Abstract
In this paper, rather than focusing on genes as an organising concept around which historical considerations of theory and practice in genetics are elucidated, we place genetic markers at the heart of our analysis. This reflects their central role in the subject of our account, livestock genetics concerning the domesticated pig, Sus scrofa. We define a genetic marker as a (usually material) element existing in different forms in the genome, that can be identified and mapped using a variety (and often combination) of quantitative, classical and molecular genetic techniques. The conjugation of pig genome researchers around the common object of the marker from the early-1990s allowed the distinctive theories and approaches of quantitative and molecular genetics concerning the size and distribution of gene effects to align (but never fully integrate) in projects to populate genome maps. Critical to this was the nature of markers as ontologically inert, internally heterogeneous and relational. Though genes as an organising and categorising principle remained important, the particular concatenation of limitations, opportunities, and intended research goals of the pig genetics community, meant that a progressively stronger focus on the identification and mapping of markers rather than genes per se became a hallmark of the community. We therefore detail a different way of doing genetics to more gene-centred accounts. By doing so, we reveal the presence of practices, concepts and communities that would otherwise be hidden.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
“One of my colleagues complained I was a geneticist who didn’t study genes”
Michael Goddard, animal geneticist and pioneer of genomic selection, 2018Footnote 1
1 Introduction
In this paper, we argue that to comprehend the practices of genetics research, one must supplement accounts centred on genes with ones that focus on genetic markers. We highlight the salience of genetic markers using the example of one area of their application, namely livestock genetics concerning the domesticated pig, Sus scrofa.Footnote 2 In that field, due to its particular concatenation of limitations, opportunities, collaborations and intended research goals, the mapping and use of markers is a dominant practice, and not exclusively or even primarily performed for the eventual discovery, investigation and use of genes.
We define a genetic marker as a (usually material) element existing in different forms in the genome, that can be identified and mapped using a variety (and often combination) of quantitative, classical and molecular genetic techniques. One of the more concise and inclusive definitions of genetic markers is that they are “specific DNA sequences with a known location on a chromosome” (Benavides and Guénet 2012, p. 65).
In historical and philosophical literature on genetics, the gene is paramount. In its various formulations from the early twentieth-century onwards, the gene concept “became central to all main branches of the life sciences and promoted unprecedented visions of controlling and directing life.” It was “something akin to the organizing principle of twentieth-century biology” (Rheinberger and Müller-Wille 2017, p. 1). From the 1970s, the ability of scientists to cut, paste and edit genes with measurable effects formed the basis of a new industrial sector: biotechnology (Rasmussen 2014; Yi 2015).
Understood either abstractly (as in classical genetics) or materially (as in post-1960s molecular genetics), genes are at the heart of debates over the nature of heredity, causal attributions concerning the development and functioning of organisms, and the nature of evolution. Indeed, the gene has been a central focus of historical and philosophical accounts that attempt to contest, complicate, deflate or historically-situate our metaphysical, epistemic and methodological understandings of the role of genetics in processes of organismal development and functioning. Much of this gene-centred work has convoluted, even fragmented, what we understand genes to be, and their role and nature in living organisms (e.g. Beurton et al. 2000; Fox Keller 2000; Griffiths and Stotz 2013; Moss 2003; Rheinberger and Müller-Wille 2017).
Previous historical work has discussed the use of markers in particular research programmes. Kaufmann (2004) detailed the work of the generation and mapping of markers at the Généthon genomics centre in Évry, France. Rajagopalan and Fujimura (2018) have traced the advent of single nucleotide polymorphisms (SNPs) as markers and their adoption in human genome mapping and later in studies of human genetic diversity. This research into the genetic variation in and of human populations used and contributed to the development of microarrays known as SNP chips. These were used in disease studies, but also could be used to map difference—variation—“as an end in itself” in the context of population genetics.
A variety of different kinds of markers have been used to indicate the existence of genes, and elucidate both the abstract and physical relationships between these genes. These markers included phenotypic indicators such as eye colour, detectable biochemicals that might be the products of metabolic processes in which a given gene or genes may be involved, and, the subject of this paper, genetic markers (Falk 2009). In medical genetics, markers were identified and mapped in the pursuit of genes from the 1980s, with marker-oriented practices not constituting an end in themselves (Harper 2008; Lindee 2005).Footnote 3 Gene-centric genetics may involve practices concerning markers, but these are subordinate to an agenda focused on genes.
Henceforth, we refer to genetic markers as ‘markers’; references to non-genetic markers will be specifically indicated (see Fig. 1 for how genetic markers relate to other types). In this paper, we explore the formulation of the marker as a common object from the outset of systematic projects to map pig genomes in the early-1990s. Before the 1980s, livestock genetics was dominated by quantitative approaches. From the 1980s, molecular biological methods became more prominent. The conjugation around the common object of the marker allowed the distinctive theories and approaches of quantitative and molecular genetics concerning the size and distribution of gene effects to align (but never fully integrate) in projects to populate genome maps with markers derived from different techniques. Though the mapping was intended to be used to enable more precise selective breeding, the existence and position of the markers themselves were not presumed to have any direct functional implications.
Diagram depicting the nested relationships concerning the overall category of markers, genetic markers as a subset of these, and within genetic markers, genetic markers that happen to also be genes. Note that the diagram is not intended to depict the relative size or scale of any of these categories
The pig genetics community can conduct interventions in populations for which they can control the breeding and, therefore, construct families and make particular crosses; options not open to the human geneticist (see Müller-Wille 2018 on the significance for scientific classificatory and data practices of being able or unable to construct populations). Also, as the economics of livestock breeding makes interventions in populations through breeding practices more viable than treating individuals, livestock geneticists do not have to find particular causative mutations that could form the basis, for instance, of pharmacotherapeutic interventions (as many human geneticists do).
Although they do not have to, livestock geneticists can therefore ignore the mechanistic underpinnings of aspects of phenotypic variation, in favour of statistical associations based on the genotypic and phenotypic data of particular populations. Thus, when a search for candidate genes exhibiting relatively large effects was stymied by the polygenic nature of variation, the transition towards a statistical quantitative marker-based approach was easy to make.
The marker as an object was valued because of its ontological inertness, its internal heterogeneity and its relational nature. Markers are ontologically inert because there are no necessary presumed structural or functional consequences dependent upon any one individual marker. They are internally heterogeneous, as the material nature of markers need not be of the same kind or congruent with other markers for them to be deemed comparable, equivalent or useful.
They are relational in nature, as the value of an individual marker relies on its relations to other markers, and associations drawn with variation in phenotypic properties of interest. To be an indicator for breeding purposes, a marker simply needs to be a better indicator than another one, or a set of them need to be better than another set.Footnote 4 Any relations or associations that are discerned will always be relative to a reference population from which such relations and associations were drawn. This implies nothing intrinsic about the nature or properties of the marker itself. The marker’s inertness, heterogeneity and relational nature allows them to perform the functions presented in this paper, revealed through a marker-centric lens.
2 Before systematic mapping projects
The disciplinary traditions of quantitative and molecular genetics that interacted in pig genome mapping projects from the 1990s onwards have distinct histories that have consequently been dealt with separately in the scholarly literature. Here we outline some of the background of these traditions, focusing in particular on their characteristic models of gene effects.
2.1 Quantitative genetics
Quantitative genetics traces its roots to the early-twentieth-century work of Ronald Aylmer Fisher (1890–1962), John Burdon Sanderson Haldane (1892–1964) and Sewall Wright (1889–1988), each of whom contributed towards the reconciliation of Mendelian genetics and biometry (Provine 1971). The Mendelian conception of heredity centred on the transmission of discrete units—genes—that could be discerned in the genotype through observed differences in the phenotype (Johannsen 1909). Biometry focused instead on continuous variation, and statistical analysis of variants of small-effect. Though there were important differences between their approaches, Fisher, Haldane and Wright’s fundamental insight was that continuous variation could be explained with reference to a multitude of Mendelian factors, each of which may only contribute a small part to the overall observed variance. This was used to explain how variation is conserved across generations, and helped to inaugurate population genetics (Provine 1971).
Wright had an interest in animal breeding; he had worked at the United States Department of Agriculture (USDA) facility at Beltsville, Maryland from 1915 to 1925. While there, he worked with livestock breeders, responded to queries from farmers, worked on problems concerning quantitative analysis posed by other researchers at the facility and devised an ancestry-based evaluation scheme to help calculate the market value of quarantined cattle (Russell 1989, pp. 5–6).
Jay Lush (1896–1982), who had worked in animal breeding at the University of Texas and was based at Iowa State College from 1930, attended Wright’s course on statistical genetics at the University of Chicago in 1931. He also absorbed Fisher’s insights when the latter lectured at Iowa State over the summers of 1931–1936. Lush was concerned almost wholly with developing the quantitative basis for informing the selection of livestock animals for breeding, to improve upon mass selection based largely on phenotypes (Hill 2014). He published voluminously, included the highly influential Animal Breeding Plans (in successive editions published in 1937, 1943 and 1945), and was recognised for his teaching and training of many future leaders of livestock genetics. The programme of research begun in Iowa soon became established in other centres worldwide.
One of the key underpinnings of quantitative genetics as applied to livestock breeding is the infinitesimal model, which was originally formulated by Fisher in 1918.Footnote 5 Simplified, the infinitesimal model posits “an effectively infinite number of unlinked loci with infinitesimal effects” (Bulmer 1980, p. 150). The power of the model does not derive from any congruence to a supposed underlying reality, but because “it has very powerful simplifying statistical properties and avoids the need to specify individual gene effects, information on which has until recently been impossible to obtain” (Hill 2014, p. 4).
Quantitative genetic approaches to livestock breeding have been produced by researchers attentive to the needs and problems of breeders: namely, the creation of methods to index animals, calculate accurate Estimated Breeding Values and conduct selection experiments to test theory.
2.2 Classical and molecular approaches to genetics
Apart from population and quantitative genetics, a distinct Mendelian tradition remained, however, instantiated in the ‘classical genetics’ founded in the work of Thomas Hunt Morgan’s laboratory at Columbia University in New York from the 1910s to the late-1920s. Working with the fruit flies Drosophila melanogaster, researchers in this laboratory identified particular mutants: flies with different characteristics to the presumed norm or ‘wild type’, for example different eye colours or wing morphologies.Footnote 6 They presumed that genic differences underlay these various mutations. In millions of flies, they measured the frequencies with which particular mutations manifested together. The laboratory then used the data on these associations between mutations—the linkages—to map the genes, which were presumed to be stable entities that were transmitted from one generation to the next (Waters 1994).Footnote 7
This linkage mapping is based on the fact that during meiosis in the production of gametes (sex cells: sperm and ova), parts of chromosomes break and then recombine. Sometimes when they do this, part of one pair of chromosomes (the part being a chromatid) can become part of the other pair. As any two loci (specific places on a chromosome) that are closer together are more likely to remain on the same chromatid than two loci that are further apart, measurements of the co-location of loci can be used to conduct a linkage analysis that uses the presence or absence of particular loci to estimate the relative positions of the loci on the chromosome (Kohler 1994).
From the 1950s, the determination of the structure of DNA helped to establish it as the material basis of genes. From this foundation, research established how DNA is transcribed and translated to produce particular proteins with specific amino acid compositions, with the aid of intermediary molecules such as messenger RNA and transfer RNA. On the basis of this work, the gene was reconceptualised from the classical conception of a unit of distant control over some aspect of the phenotype, to a conception in which the gene consists of information for the precise specification of the structure of a protein (Morange 2000).Footnote 8
While further research into the nature of the ‘molecular gene’ and its relationship to observable structure and function rapidly complicated simple models of gene expression and function, for instance through the recognition of complex webs of gene regulation and interaction (Rheinberger and Müller-Wille 2017), the molecular approach to genetics still focused on the identification and characterisation of genes. Whereas classical genetics inferred the presence of genes and different versions of them (alleles) from the existence of phenotypic variants, variants of molecular genes were associated with variation in the structure and function of macromolecules, which provide links from genetic variation to observable phenotypic variation.
This approach held the promise of prospective intervention in biological processes and mechanisms to specifically alter phenotypic outcomes. This promoted the search for genes (and variants and mutants thereof) that exhibit relatively large effects.
From the 1950s, molecular biologists and biochemists had furnished an increasing number of different kinds of biomarker, from which the presence or absence of particular genes (and alleles thereof) could be inferred. Many of these were assayed in the blood, and were presumed to be macromolecular products of these genes. Linkage could be inferred by the combinations of presence or absence of particular biomarkers, and perhaps also phenotypic indicators, as in the origins of classical genetics.
In the context of medical genetics, identifying and mapping the actual gene presumed to play a role in a physiological or pathological process of interest was of paramount importance. This still required markers to be identified, however. Research on the Human Leucocyte Antigen (HLA) system is an instructive example. The genes in this system, implicated in immune response among other functions, are densely packed and highly polymorphic. As antibodies could be used to assay the presence or absence of particular antigenic molecules coded for by specific variants of HLA genes, this region was ripe for mapping genes using the antigens as markers. Mapping of disease-related genes proceeded into the 1980s through the development of genetic markers linked to prospective genes (Harper 2008, pp. 200–211). Some of the techniques by which these markers were produced are discussed in the next section.
In the context of livestock genetics, the inference that, for a given trait of interest, a gene (or version thereof) must exist in a certain part of a given chromosome, need not culminate in the precise identification and localisation of that gene. As a therapeutic or corrective intervention into the individual animal is not necessarily the aim, it is sufficient to identify and map the genetic loci of interest, to be able to confirm that particular markers were sufficiently close to the locus of interest to serve as a proxy, so that a test could be developed to detect the marker when genotyping animals. This relied on the classical genetical insights into linkage and recombination.
2.3 Investigating halothane sensitivity: towards a candidate gene strategy
This was the approach taken by the geneticists who worked on the Halothane sensitivity locus in the 1980s. Halothane is a veterinary anaesthetic that had been identified as a test for Porcine Stress Syndrome (PSS). Pigs affected by PSS died suddenly when exposed to stress, and the meat from these pigs was of poor quality, and therefore unappealing to consumers. Prior to the full elucidation of the genetic basis of the syndrome, attempts were made to identify pre-symptomatic pigs by their response to exposure to halothane, responders becoming rigid within 3 min (Webb 1980). However, as the condition had been theorised to be recessive, requiring two copies of the version (allele) of the gene involved (Ollivier et al. 1975), only homozygous pigs carrying two copies would be symptomatic, and thus exposed by this test. Heterozygous pigs who carry one copy of the halothane-sensitivity allele would be undetected and therefore would not necessarily be excluded from breeding, enabling the lesion to be transmitted to the next generation.
This, combined with the time-consuming nature of testing herds in this way, meant that alternative tests to identify carriers as well as symptomatic pigs were sought. As with the human medical geneticists who in the mid-1980s attempted to identify and map disease genes (Harper 2008; Lindee 2005), it became a pressing matter for the livestock and breeding industries to identify, localise and characterise the locus or loci associated with PSS.Footnote 9 Then, using linked markers, animals could be genotyped with a view to removing the rogue allele from breeding populations.
The relationship of this locus with nearby loci was therefore of interest to geneticists. In the UK, Alan Archibald of the Animal Breeding Research Organisation (ABRO; one of the main institutional antecedents of the Roslin Institute: see García-Sancho 2015; Myelnikov 2017; Lowe, in review) obtained a research commission from the Ministry of Agriculture, Fisheries and Food to investigate the genetics of PSS, and in particular the group of linked loci it was deemed to be associated with. In a volume of the journal Animal Genetics edited by Archibald and Pat Imlah (of the Royal Dick School of Veterinary Studies, University of Edinburgh), papers included research on the order of the linked loci, the use of them in genotyping animals, and rates of recombination between the loci (Archibald and Imlah 1985). Data on recombination events between loci supported a consensus order of the loci, but the identity of the halothane locus was still unknown.
By 1986, Archibald recognised the potential importance of genetic markers, markers representing some section of DNA rather than a molecule circulating in the blood. He cited the use of Restriction Fragment Length Polymorphisms (RFLPs) to isolate and characterise a gene implicated in Duchenne muscular dystrophy (Archibald 1986). RFLPs were developed for use in human genetics. Linkage analyses require detectable markers of polymorphisms or discrete variants, which had been in short supply beyond serological tests for genes in the dense and polymorphic HLA region. RFLPs use restriction enzymes to cut sections of DNA; the resulting fragments are then separated by length using gel electrophoresis. The principle behind RFLP techniques is that variation or polymorphisms result in cuts being made at different places, and therefore the DNA fragments will end up in different places in the gel; on the membrane to which it is transferred; and in the resulting autoradiograph. The basis for using RFLPs was established in 1974 and 1975. In 1980, the basis for using RFLPs to construct a human genetic linkage map was outlined (Botstein et al. 1980).
From the very beginning, it was recognised that RFLPs need not include a gene. If an RFLP was linked to a gene or genes, however, this created the prospect that genes may be mapped—or tracked—without isolating or characterising the gene itself (Botstein et al. 1980, p. 317). In 1988, a gene linked to the halothane locus was mapped to a particular location on swine chromosome 6 using cDNA (complementary DNA) probes obtained from a pig genomic library (Davies et al. 1988). Following this and research on malignant hyperthermia, the human manifestation of the trait associated with the halothane locus in pigs, a candidate gene was proposed. In 1990, the malignant hyperthermia locus was mapped to a location on human chromosome 19 by inference from the linkage group established by the 1988 research on pigs to the corresponding area of the human genome. This location coincided with the position where the ryanodine receptor gene (RYR) had recently been localised (McCarthy et al. 1990).
In the same issue of Nature where this research appeared, David MacLennan and colleagues reported the results of a linkage analysis that they had conducted on families containing individuals deemed to be at risk from malignant hyperthermia. They analysed seven loci, including RYR, using seven different cDNA probes, and applied ten different RFLP enzymes to chromosome 19 DNA obtained from the family members. When they analysed the results in terms of recombination (and, by implication, co-segregation) frequencies, they found strong linkage between the malignant hyperthermia trait and RYR. This, combined with the other evidence already indicated, led them to conclude that RYR must be considered to be a candidate gene implicated in malignant hyperthermia in humans, and PSS in pigs (MacLennan et al. 1990).
It proved to be so. With the gene, and the particular mutation (RYR1) responsible for PSS identified (Fujii et al. 1991), genotyping tests could be developed to detect it in individuals (Dalens and Runavot 1993).Footnote 10 This offered breeders the opportunity to manage the responsible gene, and potentially to eradicate it (Fujii et al. 1991; O’Brien and Ball 2013; Otsu et al. 1991).
Within the space of a decade, linkage mapping had progressed to the identification and characterisation of a single gene responsible for a condition that was causing considerable losses to the livestock industry. This provided the impetus for two linked strategies. One was the identification of candidate genes related to production traits of interest, and the development methods to genotype them. The other was a more comprehensive identification and mapping of markers in the genome, deemed important in part to aid in the mapping of quantitative trait loci (QTL), locations in the genome associated with variation in traits of interest. Animals could then be genotyped using nearby markers, and therefore the livestock geneticists did not need to methodologically orient themselves towards genes.
The latter strategy underpinned support for major projects to populate pig chromosomes with maps of ordered and, in some cases, physically localised or anchored markers, as will be discussed in the next section. Major projects based on sets of reference families included PiGMaP, funded by the European Commission (1991–1996), one led by the United States Department of Agriculture Agricultural Research Service Meat Animal Research Center (USDA-ARS MARC), and a Nordic collaboration.
3 Candidate genes and markers
The aim of genome mappers in the pig genetics community was not to produce a comprehensive genome map, as it was with the Human Genome Project. Nor was it to derive a partial map through cDNA (as with the UK’s Human Genome Mapping Project), or to primarily hunt genes. Instead, the aim was to populate chromosomes with sufficiently spaced-out markers to permit the further addition of markers and assist with the locating of QTL, with progressively greater resolution between flanking markers, using statistical and computational tools developed within the community for that purpose. Estimates in the early-1990s indicate that the mappers were starting almost from scratch: 31 markers assigned to loci in 1990 (Schook et al. 1990), and 42 in 1991 (O’Brien 1991).
3.1 RFLPs and other techniques for mapping markers
PiGMaP began with two parallel mapping approaches. The first was to detect RFLP loci using homologous (mainly cDNA) probes derived from pigs and heterologous (again mainly cDNA) probes derived from humans and other non-porcine mammals. This work was done on crosses of distinct pig breeds with known pedigrees. Crossing distinct breeds helped to generate polymorphisms that could be used in linkage analyses (Archibald et al. 1995). The second approach was physical mapping, the assignment of genes to chromosomes using techniques of in situ hybridisation, first using radioactive probes, then increasingly moving to Fluorescent In Situ Hybridisation (FISH). The technical challenge presented by the development and mapping of markers meant, according to Chris Haley, a leading researcher and organiser in PiGMaP, that “there was benefit in different groups developing different markers and validating them using both physical and linkage mapping approaches.”Footnote 11
Increasingly, the number of markers used in linkage mapping (also known as genetic mapping) became dominated by hypervariable markers: first minisatellites, and then microsatellites. These are based on areas of the genome featuring repetitive sequences of DNA: particular patterns or ‘motifs’ that are repeated a number of times over. They are not genes. The difference between minisatellites and microsatellites is in the length of the motifs in base pairs, typically the former from ten to dozens of base pairs, the latter ranging up to ten. There are many more microsatellites than minisatellites in the genome, and microsatellites are more widely distributed across chromosomes. Consequently, microsatellites came to be of more importance to mapping and other applications.
The different number of repeats contained in microsatellites is valuable for linkage analysis due to the large number of alleles it may exhibit as a result, leading often to differences between individuals and heterozygosity within an individual (Gulcher 2012). This variation leads to these markers having a high polymorphic information content. It was in 1989 that the potential use of “simple sequence length polymorphisms” as polymorphic markers was recognised (Tautz 1989), and that the relatively new Polymerase Chain Reaction (PCR) could be used to amplify these variable repeats, using specially-designed custom primers (Weber and May 1989). This could then be used as a basis for linkage analysis that would be less complicated than the RFLP approach, involving fewer stages, and be faster, cheaper and more sensitive (Weber 1990).
Microsatellites, as previously noted, are found all over the genome. They disproportionately tend to be in noncoding regions, however, though the exact patterns of distribution beyond that varies among different kinds of noncoding region, and across different taxa (Tóth et al. 2000). There is, therefore, no presumed functional correlation between the presence, absence or variant of a microsatellite and any variation in a phenotypic trait.
The markers used to populate pig genome maps derived from different techniques and methodologies (see Tables 1, 2). As such, the identification, interrelation and integration of markers onto maps did not require any prior theoretical conception of the relation of the actual mapped markers to any variation in phenotypic effect: they were ontologically inert and internally heterogeneous. The use of the concept of the ‘marker’ in genetics has enabled the integration of data of distinct and independent provenance, generated against different theoretical, disciplinary and methodological backgrounds.
There is a striking contrast here with human genetics, where in the context of searching for disease-related genes, the shift from using RFLPs and microsatellites towards identifying SNPs was associated with a switch in focus from family-based genetic linkage research towards studying populations comprised of unrelated people (Rajagopalan and Fujimura 2018). Mapping markers in the service of identifying genes, a functional interest, therefore implied that the use of different kinds of markers were associated with fundamental changes in practice. In the case of pig genomics, the common project of the quantitative and molecular geneticists enabled these different kinds of markers to instead work for and with the mapping practices that they wanted to pursue, rather than the material or technical nature of the marker directing efforts towards one particular approach.
As genetic and physical mapping continued into the mid-1990s, the methodology and tools to identify and map QTL were also developed. The existence of QTL are posited by quantitative genetic theory, with a multitude of loci expected to each contribute a small degree to the overall variance in a trait. In the second iteration of the PiGMaP project (1994–1996), five different centres across Europe conducted genotyping using panels of 100 microsatellite markers derived from those mapped in PiGMaP and at USDA-ARS MARC. They also recorded phenotypic data on 2945 animals, each with known pedigrees from the third generation of reference populations. Several QTL bearing on livestock production traits, such as growth, fatness and meat quality, were detected as a result.Footnote 12 Chris Haley has observed that “building maps using different reference populations allowed a unified map to be produced and the homogeneity of that map between populations to be confirmed.”Footnote 13
Work to identify genes continued parallel to this. This included some groups who were involved in the more marker-centric work. Genes were mapped by a variety of methods, including linkage analysis, in situ hybridization and somatic cell hybrids (which will be discussed below). Candidate genes could be inferred in much the same way as RYR was for malignant hyperthermia; an alternative strategy was to discern ‘positional candidate genes’ through a presumed association with a known QTL. The existence of a gene could be inferred by the presence of a nearby QTL that had exhibited some association with variation in a given phenotypic metric. In some cases, this resulted in the identification of genes and mutated versions of them that were causally associated with observed phenotypic variations, for example IGF2 (insulin-like growth factor 2; Jeon et al. 1999; Nezer et al. 1999), which is associated with leanness, and ESR (estrogen receptor gene; Legault et al. 1996), associated with increased litter size.
The expectation that this positional candidate gene strategy would lead to a detailed mechanistic understanding of the genes and pathways involved in various traits, and that this would form the basis of genetic change in livestock, did not, however, come to pass. In academia, Max Rothschild was a prominent exponent of this strategy, and was involved in discovering genes, including ESR, which formed the basis of patents exclusively licenced to a pig breeding company, the Pig Improvement Company (PIC; Rothschild and Plastow 2002). By 2007, having commented on the range of QTL so far identified, he and his co-authors confided that, “[p]erhaps more interesting is the fact that only a limited number of these QTL have been further investigated to the point that a known causative mutation has been implicated or proven” (Rothschild et al. 2007).
The ESR research and application thereof was actually an example of Marker-Assisted Selection (MAS): the patents were for genetic markers indicative of polymorphisms of the gene, and methods for identifying these kinds of markers, rather than for the gene or any presumed causative mutation itself (Rothschild and Plastow 2002). The research and its putative application could therefore live without genes. In the context of practices already centred on the identification and mapping of markers, and the continued presence of quantitative genetics approaches, gene-hunting diversions could be readily re-routed towards a marker-centred approach amenable to quantitative genetic methods. This was made possible by the intersection of quantitative and molecular approaches. A marker-centred historiographical lens allows us to discern this dynamic in the way that a gene-centred approach, which might instead interpret the effort to find and use candidate genes as a failure, may not.
The pursuance of MAS was a key part of the strategy of PIC, a company with close links to researchers at publicly-funded research institutions. To aid in the development and implementation of MAS, as well as alternative approaches favoured by other companies, the livestock genetics community continued to find new ways of identifying and mapping markers. They also sought to develop the methods to make direct use of those markers in selective breeding, rather than just using them to identify genes. This was, for instance, a key difference with initiatives such as the Human Genome Mapping Project or the Human Genome Project, in large part because of differences in intended translational goals.
Chris Haley has attested that the unified concept of the marker, with the actual processes of detection and the exact material differences between different kinds of marker abstracted away, was key to helping people from disparate disciplinary and research traditions to work together.Footnote 14 The different markers became equivalent units, amenable to—and using the methods of—both quantitative genetics and molecular genetics. This was because the identification and localisation of markers did not require them to be imbued with the kinds of theoretical connotations, for instance concerning the nature and distribution of gene effects, that would otherwise prevent these two approaches from adopting a common orientation to these objects. The marker therefore enabled different communities to contribute towards the common task of mapping. It also functioned as a unit with which comparisons with other maps of the pig genome, and the genomes of other species, could be made.
3.2 Markers as a basis for comparison
To take advantage of the resources available from better-resourced communities working on human genetics, pig geneticists were keen to identify correspondences between the human and pig genomes. Recognising homologous regions of the respective genomes of human and pig could aid in the identification of candidate genes, as with RYR, and also in selecting which probes to use from human sources to aid the mapping of markers in the pig genome. The use of human genome data did not mean that the purposes of its use would be the same. For instance, for pig geneticists the search for markers would not necessarily be a precursor to the localisation and characterisation of genes, as was conceived in human genome research.
Comparative mapping was a key strategy of pig genomics from its inception. It required initial marker and mapping data to be able to begin to ascertain the comparative relationships on which inferences could be built. Rather than simply providing an initial skeleton on which pig-specific data would provide the flesh, it opened up an ongoing iterative process, once sufficient development of maps enabled comparisons of correspondences between pig and human genomes to be drawn. The identification of ever more precise patterns of correspondence (which included the creation of comparative maps), enabled maps to be populated with progressively more markers. This, in turn, enabled the further elaboration of relations of homology between areas of the pig and human genomes.
An example of this derives from some of the early work undertaken using labelled probes derived from human DNA libraries, then called heterologous painting or chromosome painting, now called Zoo-FISH. The probes were derived from mapped areas of the human genome, and would fluoresce when hybridised to corresponding pig DNA, providing a signal when photographed. Human genome data were also used to identify potential primers to amplify markers in particular regions of the pig’s genome; markers do not simply present themselves, but require some means of teasing them out from the genome.
These methods enabled researchers to determine areas of synteny—conserved order between chromosomes, in this case between pig and human chromosomes—and the pig chromosomes could then be sorted and identified. As well as this, a further refinement of the inferential techniques was advanced by the finding that conserved synteny did not necessarily imply conservation of gene order on the chromosome (Rettenberger et al. 1995). Markers were therefore used as indicators to assess homology relationships.Footnote 15
Synteny and the relative positions of markers were also detected through methods based on the hybridisation of porcine cells (and the DNA contained within) with those of more genetically well-characterised species such as mice, rats and hamsters: somatic cell hybrid mapping. The basic principle was to use the presence of known markers from better genetically-characterised species to identify areas of synteny and linkage, the latter based on measurements of the co-retention of markers.
One example of this, which became an important mapping method and tool from the late-1990s, was radiation hybrid mapping. In radiation hybrid mapping, cells are irradiated, and then fused with recipient cells of another species. The irradiation breaks apart the DNA, fragments of which then hybridise to corresponding parts of the recipient cell’s chromosomes. The presence or absence of particular markers can then be ascertained and measured, and used to physically map markers to particular parts of the genome, and discern linkage relationships through their co-retention (or otherwise) on individual fragments. The higher the initial dose of radiation, the smaller the fragments of DNA, and therefore the higher resolution the mapping.
The first whole genome radiation hybrid panel for mapping (the IMpRH panel) was developed in 1998 through a collaboration between teams at the Institut National de la Recherche Agronomique (INRA) institute at Castanet-Tolosan near Toulouse and at the University of Minnesota in the USA (Yerle et al. 1998). The same collaborators used the IMpRH panel to produce a map consisting of 757 markers in 128 linkage groups. They noted that this method was not as reliant upon highly polymorphic markers as previous maps, and that the search for QTL would be progressed by further “increasing the amount of comparative mapping information between the swine and human expression maps”, as well as using genome libraries to isolate new markers of interest, once the genomic region of interest is identified in this way (Hawken et al. 1999).
3.3 Using markers
If a marker is identifiable by some test, and that marker is indeed linked (technically, in ‘linkage disequilibrium’) with a gene that may be implicated in variation in a trait of interest, in principle one could use genotyping tests for this and other markers of interest to inform selective breeding decisions. Similarly, if the gene and particular variants of it have been identified, as for example RYR1, then its presence could be tested for.
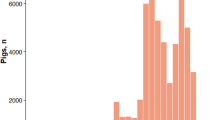
The polygenic, quantitative nature of traits of economic importance means that individual genes have small effect and are therefore themselves difficult to map. Given these features, MAS was deemed to be a promising approach, especially for low heritability traits. In order to perform MAS efficiently, however, markers would have to be mapped with considerable density across the genome (Lande and Thompson 1990). As the 1990s wore on, new techniques to identify markers of differing natures were developed and used, and the number of identified markers rapidly increased, as did the number of genes (see Table 3).
Using a new method of DNA fingerprinting first published in 1995, Amplified Fragment Length Polymorphisms (AFLPs) could also be identified (Vos et al. 1995). PIC identified AFLPs as a potential rich source of markers, for use in MAS as well as in identifying QTL for economically important traits. In collaboration with the Dutch company that developed AFLPs, Keygene NV, PIC ran a three-year European Commission-funded project as part of the BIOTECH2 programme of Framework Programme 4 to develop the means to use AFLPs in animal breeding.Footnote 16
Physical mapping also continued apace, with the use of the successive radiation hybrid panels to map increasing numbers of markers. Between April 1999 and November 2005, 7138 markers were mapped using the two INRA-maintained radiation hybrid panels.Footnote 17
MAS was thought to have the potential to be used in selection programmes: within breeds to capitalise on the variation existent in a given breed; or between breeds, for example to introduce (introgress) a desired gene variant or allele from one breed to a recipient breed. In addition to the identification and mapping of markers, extensive phenotypic data and information on the pedigree of the animals would be required to develop an effective programme of MAS (Dekkers and van der Werf 2007).
PIC claims to have implemented MAS for pork quality traits in 1998. The strategy of using a select set of markers persisted into the late-2000s. One estimate of the impact of MAS in the European Union was that in the livestock industry as a whole, MAS had been used in the breeding of between 40 and 80% of breeding females, and that the total direct economic contribution to the livestock industry had been 207–560 million euros (Papatryfon et al. 2008).Footnote 18
In the annual reports of Genus plc, the leading animal breeding company into which PIC became incorporated, the last mention of a strategy based on the identification of particular markers from the total set of genetic markers they had identified, was in 2009.Footnote 19 In the 2011 annual report, a new approach was cited: genomic selection.
Historian of animal breeding Margaret Derry has questioned the value of MAS, viewing it as a Mendelian interlude between the ‘black-box thinking’ and infinitesimal model that dominated breeding before the late-1980s, and now again in the genomic era (Derry 2015, pp. 131–159). The Mendelian characterisation is not, however, strictly true for many of the markers mapped and used to inform selective breeding decisions, at least if the Mendelian interest in transmission across generations is excluded. Where an actual gene was not the basis of selection, a marker could be used that was close enough to a gene whose presence might be inferred, but not necessarily confirmed, let alone characterised. The marker was characterised by simply existing as a marker first and foremost, with no presuppositions concerning its role in constructing the phenotype, or being a marker for a particular phenotypic effect, change or trait. As geneticist Michael Goddard told us, “for us animal breeders it doesn’t matter, we regard them all as markers.”Footnote 20
What was being searched for was a way to link the marker to some form of phenotypic variation on which there was data. Even where this is found, it may not be presumed to be causative of anything mechanistically or be the ‘real’ marker of the phenotypic trait. It may just be statistically and probabilistically associated with the phenotypic variation because it is linked to a particular stretch of DNA that is mechanistically and functionally associated. But it is not important that the ‘real’ marker or ‘causative’ gene ever be identified. The marker is neither like a classic Mendelian gene, nor a gene that is a defined sequence of DNA that is transcribed into RNA and then translated into protein (sensu Francis Crick, as explicated by Griffiths and Stotz 2013).
4 Single nucleotide polymorphisms and genomic selection
The advent of genomic selection around the turn of the twenty-first century derives from the projects, methods and theoretical models of genetics formulated by livestock geneticists. It drew on the object of the marker and the maps and resources developed using them.
Genomic selection is a set of approaches to genomic prediction predicated on the use of markers across the whole genome, potentially all detectable and measurable ones, to estimate breeding values for individual animals to inform selective breeding decisions (Haley and Visscher 1998; Meuwissen et al. 2001). With a sufficiently large and well-distributed set of markers, it could be assumed that any genes involved in a particular trait would have linked markers associated with them.
Initially, genomic selection was articulated as a development of MAS, applying the principle on a genome-wide scale, rather than with a select set of markers (e.g. Haley and Visscher 1998; Meuwissen 2007). However, there are key differences (see Table 4). The main one is that genomic selection did not seek to identify QTL of significant effect and then attribute zero effect sizes to all other loci. Instead, in genomic selection a prior distribution of single nucleotide polymorphism (single-base changes or variants in the DNA; SNP) effects is assumed, using Bayesian models. This proved to be notably accurate in estimating breeding values in simulation studies (Meuwissen et al. 2001). One model, BayesR, posits a distribution of effects that includes the attribution of zero effect to some SNPs. In calculating Estimated Breeding Values (EBVs), its accuracy equals or outperforms Best Linear Unbiased Prediction (BLUP). BLUP is an important and widely-used methodology originally developed in the late-1940s and subsequently refined; it is based on a quantitative approach assuming equal, tiny effect sizes of all loci (Erbe et al. 2012; Hill 2014, p. 5).
This approach is quite distinct from the more gene-centric approach of Genome Wide Association Studies (GWAS). Like genomic selection, GWAS aims to statistically relate data on the presence of particular markers (e.g. SNPs) and phenotypic variation in traits. The purpose of GWAS is, however, to identify potential causative loci (i.e. genes), and to be predictive about individuals (rather than about populations, as for genomic selection). Therefore, to avoid false positives, a significance threshold (set at a particular p value, typically far lower than < 0.05) is set for identifying valid effects. Rather than attributing a range of effects to markers, an effect is therefore determined to be either present or absent: potentially a strong clue as to the presence of a pertinent gene. Such attributions are the result of measures of statistical significance being applied to prevent the identification of false positives, yet it still results in the overestimation of the effects of QTL so identified, and the reduction of small but non-zero effect sizes to zero.
In genomic selection, the marker becomes ever more abstract as an entity and unit, and further divorced from presumed or inferred genes. The markers genotyped are ‘anonymous’. Their identity and position are irrelevant. If MAS represented a molecularisation (if not a Mendelianisation) of quantitative approaches to breeding, genomic selection moved away from more molecular genetic models of the distribution of gene effects, even as it came to rely on the masses of molecular data generated through sequencing projects (on the swine genome sequencing project, c. 2006–2009, see Lowe 2018). These included the identification of many hundreds of thousands of SNPs.
The vast increase in the number of available markers enabled the development of SNP chips, a kind of microarray. SNP chips are slides onto which specific DNA sequences have been affixed. These oligonucleotide probes are intended to capture a specific SNP. DNA fragments with attached coloured or fluorescent probes from the individual to be genotyped can then be introduced. Complementary sequences will hybridise to the fixed probes, and this will produce a signal that can be picked up using a detection system. The advent of SNP chips radically reduced the costs of genotyping, and therefore of the acquisition of data on the presence or absence of particular markers.
In the case of the pig, the first high-density SNP chip was produced in 2008 by Illumina, in conjunction with many of the researchers involved in the projects to map and then sequence the pig genome. After rounds of selection and validation, there were 62,121 markers included on what was called ‘PorcineSNP60’ (Ramos et al. 2009).
With the fast and relatively cheap genotyping of at least tens of thousands of markers now possible, the principles of genomic selection could be put into practice. The first step is to create a reference population. These are animals that will be genotyped, and have extensive phenotypic data collected on them. These data are then combined to produce a prediction model for calculating the genomic Estimated Breeding Values (gEBVs) of selection candidates. The reference populations should ideally be large, and be similar to the target population intended for genetic evaluation. The linkage between markers and QTL may not be present in the same way in different breeds, so a reference population for one breed may not be appropriate for generating a prediction model for a different breed. Not only are associations between markers themselves therefore relative to a particular population, but so are the associations between (constellations of) markers and variation in phenotypic traits. In the context of this marker-centric biology, markers are deeply relational in nature, as well as being ontologically inert and internally heterogeneous.
Genomic selection has had a remarkably short route from theoretical elucidation to practical implementation. The main gains for pig breeding would come from the improvement in accuracy of calculating EBVs, but this has been enough to ensure that genomic selection has been commercially implemented (e.g. Knol et al. 2016).
Downsides of genomic selection include an increase in inbreeding, due to the faster selection of a smaller pool of animals for breeding, and an enhancement of the in-built advantage of already well-populated breeds, for whom large reference populations can be constructed. Methods have been developed to deal with the inbreeding, while the development of across-breed reference populations has been mooted to counteract the disadvantage faced by smaller breeds.Footnote 21
The irony is that the development of materially and theoretically marker-dependent genomic selection now means that further addition of markers to SNP chips and prediction models is no longer fruitful, but greater numbers of animals and phenotypic data of greater depth are required instead.
Although genomic selection operates as a black-box, the marker is still assumed to be linked in some way to something causative. As linkage disequilibrium is important, this means that there is limited applicability across breeds, and that prediction models are liable to degrade in accuracy over generations. The genes themselves, while they are known to be there, are largely incidental. If they and any relevant mutations and variants were known, this would be welcomed, but if they are not known, the predictive power and accuracy of the thousands of markers suffices. Even if genes could be identified and edited to increase the proportion of advantageous alleles and remove deleterious ones, this need not replace marker-centric approaches such as genomic selection, but supplement them by modifying the variation upon which they work (as proposed in Tait-Burkard et al. 2018).
It is possible that in the future, sequencing costs will fall to such a level as to make genotyping-by-sequencing economically viable. It is not clear, however, that the gain in accuracy produced through achieving full coverage across the genome over the selective coverage provided by SNP chips would be sufficient to justify the additional investment in bioinformatics capacity required to make this work (Pérez-Enciso et al. 2017). The costs of whole genome sequencing are not restricted to the ever-declining expense of determining bases, but also sample collection, data processing, analysis and storage; these are not necessarily declining in price (Sboner et al. 2011). Furthermore, studies have suggested that to exploit the small projected improvements in accuracy by sequencing rather than using SNP chips, breeders would need to incorporate prior information concerning particular markers in whole genome sequence data anyway. They would also need to improve the collection of other data by using reference populations with far more animals and enhancing phenotypic data collection (Pérez-Enciso et al. 2017).
Finally, whole genome sequencing increases the coverage and density of what are still considered markers. We may therefore expect that the use of this in genomic selection will exhibit many of the same features as the more obviously marker-centric genomics discussed above.
5 What is distinctive about marker-centric biology?
Historian of genetics Raphael Falk has noted, in the context of a discussion of popular genetic determinism, that “[a]t the beginning of the twenty-first century even the term ‘gene’ seems to have lost its popularity to the term ‘DNA’” (Falk 2009, p. 289). It is important not to oversell this point, however. The gene as a conceptual tool, as well as a focus for experimental research, is remarkably persistent. Its mutability as a concept contributes to this, rather than detracting from it (Rheinberger and Müller-Wille 2017). To biologists, the promise of the gene is the potential for discerning causally-specific processes and mechanisms that can help them to explain, understand, experiment further and design interventions for a wide range of possible applications.
It is true, nonetheless, that the genome as a whole has now been opened up as a field of investigation. And to map this new terrain, and make use of the data generated, the means to identify and relate markers of very different kinds has become significant.
We have detailed work centred on the discernment, mapping and use of markers, a distinctive characteristic of research in livestock genetics. By doing so, we have been able to exhibit practices, concepts and communities that would otherwise be hidden in a more gene-centred account. This has enabled us to decentre prominent modes of, in particular, human genetics in the overall history of genetics and genomics, and to highlight the contributions of quantitative genetics. Our account counterposes the approaches of quantitative genetics to those of molecular genetics, but also draws attention to the importance of their intersection.
The advent of the means by which plentiful markers could be identified and mapped enabled these separate disciplinary and methodological approaches to genetics to intersect. This was aided by these markers’ ontological inertness, their abstraction from the techniques through which they were instantiated (and therefore their material nature), and their abstraction from considerations of mechanism and function. The additional abstraction of these material traces from species further enabled them to be used to construct—and be discerned from—inter-specific models of correspondence between the genomes of different species, such as pigs and humans. The alignment of quantitative and molecular genetics, which continued as separate lineages rather than fully integrating, enabled the advent of genomic selection around the turn of the twenty-first century. This has led to improved effectiveness of selective breeding in the livestock industry. Had the two approaches fully integrated on the terms of molecular genetics, the full flowering of a marker-centric approach may have been stymied in favour of using markers more predominantly for gene discovery.
Marker-centric biology has been a distinctive feature of livestock genetics due to the use of reference families and populations, the goals of intervention at the population (e.g. herd) level as opposed to the individual treatment goals of medicine, and the prominence of the quantitative genetics tradition that accepts ‘black-boxing’. Ever-denser maps of markers were intended to be of use in the pursuit of different strategies to advance livestock breeding, albeit in different ways. What united them was the absence of any presumption that the markers themselves have any necessary functional salience; that is, whether they were supposed to be linked to an inferred gene or not. Markers are different kinds of variants with different (potential) functional implications (including none). Markers are not necessarily presumed to be functional, yet held out the potential of altering function. The marker played this role in the context of producing the tools, which included maps, to map and isolate QTL. Maps became progressively more developed, aided by the addition of fresh markers, new DNA primers and probes, and the elucidation of syntenic (orthologous) relationships with (primarily) the human genome. Different types of marker reflected the multiple ways in which forms of genomic variation could be revealed, identified, created and added to the map. This, in part, helps to make the maps functional for a variety of purposes, though many had to do with livestock production and breeding. Inferences and methods to improve the efficiency of selection breeding have therefore been based on markers with no presumed mechanistic relationship to phenotypic variation.
Although they operated on different bases, in positional candidate gene, QTL, MAS and genomic selection-based strategies alike, the actual implementable product was a marker, set of markers or models based on markers. This informed and motivated the publicly-funded researchers. They did not ignore the functional, of course, and the identification and characterisation of genes remained a goal, particularly for parts of the community that were interested in the potential for transgenesis, or still saw some possibilities in ‘biological’ rather than the statistical approaches (a distinction and point recognised in Knol et al. 2016). Nevertheless, the exigencies of research promoted an agenda which focused on the means to discern and map markers, useful due to their relative abundance and highly polymorphic nature, and to develop the statistical and quantitative means with which to associate them with phenotypic variation.
As the markers have not been imbued with presuppositions concerning their relation to the phenotype, their properties are distinct from the two different meanings of the gene identified by Lenny Moss (2003): ‘gene-D’ and ‘gene-P’. Gene-Ds play developmental roles in constructing the phenotype; they are resources. Gene-Ps, on the other hand, are inferred from analysis of particular phenotypic effects, changes or traits. If links at the level of populations between markers and phenotypic variation are found, however, that does not necessarily mean that the marker operates in the same way as a gene-D or gene-P. It merely means that it is statistically and probabilistically associated with the phenotypic variation, linked as it is to a particular stretch of DNA that is mechanistically and functionally associated.
For informing human reproductive decision-making, rather than for diagnostic and medical decision-making, genotyping thousands of markers of small effect would have very little traction. One might be able to make reproductive decisions concerning Mendelian genes of large effect like Huntington’s or Tay-Sachs. However, for more complex conditions and traits, for which there would be a plethora of small probabilistic effects one way or the other, it is not really useful at all. It is for livestock breeding, though, where small improvements in various traits at the population level matter, and where there is control over the breeding in a population, rather than just single autonomous pair-bonding decisions.
We have detailed some of the biological research that took as a key object, or set of objects, the genetic marker. That the identification and use of markers was important to research in livestock genetics was undeniable, and due in large part, we contend, to presumed translational outcomes, towards which researchers in publicly-funded research institutions were working. These prompted the need to develop the resources and tools for a particular set of practices and arrangements for intervening in the variation of target objects such as the herds of breeding companies. This entailed a need to investigate and use the variation detectable and measurable in and from a particular, internally heterogeneous, set of biological elements: genetic markers. The variation was detected at the level of the genome, and the reference population. In order to make it work, the internal variation of the different material bases represented by the markers needed to be abstracted away. This in turn allowed different researchers from different backgrounds to work on what was, for the purposes of mapping, a common object.
Notes
Interview with Michael Goddard, conducted by authors, Edinburgh, 11th October 2018.
We generally refer to this species as the ‘pig’. Sometimes, however, we use the term ‘swine’ to reflect the usage of that term in certain contexts.
In principle, people conducting marker-centric genetics might view the prospect of a shift towards a more gene-centric approach as desirable and/or possible, but as our focus here is on concrete practices and outcomes rather than expectations or general plans, this is not pertinent.
Concerning pigs, markers have also been used for product tracing, confirmation of parentage, phylogenetic studies and the evaluation of biodiversity and genetic resources (Alderson and Plastow 2004).
On the articulation of ‘quantitative genetics’ as distinct from ‘population genetics’, see Derry (2015, pp. 112–114).
On the concept of the ‘wild type’, and its history, see Holmes (2017).
This required the elucidation of the mechanics of chromosomes and cell division to help them to account for anomalies they observed; it has been argued that the Morgan laboratory’s experimental programme was aimed at using the problem of mapping as an investigative strategy for tackling wider biological problems such as this (Waters 2003).
The relation of classical genetics to molecular genetics, and particularly the questions of continuity from the former to the latter, and of reduction of the former to the latter, have received considerable philosophical attention (e.g. Schaffner 1993; Waters 1994; Weber 2005). For a nuanced and historically-informed discussion, see Rheinberger and Müller-Wille (2000) and Müller-Wille and Rheinberger (2012, pp. 150–169).
This imperative was not confined to PSS. In France in the 1980s, a new method of measuring meat quality called ‘Napole’ technological yield, or ‘RTN’, was introduced. Using this measure, quantitative geneticist Jean Naveau (who worked for Pen Ar Lan, a company producing new synthetic lines of pigs) and colleagues detected the presence of a separate locus to the Halothane one that was presumed to have some effect on meat quality (Le Roy et al. 1990). This was named the Rendement Napole locus, or RN. It was established that it existed in populations derived from Hampshire breed stock. Due to the large effect of the gene, presence of either the dominant or recessive alleles could easily be established, though the locus itself was not fully localised until 1996 (Mariani 1999).
Before this, as well as the halothane sensitivity test, French scientists had developed a test to detect the presence of either of two variants of the blood enzymes phosphohexose isomerase and 6-phosphogluconate dehydrogenase in Landrace pigs. The variants detected were shown to indicate which of the alleles of two markers known to be closely linked to the Hal locus (Phi and Pgd) were present, in turn evincing which Hal allele was present (Courreau et al. 1985).
Personal communication from Chris Haley to James Lowe, 28th March 2019.
PiGMaP II final report, 1996, in ‘EC PiGMaPII—Final Report’ folder, Alan Archibald’s personal papers.
Personal communication from Chris Haley to James Lowe, 28th March 2019.
Interview with Chris Haley, conducted by authors, Edinburgh, 18th December 2017.
The most stringent criteria for assessing homology relationships were deemed to be: similar nucleotide sequence, cross-hybridisation to the same molecular probe, conserved map position as well as similar amino acid sequence of proteins and complementation of function (Archibald 1998).
https://cordis.europa.eu/project/rcn/31058/factsheet/en (Accessed 18th October 2019).
Source: dataset on mapped markers on the INRA2006 map, extracted from data provided at: http://rhdev.toulouse.inra.fr—as this website is no longer active, the dataset is available from the authors on request.
MAS was not necessarily an approach used by other breeding companies. It is cited here because the prospect of its application in one breeding company, PIC, motivated marker-centred research outside of industry, in academia for example, that has left its footprint in various projects and publications.
They had earlier disposed of patents relating to the use of markers, upon taking over Sygen (incorporating PIC) in the mid-2000s.
Interview with Michael Goddard, conducted by authors, Edinburgh, 11th October 2018. He is not alone in this assessment. For example, quantitative geneticist William Hill reported much the same sentiment; interview conducted by James Lowe, Edinburgh, 31st January 2018.
Interviews with Theo Meuwissen and Michael Goddard, conducted by authors over telephone on 6th April 2018 and in Edinburgh on 11th October 2018, respectively.
References
Alderson, G. L. H., & Plastow, G. S. (2004). Use of DNA markers to assist with product traceability and pedigree analysis and their role in breed conservation. Animal Genetic Resources, 35, 1–7.
Archibald, A. L. (1986). A molecular genetic approach to the porcine stress syndrome. In P. V. Tarrant, G. Eikelenboom, & G. Moni (Eds.), Evaluation and control of meat quality in pigs (pp. 343–357). Dordrecht: Martinus Nijhoff Publishers.
Archibald, A. L. (1998). Comparative genome mapping—the livestock perspective. In A. J. Clark (Ed.), Animal breeding: Technology for the 21st century (pp. 137–164). Amsterdam: Harwood Academic Publishers.
Archibald, A. L., Haley, C. S., Brown, J. F., Couperwhite, S., McQueen, H. A., Nicholson, D., et al. (1995). The PiGMaP consortium linkage map of the pig (Sus scrofa). Mammalian Genome, 6, 157–175.
Archibald, A. L., & Imlah, P. (1985). The halothane sensitivity locus and its linkage relationships. Animal Blood Groups and Biochemical Genetics (Special Issue), 4, 253–335.
Benavides, F. J., & Guénet, J.-L. (2012). Mouse genomics. In H. J. Hedrich (Ed.), The laboratory mouse (2nd ed., pp. 57–90). London: Elsevier.
Beurton, P. J., Falk, R., & Rheinberger, H.-J. (Eds.). (2000). The concept of the gene in development and evolution: Historical and epistemological perspectives. Cambridge: Cambridge University Press.
Botstein, D., White, R. L., Skolnick, M., & Davis, R. W. (1980). Construction of a genetic linkage map in man using restriction fragment length polymorphisms. American Journal of Human Genetics, 32, 314–331.
Bulmer, M. G. (1980). The mathematical theory of quantitative genetics. Oxford: Clarendon Press.
Courreau, J.-F., Sellier, P., Boulard, J., Breton, T., Goullieux, P., & Guérin, G. (1985). Blood markers (Phi and Pgd) and halothane sensitivity in the French Landrace pig breed. Journées de la Recherche Porcine en France, 34, 364.
Dalens, M., & Runavot, J.-P. (1993). Test moléculaire pour le dépistage du gène de la sensibilité à l’halothane chez le porc. Techni-Porc, 16, 17–20.
Davies, W., Harbitz, I., Fries, R., Stranzinger, G., & Hauge, J. G. (1988). Porcine malignant hyperthermia carrier detection and chromosomal assignment using a linked probe. Animal Genetics, 19(2), 203–212.
Dekkers, J. C. M., & van der Werf, J. H. J. (2007). Strategies, limitations and opportunities for marker-assisted selection in livestock. In E. P. Guimarães, J. Ruane, B. D. Scherf, A. Sonnino, & J. D. Dargie (Eds.), Marker-assisted selection: Current status and future perspectives in crops, livestock, forestry and fish (pp. 167–184). Rome: Food and Agriculture Organization of the United Nations.
Derry, M. E. (2015). Masterminding nature: The breeding of animals, 1750–2010. Toronto: University of Toronto Press.
Erbe, M., Hayes, B. J., Matukumalli, L. K., Goswami, S., Bowman, P. J., Reich, C. M., et al. (2012). Improving accuracy of genomic predictions within and between dairy cattle breeds with imputed high-density single nucleotide polymorphism panels. Journal of Dairy Science, 95(7), 4114–4129.
Falk, R. (2009). Genetic analysis: A history of genetic thinking. Cambridge: Cambridge University Press.
Fox Keller, E. (2000). The century of the gene. Cambridge, MA: Harvard University Press.
Fujii, J., Otsu, K., Zorzato, F., de Leon, S., Khanna, V. K., Weiler, J. E., et al. (1991). Identification of a mutation in porcine ryanodine receptor associated with malignant hyperthermia. Science, 253(5018), 448–451.
García-Sancho, M. (2015). Animal breeding in the age of biotechnology: The investigative pathway behind the cloning of Dolly the sheep. History and Philosophy of the Life Sciences, 37(3), 282–304.
Griffiths, P., & Stotz, K. (2013). Genetics and philosophy: An introduction. Cambridge: Cambridge University Press.
Gulcher, J. (2012). Microsatellite markers for linkage and association studies. Cold Spring Harbor Protocols. https://doi.org/10.1101/pdb.top068510.
Haley, C., & Visscher, P. M. (1998). Strategies to utilize marker-quantitative trait loci associations. Journal of Dairy Science, 81(2), 85–97.
Harper, P. S. (2008). A short history of medical genetics. Oxford: Oxford University Press.
Hawken, R. J., Murtaugh, J., Flickinger, G. H., Yerle, M., Robic, A., Milan, D., et al. (1999). A first-generation porcine whole-genome radiation hybrid map. Mammalian Genome, 10, 824–830.
Hill, W. G. (2014). Applications of population genetics to animal breeding, from Wright, Fisher and Lush to genomic prediction. Genetics, 196, 1–16.
Holmes, T. (2017). The wild type as concept and in experimental practice: A history of its role in classical genetics and evolutionary theory. Studies in History and Philosophy of Biological and Biomedical Sciences, 63, 15–27.
Jeon, J.-T., Carlborg, Ö., Törnsten, A., Giuffra, E., Amarger, V., Chardon, P., et al. (1999). A paternally expressed QTL affecting skeletal and cardiac muscle mass in pigs maps to the IGF2 locus. Nature Genetics, 21, 157–158.
Johannsen, W. (1909). Elemente der Exakten Erblichkeitslehre. Jena: Gustav Fischer.
Kaufmann, A. (2004). Mapping the human genome at Généthon laboratory: The French Muscular Dystrophy Association and the politics of the gene. In J.-P. Gaudillière & H.-J. Rheinberger (Eds.), From molecular genetics to genomics: The mapping cultures of twentieth-century genetics (pp. 129–157). London: Routledge.
Knol, E. F., Nielsen, B., & Knap, P. W. (2016). Genomic selection in commercial pig breeding. Animal Frontiers, 6(1), 15–22.
Kohler, R. E. (1994). Lords of the fly: Drosophila genetics and the experimental life. Chicago, IL: The University of Chicago Press.
Lande, R., & Thompson, R. (1990). Efficiency of marker-assisted selection in the improvement of quantitative traits. Genetics, 124, 743–756.
Le Roy, P., Naveau, J., Elsen, J.-M., & Sellier, P. (1990). Evidence for a new major gene influencing meat quality in pigs. Genetics Research, 55(1), 33–40.
Legault, C., Gruand, J., Lebost, J., Garreau, H., Ollivier, L., Messer, L. A., et al. (1996). Fréquence et effet sur la prolificité du gène ESR dans deux lignées Large White en France. Journées de la Recherche Porcine, 28, 9–14.
Lindee, M. S. (2005). Moments of truth in genetic medicine. Baltimore, MD: Johns Hopkins University Press.
Lowe, J. W. E. (in review). Ephemerality and indispensability: Establishing the Roslin Institute in international networks of pig genomics research. The British Journal for the History of Science (submitted to).
Lowe, J. W. E. (2018). Sequencing through thick and thin: Historiographical and philosophical implications. Studies in History and Philosophy of Biological and Biomedical Sciences, 72, 10–27.
MacLennan, D. H., Duff, C., Zorzato, F., Fujii, J., Phillips, M., Korneluk, R. G., et al. (1990). Ryanodine receptor gene is a candidate for predisposition to malignant hyperthermia. Nature, 343, 559–561.
Mariani, P. (1999). QTL identification and marker assisted selection: New solutions to old problems. In G. Piva, G. Bertoni, F. Masoero, P. Bani, & L. Calamari (Eds.), Recent progress in animal production science: Proceedings of the A.S.P.A. XIII Congress, Piacenza, June 21–24, 1999 (pp. 109–124). Milan: FrancoAngeli.
McCarthy, T. V., Healy, J. M. S., Heffron, J. J. A., Lehane, M., Deufel, T., Lehmann-Horn, F., et al. (1990). Localization of the malignant hyperthermia susceptibility locus to human chromosome 19ql2–13.2. Nature, 343, 562–564.
Meuwissen, T. (2007). Genomic selection: Marker assisted selection on a genome wide scale. Journal of Animal Breeding and Genetics, 124, 321–322.
Meuwissen, T. H. E., Hayes, B. J., & Goddard, M. E. (2001). Prediction of total genetic value using genome-wide dense marker maps. Genetics, 157, 1819–1829.
Morange, M. (M. Cobb, Transl.) (2000). A history of molecular biology. Cambridge, MA: Harvard University Press.
Moss, L. (2003). What genes can’t do. Cambridge, MA: The MIT Press.
Müller-Wille, S. (2018). Making and unmaking populations. Historical Studies in the Natural Sciences, 48(5), 604–615.
Müller-Wille, S., & Rheinberger, H.-J. (2012). A cultural history of heredity. Chicago, IL: The University of Chicago Press.
Myelnikov, D. (2017). Cuts and the cutting edge: British science funding and the making of animal biotechnology in 1980s Edinburgh. British Journal for the History of Science, 50(4), 701–728.
Nezer, C., Moreau, L., Brouwers, B., Coppieters, W., Detilleux, J., Hanset, R., et al. (1999). An imprinted QTL with major effect on muscle mass and fat deposition maps to the IGF2 locus in pigs. Nature Genetics, 21, 155–156.
O’Brien, P. J., & Ball, R. O. (2013). Porcine stress syndrome. In B. E. Straw, J. J. Zimmerman, S. D’Allaire, & D. J. Taylor (Eds.), Diseases of swine (9th ed., pp. 945–964). London: Blackwell.
O’Brien, S. J. (1991). Mammalian genome mapping: Lessons and prospects. Current Opinion in Genetics & Development, 1(1), 105–111.
Ollivier, L., Sellier, P., Monin, G., Dando, P., Vernin, P., & Talmant, A. (1975). Déterminisme génétique du syndrome d’hyperthermie maligne chez le porc de piétrain. Annales de génétique et de sélection animale, 7(2), 159–166.
Otsu, K., Khanna, V. K., Archibald, A. L., & MacLennan, D. H. (1991). Cosegregation of porcine malignant hyperthermia and a probable causal mutation in the skeletal muscle ryanodine receptor gene in backcross families. Genomics, 11, 744–750.
Papatryfon, I., Zika, E., Wolf, O., Gómez-Barbero, M., Stein, A. J., & Bock, A.-K. (2008). Consequences, opportunities and challenges of modern biotechnology for Europe. The analysis report: Contributions of modern biotechnology to European policy objectives. Luxembourg: Office for Official Publications of the European Communities.
Pérez-Enciso, M., Forneris, N., de los Campos, G., & Legarra, A. (2017). Evaluating sequence-based genomic prediction with an efficient new simulator. Genetics, 205, 939–953.
Provine, W. B. (1971). The origins of theoretical population genetics. Chicago, IL: The University of Chicago Press.
Rajagopalan, R. M., & Fujimura, J. H. (2018). Variations on a chip: Technologies of difference in human genetics research. Journal of the History of Biology, 51, 841–873.
Ramos, A. M., Crooijmans, R. P. M. A., Affara, N. A., Amaral, A. J., Archibald, A. L., Beever, J. E., et al. (2009). Design of a high density snp genotyping assay in the pig using SNPs identified and characterized by next generation sequencing technology. PLoS ONE, 4(8), e6524.
Rasmussen, N. (2014). Gene jockeys: Life science and the rise of biotech enterprise. Baltimore, MD: Johns Hopkins University Press.
Rettenberger, G., Klett, C., Zechner, U., Kunz, J., Vogel, W., & Hameister, H. (1995). Visualization of the conservation of synteny between humans and pigs by heterologous chromosomal painting. Genomics, 26, 372–378.
Rheinberger, H.-J., & Müller-Wille, S. (2000). Gene concepts. In S. Sarkar & A. Plutynski (Eds.), A companion to the philosophy of biology (pp. 3–21). Oxford: Wiley-Blackwell.
Rheinberger, H.-J., & Müller-Wille, S. (2017). The gene: From genetics to postgenomics. Chicago, IL: The University of Chicago Press.
Rothschild, M. F., Hu, Z.-L., & Jiang, Z. (2007). Advances in QTL mapping in pigs. International Journal of Biological Sciences, 3(3), 192–197.
Rothschild, M. F., & Plastow, G. S. (2002). Development of a genetic marker for litter size in the pig: A case study. In M. F. Rothschild & S. Newman (Eds.), Intellectual property rights in animal breeding and genetics (pp. 179–196). New York, NY: CABI Publishing.
Russell, E. S. (1989). Sewall Wright’s contributions to physiological genetics and to inbreeding theory and practice. Annual Review of Genetics, 23, 1–18.
Sboner, A., Mu, X. J., Greenbaum, D., Auerbach, R. K., & Gerstein, M. B. (2011). The real cost of sequencing: Higher than you think! Genome Biology, 12, 125.
Schaffner, K. F. (1993). Discovery and explanation in biology and medicine. Chicago, IL: The University of Chicago Press.
Schook, L. B., Beever, J. E., Clamp, P. A., Lewin, H. A., & McLaren, D. G. (1990). Status of the pig gene map. In J. E. Womack (Ed.), Mapping the genomes of agriculturally important animals (pp. 123–130). Cold Spring Harbor, NY: Cold Spring Harbor Laboratory Press.
Tait-Burkard, C., Doeschl-Wilson, A., McGrew, M. J., Archibald, A. L., Sang, H. M., Houston, R. D., Whitelaw, C. B, & Watson, M. (2018). Livestock 2.0 – genome editing for fitter, healthier, and more productive farmed animals. Genome Biology, 19, 204.
Tautz, D. (1989). Hypervariability of simple sequences as a general source for polymorphic DNA markers. Nucleic Acids Research, 17(6), 6463–6471.
Tóth, G., Gáspári, Z., & Jurka, J. (2000). Microsatellites in different eukaryotic genomes: survey and analysis. Genome Research, 10, 967–981.
Vos, P., Hogers, R., Bleeker, M., Reijans, M., van de Lee, T., Homes, M., et al. (1995). AFLP: A new technique for DNA fingerprinting. Nucleic Acids Research, 23(21), 4407–4414.
Waters, C. K. (1994). Genes made molecular. Philosophy of Science, 61(2), 163–185.
Waters, C. K. (2003). What was classical genetics? Studies in History and Philosophy of Science Part A, 35(4), 783–809.
Webb, A. J. (1980). The halothane test—A practical method of eliminating porcine stress syndrome. The Veterinary Record, 106, 410–412.
Weber, M. (2005). Philosophy of experimental biology. Cambridge: Cambridge University Press.
Weber, J. L. (1990). Human DNA polymorphisms and methods of analysis. Current Opinion in Biotechnology, 1(2), 166–171.
Weber, J. L., & May, P. E. (1989). Abundant class of human DNA polymorphisms which can be typed using the polymerase chain reaction. American Journal of Human Genetics, 44, 388–396.
Yerle, M., Pinton, P., Robic, A., Alfonso, A., Palvadeau, Y., Delcros, C., et al. (1998). Construction of a whole-genome radiation hybrid panel for high-resolution gene mapping in pigs. Cytogenetics and Cell Genetics, 82, 182–188.
Yi, D. (2015). The recombinant university: Genetic engineering and the emergence of stanford biotechnology. Chicago, IL: The University of Chicago Press.
Acknowledgements
We would like to thank Miguel García-Sancho, Doug Lowe and Erika Szymanski for their comments on drafts, and the two anonymous reviewers for their helpful suggestions. We would additionally like to thank all those who have participated in oral history interviews as part of our research into pig genomics, and those who have allowed us to see their personal documents. The research for this paper was conducted through the ‘TRANSGENE: Medical translation in the history of modern genomics’ project, which is funded by the European Research Council (ERC) under the European Union’s Horizon 2020 research and innovation programme under Grant Agreement No. 678757. This European research funding is deeply appreciated.
Author information
Authors and Affiliations
Corresponding author
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is distributed under the terms of the Creative Commons Attribution 4.0 International License (http://creativecommons.org/licenses/by/4.0/), which permits unrestricted use, distribution, and reproduction in any medium, provided you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made.
About this article
Cite this article
Lowe, J.W.E., Bruce, A. Genetics without genes? The centrality of genetic markers in livestock genetics and genomics. HPLS 41, 50 (2019). https://doi.org/10.1007/s40656-019-0290-x
Received:
Accepted:
Published:
DOI: https://doi.org/10.1007/s40656-019-0290-x