Abstract
To retrospectively evaluate the successful test rate and performance of non-invasive prenatal screening (NIPS) for aneuploidies and microdeletions with international transportation of samples. Blood samples from Iberian women with singleton pregnancies were sent to a US laboratory for NIPS for aneuploidy and microdeletion syndromes (22q11.2, 1p36, Cri-du-chat, Prader Willi and Angelman). The NIPS methodology involved the analysis of single nucleotide polymorphisms in cell-free DNA in maternal plasma. Women with high-risk results were offered karyotyping and/or microarray confirmatory studies. Based on 14,175 women with successful testing (98.76% of all referrals), the overall test positive rate was 2.37% (1.9% for aneuploidy and 0.47% for microdeletion syndromes). Based on cases with known outcome, the positive predictive values (PPVs) were: for trisomy 21, 98.6%; trisomy 18, 85.7%; trisomy 13, 71.4%; monosomy-X, 87.5%; other sex chromosome aneuploidies, 100%; 22q11.2 deletion, 15.4%; and other microdeletions combined, 20%. With a protocol change that involved selective use of resequencing at a higher depth of read, the PPV for 22q11.2 deletion increased to 33.3 and 75% for the other microdeletions. Effective NIPS for both aneuploidies and select microdeletion syndromes can be provided even when this involves international transportation of blood specimens.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Clinical trials for non-invasive prenatal screening (NIPS) through the analysis of cell free-DNA in maternal plasma have established this testing as the most effective approach in screening for fetal trisomies 21, 18 and 13 and monosomy-X [1]. Reports from reference laboratories have been broadly consistent with the estimates of detection rates (DRs) and false-positive rates (FPRs) established in these trials [2,3,4,5,6]. Additionally, both proof of principle [7,8,9] and data from reference laboratories [10, 11] have indicated that screening is possible for select sets of microdeletion syndromes.
Various models are emerging for the delivery of NIPS in actual practice [12]. This includes offering NIPS as a primary screen for all women who formerly received conventional serum and ultrasound based screening, selective use for those who are at high prior risk, or as a contingent-based test for women who are determined to be at high-risk based on conventional screening tests. Economic considerations and the extent of the established infra-structure for conventional screening are also important determinants for the utilization of NIPS. Because the costs of establishing a NIPS laboratory are relatively high and aspects of the technology are proprietary, local availability of testing is largely being met by referral to large reference laboratories. For many countries, this often requires international transportation of the plasma samples. Because degradation of maternal cells or other sample handling issues could materially affect achieving a result, [13] it is useful to document the experience with international shipping in actual clinical practice.
Laboratorio Echevarne is a network of laboratories in Spain that provides a broad range of services including molecular and cytogenetic analysis. To meet the need for NIPS, Echevarne contracted with a US laboratory, Natera, Inc. (San Carlos, CA, USA). Maternal plasma samples are sent to California for the testing which is based on the analysis of single nucleotide polymorphisms (SNPs) in the cell-free DNA [14]. The aim of this study was to document test success rate and, secondarily, to evaluate initial performance with this model. As far as authors know, this is the first paper to provide a NIPS actual clinical experience on unselected test referrals that includes both autosomal trisomy, sex chromosome trisomy and microdeletions on the same population.
Materials and Methods
From February 2013 to April 2016, inclusive, women throughout Spain and also in Lisbon, Portugal, were offered NIPS through a network of 45 local Echevarne laboratories. NIPS was offered for singleton pregnancies for trisomies 21, 18, 13 and sex chromosome aneuploidy with the option to also receive testing for 22q11.2 microdeletion syndrome or a panel of microdeletion syndromes (22q11.2 deletion, 1p36 deletion, Cri-du-chat, Prader Willi, and Angelman syndromes). Peripheral blood specimens were collected and sent by 2-day air transportation to the Natera laboratory without plasma separation prior to shipment. Laboratory procedures were as described elsewhere [7, 11, 15,16,17,18,19,20]. Following the NIPS analyses, all patient reports were communicated electronically to Echevarne and to the individual referring physicians and clinics. The cases included in this study did not include any that have been previously reported in an evaluation of the SNP-based NIPT for fetal aneuploidy [3]. However, a minority of the cases receiving microdeletion NIPT have been included in prior reports (from February 2014 to February 2015) [11, 20, 21].
Cytogenetic analysis and/or chromosome microarray (750K Cytoscan array, Affymetrix, Santa Clara, CA, USA) through Echevarne’s laboratory was recommended for all patients with positive NIPS. Pregnancy outcome information for women with high risk calls was gathered from the genetics laboratory and information provided by referring physician offices.
Following the reporting of cases with a high risk for a microdeletion syndrome, a test modification was introduced that was expected to significantly reduce the FPR [11, 20, 21]. This modification involved a change in the confidence level needed to call a high risk result and reflex resequencing of those microdeletion tests that were positive at the standard, low, depth of read (LDOR) with the resequencing carried out at a higher depth of read (HDOR) [20]. Only those test-positive samples where the deletion was assigned to the maternally inherited chromosome were reflexed; those with a suspected paternally inherited deletion could be identified with higher confidence based on the LDOR results alone. The results of these additional studies were not reported to patients.
The study was exempted from institutional review board approval (Ethical & Independent Review Services, Corte Madera, CA, USA; Study ID 14064–01).
Results
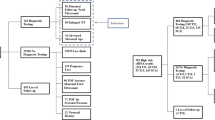
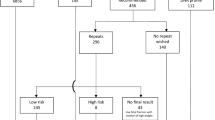
Fourteen thousand three hundred fifty three cases were referred for NIPS for fetal aneuploidy (Fig. 1). The mean age of the women tested was 36.6 years at the time of testing and the mean gestational age was 13.5 weeks. NIPT results were obtained for 14,175 (98.76%) of women; 178 (1.24%) did not receive a result. Six hundred eighty one (4.74%) required a second blood draw to obtain a result. Requests for a redraw were primarily due to low fetal fraction (582/681) (85%). Of the final 178 (1.24%) cases without results, 79 (44%) had a test cancellation after the first sample failed to provide a result. Excluding sample transportation time, the mean turn-around time was 6.49 calendar days (4.42 working days).
Table 1 summarizes the results for fetal aneuploidy. In total, 270 (1.9%) were high risk for fetal aneuploidy. Results of confirmatory cytogenetic or molecular cytogenetic studies were available for 105 (39%) cases. This included 15 cases where there was a spontaneous abortion (nine trisomy 21, two trisomy 18, three trisomy 18 and one monosomy-X). In the 165 cases where there were no documented confirmatory genetic studies carried out, referring physicians were contacted and there were no reports of discordancy between NIPT result and fetal karyotype. Under a conservative assumption that false-positives were in fact present in these cases at the same rate as that in cases with documented laboratory test confirmation, the FPR for autosomal trisomy (21, 18 and 13 combined) was 0.08%. Boundary estimates for the FPR, assuming the unknown outcome cases were either all true positives or all false positives, were 0.02–1.1%. Similarly, for sex chromosome aneuploidy (monosomy X, XXX, XYY and XXY combined), the FPR was 0.01% with boundaries of < 0.01–0.11%. The positive predictive value (PPV) for autosomal trisomy was 81/85 or 95.3% {95% CI 88.2–98.5%] and for sex chromosome aneuploidy the PPV was 19/20 or 95.0% [95% CI 74.6–100%].
Outcome information was not routinely collected for screen-negative cases. However, four false-negative cases for trisomy 21 and one false-negative for trisomy 13 were reported back to the laboratory. Maximum estimates for the DRs for trisomies 21, 18 and 13 were therefore 191/195 (98%), 24/24 (100%) and 17/18 (94%), respectively. A maximum estimate for the negative predictive value for autosomal trisomy was 13,303/13,308 or 99.96%. No cases of fetal sex inaccuracy were reported.
A total of 6831 women received testing for microdeletion syndromes (33 patients for 22q11.2 alone and 6798 patients for the full panel of disorders). Overall, 32 (0.47%) women received a high-risk call for a microdeletion. Table 2 summarizes the results of this component of the screening.
For the 22q11.2 microdeletion screening, there were 2 true positives confirmed by microarray, 11 false-positives, and 3 lost to follow-up. Boundary estimates for the FPR (assuming the 3 unknown cases were either all true positives or all false positives) were 0.16–0.20%. The PPV was 15.4% [95% CI 2.7–46.3%]. Of the 16 cases that were high risk for a 22q11.2 deletion, only 2 were known to have high prior risks (based on factors other than maternal age). One had increased nuchal translucency and one had abnormal maternal serum screening results.
For the other four microdeletion syndromes combined, there were 3 true-positives, 12 false-positives and one with no follow-up information providing a FPR of 0.19% with boundary values of 0.18–0.19% and a PPV of 20% [95% CI 5.3–48.6%]. Of the 16 cases that were high risk, only one was known to have a high prior risk indication (abnormal maternal serum screening result). The confirmatory analyses in the 1p36 false-positive case and one PWS false-positive case showed loss of heterozygosity (“haploblocks”) in the critical regions. There was one verbal report of a false-negative report for Cri-du-chat syndrome; however, the specifics of the clinical and diagnostic confirmatory testing were not provided.
The revised protocol involving the use of HDOR were carried out on samples that were considered high-risk for a maternally inherited microdeletion based on LDOR (Table 3). Two true-positive cases that were considered high-risk for 22q11.2 at LDOR remained high-risk with the revised HDOR protocol. In contrast, of the 11 cases that were established as false-positives by LDOR testing, only 4 remained positive with the revised protocol. The PPV based on the revised protocol was increased to 33.3% [95% CI 6.0–75.9%]. For the other microdeletion syndromes combined, no true-positives but 10 of 11 false-positives were removed when the HDOR protocol was applied (one sample was not available for analysis). This resulted in a revised PPV of 75% [95% CI 30.1–95.4%] for this group of microdeletion syndromes.
No cases were high risk for both an aneuploidy and a microdeletion syndrome. Combining all aneuploidies and microdeletions, the test positive rate was 2.37% (1.9% for aneuploidy and 0.47% for microdeletion syndromes) with an overall positive predictive value of 87.1% [95% CI 82.8–90.4%]. This would be revised to a test positive rate of 2.1% and PPV of 93.6% with the use of the revised protocol.
Discussion
This paper documents a retrospective experience with NIPS offered by a Spanish laboratory network with referral of maternal blood specimens to a laboratory in the US. The authors show that this can be an effective solution to the provision of service when a local NIPS laboratory is not available. Follow-up information indicated that test performance was similar to previous reports, and overall test failure rates were low (1.24%). Although transportation of specimens did add to the turn-around time, this was found to be manageable.
Studies that retrospectively assess NIPS experience are important in determining the overall value of the testing. Professional guidelines emphasize the importance of confirming positive screening results [22] and the PPV in a clinical cohort can be evaluated from the screen-positive cases with this outcome information. PPV is prevalence dependent and some standardization to a population incidence may be required [23]. In this study, the frequency of trisomy 21 was approximately 1.4%, which was somewhat higher than the approximately 1% that would be expected based solely on maternal age (mean 36.6 years) and gestational age (mean 13.5 weeks) [24, 25]. FPR can also be calculated under the assumption that the cases without follow-up information are not materially different from those cases with follow-up in terms of the proportions of true- and false-positives. In this context, authors note that the FPRs and PPVs in this study compare favorably with those documented for 17,885 women with singleton pregnancies receiving the same SNP-based NIPS for autosomal trisomies and monosomy-X [3]. The results of the screening for microdeletion syndromes are also consistent with prior studies that have evaluated performance of SNP-based screening [11, 20]. However, a minority of this cohort of test microdeletion referrals did overlap with those previously published.
A major difficulty in clinical experience studies has been that confirmation studies have been substantially incomplete [2,3,4, 11] and this study also has incomplete follow-up data. Low follow-up rates can be attributable to multiple factors including referral of confirmatory testing to different laboratories and incomplete documentation. Autosomal aneuploidies are associated with high spontaneous abortion rates and many such cases do not receive a formal confirmatory cytogenetic analysis. Other cases are associated with major fetal anatomic abnormalities visualized by ultrasound and these cases may be terminated without the recommended additional cytogenetic testing. On the other hand, milder conditions such as some sex chromosome abnormalities may not be apparent at birth and any additional cytogenetic studies may be deferred. Overall DRs cannot be established because it is not possible to obtain comprehensive follow-up for all low-risk results for all conditions. False-negative results are often reported to laboratories and these do allow an estimation of the maximum DR. Finally, DR, FPR, and PPV of NIPS can only be assessed on cases with results and it is known that test failures due to low fetal fraction have higher frequencies of trisomy 18, trisomy 13 and triploidy [26]. The present results need to be interpreted in the context of these practical limitations.
As is the case for NIPS for aneuploidy, there can be diverse reasons for false-positive and false-negative results when screening for microdeletions. The authors observed two false-positive results in which there was homozygosity in the critical regions. The technology used is dependent on the presence of heterogeneity of the SNPs within the critical region. Therefore, the absence of heterogeneity was the likely reason for the discordant results.
Another important consideration in the screening of microdeletion syndromes is the possibility that a case might arise as a result of the presence of an unbalanced translocation. This was the case for one of the confirmed Cri-du-chat high risk result. On the other hand, a breakpoint within the critical region could result in only a partial loss and a false-negative screening result.
Rapid advances continue to made in NIPS technology facilitating expansion in the scope of testing as well as improvements in performance in screening for well-established disorders. One such recent improvement is the reflex use of HDOR. In this study, the authors further document that a revised protocol that incorporated HDOR can result in a substantial reduction in the FPR and higher PPV for microdeletion syndromes. Improvements such as this provide the opportunity to expand the scope of screening to include other clinically significant copy number variants.
In summary, this study documents highly effective prenatal screening for fetal chromosome abnormalities. It provides an independent validation of the SNP-based approach to NIPS.
References
Gil MM, Quezada MS, Revello R, Akolekar R, Nicolaides KH. Analysis of cell-free DNA in maternal blood in screening for fetal aneuploidies: updated meta-analysis. Ultrasound Obstet Gynecol. 2015;45:249–66.
Futch T, Spinosa J, Bhatt S, de Feo E, Rava RP, Sehnert AJ. Initial clinical laboratory experience in noninvasive prenatal testing for fetal aneuploidy from maternal plasma DNA samples. Prenat Diagn. 2013;33:569–74.
Dar P, Curnow KJ, Gross SJ, et al. Clinical experience and follow-up with large scale single-nucleotide polymorphism-based noninvasive prenatal aneuploidy testing. Am J Obstet Gynecol. 2014;211:527.e1–17.
Bianchi DW, Parsa S, Bhatt S, et al. Fetal sex chromosome testing by maternal plasma DNA sequencing: clinical laboratory experience and biology. Obstet Gynecol. 2015;125:375–82.
Taneja PA, Snyder HL, de Feo E, et al. Noninvasive prenatal testing in the general obstetric population: clinical performance and counseling considerations in over 85,000 cases. Prenat Diagn. 2016;36:237–43.
Zhang H, Gao Y, Jiang F, et al. Non-invasive prenatal testing for trisomies 21, 18 and 13: clinical experience from 146,958 pregnancies. Ultrasound Obstet Gynecol. 2015;45:530–8.
Wapner RJ, Babiarz JE, Levy B, et al. Expanding the scope of noninvasive prenatal testing: detection of fetal microdeletion syndromes. Am J Obstet Gynecol. 2015;212:332.e1–9.
Zhao C, Tynan J, Ehrich M, et al. Detection of fetal subchromosomal abnormalities by sequencing circulating cell-free DNA from maternal plasma. Clin Chem. 2015;61:608–16.
Lefkowitz RB, Tynan JA, Liu T, et al. Clinical validation of a noninvasive prenatal test for genomewide detection of fetal copy number variants. Am J Obstet Gynecol. 2016;215:227.e1–16.
Helgeson J, Wardrop J, Boomer T, et al. Clinical outcome of subchromosomal events detected by whole-genome noninvasive prenatal testing. Prenat Diagn. 2015;35:999–1004.
Gross SJ, Stosic M, McDonald-McGinn DM, et al. Clinical experience with single-nucleotide polymorphism-based non-invasive prenatal screening for 22q11.2 deletion syndrome. Ultrasound Obstet Gynecol. 2016;47:177–83.
Cuckle H, Benn P, Pergament E. Cell-free DNA screening for fetal aneuploidy as a clinical service. Clin Biochem. 2015;48:932–41.
Wong D, Moturi S, Angkachatchai V, et al. Optimizing blood collection, transport and storage conditions for cell free DNA increases access to prenatal testing. Clin Biochem. 2013;46:1099–104.
Zimmermann B, Hill M, Gemelos G, et al. Noninvasive prenatal aneuploidy testing of chromosomes 13, 18, 21, X, and Y, using targeted sequencing of polymorphic loci. Prenat Diagn. 2012;32:1233–41.
Pergament E, Cuckle H, Zimmermann B, et al. Single-nucleotide polymorphism-based noninvasive prenatal screening in a high-risk and low-risk cohort. Obstet Gynecol. 2014;124:210–8.
Nicolaides KH, Syngelaki A, Gil M, Atanasova V, Markova D. Validation of targeted sequencing of single-nucleotide polymorphisms for non-invasive prenatal detection of aneuploidy of chromosomes 13, 18, 21, X, and Y. Prenat Diagn. 2013;33:575–9.
Nicolaides KH, Syngelaki A, del Mar Gil M, Quezada MS, Zinevich Y. Prenatal detection of fetal triploidy from cell-free DNA testing in maternal blood. Fetal Diagn Ther. 2014;35:212–7.
Samango-Sprouse C, Banjevic M, Ryan A, et al. SNP-based non-invasive prenatal testing detects sex chromosome aneuploidies with high accuracy. Prenat Diagn. 2013;33:643–9.
Ryan A, Hunkapiller N, Banjevic M, et al. Validation of an enhanced version of a single-nucleotide polymorphism-based noninvasive prenatal test for detection of fetal aneuploidies. Fetal Diagn Ther. 2016;40:219–23.
Martin K, Iyenger S, Kalyan A, et al. Clinical experience with a single-nucleotide polymorphism-based noninvasive prenatal test for five clinically significant microdeletions. Clin Genet. 2017. https://doi.org/10.1111/cge.13098.
Gross SJ, Ryan A, Benn P. Noninvasive prenatal testing for 22q11.2 deletion syndrome: deeper sequencing increases the positive predictive value. Am J Obstet Gynecol. 2015;213:254–5.
Benn P, Borrell A, Chiu RW, et al. Position statement from the chromosome abnormality screening committee on behalf of the board of the international society for prenatal diagnosis. Prenat Diagn. 2015;35:725–34.
Benn P. Expanding non-invasive prenatal testing beyond chromosomes 21, 18, 13, X and Y. Clin Genet. 2016;90:477–85.
Morris JK, Mutton DE, Alberman E. Revised estimates of the maternal age specific live birth prevalence of Down’s syndrome. J Med Screen. 2002;9:2–6.
Savva GM, Morris JK, Mutton DE, Alberman E. Maternal age-specific fetal loss rates in Down syndrome pregnancies. Prenat Diagn. 2006;26:499–504.
Revello R, Sarno L, Ispas A, Akolekar R, Nicolaides KH. Screening for trisomies by cell-free DNA testing of maternal blood: consequences of a failed result. Ultrasound Obstet Gynecol. 2016;47:698–704.
Acknowledgements
The authors thank Zach Demko (Natera, Inc.) for facilitating high depth of read analysis and some editorial input.
Funding
This study was supported by Laboratorio Echevarne. The resequencing research component was funded by Natera, Inc.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
RS, BB, and SC are employees of Laboratorio Echevarne. PB is a consultant to Natera, Inc.
Rights and permissions
Open Access This article is distributed under the terms of the Creative Commons Attribution 4.0 International License (http://creativecommons.org/licenses/by/4.0/), which permits unrestricted use, distribution, and reproduction in any medium, provided you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made.
About this article
Cite this article
Santamaria, R., Bermejo, B., Cigarrán, S. et al. A National Referral Laboratory’s Experience with the Implementation of SNP-Based Non-invasive Prenatal Screening for Fetal Aneuploidy and Select Microdeletion Syndromes. J. Fetal Med. 5, 7–12 (2018). https://doi.org/10.1007/s40556-017-0143-1
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s40556-017-0143-1