Abstract
One of the software engineering interests is quality assurance activities such as testing, verification and validation, fault tolerance and fault prediction. When any company does not have sufficient budget and time for testing the entire application, a project manager can use some fault prediction algorithms to identify the parts of the system that are more defect prone. There are so many prediction approaches in the field of software engineering such as test effort, security and cost prediction. Since most of them do not have a stable model, software fault prediction has been studied in this paper based on different machine learning techniques such as decision trees, decision tables, random forest, neural network, Naïve Bayes and distinctive classifiers of artificial immune systems (AISs) such as artificial immune recognition system, CLONALG and Immunos. We use four public NASA datasets to perform our experiment. These datasets are different in size and number of defective data. Distinct parameters such as method-level metrics and two feature selection approaches which are principal component analysis and correlation based feature selection are used to evaluate the finest performance among the others. According to this study, random forest provides the best prediction performance for large data sets and Naïve Bayes is a trustable algorithm for small data sets even when one of the feature selection techniques is applied. Immunos99 performs well among AIS classifiers when feature selection technique is applied, and AIRSParallel performs better without any feature selection techniques. The performance evaluation has been done based on three different metrics such as area under receiver operating characteristic curve, probability of detection and probability of false alarm. These three evaluation metrics could give the reliable prediction criteria together.
Similar content being viewed by others
Explore related subjects
Discover the latest articles, news and stories from top researchers in related subjects.Avoid common mistakes on your manuscript.
1 Introduction
As today’s software grows rapidly in size and complexity, the prediction of software reliability plays a crucial role in software development process [1]. Software fault is an error situation of the software system that is caused by explicit and potential violation of security policies at runtime because of wrong specification and inappropriate development of configuration [2]. According to [3], analyzing and predicting defectsFootnote 1 are needed for three main purposes, firstly, for assessing project progress and plan defect detection activities for the project manager. Secondly, for evaluating product quality and finally for improving capability and assessing process performance for process management. In fault prediction, previous reported faulty data with the help of distinct metrics identify the fault-prone modules. Important information about location, number of faults and distribution of defects are extracted to improve test efficiency and software quality of the next version of the software. Two benefits of software fault prediction are improvement of the test process by focusing on fault-prone modules and by identification the refactoring candidates that are predicted as fault-prone [4]. Numbers of different methods were used for software fault prediction such as genetic programming, decision trees, neural network, distinctive Naïve Bayes approaches, fuzzy logic and artificial immune system (AIS) algorithms. Almost all software fault prediction studies use metrics and faulty data of previous software release to build fault prediction models, which is called supervised learning approaches. Supervised machine learning classifiers consist of two phases: training and test phase; the result of training phase is a model that is applied to the testing data to do some prediction [5]. There are some other methods like clustering, which could be used when there are no previous available data; these methods are known as unsupervised learning approaches. It should be mentioned that some researchers like Koksal et al. [6] used another classification for data mining methods, which are famous as descriptive and predicative.
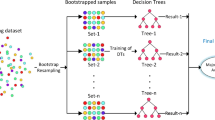
One of the main challenges in this area is how to get the data. In some works like in [7], a specific company provides the data, so the results are not fully trustable. Before 2005, more than half of the researches have used non-public datasets; however after that, with the help of PROMISE repository, the usage of public datasets reached to half [8], because the results are more reliable and not specific to a particular company. According to [8], software fault predictions are categorized based on several criteria such as metrics, datasets and methods. According to the literatures, software fault prediction models are built based on different set of metrics; method-level and class-level are two of the most important ones. Method-level metrics are suitable for both procedural and object-oriented programming style whereas class-level metrics are extracted based on object-oriented notation. It should be mentioned that compared to the other metrics, the method-level metrics is still the most dominant metrics prediction, followed by class-level metrics in fault prediction research area and machine-learning algorithms. It has been for many years that researchers work on different types of algorithms based on machine learning, statistical methods and sometimes the combination of them. In this paper, the experiments have been done on four NASA datasets with different population size using two distinct feature selection techniques that are principal component analysis (PCA) and correlation-based feature selection (CFS). We have changed the defect rates to identify what will be the effects on the predicting results. The predictability accuracy has been investigated in this paper based on two different method-level metrics, which are 21 and 37 static code attributes. The algorithms in this study are decision tree (C4.5), random forest, Naïve Bayes, back propagation neural network, decision table and various types of AIS such as AIRS1, AIRS2, AIRSParallel, Immunos1, Immunos2, Immunos99, CLONALG and clonal selection classification algorithm (CSCA). Three different performance evaluation metrics were used, area under receiver operating characteristic curve (AUC), probability of detection (PD) and probability of false alarm (PF), to give more reliable prediction analysis. Although we calculated accuracy along with above metrics, it does not have any impact on the evaluation process. Figure 1 shows the research done in this study.
We conducted four different types of experiment to answer six research questions in this paper. Research questions are listed as follows:
-
RQ1:
which of the machine learning algorithms performs best on small and large datasets when 21-method-level metrics is used?
-
RQ2:
which of the AIS algorithms performs best on small and large datasets when 21-method-level metrics is used?
-
RQ3:
which of the machine learning algorithms performs best on small and large datasets when 37-method-level metrics is used?
-
RQ4:
which of the machine learning algorithms performs best on small and large datasets when PCA and CFS applied on 21-method-level metrics?
-
RQ5:
which of the AIS algorithms performs best on small and large datasets when PCA and CFS applied on 21-method-level metrics?
-
RQ6:
which of the machine learning algorithms performs best and worst on CM1 public dataset when the rate on defected data is doubled manually?
The experiment 1 answered research question 1 (RQ1) and research question 2 (RQ2). Experiment 2 responded to research question 3 (RQ3). Experiment 3 shows the difference between the results obtained when no feature selection techniques were used. This experiment answered the research question 4 and 5. Finally, in experiment 4, to answer the last question, we doubled the defect rate of CM1 dataset to see whether it has any effect on the prediction model performances or not. This paper is organized as follows: the following section presents the related work. Section 3 explains different classifiers in AIS with its advantages and drawbacks. The feature selection and some of its methods are reviewed in Sect. 4. Experimental description and study analysis are described in Sects. 5 and 6, respectively, and finally Sect. 7 would be the results.
2 Related works
According to Catal [9], software fault prediction became one of the noteworthy research topics since 1990, and the number of research papers is almost doubled until year 2009. Many different techniques were used for software fault prediction such as genetic programming [10], decision trees [11] neural network [12], Naïve Bayes [13], case-based reasoning [14], fuzzy logic [15] and the artificial immune recognition system algorithms in [16–18]. Menzies et al. [13] have conducted an experiment based on public NASA datasets using several data mining algorithms and evaluated the results using probability of detection, probability of false alarm and balance parameter. They used log-transformation with Info-Gain filters before applying the algorithms and they claimed that fault prediction using Naïve Bayes performed better than the J48 algorithm. They also argued that since some models with low precision performed well, using it as a reliable parameter for performance evaluation is not recommended. Although Zhang et al. [19] criticized the paper but Menzies et al. defended their claim in [20]. Koru and Liu [21] have applied the J48, K-Star and random forest algorithms on public NASA datasets to construct fault prediction model based on 21 method-level. They used F-measures as an evaluation performance metrics. Shafi et al. [22] used two other datasets from PROMISE repository, JEditData and AR3; they applied 30 different techniques on them, and showed that classification via regression and locally weighted learning (LWL) are better than the other techniques; they chose precision, recall and accuracy as an evaluation performance metrics. Catal and Diri [4] have used some machine learning techniques like random forest; they also applied artificial immune recognition on five NASA datasets and used accuracy and area under receiver operating characteristic curves as evaluation metrics. Turhan and Bener have used probability of detection, probability of false alarm and balance parameter [9, 23]; the results indicate that independence assumption in Naïve Bayes algorithm is not detrimental with principal component analysis (PCA) pre-processing. Alsmadi and Najadat [24] have developed the prediction algorithm based on studying statistics of the whole dataset and each attributes; they proposed a technique to evaluate the correlation between numerical values and categorical variables of fault prone dataset to automatically predict faulty modules based on software metrics. Parvinder et al. [25] claimed that, the prediction of different level of severity or impact of faults in object oriented software systems with noise can be done satisfactory using density-based spatial clustering; they used KC1 from NASA public dataset. Burak et al. [26] analyzed 25 projects of the largest GSM operator in Turkey, Turkcell to predict defect before the testing phase, they used a defect prediction model that is based on static code attributes like lines of code, Halstead and McCabe. They suggested that at least 70 % of the defects can be detected by inspecting only 6 % of the code using a Naïve Bayes model and 3 % of the code using call graph-based ranking (CGBR) framework.
3 Artificial immune system
In late 1990, a new artificial intelligence branch that was called AIS was introduced. AIS is a technique to the scene of biological inspired computation and artificial intelligence based on the metaphor and abstraction from theoretical and empirical knowledge of the mammalian immune system. The immune system is known to be distributed in terms of control, parallel in terms of operation, and adaptive in terms of functions, all the features of which are desirable for solving complex or intractable problems faced in the field of artificial intelligence [27].
AISs embody the principles and advantages of vertebrate immune system. The AIS has been used in intrusion detection, classification, optimization, clustering and search problems [4].
In the AIS, the components are artificial cells or agents which flow through a computer network and process several tasks to identify and prevent attacks from intrusions. Therefore, the artificial cells are equipped with the same attributes as the human immune system. The artificial cells try to model the behavior of the immune-cells of the human immune system. Network security, optimization problems and distributed computing are some of the AIS’s applications.
There are several classifiers available based on AIS paradigm, some of them are as follows: AIRS, CLONALG, and IMMUNOS81, each one of them is reviewed in the following subsections.
3.1 Artificial intelligence recognition system (AIRS)
Artificial intelligence recognition system is one of the first AIS techniques designed specifically and applied to classification problems. It is a novel immune inspired supervised learning algorithm [28, 29].
AIRS has five steps: initialization, antigen training, competition for limited resources, memory cell selection and classification [4, 27]. These five steps are summarized in Fig. 2.
First, the dataset is normalized, and then based on the Euclidian formula distances between antigens, which is called affinity, are calculated. Affinity threshold that is the user-defined value is calculated. Antibodies that are present in the memory pool are stimulated with a specific infected antigen, and the stimulated value is assigned to each cell. The cell, which has the highest stimulation value, is chosen as the finest memory cell. Afterwards, the best match from the memory pool is selected and added to the artificial recognition ball (ARB) pool. This pool contains both antigen and antibodies with their stimulation value along with some other information related to them. Next, the numbers of clones are calculated and cloning starts. These clones are also added to the ARB pool. After that, competition for the finite resources begins. Again, ARB pool is stimulated with the antigens, and limited resources are assigned to them based on derived stimulation values. This is a recursive task until the stopping condition happens that is, if the stimulation level between ARB and antigen is less than the affinity threshold, it stops; otherwise, it goes on. After that, the ARB with the highest stimulation value is selected as a candidate to be a memory cell. The stimulation value is compared to the best previous matching value, if it is better; it is going to be replaced by the old one. Another explanation of the AIRS is shown in Fig. 3 and is described completely in [27].
The activity diagram of AIRS algorithm based on [27]
In a simple word, each input vector is a representative of an antigen, and all attributes of this vector are the epitope of the antigens, which is recognizable by the antibodies. So each antigen with \(M\) epitope is like Ag \(=\) \([\mathrm{ep}_{1}, \mathrm{ep}_{2}, \mathrm{ep}_{3},\ldots ]\).
For each epitope, one antibody is considered (Ab). The affinity between each pair of epitope of each antigen and antibody is calculated in Eqs. 1 and 2 as follows:
AIRSv1 is the first version of this algorithm. AIRSv1 (AIRS1) treats the ARB pool as a persistent resource during the entire training process whereas ARB pool is used as a temporary resource for each antigen in AIRSv2 (AIRS2). In another word, ARB’s leftovers for past antigens from the previous ARB refinement step are maintained and are participated in a competition for limited resources. According to [28] this cause more time spending in rewarding and refining ARBs that belong to the same class of the antigen in question. In order to solve this problem, a user defined stimulation value is raised in AIRS2 and only clones of the same class as the antigen are considered in ARB pool. The other difference between these two algorithms is how mutation is done. In AIRS1, the mutate rate is a user defined parameter and shows the mutate degree for producing a clone; mutate rate is simply replaced with normalized randomly generated value. Mutate rate in AIRS2 is identified as proportional to its affinity to antigen in question. This approach performs a better search when there is a tight affinity. Both AIRS1 and AIRS2 show similar accuracy performance behavior except that, AIRS2 is a simpler algorithm and is also show better generalization capability in terms of improved data reduction of the training dataset [4, 27]. Watkins [29] introduced the parallel implementation in 2005. The model shows the distributed nature and parallel processing attributes exhibited in mammalian immune system. The approach is simple and we have to add the following step to the standard AIRS training schema. If the dataset is not partitioned too widely, then the training speed is observable. AIRS Parallel have the following steps [4, 27, 29]:
-
Divide the training dataset into np Footnote 2 partitions.
-
Allocate training partitions to processes.
-
Combine np number of memory pools.
-
Use a merging approach to create the combined memory pool.
Acceptance of continuous and nominal variables, capacity to learn and recall large numbers of patterns, experienced-based learning, supervised learning, classification accuracy, user parameter and the ability to predict the training times are some of the design goals that could be noted for an AIRS-like supervised learning system.
3.2 CLONALG
The theory specifies that the organism has a pre-existing pool of heterogeneous antibodies that can recognize all antigens with some level of specificity [30].
As you may see in Fig. 4, when matching occurs, the candidate cell undergoes mitosis and produces \(B\) lymphoblast that could be one of the following:
-
Plasma cell that produces antibody as an effector of the immune response.
-
Long-lived memory cell, in case a similar antigen appears.
The simple overview of the clonal selection process, image taken from [32]
CLONal selection ALGorithm is inspired by the clonal selection theory of acquired immunity, previously known as CSA. A new clonal selection inspired classification algorithm is called CSCA.
CLONALG inspires some features from clonal selection theory, which is mentioned above. The goal here is to develop a memory pool containing best antigen matching antibodies that represent a solution to engineering problems.
The algorithm provides two searching mechanism for the desired final pool of memory antibodies. These two are noted in [30] as follows:
-
Local search, provided via affinity maturation of cloned antibodies. More clones are produced for better-matched antibodies.
-
A search that provides a global scope and involves the insertion of randomly generated antibodies to be inserted into the population to further increase the diversity and provide a means for potentially escaping local optima.
Figure 5 shows an algorithm based on GLONALG Theory. A CLONALG technique has a lower complexity and smaller number of user parameters compared to other AIS systems such as AIRS [31]. CLONALG algorithm is mainly used in three engineering problem domains: pattern recognition, function optimization and combinatorial optimization [30].
Parallel CLONALG works like a distributed system. The problem is divided into number of processes. The task of each one is preparation of themselves as antigen pools. After completion, all results will be sent to the root and the memory pool forms based on the combination of them.
There are some other classifications based on clonal selection algorithms such as CLONCLAS (CLONal selection algorithm for CLASsification) which is mostly used in character recognition. Here, there is a concept called class that contains an antibody and the antigen exposed to a class of specific antibody.
To improve and maximize the accuracy of the classification and also minimize the wrong classification, CSCA came to the picture. CSCA or clonal selection classifier algorithm has four distinct steps, which is started with Initialization like every other AIS algorithm followed by repetition (loop) of Selection and Pruning, Cloning and Mutation and Insertion until the stopping point, and at the end it has Final Pruning and Classification.
In Initialization step, an antibody pool is created based on randomly selected antigen which has size \(S\). In loop phase, there is a Selection and Pruning step that shows and scoring the antibody pool to each antigen set, which could be either correct classification score or misclassification score. After that selection rules are applied, antibodies with a misclassification score of zero are eliminated or antibodies with the fitness scoring of less than epsilon are removed from the chosen set and from the original antibody set as well. After that, all remaining antibodies in the selected set are cloned, mutated, and inserted to the main antibody set. When the loop condition is fulfilled, the Final Pruning step starts which exposes the final antibodies set to each antigen and calculates fitness scoring exactly like the loop step. Finally, the set of exemplar is ready in antibody set, so in case of any unclassified data instances that are exposed to the antibodies set, the affinities between each matches are calculated and selected and according to this result, unclassified data set could be classified. Figure 6 shows the CSCA steps.
Overview of the CSCA algorithm taken from [30]
3.3 Immunos81
Most of the issues that described in AIS are close to biological metaphor but the goal for Immunos81 is to reduce this part and focus on the practical application. Some of the terminologies are listed as below:
-
T-Cell, both partitioning learned information and decisions about how new information is exposed to the system are the duty of this cell, each specific antigen has a T-Cell, and it has one or more groups of B-Cells.
-
B-Cell, there is an instance of a group of antigens.
-
Antigen, this is a defect; it has a data vector of attributes where the nature of each attribute like name and data type is known.
-
Antigen-Type, depends on the domain, antigens are identified by their names and series of attributes, which is a duty of T-Cell.
-
Antigen group/clone based on the antigen’s type or a specific classification label forms a group that is called clone of B-Cell and as mentioned above, are controlled by a T-Cell.
-
The recognition part of the antibody is called paratope, which is bound to the specific part of the antigen, epitopes (attributes of the antigen).
This algorithm has three main steps: initialization, training and classification [33]; the general idea behind the Immunos81 is shown in Fig. 7.
To calculate the affinity values for each paratope across all B-Cells, Eq. 3 is used, \(p_{i}\) is the paratope affinity for the \(i{\mathrm{th}}\) paratope, \(k\) is a scale factor and \(S\) is total number of B-Cell, \(j{\mathrm{th}}\) B-Cell affinity in a clone set is shown by \(a_{i}\).
There is another concept called Avidity, which is sum of affinity values scaled both by the size of the clone population and additional scale parameter, according to Eq. 4, ca is a clone avidity for the \(i{\mathrm{th}}\), \(k_{2i}\) defines by user, \(N\) is the total paratopes and total number of B-Cells in the \(i{\mathrm{th}}\) clone set is shown by \(S_{i}\).
There are two basic implementations for Immunos8 that are known as naïve immunos algorithms; they are called, Immunos1 and Immunos2. There is no data reduction in Immunos1, and it is similar to the k-nearest neighbors. The primary difference is obviously that the training population is partitioned and \(k\) is set to one for each partition; multiple-problem support can be provided with simpler mechanism that uses classifier for each problem, and each classifier has its own management mechanism. The Immunos2 implementation is the same as Immunos1 only seeks to provide some form of primary generalization via data reduction, so the closer representation to basic Immunos [33].
Immunos99 could be identified as a combination of Immunos81 and CSCA which some user-defined parameters are either fixed or removed from the CSCA. Immunos99 is very different from the AIR classifiers such as AIRS and CLONALG; they all do competition in a training phase so the size of the group set has some affection on affinity and avidity calculation. There is another distinguishing point also; there is a single exposure to the training set (only in CLONALG).
As mentioned before, there is a classification called antigen-group and antigen-type. If the algorithm could identify some groups of B-Cell that are able to identify a specific type of antigens which these antigens might be also in different forms, then we can conclude that, how well any B-Cells could respond to its designed class of antigens compared to any other classes. The training step is composed of four basic levels; first data is divided into antigen-groups, and then the B-Cell population is set for each antigen-group (same as the immunos). For user, defined number of times, the B-Cell is shown to the antigens from all groups and fitness value is calculated; population pruning is done after that, two affinity maturations based on cloning and mutations are performed and some random selected antigens from the same group are inserted to the set. The loop is finished here when it is fulfilled the stopping condition. After last pruning for each B-Cell population, the final B-Cell population is introduced as a classifier. The usefulness or fitness formula for each B-Cell is shown in Eq. 5.
Correct means, sum of antigen ranked based score of the same group and incorrect means, sum of them in different groups of B-Cell. Here, all B-Cells have correct and incorrect scores, and also the antigen-group could not be changed; this is different from CSCA method.
4 Feature selection
Feature selection identifies and extracts the most useful features of the dataset for learning, and these features are very valuable for analysis and future prediction. So by removing less important and redundant data, the performance of learning algorithm could be improved. The nature of the training data plays the major role in classification and prediction. If the data fail to exhibit the statistical regularity that machine learning algorithms exploit, then learning will fail, so one of the important tasks here is removing the redundant data from the training set; it will, afterwards make the process of discovering regularity much easier, faster and more accurate.
According to [34], Feature selection has four different characteristics which are starting point, search organization, evaluation strategy, and stopping criterion. Starting point means from where the research should begin, it could be either begin with no feature and add a feature as you proceed forward, or it could be a backward process; you start with all attributes, and as you proceed, you do the feature elimination, or it could start from somewhere in the middle on the training set. In search organization step, suppose the dataset has \(N\) number of features, so there will be 2 \(N\) number of subsets. So the search could be either exhausted or heuristic.
Evaluation strategy is divided into two main categories, which are wrapper, and filters. Wrapper evaluated the importance of the features based on the learning algorithm, which could be applied on data later. It uses the search algorithm to search the entire feature’s population, run the model on them, and evaluate each subset based on that model. This technique could be computationally expensive; it has been seen that it may have suffered from over fitting to the model. Cross validation is being used to estimate the final accuracy of the feature subset. The goal of cross-validation is to estimate the expected level of fit of a model to a data set that is independent of the data, which were used to train the model. In filters, the evaluation is being done based on heuristic’s data and general characteristics of the data come to the picture. Searching algorithm in both techniques are similar but filter is faster and more practical when the populations of the features are high because the approach is based on general characteristic of heuristic data rather than a method with a learning algorithm to evaluate the merit of a feature subset.
Feature selection algorithms typically fall into two categories; it could be either feature ranking or subset ranking. If the ranking is done based on metric and all features that do not achieve a sufficient score are removed, it is called feature ranking but subset selection searches the set of possible features for the optimal subset, which includes wrapper and filter.
4.1 Principal component analysis
Principal component analysis is a mathematical procedure, the aim of which is reducing the dimensionality of the dataset. It is also called an orthogonal linear transformation that transforms the data to a new coordinate system. In fact, PCA is a feature extraction technique rather than a feature selection method. The new attributes are obtained by a linear combination of the original attributes. Here, the features with the highest variance are kept to do the reduction. Some papers like [35] used PCA for improving their experiments’ performance. According to [36], the PCA technique transforms \(n\) vector \(\{ {x_1 ,x_2 ,\ldots ,x_n }\}\) from the \(d\)-dimensional space to \(n\) vectors \(\{{x}_1^{\prime } ,{x}_2^{\prime } ,\ldots ,{x}_n^{\prime }\}\) in a new \({d}^{\prime }\) dimensional space.
where \(\mathtt{e }_{\mathrm{k}}\) are eigenvectors corresponding to \({d}^{\prime }\) largest eigen vectors for the scatter matrix \(S\) and \(a_{k,i} \) are the projections (principal components original data sets) of the original vectors \(x_{i}\) on the eigenvectors \(\mathtt{e }_{{k}} \).
4.2 Correlation-based feature selection
Correlation-based feature selection is an automatic algorithm, which does not need user-defined parameters like the number of features that need to be selected. CFS is categorized aa filter.
According to [37], feature \(V_{i}\) is said to be relevant, If there exists some \(v_{i}\) and \(c\) for which \(p\) (\(V_{i}~=~v_{i})\) \(>\) 0 such that in Eq. 7.
According to research on feature selection experiments, irrelevant features should be removed along with redundant information. A feature is said to be redundant if one or more of the other features are highly correlated with it [38]. As it was mentioned before, all redundant attributes should be eliminated, so if any features’ prediction ability could be covered by another, then it can be removed. CFS computes a heuristic measure of the “merit” of a feature subset from pair-wise feature correlations and a formula adapted from test theory. Heuristic search is used to traverse the space of feature subsets in reasonable time; the subset with the highest merit found during the search is reported. This method also needs discretizing the continuous features.
5 Experiment description
5.1 Dataset selection
Here, four datasets from REPOSITORY of NASA [39, 40] are selected. These datasets are different in number of rows and rate of defects. The largest dataset is JM1 with 10,885 rows, which belongs to real time predictive ground system project; 19 % of these data are defected. The smallest dataset, CM1, belongs to NASA spacecraft instrument project and it has 498 modules and 10 % of the data are defected. KC1 is another dataset which belongs to storage management project for receiving and processing ground data with 2,109 modules. 15 % of the KC1 modules are defected. PC3 is the last dataset that has 1,563 modules; 10 % of the data are defected and it belongs to flight software for earth orbiting satellite [39]. It should mention that PC3 is used only in first two experiments.
5.2 Variable selection
Predictability performance is calculated based on two distinct method-level metrics 21 and 37. In this work, experiments that have not been performed in [4] are studied. All 21 metrics which are the combination of McCabe’s and Halstead’s attributes are listed in Table 1. McCabe’s and Halstead’s metrics are called module or method level metrics and the faulty or non-faulty label is assigned to each one of the modules. These set of metrics are also called static code attributes and according to [13] they are useful, easy to use and widely used. Most of these static codes could be collected easily, cheaply and automatically. Many researchers and verification and validation text books such as [13, 41] suggest using complexity metrics to decide about which module is worthy of manual inspection. NASA researchers like Menzies et al. [13] with lots of experience about large governmental software have declared that they will not review software modules unless tools like McCabe predict that they are fault prone. Nevertheless, some researchers such as Shepperd and Ince [42] and Fenton and Pfleeger [43] argued that static codes such as McCabe are useless metrics, but Menzies et al. in [13, 20] proved that prediction based on the selected dataset with static code metrics performed very well and based on their studies they built prediction model with higher probability of detection and lower probability of false alarm which was in contrast with Shepherd and Ince [42] and Fenton and Pfleeger [43] beliefs.
As it is shown in Table 1, the attributes mainly consist of two different types, McCabe and Halstead. McCabe argued that codes with complicated pathways are more error-prone. His metrics, therefore, reflects the pathways within a code module but Halstead argued that, code that is hard to read, is more likely to be fault prone.
5.3 Simulator selection
All the experiments have been done in WEKA, which is open-source software and implemented in JAVA; it is developed in the University of Waikato and it is used for machine learning studies [44].
5.4 Performance measurements criteria
We used tenfold cross validations and all experiments were repeated five times. According to Menzies et al. [13, 20] since some models with low precision performed well, using it as a reliable parameter for performance evaluation is not good. They also mentioned that if the target class (faulty/non-faulty) is in the minority, accuracy is a poor measure as for example, a classifier could score 90 % accuracy on a dataset with 10 % faulty data, even if it predicts that all defective modules are defect free. Hence in this study, area under receiver operating characteristic curve values were used for benchmarking especially when the dataset is unbalanced. Other performance evaluation metrics that were used are: PD which is the number of fault-prone modules that are classified correctly and PF that is the number of not fault-prone modules that are classified incorrectly as defected, Table 2, Eqs. 8, 9 and 10 show all details about calculation of performance evaluation metrics. As a brief explanation, true negative (TN) means that the module is predicted as non-faulty correctly, whereas false negative (FN) means that the module is predicted as non-faulty wrongly. On the other hand, false positive (FP) means that the module is estimated as faulty incorrectly and true positive (TP) denotes that the module is predicted as faulty correctly.
6 Analysis of the experiment
6.1 Experiment 1
Twenty-one method-level metrics were used for this experiment. All 13 machine-learning techniques were applied on four different NASA datasets, and the results were compared. Table 3 shows accuracy and AUC value of algorithms and Table 4 presents PD and PF values for this experiment. This experiment has been done to answer research question 1 and 2. Notable values obtained after applying each algorithm are specified in bold.
As mentioned earlier and according to Menzies et al. [13, 20], a good prediction should have high AUC and PD values as well as low PF value, so with this consideration, we have evaluated the results. Figure 8 presents the performance comparison in terms of AUC among four different NASA projects. According to the figure, both random forest and decision table performed best compared to other algorithms for JM1. For KC1, Naïve Bayes also performed well along with random forest and decision table. Random forest performed best when it comes to CM1 as well. Figures 9, 10 and 11 have been drawn to show different evaluation values based on PF, PD and AUC for easier comparison between prediction models.
According to Fig. 9, both random forest and decision table have high AUC value but when we consider PF and PD values, random forest has better combination of low PF and high PD. Among AIS classifiers, although Immunos1 has higher AUC and PD compared to the others but PF value is high and not acceptable. Here AIRSParallel performed better. Figure 10 shows the comparison results for KC1, here also random forest, Naïve Bayes and decision tree have higher AUC values, but if we consider PF and PD values, decision tree will be eliminated. In AIS algorithms, both AIRSParallel and CSCA classifiers performed better than the others. Results based on PC3 dataset are also similar to KC1 as they have similarities in number of modules and defect rate. According to Fig. 11, random forest followed by Naïve Bayes has highest AUC value for CM1, but Naïve Bayes performed better when PD and PF values are also considered. AIRS1 and AIRSParallel have better results compared to the other AIS algorithms.
From Tables 3 and 4, we could conclude that, random forest and decision table are the best classifiers when the size of the dataset is not that much small; of course Naïve Bayes is an acceptable algorithm as it performs well among the others for all four datasets. AIRSParallel is better than the others when the size of dataset is not so large, and CSCA performs well when the size of dataset is not small. It seems that if the algorithms execute best in big datasets, there is a high chance that they perform well with smaller datasets as well.
6.2 Experiment 2
As only for PC3, 37 method-level metrics was available; we applied 13 algorithms on PC3 dataset. The attributes and the results are shown in Tables 5 and 6, respectively.
To have a reliable results, this experiment has been repeated ten times but no significant changes are observed compared to the results obtained from 21 method-level metrics in experiment 2. There is only a slight change in accuracy, PD and PF values when AIRSParallel [27] classifier is used, so it seems that other 37 variables except common variables with 21 method-level do not have major effect on building a prediction model and it only increases the training time.
6.3 Experiment 3
In this part, different feature selection techniques were applied on datasets to see their probable effects on the results and evaluation metrics. As it has been explained before, by eliminating the redundant attributes, performance of the model could be improved. Here two feature selection techniques, PCA and CFS, were used. When PCA applied on the 21 method-level metrics, the number of attributes is reduced to 7, 8 and 8 for CM1, PC1 and JM1, respectively. The results are shown in Tables 7 and 8. This experiment has been done to answer the research questions 4 and 5.
Using PCA as feature selection techniques and with the consideration of high AUC and PD values as well as low PF value, random forest and Naïve Bayes perform well for CM1 dataset compared to the others. Among AIS classifiers, Immunos1 and Immunos99 are the finest. RF and Decision Table are best for both JM1 and KC1 datasets with AUC, PD and PF as a performance evaluation metrics; also Immunos99 performs best among the other AIS algorithms for JM1.
After applying CFS with the best first classifier, some of the attributes were eliminated from the 21 of total attributes; seven attributes remain from CM1, eight from KC1 and JM1. The results are also shown in Tables 9 and 10. The remaining attributes form JM1, KC1 and CM1 after applying CFS, best first are listed below:
- CM1::
-
loc, iv(g), i, LOComment, LOBlank, uniq_Op, Uniq_Opnd
- KC1::
-
v, d, i, LOCode, LOComment, LOBlank, Uniq_Opnd, branchcout
- JM1::
-
loc, v(g), ev(g), iv(g), i, LOComment, LOBlank, locCodeAndComment
It should be mentioned here, after applying CFS, Best First, there are only slight changes observed in the results, so it means that there is no considerable difference in the results after applying distinct feature selection techniques. The main change would be the execution time reduction. As it is shown in Tables 9 and 10, the best performance for JM1, which is the largest dataset, belongs to decision tree followed by random forest. By checking the AUC, PF and PD values, naïve bayes perform greatly on CM1 dataset as well as the other three. CSCA also performed well among the other AIS classifiers, but the problem with this algorithm is long execution time.
We also applied CFS, random search to see whether it has a considerable change in the results or not. It was found that there is no significant difference between selected attributes for KC1 after applying feature selection methods. There were only six attributes selected for CM1, which is less than the best first method. The number of selected attributes for JM1 is increased by one compared to the best first as well. Since no noticeable differences observed in performance evaluation metrics after building prediction model based on CFS, random search, we do not show the results in this section. However, the remaining attributes from JM1, KC1 and CM1 after applying CFS, random search are presented as follows:
- CM1::
-
loc, iv(g), i, b, LOComment, uniq_Op
- KC1::
-
v, d, i, LOCode, LOComment, LOBlank, Uniq_Opnd, branchcout
- JM1::
-
loc, v(g), ev(g), iv(g), n, i, LOComment, LOBlank, locCodeAndComment
6.4 Experiment 4
As we noted earlier, each of these datasets has a different rate of defected data; 19 % of the JM1, 10 % of the CM1 and 15 % of KC1 are defected. So in this experiment, the rate of defected data was doubled in CM1 to identify whether any of the classifiers shows any distinctive changes in results compared to the previous trials; this experiment uses 21 method-level metrics to answer the research question 6. We show the results in Table 11.
According to Table 11, the accuracy rate of all algorithms is decreased except for Immunos1 and Immunos99. Therefore, it means that the findings could be a challenging fact because it shows that the performance of each classifier could tightly related to defect rate in the datasets. So previously suggested classifiers may not be the best choices anymore to build a prediction model when the defect rate is changed. Accuracy in almost all the algorithms is fallen down, but if we consider AUC for evaluation, we see that this value grows in two other algorithms that are basically from one category, GLONALG and CSCA [30]. It could be concluded that by increasing the defect rate, the artificial immune classifiers perform better and gives a finest prediction model compared to others.
7 Summary and conclusion
In this paper, we identified fault prediction algorithms based on different machine learning classifiers and distinct feature selection techniques. Since the accuracy rate is not a reliable metrics for performance evaluation, three other metrics were used, AUC, PD and PF which were not used together in other experiments before. If we consider high AUC and PD values along with low PF value as a well-performed benchmark, random forest performs best on both small and big datasets. According to [4], AIRSParallel performs better than the other AIS, but according to experiment 1 and 3, Immunos99 is the best among the other AIS classifiers. This study shows that applying different feature selection techniques does not have that much effect on the results; they mainly reduce the execution time. Experiment 4 shows that the prediction rate reduced in CM1 dataset when the defected data were doubled manually except for AIRSParallel, CLONALG and mostly CSCA, so it seems that when the rate of defected modules was increased, the mentioned AIS classifiers perform best among the others. We can conclude here that different kinds of feature selection and method-level metrics do not have a considerable effect on the performance of the algorithm, and the most important factor here is the type of algorithm itself; therefore, it is better to improve the algorithms to get better prediction results. In addition, building the model based on large datasets like JM1 or even smaller ones consumes lots of time when CSCA is used compared to the other algorithms but the results are relatively acceptable, especially when we consider AIS classifiers for building models. The results in this study show that AUC, PD and PF could be used as three performance evaluation metrics together for more reliable performance analysis.
Notes
Defects and faults have the same meaning in this paper.
np is the number of partitions.
References
Zheng, J.: Predicting software reliability with neural network ensembles. J. Expert Syst. Appl. 36, 2116–2122 (2009)
Dowd, M., MC Donald, J., Schuh, J.: The Art of Software Security Assessment: Identifying & Preventing Software Vulnerabilities. Addison-Wesley, Boston (2006)
Clark, B., Zubrow, D.: How Good is the Software: A Review of Defect Prediction Techniques. In: Software Engineering Symposium, Carreige Mellon University (2001)
Catal, C., Diri, B.: Investigating the effect of dataset size, metrics sets, and feature selection techniques on software fault prediction problem. Inf. Sci. 179(8), 1040–1058 (2009)
Xie, X., Ho, J.W.K., Murphy, C., Kaiser, G., Xu, B., Chen, T.Y.: Testing and validating machine learning classifiers by metamorphic testing. J. Syst. Softw. 84, 544–558 (2011)
Koksal, G., Batmaz, I., Testik, M.C.: A review of data mining applications for quality improvement in manufacturing industry. J. Expert Syst. Appl. 38, 13448–13467 (2011)
Hewett, R.: Minig Software defect Data to Support Software testing Management. Springer Science + Business Media, LLC, Berlin (2009)
Catal, C., Diri, B.: A systematic review of software fault prediction. J. Expert Syst. Appl. 36, 7346–7354 (2009)
Catal, C.: Software fault prediction: a literature review and current trends. J. Expert Syst. Appl. 38, 4626–4636 (2011)
Evett, M., Khoshgoftaar, T., Chien, P., Allen, E.: GP-based software quality prediction. In: Proceedings of the Third Annual Genetic Programming Conference, San Francisco, CA, pp. 60–65 (1998)
Koprinska, I., Poon, J., Clark, J., Chan, J.: Learning to classify e-mail. Inf. Sci. 177(10), 2167–2187 (2007)
Thwin, M.M., Quah, T.: Application of neural networks for software quality prediction using object-oriented metrics. In: Proceedings of the 19th International Conference on Software Maintenance, Amsterdam, The Netherlands, pp. 113–122 (2003)
Menzies, T., Greenwald, J., Frank, A.: Data mining static code attributes to learn defect predictors. IEEE Trans. Softw. Eng. 33(1), 2–13 (2007)
El Emam, K., Benlarbi, S., Goel, N., Rai, S.: Comparing case-based reasoning classifiers for predicting high risk software components. J. Syst. Softw. 55(3), 301–320 (2001)
Yuan, X., Khoshgoftaar, T.M., Allen, E.B., Ganesan, K.: An application of fuzzy clustering to software quality prediction. In: Proceedings of the Third IEEE Symposium on Application-Specific Systems and Software Engineering Technology. IEEE Computer Society, Washington, DC (2000)
Catal, C., Diri, B.: Software fault prediction with object-oriented metrics based artificial immune recognition system. In: Proceedings of the 8th International Conference on Product Focused Software Process Improvement. Lecture Notes in Computer Science, pp. 300–314. Springer, Riga (2007)
Catal, C., Diri, B.: A fault prediction model with limited fault data to improve test process. In: Proceedings of the Ninth International Conference on Product Focused Software Process Improvements. Lecture Notes in Computer Science, pp. 244–257. Springer, Rome (2008)
Catal, C., Diri, B.: Software defect prediction using artificial immune recognition system. In: Proceedings of the Fourth IASTED International Conference on Software Engineering, pp. 285–290. IASTED, Innsburk (2007)
Zhang, H., Zhang, X.: Comments on data mining static code attributes to learn defect predictors. IEEE Trans. Softw. Eng. (2007)
Menzies, T., Dekhtyar, A., Di Stefano, J., Greenwald, J.: Problems with precision: a response to comments on data mining static code attributes to learn defect predictors. IEEE Trans. Softw. Eng. 33(7), 637–640 (2007)
Koru, G., Liu, H.: Building effective defect prediction models in practice. IEEE Softw. 22(6), 23–29 (2005)
Shafi, S, Hassan, S.M., Arshaq, A., Khan, M.J., Shamail, S.: Software quality prediction techniques: a comparative analysis. In: Fourth International Conference on Emerging Technologies, pp. 242–246 (2008)
Turhan, B., Bener, A.: Analysis of Naïve Bayes assumption on software fault data: an empirical study. Data Knowl. Eng. 68(2), 278–290 (2009)
Alsmadi, I., Najadat, H.: Evaluating the change of software fault behavior with dataset attributes based on categorical correlation. Adv. Eng. Softw. 42, 535–546 (2011)
Sandhu, P.S., Singh, S., Budhija, N.: Prediction of level of severity of faults in software systems using density based clustering. In: 2011 IEEE International Conference on Software and Computer Applications. IPCSIT, vol. 9 (2011)
Turhan, B., Kocak, G., Bener, A.: Data mining source code for locating software bugs; a case study in telecommunication industry. J. Expert Syst. Appl. 36, 9986–9990 (2009)
Brownlee, J.: Artificial immune recognition system: a review and analysis. Technical Report 1–02, Swinburne University of Technology (2005)
Watkins, A.: A Resource Limited Artificial Immune Classifier. Master’s thesis, Mississippi State University (2001)
Watkins, A.: Exploiting immunological metaphors in the development of serial, parallel, and distributed learning algorithms. PhD thesis, Mississippi State University (2005)
Brownlee, J.: Clonal selection theory & CLONALG. The clonal selection classification algorithm. Technical Report 2–02, Swinburne University of Technology (2005)
Watkins, A., Timmis, J., Boggess, L.: Artificial Immune Recognition System (AIRS): An Immune-Inspired Supervised Learning Algorithm. Genetic Programming and Evolvable Machines, vol. 5, pp. 291–317 (2004)
http://users.rcn.com/jkimball.ma.ultranet/BiologyPages/C/ClonalSelection.html. Retrieved 1 Nov 2013
Brownlee, J.: Immunos-81—The Misunderstood Artificial Immune System. Technical Report 3–01. Swinburne University of Technology (2005)
Langley, P.: Selection of relevant features in machine learning. In: Proceedings of the AAAI Fall Symposium on Relevance. AAAI Press, California (1994)
Khoshgoftaar, T.M., Seliya, N., Sundaresh, N.: An empirical study of predicting software faults with case-based reasoning. Softw. Qual. J. 14(2), 85–111 (2006)
Malhi, A.: PCA-Based feature selection scheme for machine defect classification. IEEE Trans. Instrum. Meas. 53(6) (2004)
Kohavi, R., John, G.: Wrappers for feature subset selection. Artif. Intell. Special Issue Relev. 97(1–2), 273–324 (1996)
Hall, M.A.: Correlation-based Feature Subset Selection for Machine Learning. PhD dissertation, Department of Computer Science, University of Waikato (1999)
http://promise.site.uottawa.ca/SERepository/datasets. Retrieved 01 Dec 2011
http://promisedata.org/?cat=5. Retrieved 01 Dec 2011
Rakitin, S.: Software Verification and Validation for Practitioners and Managers, 2nd edn. Artech House, London (2001)
Shepperd, M., Ince, D.: A critique of three metrics. J. Syst. Softw. 26(3), 197–210 (1994)
Fenton, N.E., Pfleeger, S.: Software Metrics: A Rigorous and Practical Approach. Int’l Thompson Press, New York (1997)
http://www.cs.waikato.ac.nz/ml/weka. Retrieved 01 Nov 2011
http://promisedata.org/repository/data/pc3/pc3.arff. Retrieved 01 Dec 2011
Acknowledgments
The authors thank to Universiti Teknologi Malaysia (UTM) for some of the facilities and supports during the course of this research under vot 03H02. The Ministry of Science, Technology & Innovation (MOSTI) is also acknowledged for supporting the research under vot 4S062. The authors wish to thank the anonymous reviewers for their comments in improving the manuscript.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
Open Access This article is distributed under the terms of the Creative Commons Attribution 2.0 International License (https://creativecommons.org/licenses/by/2.0), which permits unrestricted use, distribution, and reproduction in any medium, provided the original work is properly cited.
About this article
Cite this article
Abaei, G., Selamat, A. A survey on software fault detection based on different prediction approaches. Vietnam J Comput Sci 1, 79–95 (2014). https://doi.org/10.1007/s40595-013-0008-z
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s40595-013-0008-z