a few spreadsheets with field measurements, the prospect of switching to methods that regularly accumulate hundreds of gigabytes of data could be daunting. Technologies, such as ULS have advantages here as they collect less dense data; however, further work on this technology is required to ensure that the required branch characteristics can be derived from the data.
The advent of QSM has made branch-level tree modelling accessible to domain experts. Nevertheless, the manual extraction of precise metrics from point cloud data remains a significant challenge. This hinders the uptake of these technologies by operational users such as tree breeders, foresters, and mensuration technicians, who require reliable, accurate and repeatable methods for assessing tree form. Thus, to promote widespread adoption of these technologies, it is crucial to introduce fully automated solutions that can transform input point clouds into tree metrics without the need for excessive parameterisation. DL is emerging as a technology with the potential to automate specific manual aspects of this workflow. The implementation of fully automated methods for digital branch characterisation through DL could substantially reduce the entry barrier into this field.
Conclusion
The literature highlights a rapidly growing body of research focused on branch characterisation utilising point cloud-generating technologies, notably laser scanners. This review has evaluated the advantages and disadvantages of these various technologies and highlighted key research trends. The review showed that there are a wide range of methods available for characterising branch structure, with QSM playing a significant role in promoting the use of point cloud-based branch characterisation in applied research. This trend infers that branch-level characterisation has moved beyond the exclusive capabilities of programming experts and is becoming more accessible to domain experts, expanding its utility in various scientific fields. Previous studies have examined branch characterisation across a diverse range of forest types, uncovering areas for potential future investigation. This review has identified a number of challenges, such as the persistent issue of occlusion, large data sizes, and the noise and uncertainty present in some of technologies, such as PLS. Conversely, a number of opportunities in this field have been identified, including the emergence of deep learning methodologies to streamline processing pipelines, and open source databases for archiving point cloud datasets, making them accessible to other researchers. It is hoped that this review can guide forest growers, tree breeders and the 3D tree modelling community toward operational advancements.
Data availability
No datasets were generated or analysed during the current study.
References
Papers of particular interest have been highlighted as: • Of importance •• Of major importance
Sellier D, Fourcaud T, Lac P. A finite element model for investigating effects of aerial architecture on tree oscillations. Tree Physiol. 2006;26(6):799–806.
Damesin C, Ceschia E, Le Goff N, Ottorini JM, Dufrêne E. Stem and branch respiration of beech: from tree measurements to estimations at the stand level. New Phytol. 2002;153(1):159–72.
Malhi Y, Jackson T, Patrick Bentley L, Lau A, Shenkin A, Herold M, et al. New perspectives on the ecology of tree structure and tree communities through terrestrial laser scanning. Interface Focus. 2018;8(2):20170052.
Fleck S, Niinemets Ü, Cescatti A, Tenhunen JD. Three-dimensional lamina architecture alters light-harvesting efficiency in Fagus: a leaf-scale analysis. Tree Physiol. 2003;23(9):577–89.
Niinemets Ü. A review of light interception in plant stands from leaf to canopy in different plant functional types and in species with varying shade tolerance. Ecol Res. 2010;25:693–714.
Grace J, Pont D, Goulding C, Rawley B. Modelling branch development for forest management. N Z J For Sci. 1999;29(3):391–408.
Rais A, Poschenrieder W, Pretzsch H, van de Kuilen J-WG. Influence of initial plant density on sawn timber properties for Douglas-fir (Pseudotsuga menziesii (Mirb.) Franco). Ann Forest Sci. 2014;71(5):617–26.
Hartley RJ, Jayathunga S, Massam PD, De Silva D, Estarija HJ, Davidson SJ, et al. Assessing the potential of backpack-mounted mobile laser scanning systems for tree phenotyping. Remote Sens. 2022;14(14):3344.
Huuskonen S, Hakala S, Mäkinen H, Hynynen J, Varmola M. Factors influencing the branchiness of young Scots pine trees. Forestry. 2014;87(2):257–65.
Watt M, Moore J, McKinlay B. The influence of wind on branch characteristics of Pinus radiata. Trees. 2005;19(1):58–65.
McCallum D, Mason E, Whitley B. Influence of exposure and elevation on radiata pine branch size, log velocity, sweep, taper and value. N Z J For. 2007;52(3):10.
Mead D. Response of young Pinus radiata to cultivation and fertiliser Near Motueka, New Zealand. N Z J For Sci. 1990;20(3):268–78.
Carson M, Inglis C. Genotype and location effects on internode length of Pinus radiata in New Zealand. N Z J For Sci. 1988;18(3):267279.
Siemon G, Wood G, Forrest W. Effects of thinning on crown structure in radiata pine. N Z J For Sci. 1976;6(1):57–66.
Bollmann M, Sweet G. Bud morphogenesis of Pinus radiata in New Zealand. 1: the initiation and extension of the leading shoot of one clone at two sites. N Z J For Sci=. 1976;6(3):376–92.
Ceulemans R, Stettler R, Hinckley T, Isebrands J, Heilman P. Crown architecture of Populus clones as determined by branch orientation and branch characteristics. Tree Physiol. 1990;7(1-2-3–4):157–67.
Nelson ND, Burk T, Isebrands J. Crown architecture of short-rotation, intensively cultured Populus.: I. Effects of clone and spacing on first-order branch characteristics. Can J For Res. 1981;11(1):73–81.
Sillett SC, Van Pelt R, Carroll AL, Campbell-Spickler J, Antoine ME. Aboveground biomass dynamics and growth efficiency of Sequoia sempervirens forests. For Ecol Manag. 2020;458:117740.
Kramer RD, Sillett SC, Van Pelt R. Quantifying aboveground components of Picea sitchensis for allometric comparisons among tall conifers in North American rainforests. For Ecol Manag. 2018;430:59–77.
Niknejad N, Bidese-Puhl R, Bao Y, Payn KG, Zheng J. Phenotyping of architecture traits of loblolly pine trees using stereo machine vision and deep learning: Stem diameter, branch angle, and branch diameter. Comput Electron Agric. 2023;211:107999.
Poland JA, Nelson RJ. In the eye of the beholder: the effect of rater variability and different rating scales on QTL mapping. Phytopathology. 2011;101(2):290–8.
Enquist BJ, West GB, Brown JH. Extensions and evaluations of a general quantitative theory of forest structure and dynamics. Proc Natl Acad Sci. 2009;106(17):7046-51.
Enquist BJ, West GB, Charnov EL, Brown JH. Allometric scaling of production and life-history variation in vascular plants. Nature. 1999;401(6756):907–11.
Shinozaki K, Yoda K, Hozumi K, Kira T. A quantitative analysis of plant form-the pipe model theory: II. Further evidence of the theory and its application in forest ecology. Jpn J Ecol. 1964;14(4):133–9.
Dassot M, Fournier M, Deleuze C. Assessing the scaling of the tree branch diameters frequency distribution with terrestrial laser scanning: methodological framework and issues. Ann Forest Sci. 2019;76(3). https://doi.org/10.1007/s13595-019-0854-7.
Åkerblom M, Kaitaniemi P. Terrestrial laser scanning: A new standard of forest measuring and modelling? Ann Bot. 2021;128(6):653–62. https://doi.org/10.1093/aob/mcab111.
Duncanson L, Kellner JR, Armston J, Dubayah R, Minor DM, Hancock S, et al. Aboveground biomass density models for NASA’s Global Ecosystem Dynamics Investigation (GEDI) lidar mission. Remote Sens Environ. 2022;270:112845.
Bombrun M, Dash JP, Pont D, Watt MS, Pearse GD, Dungey HS. Forest-scale phenotyping: productivity characterisation through machine learning. Front Plant Sci. 2020;11(99). https://doi.org/10.3389/fpls.2020.00099.
Côté JF, Fournier RA, Frazer GW, Olaf NK. A fine-scale architectural model of trees to enhance LiDAR-derived measurements of forest canopy structure. Agric For Meterol. 2012;166–167:72–85. https://doi.org/10.1016/j.agrformet.2012.06.007.
Andersen H-E, Reutebuch SE, McGaughey RJ. A rigorous assessment of tree height measurements obtained using airborne lidar and conventional field methods. Can J Remote Sens. 2006;32(5):355–66.
•Brede B, Calders K, Lau A, Raumonen P, Bartholomeus HM, Herold M, et al. Non-destructive tree volume estimation through quantitative structure modelling: comparing UAV laser scanning with terrestrial LIDAR. Remote Sens Environ. 2019;233. https://doi.org/10.1016/j.rse.2019.111355. Study comparing above and below canopy close-range laser scanning approached for tree structure characterisation.This paper demonstrates the exploration of new technologies to address the occlusion issues associated with TLS.
Calders K, Adams J, Armston J, Bartholomeus H, Bauwens S, Bentley LP, et al. Terrestrial laser scanning in forest ecology: expanding the horizon. Remote Sens Environ. 2020;251:112102.
Lau A, Bentley LP, Martius C, Shenkin A, Bartholomeus H, Raumonen P, et al. Quantifying branch architecture of tropical trees using terrestrial LiDAR and 3D modelling. Trees Struct Funct. 2018;32(5):1219–31. https://doi.org/10.1007/s00468-018-1704-1.
Lambeth CC. Juvenile-mature correlations in Pinaceae and implications for early selection. For Sci. 1980;26(4):571–80.
Doede D, Adams W. The genetics of stem volume, stem form, and branch characteristics in sapling noble fir. Silvae Genet. 1998;47(4):177–82.
Codesido V, Fernández-López J. Juvenile genetic parameter estimates for vigour, stem form, branching habit and survival in three radiata pine (Pinus radiata D.Don) progeny tests in Galicia, NW Spain. Eur J For Res. 2008;127(4):315–25. https://doi.org/10.1007/s10342-008-0207-9.
Zhang H, Huang M, Qing X, Li G, Tian C. Bibliometric analysis of global remote sensing research during 2010–2015. ISPRS Int J Geo-Inf. 2017;6(11):332.
Kelly J, Sadeghieh T, Adeli K. Peer review in scientific publications: benefits, critiques, & a survival guide. Ejifcc. 2014;25(3):227.
Shelbourne C. Genetic improvement in different tree characteristics of Pinus radiata and the consequences for silviculture and utilisation. Pruning Thinning Pract. 1970;2:44–58.
Zobel B, Talbert J. Applied forest tree improvement. Wiley; 1984.
Calders K, Origo N, Burt A, Disney M, Nightingale J, Raumonen P, et al. Realistic forest stand reconstruction from terrestrial LiDAR for radiative transfer modelling. Remote Sens. 2018;10(6):933.
Côté JF, Fournier RA, Luther JE, van Lier OR. Fine-scale three-dimensional modeling of boreal forest plots to improve forest characterization with remote sensing. Remote Sens Environ. 2018;219:99–114. https://doi.org/10.1016/j.rse.2018.09.026.
Gonzalez de Tanago J, Lau A, Bartholomeus H, Herold M, Avitabile V, Raumonen P, et al. Estimation of above-ground biomass of large tropical trees with terrestrial LiDAR. Methods Ecol Evol. 2018;9(2):223–34.
Stovall AEL, Vorster AG, Anderson RS, Evangelista PH, Shugart HH. Non-destructive aboveground biomass estimation of coniferous trees using terrestrial LiDAR. Remote Sens Environ. 2017;200:31–42. https://doi.org/10.1016/j.rse.2017.08.013.
Van de Peer T, Verheyen K, Kint V, Van Cleemput E, Muys B. Plasticity of tree architecture through interspecific and intraspecific competition in a young experimental plantation. For Ecol Manag. 2017;385:1–9.
Pérez-Cruzado C, Kleinn C, Magdon P, Álvarez-González JG, Magnussen S, Fehrmann L, et al. The horizontal distribution of branch biomass in European beech: a model based on measurements and TLS based proxies. Remote Sens. 2021;13(5):1041.
Osborne NL, Maguire DA. Modeling knot geometry from branch angles in Douglas-fir (Pseudotsuga menziesii). Can J For Res. 2016;46(2):215–24.
Kint V, Hein S, Campioli M, Muys B. Modelling self-pruning and branch attributes for young Quercus robur L. and Fagus sylvatica L. trees. For Ecol Manag. 2010;260(11):2023–34.
Raymond CA, Cotterill PP. Methods of assessing crown form of Pinus radiata. Silvae Genet. 1990;39(2):67–71.
Karkee M, Adhikari B, Amatya S, Zhang Q. Identification of pruning branches in tall spindle apple trees for automated pruning. Comput Electron Agric. 2014;103:127–35.
Nguyen TT, Slaughter DC, Max N, Maloof JN, Sinha N. Structured light-based 3D reconstruction system for plants. Sensors. 2015;15(8):18587–612.
Tabb A. Three-dimensional reconstruction of fruit trees by a shape from silhouette method. In 2009 Proceedings of the ASABE Annual International Meeting, Reno, Nevada, ASABE; 2009. pp. 1. https://doi.org/10.13031/2013.27064.
Dutagaci H, Rasti P, Galopin G, Rousseau D. ROSE-X: an annotated data set for evaluation of 3D plant organ segmentation methods. Plant Methods. 2020;16:1–14.
Song J, Brendel O, Bodénès C, Plomion C, Kremer A, Colin F. X-ray computed tomography to decipher the genetic architecture of tree branching traits: oak as a case study. Tree Genet Genomes. 2017;13:1–15.
Wilkes P, Shenkin A, Disney M, Malhi Y, Bentley LP, Vicari MB. Terrestrial laser scanning to reconstruct branch architecture from harvested branches. Methods Ecol Evol. 2021;12(12):2487–500. https://doi.org/10.1111/2041-210X.13709.
Dassot M, Colin A, Santenoise P, Fournier M, Constant T. Terrestrial laser scanning for measuring the solid wood volume, including branches, of adult standing trees in the forest environment. Comput Electron Agric. 2012;89:86–93. https://doi.org/10.1016/j.compag.2012.08.005.
Bournez E, Landes T, Najjar G, Kastendeuch P, Ngao J, Saudreau M. Sensitivity of simulated light interception and tree transpiration to the level of detail of 3D tree reconstructions. Urban For Urban Greening. 2019;38:1–10. https://doi.org/10.1016/j.ufug.2018.10.016.
Sinoquet H, Rivet P, Godin C. Assessment of the three-dimensional architecture of walnut trees using digitising. Silva fennica. 1997;31(3):265–73. https://doi.org/10.14214/sf.a8525.
Sellier D, Fourcaud T. A mechanical analysis of the relationship between free oscillations of Pinus pinaster Ait. saplings and their aerial architecture. J Exp Botany. 2005;56(416):1563–73.
Morgenroth J, Gomez C. Assessment of tree structure using a 3D image analysis technique-a proof of concept. Urban For Urban Greening. 2014;13(1):198–203. https://doi.org/10.1016/j.ufug.2013.10.005.
Dong Y, Fan G, Zhou Z, Liu J, Wang Y, Chen F. Low cost automatic reconstruction of tree structure by adqsm with terrestrial close-range photogrammetry. Forests. 2021;12(8). https://doi.org/10.3390/f12081020.
Miller J, Morgenroth J, Gomez C. 3D modelling of individual trees using a handheld camera: accuracy of height, diameter and volume estimates. Urban For Urban Greening. 2015;14(4):932–40. https://doi.org/10.1016/j.ufug.2015.09.001.
Bucksch A, Fleck S. Automated detection of branch dimensions in woody skeletons of Fruit tree canopies. Photogramm Eng Remote Sens. 2011;77(3):229–40. https://doi.org/10.14358/PERS.77.3.229.
Raumonen P, Casella E, Calders K, Murphy S, Åkerblom M, Kaasalainen M. Massive-scale tree modelling from TLS data. ISPRS Ann Photogramm Remote Sens Spat Inform Sci. 2015;2(3):189.
•Hackenberg J, Spiecker H, Calders K, Disney M, Raumonen P. SimpleTree - an efficient open source tool to build tree models from TLS clouds. Forests. 2015;6(11):4245–94. https://doi.org/10.3390/f6114245. One of the early, fully open source QSM tool sets.
Guo J, Xu S, Yan DM, Cheng Z, Jaeger M, Zhang X. Realistic procedural plant modeling from multiple view images. IEEE Trans Visual Comput Graphics. 2020;26(2):1372–84. https://doi.org/10.1109/TVCG.2018.2869784.
•Harikumar A, Bovolo F, Bruzzone L. An internal crown geometric model for conifer species classification with high-density LiDAR Data. IEEE Trans Geosci Remote Sens. 2017;55(5):2924–40. https://doi.org/10.1109/TGRS.2017.2656152. Early, novel methodology deriving internal branch characteristics from ALS.
••du Toit F, Coops NC, Goodbody TR, Stoehr M, El-Kassaby YA. Deriving internal crown geometric features of Douglas-fir from airborne laser scanning in a realized-gain trial. Forestry: Int J For Res. 2021;94(3):442–54. https://doi.org/10.1093/forestry/cpaa046. Expanding on earlier methods to characterise branching from ALS into forest environments. This study demonstrates that branchcharacterisation could be scaled over large areas using ALS.
Briggs DG, Kantavichai R, Turnblom EC. Effect of precommercial thinning followed by a fertilization regime on branch diameter in coastal United States Douglas-fir plantations. Can J For Res. 2008;38(6):1564–75. https://doi.org/10.1139/X07-199.
Briggs DG, Kantavichai R, Turnblom EC. Predicting the diameter of the largest breast-height region branch of Douglas-fir trees in thinned and fertilized plantations. For Prod J. 2010;60(4):322–30. https://doi.org/10.13073/0015-7473-60.4.322.
Ko C, Sohn G, Remmel TK. Tree genera classification with geometric features from high-density airborne LiDAR. Can J Remote Sens. 2013;39(sup1):S73–85.
Pyörälä J, Saarinen N, Kankare V, Coops NC, Liang X, Wang Y, et al. Variability of wood properties using airborne and terrestrial laser scanning. Remote Sens Environ. 2019;235. https://doi.org/10.1016/j.rse.2019.111474.
du Toit F, Coops NC, Ratcliffe B, El-Kassaby YA, Lucieer A. Modelling internal tree attributes for breeding applications in Douglas-fir progeny trials using RPAS-ALS. Sci Remote Sens. 2023;7:100072.
Qi Y, Coops NC, Daniels LD, Butson CR. Assessing the effects of burn severity on post-fire tree structures using the fused drone and mobile laser scanning point clouds. Front Environ Sci. 2022;10:949442. https://doi.org/10.3389/fenvs.2022.949442.
Qi Y, Coops NC, Daniels LD, Butson CR. Comparing tree attributes derived from quantitative structure models based on drone and mobile laser scanning point clouds across varying canopy cover conditions. ISPRS J Photogramm Remote Sens. 2022;192:49–65.
Hu S, Li Z, Zhang Z, He D, Wimmer M. Efficient tree modeling from airborne LiDAR point clouds. Comput Graphics (Pergamon). 2017;67:1–13. https://doi.org/10.1016/j.cag.2017.04.004.
Li J, Wu H, Xiao Z, Lu H. 3D modeling of laser-scanned trees based on skeleton refined extraction. Int J Appl Earth Obs Geoinformation. 2022;112. https://doi.org/10.1016/j.jag.2022.102943.
Calders K, Newnham G, Burt A, Murphy S, Raumonen P, Herold M, et al. Nondestructive estimates of above-ground biomass using terrestrial laser scanning. Methods Ecol Evol. 2015;6(2):198–208.
Xu H, Gossett N, Chen B. Knowledge and heuristic-based modeling of laser-scanned trees. ACM Trans Graphics. 2007;26(4). https://doi.org/10.1145/1289603.1289610.
Li Y, Su Y, Zhao X, Yang M, Hu T, Zhang J, et al. Retrieval of tree branch architecture attributes from terrestrial laser scan data using a Laplacian algorithm. Agric For Meterol. 2020;284. https://doi.org/10.1016/j.agrformet.2019.107874.
•Pyorala J, Liang X, Vastaranta M, Saarinen N, Kankare V, Wang Y, et al. Quantitative assessment of scots pine (Pinus Sylvestris L.) whorl structure in a forest environment using terrestrial laser scanning. IEEE J Sel Top Appl Earth Obs Remote Sens. 2018;11(10):3598–607. https://doi.org/10.1109/JSTARS.2018.2819598. Detailed study representing an early attempt to describe branch structure from TLS for timber quality in a forestry setting.
Hackenberg J, Wassenberg M, Spiecker H, Sun D. Non destructive method for biomass prediction combining TLS derived tree volume and wood density. Forests. 2015;6(4):1274–300. https://doi.org/10.3390/f6041274.
Hu M, Pitkänen TP, Minunno F, Tian X, Lehtonen A, Mäkelä A. A new method to estimate branch biomass from terrestrial laser scanning data by bridging tree structure models. Ann Bot. 2021;128(6):737–52. https://doi.org/10.1093/aob/mcab037.
Wilkes P, Lau A, Disney M, Calders K, Burt A, Gonzalez de Tanago J, et al. Data acquisition considerations for terrestrial laser scanning of forest plots. Remote Sens Environ. 2017;196:140–53. https://doi.org/10.1016/j.rse.2017.04.030.
Aiteanu F, Klein R. Exploring shape spaces of 3D tree point clouds. Comput Graphics (Pergamon). 2021;100:21–31. https://doi.org/10.1016/j.cag.2021.07.013.
Vandendaele B, Martin-Ducup O, Fournier RA, Pelletier G, Lejeune P. Mobile laser scanning for estimating tree structural attributes in a temperate hardwood forest. Remote Sens. 2022;14(18):4522.
Liu Y, Guo J, Benes B, Deussen O, Zhang X, Huang H. TreePartNet: neural decomposition of point clouds for 3D tree reconstruction. In: ACM Transactions on Graphics (Proceedings of SIGGRAPH ASIA). ACM; 2021;40(6):1–16. https://doi.org/10.1145/3478513.3480486.
Åkerblom M, Raumonen P, Casella E, Disney MI, Danson FM, Gaulton R, et al. Non-intersecting leaf insertion algorithm for tree structure models. Interface Focus. 2018;8(2):20170045.
Åkerblom M, Raumonen P, Kaasalainen M, Casella E. Analysis of geometric primitives in quantitative structure models of tree stems. Remote Sens. 2015;7(4):4581–603. https://doi.org/10.3390/rs70404581.
Åkerblom M, Raumonen P, Mäkipää R, Kaasalainen M. Automatic tree species recognition with quantitative structure models. Remote Sens Environ. 2017;191:1–12. https://doi.org/10.1016/j.rse.2016.12.002.
Arseniou G, Macfarlane DW, Seidel D. Woody surface area measurements with terrestrial laser scanning relate to the anatomical and structural complexity of urban trees. Remote Sens. 2021;13(16). https://doi.org/10.3390/rs13163153.
Bayer D, Reischl A, Rötzer T, Pretzsch H. Structural response of black locust (Robinia pseudoacacia L.) and small-leaved lime (Tilia cordata Mill.) to varying urban environments analyzed by terrestrial laser scanning: Implications for ecological functions and services. Urban For Urban Greening. 2018;35:129–38. https://doi.org/10.1016/j.ufug.2018.08.011.
Bayer D, Seifert S, Pretzsch H. Structural crown properties of Norway spruce (Picea abies [L.] Karst.) and European beech (Fagus sylvatica [L.]) in mixed versus pure stands revealed by terrestrial laser scanning. Trees Struct Funct. 2013;27(4):1035–47. https://doi.org/10.1007/s00468-013-0854-4.
Beyer RM, Basler D, Raumonen P, Kaasalainen M, Pretzsch H. Do trees have constant branch divergence angles? J Theor Biol. 2021;512. https://doi.org/10.1016/j.jtbi.2020.110567.
Bittner S, Legner N, Beese F, Priesack E. Individual tree branch-level simulation of light attenuation and water flow of three F. sylvatica L. trees. J Geophys Res: Biogeosci. 2012;117(1). https://doi.org/10.1029/2011JG001780.
Bohn Reckziegel R, Mbongo W, Kunneke A, Morhart C, Sheppard JP, Chirwa P, et al. Exploring the branch wood supply potential of an agroforestry system with strategically designed harvesting interventions based on terrestrial LiDAR data. Forests. 2022;13(5). https://doi.org/10.3390/f13050650.
Bohn Reckziegel R, Sheppard JP, Kahle HP, Larysch E, Spiecker H, Seifert T, et al. Virtual pruning of 3D trees as a tool for managing shading effects in agroforestry systems. Agrofor Syst. 2022;96(1):89–104. https://doi.org/10.1007/s10457-021-00697-5.
•Boudon F, Preuksakarn C, Ferraro P, Diener J, Nacry P, Nikinmaa E, et al. Quantitative assessment of automatic reconstructions of branching systems obtained from laser scanning. Ann Bot. 2014;114(4):853–62. https://doi.org/10.1093/aob/mcu062. One of the only attempts to create a framework to assess the fidelity of tree structural models to the original tree.
Bournez E, Landes T, Saudreau M, Kastendeuch P, Najjar G. From TLS point clouds to 3D models of trees: a comparison of existing algorithms for 3D tree reconstruction. In: The International Archives of the Photogrammetry, Remote Sensing and Spatial Information Sciences, Nafplio, Greece. ISPRS; 2017;42(2/W3):113–20. https://doi.org/10.5194/isprs-archives-XLII-2-W3-113-2017.
Bremer M, Rutzinger M, Wichmann V. Derivation of tree skeletons and error assessment using LiDAR point cloud data of varying quality. ISPRS J Photogramm Remote Sens. 2013;80:39–50. https://doi.org/10.1016/j.isprsjprs.2013.03.003.
Bremer M, Wichmann V, Rutzinger M. Calibration and validation of a detailed architectural canopy model reconstruction for the simulation of synthetic hemispherical images and airborne LiDAR data. Remote Sens. 2017;9(3). https://doi.org/10.3390/rs9030220.
Bremer M, Wichmann V, Rutzinger M. Multi-temporal fine-scale modelling of Larix decidua forest plots using terrestrial LiDAR and hemispherical photographs. Remote Sens Environ. 2018;206:189–204. https://doi.org/10.1016/j.rse.2017.12.023.
Bucksch A, Lindenbergh R. CAMPINO—a skeletonization method for point cloud processing. ISPRS J Photogramm Remote Sens. 2008;63(1):115–27.
Burkardt K, Annighöfer P, Seidel D, Ammer C, Vor T. Intraspecific competition affects crown and stem characteristics of non-native Quercus rubra L. stands in Germany. Forests. 2019;10(10). https://doi.org/10.3390/f10100846.
Burkardt K, Pettenkofer T, Ammer C, Gailing O, Leinemann L, Seidel D, et al. Influence of heterozygosity and competition on morphological tree characteristics of Quercus rubra L.: a new single-tree based approach. New For. 2021;52(4):679–95. https://doi.org/10.1007/s11056-020-09814-1.
Burt A, Disney M, Raumonen P, Armston J, Calders K, Lewis P. Rapid characterisation of forest structure from TLS and 3D modelling. In: Proceedings of the IEEE international geoscience and remote sensing symposium (IGARSS): IEEE; 2013. pp. 3387–90. https://doi.org/10.1109/IGARSS.2013.6723555.
Calders K, Burt A, Newnham G, Disney M, Murphy S, Raumonen P, et al. Reducing uncertainties in above-ground biomass estimates using terrestrial laser scanning. In: Proc Silvilaser, La Grande Motte, France. Silvilaser; 2015. pp. 197–9.
Chaudhury A, Godin C. Skeletonization of plant point cloud data using stochastic optimization framework. Front Plant Sci. 2020;11. https://doi.org/10.3389/fpls.2020.00773.
Côté J-F, Widlowski J-L, Fournier RA, Verstraete MM. The structural and radiative consistency of three-dimensional tree reconstructions from terrestrial lidar. Remote Sens Environ. 2009;113(5):1067–81.
Côté JF, Fournier RA, Egli R. An architectural model of trees to estimate forest structural attributes using terrestrial LiDAR. Environ Model Softw. 2011;26(6):761–77. https://doi.org/10.1016/j.envsoft.2010.12.008.
Côté JF, Fournier RA, Luther JE. Validation of L-Architect model for balsam fir and black spruce trees with structural measurements. Can J Remote Sens. 2013;39(SUPPL.1):S41–59. https://doi.org/10.5589/m13-014.
Côté JF, Luther JE, Lenz P, Fournier RA, van Lier OR. Assessing the impact of fine-scale structure on predicting wood fibre attributes of boreal conifer trees and forest plots. For Ecol Manag. 2021;479. https://doi.org/10.1016/j.foreco.2020.118624.
Demol M, Calders K, Verbeeck H, Gielen B. Forest above-ground volume assessments with terrestrial laser scanning: a ground-truth validation experiment in temperate, managed forests. Ann Bot. 2021;128(6):805–19. https://doi.org/10.1093/aob/mcab110.
Demol M, Verbeeck H, Gielen B, Armston J, Burt A, Disney M, et al. Estimating forest above-ground biomass with terrestrial laser scanning: current status and future directions. Methods Ecol Evol. 2022;13(8):1628–39.
Demol M, Wilkes P, Raumonen P, Moorthy SMK, Calders K, Gielen B, et al. Volumetric overestimation of small branches in 3D reconstructions of Fraxinus excelsior. Silva Fenn. 2022;56(1). https://doi.org/10.14214/sf.10550.
Dorji Y, Annighöfer P, Ammer C, Seidel D. Response of beech (Fagus sylvatica L.) trees to competition-new insights from using fractal analysis. Remote Sens. 2019;11(22). https://doi.org/10.3390/rs11222656.
Du S, Lindenbergh R, Ledoux H, Stoter J, Nan L. AdTree: Accurate, detailed, and automatic modelling of laser-scanned trees. Remote Sens. 2019;11(18). https://doi.org/10.3390/rs11182074.
Dutcă I, Cernat A, Stăncioiu PT, Ioraș F, Niță MD. Does Slope Aspect affect the aboveground tree shape and volume allometry of european beech (Fagus sylvatica L.) trees? Forests. 2022;13(7). https://doi.org/10.3390/f13071071.
Eysn L, Pfeifer N, Ressl C, Hollaus M, Grafl A, Morsdorf F. A practical approach for extracting tree models in forest environments based on equirectangular projections of terrestrial laser scans. Remote Sens. 2013;5(11):5424–48. https://doi.org/10.3390/rs5115424.
Fan G, Nan L, Chen F, Dong Y, Wang Z, Li H, et al. A new quantitative approach to tree attributes estimation based on LiDAR point clouds. Remote Sens. 2020;12(11). https://doi.org/10.3390/rs12111779.
Fan G, Nan L, Dong Y, Su X, Chen F. AdQSM: A new method for estimating above-ground biomass from TLS point clouds. Remote Sens. 2020;12(18). https://doi.org/10.3390/RS12183089.
Fang R, Strimbu BM. Comparison of mature Douglas-firs' crown structures developed with two quantitative structural models using TLS point clouds for neighboring trees in a natural regime stand. Remote Sens. 2019;11(14). https://doi.org/10.3390/rs11141661.
Ferrara R, Virdis SGP, Ventura A, Ghisu T, Duce P, Pellizzaro G. An automated approach for wood-leaf separation from terrestrial LIDAR point clouds using the density based clustering algorithm DBSCAN. Agric For Meterol. 2018;262:434–44. https://doi.org/10.1016/j.agrformet.2018.04.008.
Gorte B, Pfeifer N. Structuring laser-scanned trees using 3D mathematical morphology. Int Arch Photogramm Remote Sens. 2004;35(B5):929–33.
Guillemot J, Kunz M, Schnabel F, Fichtner A, Madsen CP, Gebauer T, et al. Neighbourhood-mediated shifts in tree biomass allocation drive overyielding in tropical species mixtures. New Phytol. 2020;228(4):1256–68. https://doi.org/10.1111/nph.16722.
Hackenberg J, Bontemps JD. Improving quantitative structure models of trees inspired by pipe and metabolic scaling theory. BioRxiv; 2022. pp. 1–32. https://doi.org/10.1101/2022.10.31.514601.
Hackenberg J, Disney M, Bontemps J-D. Gaining insight into the allometric scaling of trees by utilizing 3d reconstructed tree models-a SimpleForest study. BioRxiv; 2022. pp. 1–18. https://doi.org/10.1101/2022.05.05.490069.
Hackenberg J, Morhart C, Sheppard J, Spiecker H, Disney M. Highly accurate tree models derived from terrestrial laser scan data: a method description. Forests. 2014;5(5):1069–105. https://doi.org/10.3390/f5051069.
Harikumar A, Liang X, Bovolo F, Bruzzone L. Void-volume-based stem geometric modeling and branch-knot localization in terrestrial laser scanning data. IEEE J Sel Top Appl Earth Obs Remote Sens. 2022;15:3024–40. https://doi.org/10.1109/JSTARS.2022.3163404.
Hauglin M, Astrup R, Gobakken T, Næsset E. Estimating single-tree branch biomass of Norway spruce with terrestrial laser scanning using voxel-based and crown dimension features. Scand J For Res. 2013;28(5):456–69. https://doi.org/10.1080/02827581.2013.777772.
He G, Yang J, Behnke S. Research on geometric features and point cloud properties for tree skeleton extraction. Pers Ubiquitous Comp. 2018;22(5–6):903–10. https://doi.org/10.1007/s00779-018-1153-2.
Heidenreich MG, Seidel D. Assessing forest vitality and forest structure using 3D data: a case study from the Hainich National Park, Germany. Front For Glob Change. 2022;5. https://doi.org/10.3389/ffgc.2022.929106.
Henning JG, Radtke PJ. Detailed stem measurements of standing trees from ground-based scanning lidar. For Sci. 2006;52(1):67–80.
Hess C, Bienert A, Härdtle W, Von Oheimb G. Does tree architectural complexity influence the accuracy of wood volume estimates of single young trees by terrestrial laser scanning? Forests. 2015;6(11):3847–67. https://doi.org/10.3390/f6113847.
Hildebrand M, Perles-Garcia MD, Kunz M, Härdtle W, von Oheimb G, Fichtner A. Reprint of: tree-tree interactions and crown complementarity: the role of functional diversity and branch traits for canopy packing. Basic Appl Ecol. 2021;55:53–63. https://doi.org/10.1016/j.baae.2021.01.010.
Hosoi F, Nakai Y, Omasa K. 3-D voxel-based solid modeling of a broad-leaved tree for accurate volume estimation using portable scanning lidar. ISPRS J Photogramm Remote Sens. 2013;82:41–8. https://doi.org/10.1016/j.isprsjprs.2013.04.011.
Höwler K, Annighöfer P, Ammer C, Seidel D. Competition improves quality-related external stem characteristics of Fagus sylvatica. Can J For Res. 2017;47(12):1603–13.
Höwler K, Vor T, Seidel D, Annighöfer P, Ammer C. Analyzing effects of intra-and interspecific competition on timber quality attributes of Fagus sylvatica L.—from quality assessments on standing trees to sawn boards. Eur J For Res. 2019;138(2):327–43.
Huang Z, Huang X, Fan J, Eichhorn M, An F, Chen B, et al. Retrieval of aerodynamic parameters in rubber tree forests based on the computer simulation technique and terrestrial laser scanning data. Remote Sens. 2020;12(8). https://doi.org/10.3390/RS12081318.
Hui Z, Cai Z, Liu B, Li D, Liu H, Li Z. A self-adaptive optimization individual tree modeling method for terrestrial LiDAR point clouds. Remote Sens. 2022;14(11). https://doi.org/10.3390/rs14112545.
Indirabai I, Nair MVH, Jaishanker RN, Nidamanuri RR. Terrestrial laser scanner based 3D reconstruction of trees and retrieval of leaf area index in a forest environment. Ecol Inform. 2019;53. https://doi.org/10.1016/j.ecoinf.2019.100986.
Janoutová R, Homolová L, Malenovskỳ Z, Hanuš J, Lauret N, Gastellu-Etchegorry JP. Influence of 3D spruce tree representation on accuracy of airborne and satellite forest reflectance simulated in DART. Forests. 2019;10(3). https://doi.org/10.3390/f10030292.
Janoutová R, Homolová L, Novotný J, Navrátilová B, Pikl M, Malenovský Z. Detailed reconstruction of trees from terrestrial laser scans for remote sensing and radiative transfer modelling applications. In Silico Plants. 2021;3(2) diab026:1–22. https://doi.org/10.1093/insilicoplants/diab026.
Jin S, Zhang W, Shao J, Wan P, Cheng S, Cai S, et al. Estimation of larch growth at the stem, crown, and branch levels using ground-based LiDAR Point Cloud. J Remote Sens. 2022;2022.
Juchheim J, Annighöfer P, Ammer C, Calders K, Raumonen P, Seidel D. How management intensity and neighborhood composition affect the structure of beech (Fagus sylvatica L.) trees. Trees Struct Funct. 2017;31(5):1723–35. https://doi.org/10.1007/s00468-017-1581-z.
Kaasalainen S, Krooks A, Liski J, Raumonen P, Kaartinen H, Kaasalainen M, et al. Change detection of tree biomass with terrestrial laser scanning and quantitative structure modelling. Remote Sens. 2014;6(5):3906–22. https://doi.org/10.3390/rs6053906.
Kankare V, Holopainen M, Vastaranta M, Puttonen E, Yu X, Hyyppä J, et al. Individual tree biomass estimation using terrestrial laser scanning. ISPRS J Photogramm Remote Sens. 2013;75:64–75. https://doi.org/10.1016/j.isprsjprs.2012.10.003.
Kankare V, Joensuu M, Vauhkonen J, Holopainen M, Tanhuanpää T, Vastaranta M, et al. Estimation of the timber quality of scots pine with terrestrial laser scanning. Forests. 2014;5(8):1879–95. https://doi.org/10.3390/f5081879.
Kędra K, Barbeito I, Dassot M, Vallet P, Gazda A. Single-image photogrammetry for deriving tree architectural traits in mature forest stands: a comparison with terrestrial laser scanning. Ann Forest Sci. 2019;76(1). https://doi.org/10.1007/s13595-018-0783-x.
Knapp-Wilson J, Bohn Reckziegel R, Thapa Magar S, Bucksch A, Chavez DJ. Three-dimensional phenotyping of peach tree-crown architecture utilizing terrestrial laser scanning. Plant Phenome J. 2023;6(1):e20073.
Koma Z, Rutzinger M, Bremer M. Automated segmentation of leaves from deciduous trees in terrestrial laser scanning point clouds. IEEE Geosci Remote Sens Lett. 2018;15(9):1456–60. https://doi.org/10.1109/LGRS.2018.2841429.
•Kretschmer U, Kirchner N, Morhart C, Spiecker H. A new approach to assessing tree stem quality characteristics using terrestrial laser scans. Silva Fenn. 2013;47(5). https://doi.org/10.14214/sf.1071. An original and alternative method for branch characterisation in 2D, introducing the concept of the bark surface model (BSM).
Kunz M, Hess C, Raumonen P, Bienert A, Hackenberg J, Maas H, et al. Comparison of wood volume estimates of young trees from terrestrial laser scan data. iForest Biogeosci For. 2017;10(2):451–8.
Lau A, Martius C, Bartholomeus H, Shenkin A, Jackson T, Malhi Y, et al. Estimating architecture-based metabolic scaling exponents of tropical trees using terrestrial LiDAR and 3D modelling. For Ecol Manag. 2019;439:132–45. https://doi.org/10.1016/j.foreco.2019.02.019.
Lecigne B, Delagrange S, Messier C. Exploring trees in three dimensions: VoxR, a novel voxel-based R package dedicated to analysing the complex arrangement of tree crowns. Ann Bot. 2018;121(4):589–601. https://doi.org/10.1093/aob/mcx095.
Lecigne B, Delagrange S, Messier C. Crown reaction and acclimation to cyclical V-trimming of city trees: an analysis using terrestrial laser scanning. Urban For Urban Greening. 2018;29:183–91. https://doi.org/10.1016/j.ufug.2017.11.012.
Lengauer S, Houska P, Preiner R. Efficient point cloud skeletonization with locally adaptive L1-medial projection. J WSCG. 2022;2022(CSRN3201):38–47. https://doi.org/10.24132/CSRN.3201.6.
Li Y, Hess C, Von Wehrden H, Härdtle W, Von Oheimb G. Assessing tree dendrometrics in young regenerating plantations using terrestrial laser scanning. Ann Forest Sci. 2014;71(4):453–62. https://doi.org/10.1007/s13595-014-0358-4.
Liu W, Atherton J, Mõttus M, Gastellu-Etchegorry JP, Malenovský Z, Raumonen P, et al. Simulating solar-induced chlorophyll fluorescence in a boreal forest stand reconstructed from terrestrial laser scanning measurements. Remote Sens Environ. 2019;232. https://doi.org/10.1016/j.rse.2019.111274.
Martin-Ducup O, Mofack G, Wang D, Raumonen P, Ploton P, Sonké B, et al. Evaluation of automated pipelines for tree and plot metric estimation from TLS data in tropical forest areas. Ann Bot. 2021;128(6):753–66.
Martin-Ducup O, Ploton P, Barbier N, Momo Takoudjou S, Mofack G II, Kamdem NG, et al. Terrestrial laser scanning reveals convergence of tree architecture with increasingly dominant crown canopy position. Funct Ecol. 2020;34(12):2442–52. https://doi.org/10.1111/1365-2435.13678.
Mei J, Zhang L, Wu S, Wang Z, Zhang L. 3D tree modeling from incomplete point clouds via optimization and L1-MST. Int J Geogr Inf Sci. 2017;31(5):999–1021. https://doi.org/10.1080/13658816.2016.1264075.
Momo Takoudjou S, Ploton P, Sonké B, Hackenberg J, Griffon S, de Coligny F, et al. Using terrestrial laser scanning data to estimate large tropical trees biomass and calibrate allometric models: a comparison with traditional destructive approach. Methods Ecol Evol. 2018;9(4):905–16. https://doi.org/10.1111/2041-210X.12933.
Moravčík L, Vincúr R, Rózová Z. Analysis of the static behavior of a single tree on a finite element model. Plants. 2021;10(7). https://doi.org/10.3390/plants10071284.
Nguyen VT, Constant T, Colin F. An innovative and automated method for characterizing wood defects on trunk surfaces using high-density 3D terrestrial LiDAR data. Ann Forest Sci. 2021;78(2). https://doi.org/10.1007/s13595-020-01022-3.
Nguyen VT, Constant T, Kerautret B, Debled-Rennesson I, Colin F. A machine-learning approach for classifying defects on tree trunks using terrestrial LiDAR. Comput Electron Agric. 2020;171. https://doi.org/10.1016/j.compag.2020.105332.
Nock CA, Greene D, Delagrange S, Follett M, Fournier R, Messier C. In situ quantification of experimental ice accretion on tree crowns using terrestrial laser scanning. PLoS ONE. 2013;8(5). https://doi.org/10.1371/journal.pone.0064865.
Nock CA, Lecigne B, Taugourdeau O, Greene DF, Dauzat J, Delagrange S, et al. Linking ice accretion and crown structure: towards a model of the effect of freezing rain on tree canopies. Ann Bot. 2016;117(7):1163–73. https://doi.org/10.1093/aob/mcw059.
Olschofsky K, Mues V, Köhl M. Operational assessment of aboveground tree volume and biomass by terrestrial laser scanning. Comput Electron Agric. 2016;127:699–707. https://doi.org/10.1016/j.compag.2016.07.030.
Preuksakarn C, Boudon F, Ferraro P, Durand J-B, Nikinmaa E, Godin C. Reconstructing plant architecture from 3D laser scanner data. 6th International Workshop on Functional-Structural Plant Models2010. pp. 12-7.
••Puliti S, McLean JP, Cattaneo N, Fischer C, Astrup R. Tree height-growth trajectory estimation using uni-temporal UAV laser scanning data and deep learning. Foresty Int J For Res. 2023;96(1):37–48. https://doi.org/10.1093/forestry/cpac026. Important paper demonstrating the efficacy of deep learning techniques for characterising branches from laser scanner data. Deep learning offers advantages over traditional heuristic approaches, due to adaptability, capacity for continual learning and reduced manual intervention.
Pyörälä J, Liang X, Saarinen N, Kankare V, Wang Y, Holopainen M, et al. Assessing branching structure for biomass and wood quality estimation using terrestrial laser scanning point clouds. Can J Remote Sens. 2018;44(5):462–75. https://doi.org/10.1080/07038992.2018.1557040.
Rais A, Jacobs M, van de Kuilen JWG, Pretzsch H. Crown structure of european beech (Fagus sylvatica): a noncausal proxy for mechanical–physical wood properties. Can J For Res. 2021;51(6):834–41. https://doi.org/10.1139/cjfr-2020-0382.
••Raumonen P, Kaasalainen M, Markku A, Kaasalainen S, Kaartinen H, Vastaranta M, et al. Fast automatic precision tree models from terrestrial laser scanner data. Remote Sens. 2013;5(2):491–520. https://doi.org/10.3390/rs5020491. The original paper that introduced the concept of QSM. This paper and associated software, TreeQSM, arguably catalysed the expansion and accessibility of branch characterisation from laser scanned point clouds.
Schilling A, Schmidt A, Maas H-G. Tree topology representation from TLS point clouds using depth-first search in voxel space. Photogramm Eng Remote Sens. 2012;78(4):383–92.
Schütt C, Aschoff T, Winterhalder D, Thies M, Kretschmer U, Spiecker H. Approaches for recognition of wood quality of standing trees based on terrestrial laser scanner data. In: Proceedings of ISPRS WG VIII/2 The International Archives of the Photogrammetry, Remote Sensing and Spatial Information Sciences, Freiburg, Germany. ISPRS; 2004;36(8/W2):179–182.
Seidel D, Ehbrecht M, Dorji Y, Jambay J, Ammer C, Annighöfer P. Identifying architectural characteristics that determine tree structural complexity. Trees Struct Funct. 2019;33(3):911–9. https://doi.org/10.1007/s00468-019-01827-4.
Sheppard J, Morhart C, Hackenberg J, Spiecker H. Terrestrial laser scanning as a tool for assessing tree growth. IForest. 2017;10(1):172–9. https://doi.org/10.3832/ifor2138-009.
Sloup P. Automatic tree reconstruction from its laser scan. Master’s thesis, Faculty of Informatics, Masaryk University, Brno, Czech Republic, 2013. Available online https://is.muni.cz/th/nzqo6/thesis.pdf. Accessed 21 Nov 2022.
Stängle SM, Brüchert F, Kretschmer U, Spiecker H, Sauter UH. Clear wood content in standing trees predicted from branch scar measurements with terrestrial LiDAR and verified with X-ray computed tomography. Can J For Res. 2014;44(2):145–53.
Sun J, Wang P, Li R, Zhou M, Wu Y. Fast tree skeleton extraction using voxel thinning based on tree point cloud. Remote Sens. 2022;14(11). https://doi.org/10.3390/rs14112558.
Tao S, Guo Q, Xu S, Su Y, Li Y, Wu F. A geometric method for wood-leaf separation using terrestrial and simulated lidar data. Photogramm Eng Remote Sens. 2015;81(10):767–76. https://doi.org/10.14358/PERS.81.10.767.
Terryn L, Calders K, Åkerblom M, Bartholomeus H, Disney M, Levick S, et al. Analysing individual 3D tree structure using the R package ITSMe. Methods Ecol Evol. 2023;14(1):231–41.
Terryn L, Calders K, Disney M, Origo N, Malhi Y, Newnham G, et al. Tree species classification using structural features derived from terrestrial laser scanning. ISPRS J Photogramm Remote Sens. 2020;168:170–81.
Tomșa VR, Curtu AL, Niță MD. Tree shape variability in a mixed oak forest using terrestrial laser technology: Implications for mating system analysis. Forests. 2021;12(2):1–14. https://doi.org/10.3390/f12020253.
Van Den Berge S, Vangansbeke P, Calders K, Vanneste T, Baeten L, Verbeeck H, et al. Biomass expansion factors for hedgerow-grown trees derived from terrestrial LiDAR. Bioenergy Res. 2021;14(2):561–74. https://doi.org/10.1007/s12155-021-10250-y.
Wang D, Momo Takoudjou S, Casella E. LeWoS: a universal leaf-wood classification method to facilitate the 3D modelling of large tropical trees using terrestrial LiDAR. Methods Ecol Evol. 2020;11(3):376–89. https://doi.org/10.1111/2041-210X.13342.
Wang Z, Zhang L, Fang T, Mathiopoulos PT, Qu H, Chen D, et al. A structure-aware global optimization method for reconstructing 3-D tree models from terrestrial laser scanning data. IEEE Trans Geosci Remote Sens. 2014;52(9):5653–69. https://doi.org/10.1109/TGRS.2013.2291815.
Wu B, Zheng G, Chen Y, Yu D. Assessing inclination angles of tree branches from terrestrial laser scan data using a skeleton extraction method. Int J Appl Earth Obs Geoinformation. 2021;104:102589.
Wu S, Xiao B, Guo X, Wen W, Zhao C. An accurate fruit tree canopy reconstruction method based on dense point cloud. ICIC Express Lett Part B Appl. 2017;8(1):159–66.
Xi Z, Hopkinson C, Chasmer L. Filtering stems and branches from terrestrial laser scanning point clouds using deep 3-D fully convolutional networks. Remote Sens. 2018;10(8). https://doi.org/10.3390/rs10081215.
Xu S, Zhou K, Sun Y, Yun T. Separation of wood and foliage for trees from ground point clouds using a novel least-cost path model. IEEE J Sel Top Appl Earth Obs Remote Sens. 2021;14:6414–25. https://doi.org/10.1109/JSTARS.2021.3090502.
Yan D-M, Wintz J, Mourrain B, Wang W, Boudon F, Godin C. Efficient and robust reconstruction of botanical branching structure from laser scanned points. In: Proceedings of the 11th IEEE international conference on computer-aided design and computer graphics (CAD/Graphics 2009). IEEE; 2009. pp. 572–5.
Yang J, Wen X, Wang Q, Ye J-S, Zhang Y, Sun Y. A novel algorithm based on geometric characteristics for tree branch skeleton extraction from LiDAR point cloud. Forests. 2022;13(10):1534.
Yépez-Rincón FD, Luna-Mendoza L, Ramírez-Serrato NL, Hinojosa-Corona A, Ferriño-Fierro AL. Assessing vertical structure of an endemic forest in succession using terrestrial laser scanning (TLS). Case study: Guadalupe Island. Remote Sens Environ. 2021;263. https://doi.org/10.1016/j.rse.2021.112563.
Zhang C, Jiang Y, Xu B, Li X, Zhu Y, Lei L, et al. Apple tree branch information extraction from terrestrial laser scanning and backpack-LiDAR. Remote Sens. 2020;12(21):1–17. https://doi.org/10.3390/rs12213592.
Zhang X, Li H, Dai M, Ma W, Quan L. Data-driven synthetic modeling of trees. IEEE Trans Visual Comput Graphics. 2014;20(9):1214–26. https://doi.org/10.1109/TVCG.2014.2316001.
Zuleta D, Krishna Moorthy SM, Arellano G, Verbeeck H, Davies SJ. Vertical distribution of trunk and crown volume in tropical trees. For Ecol Manag. 2022;508. https://doi.org/10.1016/j.foreco.2022.120056.
Rutzinger M, Pratihast AK, Oude Elberink SJ, Vosselman G. Tree modelling from mobile laser scanning data-sets. Photogramm Rec. 2011;26(135):361–72. https://doi.org/10.1111/j.1477-9730.2011.00635.x.
Lin Y, Hyyppa J. Multiecho-recording mobile laser scanning for enhancing individual tree crown reconstruction. IEEE Trans Geosci Remote Sens. 2012;50(11 PART1):4323–32. https://doi.org/10.1109/TGRS.2012.2194503.
Livny Y, Yan F, Olson M, Chen B, Zhang H, El-Sana J. Automatic reconstruction of tree skeletal structures from point clouds. ACM SIGGRAPH Asia 2010 papers. 2010. pp. 1–8.
Méndez V, Rosell-Polo JR, Sanz R, Escolà A, Catalán H. Deciduous tree reconstruction algorithm based on cylinder fitting from mobile terrestrial laser scanned point clouds. Biosyst Eng. 2014;124:78–88. https://doi.org/10.1016/j.biosystemseng.2014.06.001.
••Winberg O, Pyörälä J, Yu X, Kaartinen H, Kukko A, Holopainen M, et al. Branch information extraction from Norway spruce using handheld laser scanning point clouds in Nordic forests. ISPRS Open J Photogramm Remote Sens. 2023;9:100040. https://doi.org/10.1016/j.ophoto.2023.100040. One of the first papers progressing TLS branch characterisation methodologies to MLS, representing a new approach to overcome the issues of occlusion common to TLS.
Xu J, Shan J, Wang G. Hierarchical modeling of street trees using mobile laser scanning. Remote Sens. 2020;12(14). https://doi.org/10.3390/rs12142321.
Lin W, Fan W, Liu H, Xu Y, Wu J. Classification of handheld laser scanning tree point cloud based on different KNN algorithms and random forest algorithm. Forests. 2021;12(3):1–36. https://doi.org/10.3390/f12030292.
Lowe T, Pinskier J. Tree reconstruction using topology optimisation. Remote Sens. 2022;15(1):172.
Wang M, Wong MS, Abbas S. Tropical species classification with structural traits using handheld laser scanning data. Remote Sens. 2022;14(8). https://doi.org/10.3390/rs14081948.
Xu S, Li X, Yun J, Xu S. An effectively dynamic path optimization approach for the tree skeleton extraction from portable laser scanning point clouds. Remote Sens. 2022;14(1). https://doi.org/10.3390/rs14010094.
Polhemus Inc. 3SPACE® FASTRAK® User Manual. Rev. G ed: Polhemus Inc., Colchester, Vermont, USA; 2012. Available online https://polhemus.com/_assets/img/FASTRAK_User_Manual_OPM00PI002-G.pdf. Accessed 22 Mar 2023.
White N, Hanan J. Use of functional-structural plant modelling in horticulture. Agri-science Queensland, Department of Agriculture, Fisheries and Forestry. 2012. Available online https://www.researchgate.net/publication/230877125_Use_of_Functional-Structural_Plant_Modelling_in_Horticulture. Accessed 22 Mar 2023.
Scher CL, Griffoul E, Cannon CH. Drone-based photogrammetry for the construction of high-resolution models of individual trees. Trees Struct Funct. 2019;33(5):1385–97. https://doi.org/10.1007/s00468-019-01866-x.
Iglhaut J, Cabo C, Puliti S, Piermattei L, O’Connor J, Rosette J. Structure from motion photogrammetry in forestry: a review. Curr For Rep. 2019;5(3):155–68.
Liang X, Kukko A, Balenović I, Saarinen N, Junttila S, Kankare V, et al. Close-Range Remote Sensing of Forests: The state of the art, challenges, and opportunities for systems and data acquisitions. IEEE Geoscience and Remote Sensing Magazine (GRSM). 2022;10(3):32–71. https://doi.org/10.1109/MGRS.2022.3168135.
Teng P, Zhang Y, Yamane T, Kogoshi M, Yoshida T, Ota T, et al. Accuracy evaluation and branch detection method of 3D modeling using backpack 3D Lidar SLAM and UAV-SfM for peach trees during the pruning period in winter. Remote Sens. 2023;15(2):408.
Culvenor DS, Newnham GJ, Mellor A, Sims NC, Haywood A. Automated in-situ laser scanner for monitoring forest leaf area index. Sensors. 2014;14(8):14994–5008.
Griebel A, Bennett LT, Culvenor DS, Newnham GJ, Arndt SK. Reliability and limitations of a novel terrestrial laser scanner for daily monitoring of forest canopy dynamics. Remote Sens Environ. 2015;166:205–13.
Portillo-Quintero C, Sanchez-Azofeifa A, Culvenor D. Using VEGNET in-situ monitoring LiDAR (IML) to capture dynamics of plant area index, structure and phenology in aspen parkland forests in Alberta, Canada. Forests. 2014;5(5):1053–68.
Kutila M, Pyykönen P, Holzhüter H, Colomb M, Duthon P. Automotive LiDAR performance verification in fog and rain. In: Proceedings of the 21st International conference on intelligent transportation systems (ITSC 2018). IEEE; 2018. pp. 1695–1701.
Wulder MA, White JC, Nelson RF, Næsset E, Ørka HO, Coops NC, et al. Lidar sampling for large-area forest characterization: a review. Remote Sens Environ. 2012;121:196–209.
Zhao Y, Ma Y, Quackenbush LJ, Zhen Z. Estimation of individual tree biomass in natural secondary forests based on ALS data and WorldView-3 imagery. Remote Sens. 2022;14(2):271.
Laes D, Reutebuch S, McGaughey R, Mitchell B. Guidelines to estimate forest inventory parameters from LiDAR and field plot data. Companion document to the Advanced LiDAR Applications-Forest Inventory Modeling class. US Forest Service, Salt Lake City, USA. 2011. Available online https://fsapps.nwcg.gov/gtac/CourseDownloads/Reimbursables/FY21/Lidar_Material/GTAC_Guidelines%20to%20estimate%20forest%20inventory%20parameters%20from%20lidar%20and%20field%20plot%20data.pdf. Accessed 9 Apr 2024.
Goodbody TR, Coops NC, Luther JE, Tompalski P, Mulverhill C, Frizzle C, et al. Airborne laser scanning for quantifying criteria and indicators of sustainable forest management in Canada. Can J For Res. 2021;51(7):972–85.
Hartley RJ, Davidson SJ, Watt MS, Massam PD, Aguilar-Arguello S, Melnik KO, et al. A Mixed methods approach for fuel characterisation in gorse (Ulex europaeus L.) scrub from high-density UAV laser scanning point clouds and semantic segmentation of UAV imagery. Remote Sens. 2022;14(19):4775.
Kellner JR, Armston J, Birrer M, Cushman K, Duncanson L, Eck C, et al. New opportunities for forest remote sensing through ultra-high-density drone lidar. Surv Geophys. 2019;40(4):959–77.
Hu T, Sun X, Su Y, Guan H, Sun Q, Kelly M, et al. Development and performance evaluation of a very low-cost UAV-LiDAR system for forestry applications. Remote Sens. 2020;13(1):77.
Hartley RJaL, Henderson IL, Jackson CL. BVLOS unmanned aircraft operations in forest environments. Drones. 2022;6(7):167.
Liang X, Kankare V, Hyyppä J, Wang Y, Kukko A, Haggrén H, et al. Terrestrial laser scanning in forest inventories. ISPRS J Photogramm Remote Sens. 2016;115:63–77.
Lovell J, Jupp DL, Culvenor D, Coops N. Using airborne and ground-based ranging lidar to measure canopy structure in Australian forests. Can J Remote Sens. 2003;29(5):607–22.
Hopkinson C, Chasmer L, Young-Pow C, Treitz P. Assessing forest metrics with a ground-based scanning lidar. Can J For Res. 2004;34(3):573–83.
Thies M, Pfeifer N, Winterhalder D, Gorte BG. Three-dimensional reconstruction of stems for assessment of taper, sweep and lean based on laser scanning of standing trees. Scand J For Res. 2004;19(6):571–81.
Watt PJ, Donoghue DNM. Measuring forest structure with terrestrial laser scanning. Int J Remote Sens. 2005;26(7):1437–46. https://doi.org/10.1080/01431160512331337961.
Pfeifer N, Gorte B, Winterhalder D. Automatic reconstruction of single trees from terrestrial laser scanner data. In: Proceedings of the 20th ISPRS Congress, Istanbul, Turkey. ISPRS; 2004;35(B5). pp. 114–9.
Torralba J, Carbonell-Rivera JP, Ruiz LÁ, Crespo-Peremarch P. Analyzing TLS scan distribution and point density for the estimation of forest stand structural parameters. Forests. 2022;13(12):2115.
Donager JJ, Sankey TT, Sankey JB, Sanchez Meador AJ, Springer AE, Bailey JD. Examining forest structure with terrestrial lidar: Suggestions and novel techniques based on comparisons between scanners and forest treatments. Earth Space Sci. 2018;5(11):753–76.
Berger M, Tagliasacchi A, Seversky LM, Alliez P, Guennebaud G, Levine JA, et al. A survey of surface reconstruction from point clouds. Computer Graphics Forum. 2017;36(1):301–29. https://doi.org/10.1111/cgf.12802.
Attene M, Campen M, Kobbelt L. Polygon mesh repairing: an application perspective. ACM Computing Surv (CSUR). 2013;45(2):1–33.
Fekry R, Yao W, Cao L, Shen X. Ground-based/UAV-LiDAR data fusion for quantitative structure modeling and tree parameter retrieval in subtropical planted forest. For Ecosyst. 2022;9:100065.
Yang M, Wan Y, Liu X, Xu J, Wei Z, Chen M, et al. Laser data based automatic recognition and maintenance of road markings from MLS system. Opt Laser Technol. 2018;107:192–203.
Balenović I, Liang X, Jurjević L, Hyyppä J, Seletković A, Kukko A. Hand-held personal laser scanning–current status and perspectives for forest inventory application. Croat J For Eng: J Theory Appl For Eng. 2021;42(1):165–83.
Chen S, Liu H, Feng Z, Shen C, Chen P. Applicability of personal laser scanning in forestry inventory. PLoS ONE. 2019;14(2):e0211392.
Ko C, Lee S, Yim J, Kim D, Kang J. Comparison of forest inventory methods at plot-level between a backpack personal laser scanning (BPLS) and conventional equipment in Jeju Island, South Korea. Forests. 2021;12(3):308.
Ruhan A, Du W, Ying H, Wei B, Shan Y, Dai H. Estimation of aboveground biomass of individual trees by backpack LiDAR based on parameter-optimized quantitative structural models (AdQSM). Forests. 2023;14(3):475.
O’Sullivan H, Raumonen P, Kaitaniemi P, Perttunen J, Sievänen R. Integrating terrestrial laser scanning with functional–structural plant models to investigate ecological and evolutionary processes of forest communities. Ann Bot. 2021;128(6):663–84.
Dobbs H, Batchelor O, Green R, Atlas J. Smart-Tree: Neural Medial axis approximation of point clouds for 3D tree skeletonization. In: Proceedings of the Iberian conference on pattern recognition and image analysis: Springer, Berlin; 2023. pp. 351–62. Available online https://arxiv.org/pdf/2303.11560.
Halupka K, Garnavi R, Moore S. Deep semantic instance segmentation of tree-like structures using synthetic data. In: Proceedings of the IEEE winter conference on applications of computer vision (WACV): IEEE; 2019. pp. 1713–22. Available online: https://arxiv.org/pdf/1811.03208.
Disney MI, Boni Vicari M, Burt A, Calders K, Lewis SL, Raumonen P, et al. Weighing trees with lasers: advances, challenges and opportunities. Interface Focus. 2018;8(2):20170048.
Fu L, Liu J, Zhou J, Zhang M, Lin Y. Tree skeletonization for raw point cloud exploiting cylindrical shape prior. IEEE Access. 2020;8:27327–41. https://doi.org/10.1109/ACCESS.2020.2971549.
Bucksch A, Lindenbergh R, Menenti M. SkelTre. Vis Comput. 2010;26(10):1283–300.
Lin Y, Wiegand K. Towards 3D tree spatial pattern analysis: Setting the cornerstone of LiDAR advancing 3D forest structural and spatial ecology. Int J Appl Earth Obs Geoinformation. 2021;103. https://doi.org/10.1016/j.jag.2021.102506.
Wu J, Cawse-Nicholson K, van Aardt J. 3D tree reconstruction from simulated small footprint waveform Lidar. Photogramm Eng Remote Sens. 2013;79(12):1147–57. https://doi.org/10.14358/PERS.79.12.1147.
Windrim L, Bryson M. Detection, segmentation, and model fitting of individual tree stems from airborne laser scanning of forests using deep learning. Remote Sens. 2020;12(9):1469.
Krisanski S, Taskhiri MS, Gonzalez Aracil S, Herries D, Muneri A, Gurung MB, et al. Forest structural complexity tool—an open source, fully-automated tool for measuring Forest point clouds. Remote Sens. 2021;13(22):4677.
Dijkstra EW. A note on two problems in connexion with graphs. Numerische Mathematik. 1959;1:269–271.
Li Y, Wang P, Sun J, Gan X. Simulation of tree point cloud based on the ray-tracing algorithm and three-dimensional tree model. Biosyst Eng. 2020;200:259–71. https://doi.org/10.1016/j.biosystemseng.2020.10.007.
Höfle B, Qu J, Winiwarter L, Weiser H, Zahs V, Schäfer J, et al. pytreedb: library for point clouds of tree vegetation objects. In: Zenodo, editor. 1.0.0 ed. GitHub [code]. https://doi.org/10.5281/zenodo.75513102023.
Weiser H, Schäfer J, Winiwarter L, Krašovec N, Fassnacht FE, Höfle B. Individual tree point clouds and tree measurements from multi-platform laser scanning in German forests. Earth Syst Sci Data. 2022;14(7):2989–3012.
Hyyppä E, Hyyppä J, Hakala T, Kukko A, Wulder MA, White JC, et al. Under-canopy UAV laser scanning for accurate forest field measurements. ISPRS J Photogramm Remote Sens. 2020;164:41–60.
Hyyppä J, Yu X, Hakala T, Kaartinen H, Kukko A, Hyyti H, et al. Under-canopy UAV laser scanning providing canopy height and stem volume accurately. Forests. 2021;12(7):856.
Wang Y, Kukko A, Hyyppä E, Hakala T, Pyörälä J, Lehtomäki M, et al. Seamless integration of above-and under-canopy unmanned aerial vehicle laser scanning for forest investigation. For Ecosyst. 2021;8:1–15.
Hackenberg J, Calders K, Demol M, Raumonen P, Piboule A, Disney M. SimpleForest-a comprehensive tool for 3d reconstruction of trees from forest plot point clouds. BioRxiv; 2021. pp. 1–25. https://doi.org/10.1101/2021.07.29.454344.
Douglas ES, Martel J, Li Z, Howe G, Hewawasam K, Marshall RA, et al. Finding leaves in the forest: the dual-wavelength Echidna lidar. IEEE Geosci Remote Sens Lett. 2014;12(4):776–80.
Mark Danson F, Sasse F, Schofield LA. Spectral and spatial information from a novel dual-wavelength full-waveform terrestrial laser scanner for forest ecology. Interface Focus. 2018;8(2):20170049.
Li D, Wang C, Jiang H, Peng Z, Yang J, Su Y, et al. Monitoring litchi canopy foliar phosphorus content using hyperspectral data. Comput Electron Agric. 2018;154:176–86.
Wang Z, Chen Y, Li C, Tian M, Zhou M, He W, et al. A hyperspectral LiDAR with eight channels covering from VIS to SWIR. In: Proceedings of the IEEE international geoscience and remote sensing symposium (IGARSS 2018): IEEE; 2018. pp. 4293–6. https://doi.org/10.1109/IGARSS.2018.8517741.
Chen Y, Li W, Hyyppä J, Wang N, Jiang C, Meng F, et al. A 10-nm spectral resolution hyperspectral LiDAR system based on an acousto-optic tunable filter. Sensors. 2019;19(7):1620.
Axelsson A, Lindberg E, Olsson H. Exploring multispectral ALS data for tree species classification. Remote Sens. 2018;10(2):183.
Dalponte M, Ene LT, Gobakken T, Næsset E, Gianelle D. Predicting selected forest stand characteristics with multispectral ALS data. Remote Sens. 2018;10(4):586.
Xu X, Iuricich F, Calders K, Armston J, De Floriani L. Topology-based individual tree segmentation for automated processing of terrestrial laser scanning point clouds. Int J Appl Earth Obs Geoinformation. 2023;116:103145.
Hou K, Chio S. Plane-based range calibration method for geoslam zeb-horizon handheld lidar instrument. In: Proceedings of the international symposium on remote sensing (ISRS 2021). ISRS virtual conference; 2021. pp. 26–8.
Martin-Ducup O, Lecigne B. ARchi: quantitative structural model (‘QSM’) treatment for tree architecture. 2022. Available online: https://CRAN.R-Project.Org/Package=aRchi. Accessed 12 Apr 2024.
Computree Core Team. Computree platform. 2024. https://computree.onf.fr: Computree group. Accessed 12 Apr 2024.
Schlegel P. Skeletor. 2018. Available online: https://github.com/schlegelp/skeletor. Accessed 12 Apr 2024.
Boudon F. PlantScan3D. 2014. Available online: https://github.com/fredboudon/plantscan3d. Accessed 12 Apr 2024.
Van Rossum G. Python programming language. In: USENIX annual technical conference: Santa Clara, CA: USENIX; 2007. pp. 1–36.
Cabo C, Laino D. 3D Forest Inventory (3DFIN). 2023. Available online: https://github.com/3DFin/3DFin. Accessed 12 Apr 2024.
QGIS Development Team. QGIS geographic information system. 2024. Available online: https://www.qgis.org. QGIS Association. Accessed 12 Apr 2024.
Girardeau-Montaut D. CloudCompare. Available online: https://cloudcompare.org/2024. Accessed 12 Apr 2024.
Acknowledgements
The authors would like to express our sincere gratitude to the three anonymous reviewers for their constructive feedback and valuable suggestions that helped make some significant improvements to this manuscript. We would also like to thank Dr Neil White and the Queensland Government Department of Agriculture and Fisheries for providing the image for Figure 2a and permission to reproduce it in this review.
Funding
Open Access funding enabled and organized by CAUL and its Member Institutions The authors would like to acknowledge Scion for funding this research with Strategic Science Investment Funding (SSIF) provided by the Ministry of Business Innovation and Employment (MBIE). Additional co-funding was provided by the Forest Growers Levy Trust through the Resilient Forests research programme and the MBIE Transforming Tree Phenotyping programme (C04X2101). The lead author would also like to extend his gratitude to Scion for providing additional financial support.
Author information
Authors and Affiliations
Contributions
R.J.L.H. was responsible for the conception of this review, as well as carrying out all analysis and writing of the original draft. Review and editing were carried out by all other authors.
Corresponding author
Ethics declarations
Conflict of Interest
NZ Ministry of Business Innovation and Employment (MBIE) |Strategic Science Investment Funding (SSIF) funded the hours for conducting the literature review. The Forest Growers Levy Trust provided additional co-funding of hours for the review through the Resilient Forests research programme. MBIE provided some additional hours for working on this paper through the Transforming Tree Phenotyping research programme (C04X2101).
Human and Animal Rights and Informed Consent
This article does not contain any studies with human or animal subjects performed by any of the authors.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Hartley, R.J.L., Jayathunga, S., Morgenroth, J. et al. Tree Branch Characterisation from Point Clouds: a Comprehensive Review. Curr. For. Rep. (2024). https://doi.org/10.1007/s40725-024-00225-5
Accepted:
Published:
DOI: https://doi.org/10.1007/s40725-024-00225-5