Abstract
Tomato yellow leaf curl virus (TYLCV) is one of the most destructive pathogens for tomato crops. The development of TYLCV-tolerant tomato lines (TLs) requires a thorough understanding of their genetic variability and relationships among lines and in traits of interest. Twelve F9 TLs were evaluated for phenotypic TYLCV tolerance, vegetative growth, yield, and fruit quality during the 2018 and 2019 fall seasons to identify elite breeding lines. TLs were selected by a bulk selection method from segregating generations of the commercial F1 hybrids TH99802 (TLs 1–6) and TH99806 (TLs 7–12). TLs exhibited either mild or no symptoms. The TYLCV titer varied between 0.8 × 105 and 3.9 × 105 in symptomless TLs plants compared to 56.7 × 105 in severely symptomatic plants of susceptible ‘Castlerock.’ Across both seasons, TL-3, TL-5, and TL-8 exhibited the highest total and marketable plant yields, TYLCV tolerance, and acceptable fruit quality. Most traits had high estimates of genetic variance, genotypic coefficient of variance, and broad sense heritability. Our results indicated that there was sufficient genetic variability for selection of the best lines. Principal component analysis and hierarchical cluster analysis indicated that the TLs were highly diverse of the evaluated traits and could be divided into three clusters. Cluster 1, which included TL-1, TL-3, and TL-8, performed better for TYLCV tolerance and economically important traits. Clusters 1 and 2 showed the greatest degree of dissimilarity. Therefore, crossing parents from Cluster 1 with 2 is predicated to maximum recombination for improve genotypes.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
1 Introduction
Tomato (Solanum lycopersicum L.) is an important vegetable crop. Egypt is the fifth largest producer worldwide, with 6,245,787 tons produced in 2021 from 357,259.4 fedans, averaging 17.5 tons fed−1 (< http://faostat.fao.org >). Tomato yellow leaf curl disease (TYLCD) is one of the world’s most devastating diseases for tomato producers (Lefeuvre et al. 2010; Moriones and Navas-Castillo 2000). TYLCD symptoms include yellowing, curling, and cupping of leaves, severe stunting and abortion of flowers and fruits, which can result in yield reduction of up to 100% (Abhary et al. 2007). Tomato yellow leaf curl virus (TYLCV) is the causative agent of TYLCD and belongs to the genus Begomovirus in the family Geminiviridae. TYLCV is a monopartite DNA virus with a circular genome that contains six genes, with two genes on the viral strand (V1 and V2) and four genes on the complementary sense strand (C1 to C4) (Abhary et al. 2007; Gronenborn 2007). TYLCV is exclusively transmitted in a persistent-circulative manner by the sweet potato whitefly, Bemisia tabaci (Genn. 1889) (Homoptera; Aleyrodidae) (Gronenborn 2007). B. tabaci is an invasive pest found in over 175 countries worldwide (Ramos et al. 2018). The virus has spread as a result of the tremendous proliferation of the whitefly globally. The number of tomato-producing regions reporting TYLCD pandemic breakouts has increased to up to 70 nations since its discovery in the Jordan Valley in 1930s (Mabvakure et al. 2016).
TYLCVD control is a laborious, costly, and difficult task. The primary TYLCD control strategy is the use of insecticides to eradicate the B. tabaci viral vector (Lapidot et al. 2014). However, insecticides treatment can be expensive and environmentally risky. Furthermore, because B. tabaci develops resistance to insecticides, they can become ineffectual (Palumbo et al. 2001). The simplest, safest, most practical, and ecologically friendly approach to controlling TYLCD, minimizing yield losses, and reducing viral transmission is to use TYLCD-resistant cultivars. As a result, TYLCD resistance has become one of the most important goals in tomato breeding.
Initially, the cultivated tomato lacked TYLCD resistance (Hassan et al. 2009). Various accessions of wild tomato species were found to be TYLCD resistant (Ji et al. 2007; Hassan et al. 2009; Yan et al. 2018). TYLCD resistance has been successfully introgressed into the cultivated tomato from resistant wild relatives including S. chilense (Dunal) Reiche, S. habrochaites S. Knapp & D.M Spooner, S. pennellii Correll, S. peruvianum L., and S. pimpinellifolium L. (Vidavski 2007; Singh et al. 2019). As a result, several TYLCD-tolerant cultivars/F1 hybrids have been released for commercial cultivation by global seed companies (Vidavski 2007; Vidavski et al. 2008; Dhaliwal et al. 2020).
The continuous development of new tomato lines/cultivars is important to improve TYLCD resistance/tolerance, productivity, and to overcome virulence genes developed by the virus. The selection and development of elite TYLCD-resistant/tolerant lines require a thorough understanding of the genetic variability and relationships among lines and between traits of interest (Lynch and Walsh 1998; Bernardo 2010). Plant breeders use morphological traits to estimate genetic variability because they are simple to score, rapid to analysis, and inexpensive assessment.
The phenotypic (PCV) and genotypic (GCV) coefficients of variation and heritability are essential biometric tools used to assess the genetic variability among the genotypes (Singh and Chaudhary 1985; Kumar et al. 2013). Heritability governs both the potential for improvement through selection and the effect of the environment on the expression of traits (Robinson et al. 1949). According to Burton and De Vane (1953), combining GCV with h2b estimates produces a trustworthy measure of the projected level of potential improvement via selection.
Traits correlation determines how well genetic variability may be exploited by selection. Depending on the correlation, selection for one trait may either increase or decrease the expression of another (Bernardo 2010). Phenotypic correlation is the result of the interplay between genetic and environmental factors. Genetic correlation is the only method used to direct breeding, since it is the only aspect of inheritable nature (Lynch and Walsh 1998).
Several statistical techniques can be used to estimate genetic variability and the levels of similarity/dissimilarity among genotypes (Mohammadi and Parasanna 2003). These analyses are extremely helpful for planning crossing, allocating lines to specific heterotic groupings, and precise identification with regard to plant variety preservation (Hallauer et al. 2010). Since data from morphological characterization are collected from a sizable dataset containing both qualitative and quantitative traits, multivariate analyses are highly suited to the classification and ranking of genotypes. Furthermore, genetic variability has been estimated using multivariate analyses (Mohammadi and Prasanna 2003). Principal component analysis (PCA) and cluster analysis are the best multivariate approaches for morphological characterization of genotypes (Mohammadi and Prasanna 2003; Reich et al. 2008).
This study was conducted to inform the development of TYLCV-tolerant lines. Twelve F9 tomato lines (TLs) were selected from a tomato breeding program and cultivated under TYLCV-infected conditions to evaluate their phenotypic TYLCD tolerance, vegetative growth, and fruit quality. Estimation of genetic variability, broad sense heritability, and genotypic correlations were performed, and the TLs were categorized and ranked using multivariate analyses in order to select the elite lines.
2 Materials and methods
2.1 Tomato lines
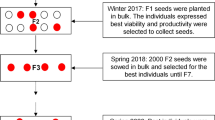
Tomato cultivar TH99802 (Yassamen F1, Syngenta) is characterized by a determinate growth habit; vigorous vegetative growth; medium-early ripening; fruit firmness; spherical fruits with an average weight 160–180 g; high tolerance to verticillium (V), fusarium (Fol 0–1), and Stemphylium (S/Ss); and moderate tolerance to TYLCD. The tomato cultivar ‘TH99806’ (Nirouz F1, Syngenta) is characterized by a determinate growth habit; vigorous vegetative growth; early ripening; heat tolerance; spherical fruits with average weight 130–150 g; tolerance to fruit cracking; highly tolerance to tobacco mosaic virus (ToMV0-2), verticillium (V), and fusarium (Fol1-2); and moderate tolerance to TYLCD. Based on these hybrids, a bulk selection program was organized for TYLCD resistance and high productivity at the Agricultural Experimental Station (AES), Faculty of Agriculture, Cairo University, Giza, Egypt (30°01′07′′N; 31°12′28′′E). Symptomless plants with vigor vegetative growth, average fruit weight > 80 g, and high fruit set were selected from segregated generations during the fall seasons. Twelve F9 lines were selected and evaluated their TYLCD tolerance, vegetative growth, yield, and fruit quality traits during the 2018 and 2019 fall seasons. Evaluation conducted at AES under TYLCD infection field conditions, where the climatic conditions are suitable for the widely flourishing of viruliferous whiteflies (Fig. 1).
Average monthly maximum and minimum temperatures and relative humidity during the period from July to December in the 2018 and 2019 fall seasons (https://power.larc.nasa.gov/data-access-viewer/)
2.2 Planting and experimental design
Seeds of TLs were sown on the 1 July of 2018 and 2019, in seedling trays (209 cells) filled with a mixture of peatmoss and vermiculite (volume 1:1) enriched with macro and microelements under greenhouse conditions. The greenhouse was covered with a black saran fabric with narrow holes to prevent the entry and exit of insects. Five-week-old seedlings were field-transplanted in a randomized complete block design (RCBD) (Singh and Choudhary 1985) with three replicates. Each experimental unit (EU) consisted of two rows/line. Each row was 1 m wide and 3 m long. Plants were set 50 cm apart and subjected to common agricultural practices without applying insecticides.
2.3 TYLCV inoculation
Whiteflies is flourishes in Egypt from April through November, with a peak from August to October (Abd-Rabou and Evans 2020). Therefore, viral inoculation depended on natural infestation with viruliferous whiteflies in both the nursery and field. To encourage TYLCV infection, highly symptomatic plants of the susceptible cultivar ‘Castlerock’ were grown in seedling greenhouse and had an abundance of whiteflies. A row of ‘Castlerock’ plants were cultivated between EUs as a source of TYLCV infection and a guide to TYLCV symptom severity.
2.4 TYLCD tolerance
Phenotypical TYLCD tolerance was evaluated based on the severity of TYLCD symptoms in TLs 3 months after transplanting (3MAT) in both seasons. Symptoms severity was assessed for individual plants of each line using a 1–5 scale as described by Mahmoud (2015): 1, no symptoms appear on the plant; 2, slight symptoms on plant top; 3, moderate symptoms; 4, severe symptoms on the entire plant; and 5, severe symptoms and plant stunting. The individual plant ratings were summed for each line and divided by the number of evaluated plants to calculate the TYLCD-mean score (TYLCD-MS).
TYLCV presence has been shown by polymerase chain reaction (PCR) using universal Begomovirus primers AVcore/ACcore (Brown et al. 2001) and the specific primers TYLC2C3F/TYLC2C3R (Table 1) for species TYLCV, TYLCV-Mld, TYLCVMalv, TYLCSV-ES[2], and TYLCSV, which are found throughout the Mediterranean region and Middle East (Anfoka et al. 2008). Total genomic DNA was isolated using the CTAB method (Murray and Thompson 1980) from the young leaves of both symptomless TLs and severely symptomatic ‘Castlerock’ plants at 3MAT. Five plants from each line were used to sample the young leaves. PCR cycle parameters are described in Table 1. All PCRs were performed in a programmable thermocycler (Mastercycler ep gradient 5, Eppendorf, Hamburg, Germany). The PCR products were resolved in 1.5% agarose gel in 1 × Tris–acetate-EDTA buffer. DNA bands visualized with ethidium bromide staining (0.5 μg mL−1) and photographed under UV light using gel documentation system (Bio-Rad® Gel Doc-2000). One-kb ladder DNA was used as the molecular weight size marker.
Vegetative, yield, and fruit quality traits
Vegetative traits, i.e., plant length (PL), number of plant leaves (NPL), and the area of the fifth leaf from the apex (LA), were measured on five randomly selected plants for each EU at 3MAT, excluding plants from row edges. The LA was measured using the leaf weighting method, as described by Pandey and Singh (2011). Yield [early (EY): the first three harvests; total (TY): all collected fruits; and marketable (MY): all normal collected fruits] and fruit quality traits [average fruit weight (AFW); fruit firmness (FF); fruit shape index (FSI); contents of total soluble solid content (TSS); and titratable acidity (TA)] were measured. Samples of 20 fully red-ripe fruits from each EU were harvested at the peak harvesting time, weighed to estimate AFW, and washed with distilled water to analyze fruit traits. FSI was calculated as the ratio between the polar and equatorial diameters of fruit according to Yeager (1937), where FSI is > 1.2 in oval fruits, 0.95–1.2 in round shape, and < 0.95 in oblate fruits. FF was determined using a food pressure tester (Force Gauge Model M4-200-Series 4; Mark-10 Corp., Copiague, NY, USA) Mark-10 (Series 4). Then, fruit extract was obtained by blending and filtering the flesh. TSS was determined using a hand refractometer. TA was ascertained using 0.1 N NaOH solution and phenolphthalein as indicators (AOAC 1990). The taste index (TI) and maturity (M) were also estimated as indications of tomato flavor and quality to assess consumer acceptance and distinguish between lines. The taste index [TI = (◦Brix/(20 × TA)) + TA] and the maturity (M = ◦Brix/TA) were calculated using equations described by Navez et al. (1999).
2.5 Biometrical analyses
Estimation of the evaluated TLs’ genetic diversity and their classification based on phenotypic traits was performed using the following statistical methods (Mohammadi and Prasanna 2003). Independent analysis of variance (ANOVA) is performed for each trait with estimate of PCV, GCV, phenotypic correlation coefficient (rph), genotypic correlation coefficient (rg), and h2b. Multivariate analyses were performed for all estimated traits using PCA and hierarchical cluster analysis (HCA).
ANOVA and ANCOVA
The collected phenotypic data were checked for the normality using the Shapiro—Wilk test (Shapiro and Wilk 1965), and data for TYLCD-MS, EY, TY, AFW, FSI, TA, TI, and M were arcsin square root transformed (Wickens and Keppek 2004). ANOVA of the RCBD was performed for each seasons according to Wickens and Keppek (2004). When Bartlett’s homogeneity test was nonsignificant, a combined ANOVA over the two seasons was also performed (Bartleet 1937). Significant differences between the combined means were calculated using the Duncan’s multiple range test at a 5% probability level (Duncan 1955). A combined analysis of covariance (ANCOVA) was also performed for the RCBD over the two seasons to estimate genotypic and phenotypic correlations between traits (Wickens and Keppek 2004). The ANOVA, ANCOVA, and mean comparisons were conducted using MSTATc v.2.1 (Michigan State University, Michigan, USA; Freed et al. 1989).
Estimation of phenotypic and genotypic variability and heritability
The components of variance attributable to differences among TLs were estimated by utilizing the mean squares (Supplementary Table 1; Singh and Chaudhary 1985). The genotypic (δ2g), phenotypic (δ2ph), genotype × year (δ2gy), and pooled error (δ2e) variances components were calculated according to Tessema et al. (2022) (Supplementary Table 1).The GCV and PCV were estimated according to Burton (1952) as follows: GCV = \((\sqrt{{\delta }_{\mathrm{g}}^{2}}/\overline{x })\times 100\) and PCV = \((\sqrt{{\delta }_{\mathrm{g}}^{2}}/\overline{x })\times 100\), where \(\overline{x }\) = grand mean of the trait. The GCV and PCV are classified as low (< 10%), moderate (10–20%), and high (> 20%) as suggested by Johnson et al. (1955). Broad sense heritability (h2b) for a trait was estimated according to Johnson et al. (1955), as follows: h2b = (δ2g/δ2ph) × 100. The h2b is classified as low (< 30%), moderate (30–60%), and high (> 60%).
Estimation of genotypic and phenotypic correlation coefficients
Covariances were computed in a similar manner as shown in Table 2 (Singh and Chaudhary 1985). These covariance components were substituted in the following formulae to compute rg and rph (Johnson et al. 1955): rg = \({\mathrm{COV}}_{g(x1x2)}/\sqrt{{\delta }_{g(x1)}^{2}{\delta }_{g(x2)}^{2}}\), where \({\mathrm{COV}}_{g(x1x2)}\) is the genotypic covariance between a given pair of traits (× 1 and × 2) and \({\delta }_{g(x1)}^{2}{\delta }_{g(x2)}^{2}\) are the genotypic variances of × 1 and × 2, respectively. rph = \({\mathrm{COV}}_{\mathrm{ph}(x1x2)}/\sqrt{{\delta }_{\mathrm{ph}(x1)}^{2}{\delta }_{\mathrm{ph}(x2)}^{2}}\), where COVph(x1x2) is the phenotypic covariance between two traits (× 1 and × 2) and \({\delta }_{\mathrm{ph}(x1)}^{2}\) and \({\delta }_{\mathrm{ph}(x2)}^{2}\) are phenotypic variances for each trait. Significance for rg and rph was performed according to Yassin (1973).
Multivariate analysis
The pooled data for all traits were standardized using Z-scores to avoid the effect of scale differences before the multivariate analysis. PCA with varimax rotation was applied (Sharma 1996). The latent root criterion (eigenvalue > 1) and parallel analysis were used to determine the number of statistically significant components (Johnson and Wichern 1988). The biplot between the top two PCs that adequately explained a significant percentage of the total variance was created to assist in identifying the relationships among PCs and traits, PCs and lines, lines and their traits, and among the different traits (Yan and Rajcan 2002; Yan and Kang 2003). The correlation coefficient between any two traits can be estimated using the cosine of the angle between the vectors (Yan and Kang 2003). The vectors are positively correlated if the angle between them is < 90°, negatively correlated if the angle is > 90°, and independent if the angle is exactly 90° (Yan and Rajcan 2002). HCA was performed to construct a dendrogram using the squared Euclidean distance and Ward’s joining method. All multivariate statistical analyses were performed using IBM SPSS software version 26.0.0 (SPSS Inc., Chicago, IL, USA; ÓConnor 2000) and XLSTAT software version 2019 (Addinsoft, Paris, France).
3 Results and discussion
3.1 ANOVA
Table 2 shows the combined ANOVA results of a RCBD for the 2018 and 2019 fall seasons for the estimated fourteen traits of the twelve evaluated TLs. There were no significant differences between years (MSy) for the estimated traits, except LA and AFW. These results demonstrated the environmental similarities between the two season and their influence on the evaluated traits (Falconer 1952). Meteorological information on minimum and maximum temperatures and air relative humidity (Fig. 1) support this result. There were significant differences in genotypes [MSg; P < 0.001 for all traits, except TI (P < 0.01) and M (P < 0.05)]. Also, genotype had the highest incidence (> 50%) on total pooled variance for all traits, except TA (22.2%). These findings showed that TLs had broad genetic variance, which may improve TYLCD tolerance and yield-related traits (Schouten et al. 2019; Hassan et al. 2022). Mean squares due to G × Y interaction (MSgy) were nonsignificant for the most of estimated traits (9 out of 14 traits). Significant differences with MSgy were seen in PL, TY, TA (P < 0.05), TSS (P < 0.01), and LA (P < 0.001). These findings indicated the genotype stability of TLs during both seasons (Singh 2001). Hence, emphasis was placed on determining the specific effects of genotypes and comparing their means for the estimated traits (Wickens and Keppek 2004).
3.2 Phenotypic performance
The phenotypic responses of the evaluated TLs under TYLCV infection for the TYLCV-mean score, vegetative, yield, and fruit quality traits are shown in Table 3.
TYLCD tolerance
TLs showed mild to moderate TYLCD symptoms at 3 MAT (Table 3). TYLCD-MS ranged between 1.15 for TL-1 and 1.82 for TL-12 across both seasons. The evaluated TLs could be divided into three groups according to significant differences in their symptom severity (TYLCD-MS). The first group had the lowest TYLCD-MS (1.15–1.34) and was represented by TL-1 to TL-3 and TL-5 to TL-8; the second group was only represented by TL-10 (1.52); and the third group had the highest TYLCD-MS (1.69–1.82) and was represented by TL-9, TL-11, and TL-12 (Table 4).
The presence of TYLCV-DNA was determined by a PCR assay in symptomless plants of TLs, as compared with the susceptible cultivar Castlerock (Fig. 2). Electrophoresis analysis revealed a single 550 bp fragment in all the DNA samples examined with the AVcore/ACcore universal primer pair (Fig. 2a). A 500-bp PCR product was produced in all of DNA samples using the specific primer pair TYLC2C3F/TYLC2C3R (Fig. 2b). These findings indicate TYLCD tolerance in TLs, where plants harbor viral DNA (Fig. 2) but exhibit either mild or no symptoms (Table 3).
Detection of TYLCV viral DNA in symptomless plants of tomato inbred lines grown in natural infectious field compared to TYLCV-infected plants of susceptible cultivar Castlerock using primers AVcore/ACcore (a) and TYLC2C3F/TYLC2C3R (b) in a PCR assay. Lanes M: 1 kbp DNA marker; 1–12: samples of tweleve tomato lines; and 13: sample of susceptible cultivar Castlerock
Vegetative traits
The results for vegetative growth traits (PL, NPL, and LA) for TLs at 3MAT are shown in Table 3. PL ranged from 0.63 m for TL-4 to 1.49 m for TL-3. TL-2 and TL-3 were the tallest plants (1.49 and 1.43 m, respectively) across both seasons, followed by TL-12, TL-11, and TL-8 (1.36, 1.29, and 1.27 m, respectively) with no significant differences between them. The shortest plants were TL-4, TL-5, and TL-9 (0.63, 0.70, and 0.73, respectively) with no significant differences among them (Table 3).
NPL generated by TLs at 3MAT ranged from approximately 37 in TL-9 to 54 in TL-8 across both seasons (Table 3). TL-8 had the highest NPL (54.3) across both seasons, followed by TL-3 (50.36), TL-2 (47.95), TL-12 (47.02), TL-11 (45.87), and TL-6 (45.66). NPL was lowest in TL-9 and TL-4 (36.91 and 39.21, respectively), with no significant differences between them (Table 4). The LA (cm2) of TLs at 3MAT ranged from 34.17 in TL-5 to 46.27 in TL-12 across both seasons (Table 3). TL-12 and TL-11 produced the largest LAs across both seasons (46.27 and 45.76, respectively). The smallest LAs were generated by TL-7, TL-5, and TL-4 (33.59, 34.17, and 35.01, respectively) across both seasons with no significant differences between them (Table 3).
Plant yield
EY (kg plant−1) for the TLs was between 0.36 in TL-9 and 0.93 in TL-8 across both seasons (Table 3). TL-8 had the greatest EY (0.93) across both seasons, followed by TL-3, TL-5, and TL-6 (0.79, 0.79, and 0.74, respectively) with no significant differences between them. TL-9 and TL-10 had the lowest EY (0.36 and 0.41, respectively) with no significant differences between them.
Across both seasons, TY (kg plant−1) values ranged from 1.94 in TL-7 to 2.67 in TL-5 (Table 3). The most promising lines for TY were TL-5 (2.67), TL-3 (2.67). TL-1, TL-7, TL-9, and TL-10 produced the lowest TY (1.97, 1.94, 1.98, and 2.03) with no significant differences from. MY (kg plant−1) ranged between 1.65 in TL-9 and 2.47 in TL-5 across both seasons (Table 3). TL-5, TL-3, and TL-8 yielded the greatest MY (2.47, 2.39, and 2.28 kg, respectively) with no significant differences among them. TL-9, TL-10, TL-1, and TL-7 yielded the lowest MY (1.65, 1.79, 1.84, and 1.81 kg, respectively) with no significant differences among them (Table 3).
Fruit quality traits
The AFW (g) of the TLs ranged from 70.82 in TL-5 to 126.02 in TL-3 across both seasons (Table 3). TL-3 and TL-8 yielded the highest AFW (126.02 and 124.95, respectively), followed by TL-1 (116.27), TL-12, and TL-11 (99.32 and 98.21, respectively). TL-5 and TL-4 had the lowest AFW (70.82 and 72.60, respectively; Table 3). Across both seasons, FF (kg cm-2) ranged from 0.34 in TL-5 to 0.69 in TL-2 (Table 3). TL-2 had the most FF, followed by TL-12 (0.60), TL-1 (0.58), TL-11 (0.53), TL-3 (0.52), and TL-7 (0.52). TL-5 (0.34), TL-3 (0.39), TL-9 (0.39), and TL-10 (0.41) had the lowest FF with no significant differences among them. FSI varied among TLs (Table 3). TL-2 and TL-9-TL-12 had oval fruits (FSI ranged from 1.31 to 1.53); TL-4-TL-7 had round fruits (FSI ranged from 0.97 to 1.11); and TL-1, TL-3, and TL-8 had oblate fruits (FSI ranged from 0.68 to 0.80) (Table 3).
Across both seasons, fruit TSS content (°Brix) ranged from 3.87 in TL-9 to 5.57 in TL-2 (Table 3). The highest TSS was seen in TL-2 (5.57), followed by TL-12 (5.17), TL-1 (4.91), TL-8 (4.89), TL-4 (4.79), TL-3 (4.76), and TL-11 (4.62) (Table 3). TA values (mg citric acid 100 g−1 FW) ranged from 0.36 in TL-8 to 0.47 in TL-12 across both seasons (Table 3). TL-12, TL-2—TL-7, and TL-9 -TL-11 showed the greatest TA. TL-8 and TL-1 had the lowest TA. TI and M were determined using the TSS and TA values. These indices are typically a more accurate indicator of a fruit’s flavor than just its TSS or acidity. TI across both seasons ranged from 0.89 in TL-9 to 1.14 in TL-2 (Table 3). TL-2 fruits had the highest TI (1.14), followed by the other lines (0.95–1.06), except for TL-9 (0.89), which was ranked third. The M values ranged from 9.11 in TL-9 to 14.35 in TL-2 (Table 3). The highest M values were observed in TL-2 (14.35), TL-8 (14.18), TL-1 (12.91), and TL-4 (11.89) with no significant differences. TL-9 and TL-3 had the lowest M values (9.11 and 9.81, respectively) with no significant differences among them (Table 3). Navez et al. (1999) stated that tomato fruits are deemed tasty with TI values > 0.7 and maturity > 10. Accordingly, the fruits of the evaluated TLs are considered to be good for fresh consumption.
According to the phenotypic evaluation results, lines TL-3, TL-5, and TL-8 had the highest TY, MY, and TYLCD tolerance, as well as acceptable fruit quality and AFW > 80 g, except TL-5, which had an AFW of approximately 71 g.
3.3 Heritability and phenotypic and genotypic variation
Tomato breeding aimed at selecting desired genotypes is linked with GCV and heritability estimates and other genetic parameters for important traits (Bernardo et al. 2010; Dhaliwal et al. 2020). The results for variability components (δ2g, δ2gy, δ2e, δ2ph, PCV, and GCV) and h2b for fourteen phenotypic traits of the evaluated TLs are presented in Table 2. These parameters are crucial for an efficient tomato breeding program for TYLCD tolerance (Falconer and Mackay 1996). The δ2g accounted for a greater proportion of the variability than δ2ph for the majority of traits and was 3.0–135.5 times that of δ2gy. PCV% values varied from 0.2 (AFW) to 31.8 (EY), whereas GCV% values ranged from 0.2% (AFW) to 31.6% (EY) (Table 5). EY, FF, and PL had high PCV% (31.8, 20.4, and 20.1, respectively). Medium PCV% were observed for FSI (19.0), M (19.0), MY (12.3), TYLCD-MS (11.8), LA (11.2), TSS (11.3), and NPL (10.9). Low PCV% were observed for TA (5.6), TY (4.6), TI (0.7), and AFW (0.2; Table 3). Only EY had a high GCV%, while TSS, TA, TY, TI, and AFW had low GCV% (6.7, 4.8, 3.9, 0.3, and 0.2, respectively). The other traits had medium GCV%, ranging from 10.2 (LA) to 19.9 (FF). High (> 20%) or moderate (10–20%) PCV and GCV suggest a high level of variability in EY, TYLCD-MS, PL, NPL, LA, MY, FSI, and M traits. High variability shows the potential for effective selection for trait improvement (Singh 2001). Low PCV and GCV values (< 10%) for TY, AFW, TA, and TI indicate that lower genetic variability exists for these traits. The slightly higher PCV than GCV indicates a lesser environmental effect. Low GCV and moderate PCV values for TSS suggest that the environment had the greatest influence on this trait.
All estimated traits had δ2ph > δ2g and PCV > GCV (Table 3), indicating additional environmental influences on trait expression (Singh 2001). However, δ2g represented a larger proportion of δ2ph (Table 3), and the difference between the PCV and the relevant GCV was negligible for all traits (PCV:GCV ratios ranged from 1.00 to 1.17), except TSS and TI (PCV:GCV ratios were 1.69 and 2.33, respectively; Table 3). These traits are, therefore, fairly stable and highly heritable (Singh 2001). This was confirmed by h2b estimates, where h2b was very high (70.8–99.1%) for most traits, with the exception of TSS and TI (35.7 and 117.6%, respectively) (Johnson et al. 1955). Heritability is used to indicate the relative degree to which a character is transmitted from parent to offspring. In improved tomato lines and cultivars, estimates of the h2b of TYLCD resistance/tolerance were high (Abdel-Ati et al. 2005; Mazyed et al. 2007; Hassan et al. 2022). The magnitude of such estimates also suggests the extent to which improvement is possible through selection. PCV and GCV estimation with h2b provides an accurate indicator of the heritable component of variance (Bello et al. 2012). High heritability traits can be improved by various phenotypic selection techniques (Singh 2001). Moderate heritability suggests non-additive gene action in their control; therefore, complex breeding methods may be recommended to improve TSS and TI (Singh 2001). So, phenotypic selection based on TYLCD tolerance (TYLCD-MS), vegetative growth (PL, NPL, and LA), plant yield (EY and MY), and fruit quality traits (FF, FSI, TA, and M) can be trusted to select the best lines among the evaluated TLs.
3.4 Phenotypic and genotypic correlation coefficients
Phenotypic (rph) and genotypic (rg) correlation coefficients for all study traits are presented in Table 4. Successful exploitation of genetic diversity through selection is determined by the correlation of traits. Selection for a particular trait may either increase or decrease the expression of another trait, depending on how closely they are correlated (Bernardo 2010). For most of the estimated traits, rg values were higher than rph values, indicating a moderately string inherent relationship between traits (Bernardo 2010). Traits that showed significant rg to each other generally showed significant rph.
Several significant phenotypic and genotypic correlations were detected. Here, we focus on correlations between TYLCD tolerance (TYLCD-MS), yield, and its components. TYLCD-MS showed positive genotypic and phenotypic correlations with FSI, but negative genotypic association with TI and M. Positive rg and rph values were found for EY and each of NPL, TY, and MY, but negative rg and rph values were found between EY and FSI. Only EY and M showed positive genotypic correlation. TY showed positive rg and rph values with each of NPL, EY, and MY, and positive rg value only with TA. MY showed significant and positive rg and rph values with each of NPL, EY, and TY, and positive and negative correlations with TA and FSI, respectively. AFW had positive rg and rph values with each of PL and NPL, as well as positive rg value with LA (Table 4).
In accordance with our findings, Zengin et al. (2020) reported a negative correlation between fruit weight and length and TYLCD resistance provided by the Ty-3a gene in F3 families. Under TYLCD natural infection, direct selection of lines with the largest NPL will maximize the plant yield. When PL, NPL, and LA increase, the AFW (a component of plant yield) also increases (Table 4).
3.5 Multivariate analysis
PCA has been widely used to evaluate tomato genotypes, identify plant traits that have contributed most to the observed variance among genotypes, and select parental lines for breeding purposes (Merk et al. 2012; Chávez-Servia et al. 2018; Figás et al. 2018; Tembe et al. 2018; Jin et al. 2019; Tripodi et al. 2021). PCA reduced the dimensions of the 14 traits to four PCS (Table 5), which represented 88.34% of the total variance based on Kaiser’s criteria (eigenvalue > 1; Fig. 3) (Johnson and Wichern 1988). According to Brejda et al. (2000), eigenvalues are considered to be the best representation of system attributes across the main components as they explain variance of at least 10%. All estimated traits had a significant effect on the first four PCs. PC1 accounted for 42.93% of the total variation (Fig. 3) and was positively influenced by NPL, PL, TI, TSS, EY, MY, and TY and negatively influenced by TYLCD-MS (Table 5). PC2 was positively correlated with FF, FSI, and LA, accounting for 19.13% of the total variation. PC3 accounted for 16.75% of the total variation and was positively correlated with TA and negatively with M. PC4 was positively correlated with AFW, accounting for 9.54% of the total variation. These findings indicate that traits representing TYLCD tolerance, vegetative, yield, and fruit quality can be used to create groups (Reich et al. 2008). These findings are consistent with those of Shteinberg et al. (2021), who found that PCA revealed a strong separation between tomato genotypes based on TYLCD tolerance. Other studies have also reported that some vegetative growth, yield, and fruit quality traits significantly differed among evaluated tomato genotypes, according to PCA findings (Agong 2001; Glogovac et al. 2012; Merk et al. 2012; Chernet et al. 2014; Iqbal et al. 2014; Figás et al. 2018; Tembe et al. 2018; Jin et al. 2019; Sehgal et al. 2021; Islam et al. 2022). Future evaluations may be based on fewer traits with little information loss, which could reduce the labor, time, and money required to discriminate and define different genotypes (Reich et al. 2008).
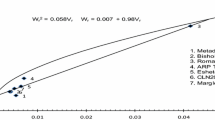
The first two PCs accounted for 11 of the 14 estimated traits and explained a high percentage (62.06%) of the variance (Table 5). Therefore, a biplot between the first two PCs (Fig. 4) was created to assist in identifying the relationships among PCs and each of the traits and lines (Yan and Rajcan 2002; Yan and Kang 2003). TYLCD-MS, vegetative growth, yield, and fruit quality traits are represented by vectors, and anywhere parallel vectors (going in the same direction) reveal a strong positive correlation among these traits while the vectors to the sides show a slight correlation between features, those at 180° or almost opposite demonstrate a strongly negative correlation. The PCA biplot divided the estimated traits into three groups. The first group included PL, LA, AFW, FF, TSS, TI, and M, which were positively correlated with the first two PCs. The second group included NPL, EY, TY and MY, which were positively correlated with PC1 and negatively correlated with PC2. The third group included TYLCD-MS, FSI, and TA, which were negatively correlated with PC1 and positively correlated with PC2. TYLCD-MS was positively correlated with FSI and TA and negatively every EY, TY, MY, and LN. The biplot correlations results corroborate those obtained from rg and rph estimates (Table 4). TA, M, and AFW traits contributed less to the total genetic diversity (short arrows) (Fig. 4). TLs separated into all quarters of the biplot between PC1 and PC2 (Fig. 4), indicating a high level of genetic diversity within and among groups (Reich et al. 2008). Lines with higher values of a given trait were plotted closer to the vector line and further in the direction of that particular vector, often on the vertices of the convex hull. Thus, TL-3, TL-5, and TL-8 were considered superior for most evaluated traits. TL-1, TL-2, TL-11, and TL-12 were closest to the biplot origin, indicating that these lines have the least variability for the estimated traits (Fig. 4).
Biplot among the first two principal components (PC1 and PC2) with fourteen phenotypic traits and twelve tomato lines (TLs) grown under field TYLCV infection conditions over the 2018 and 2019 fall seasons. Phenotypic traits were TYLCV-mean scores (TYLCV), plant length (PL), plant leaves number (LN), leaf area (LA), yield (early: EY; total: TY; and marketable: MY), average fruit weight (AFW), fruit firmness (FF), fruit shape index (FSI), fruit taste index, and fruit maturity index
Tomato lines were divided into three clusters using HCA based on the TYLCD-MS, vegetative growth, yield, and fruit quality traits as illustrated in Fig. 5 and Table 6. This result suggests significant genetic diversity within and among clusters (Table 6). HCA is particularly effective at characterizing tomato genotypes with the greatest degree of similarity/dissimilarity based on morphological traits (Iqbal et al. 2014; Chernet et al. 2014; Bhattarai et al. 2016; Chávez-Servia et al. 2018; Hussain et al. 2018; Tembe et al. 2018; Grozeva et al. 2021; Ene et al. 2022). Cluster 1 comprised TL-1, TL-3, and TL-8 (Fig. 5), which had low TYLCD-MS (highly tolerance), FF, FSI, and M; moderate LA; and high PL, NPL, EY, TY, and MY, AFW, TSS, and TA (Table 6). Six lines made up Cluster 2, which was separated into two groups. The first group included TL-2, TL-6, TL-7, and TL-9, while the second group comprisedTL-4 and TL-5 (Fig. 5). This cluster had the lowest PL, NPL, LA, EY, TY, MY, AFW, and FF values; moderate TYLCD-MS (moderate tolerance) and FSI values; and the highest TI and M (Table 6). Cluster 3 comprised TL-10, TL-11, and TL-12 (Fig. 5) and had low TA and TI values; moderate PL, NPL, EY, TY, MY, AFW, FF, TSS, and M; and high TYLCD-MS (low tolerance), and FSI (Table 6). Variance within the clusters ranged from 15.36 (Cluster 3) to 87.53 (Cluster 2) (Table 6). The maximum distance of Cluster 2 from the centroids was 10.61 and the minimum distance of Cluster 3 (1.53) (Table 6). The distance between cluster centroids ranged from 18.56 to 42.99 (Table 7). The lowest distances were found in Cluster 2 and Cluster 3 (18.56), and the highest were found in Cluster 1 and Cluster 2 (42.99) (Table 7). Cluster 1 performed better in terms of TYLCD tolerance, vegetative growth, yield, and fruit quality performance relative to Clusters 2 and 3, including the population means (Table 6). These results imply that this particular cluster would be more responsive to selection than the others, assuming that TYLCD tolerance and fruit yield are the target traits. Select lines from different clusters, notably Clusters 1 and 2, should be crossed to create custom cultivars/hybrids with beneficial TYLCD tolerance and yield traits.
4 Conclusion
Tomato lines in Clusters 1 and 2 show promise for TYLCD tolerance and economically important traits. TLs in these clusters contain useful breeding material, which could be used as parental genotypes or pre-breeding material for the development of future varieties for TYLCD tolerance, vigorous vegetative growth, productivity, unique fruit shape and size, and flavor desirable for different markets. TYLCD tolerance and productivity can be improved by crossing lines from Clusters 1 and 2.
Data availability
Data generated or analyzed during this study are provided in the manuscript.
References
Abd-Rabou S, Evans GA (2020) The whiteflies in Egypt (Hemiptera: Aleyrodidae). Acta Phytop Entomol Hungarica 55:193–205. https://doi.org/10.1556/038.55.2020.014
Abdel-Ati KEA, Hassan AA, Moustafa SES, Mohammed AA (2005) Inheritance of TYLCV resistance in some locally-bred tomato lines. In: Proceedings 3rd conference on recent technologies in agriculture, Cairo University, Egypt, pp 601–606
Abhary M, Patil BL, Fauquet CM (2007) Molecular biodiversity, taxonomy, and nomenculature of Tomato Yellow Leaf Curl-like viruses. In: Czosnek H (ed) Tomato yellow leaf curl disease: management, molecular biology, breeding for resistance. Springer, Dordrecht, pp 85–118. https://doi.org/10.1007/978-1-4020-4769-5_6
Abhary MK, Anfoka GH, Nakhla MK, Maxwell DP (2006) Post-transcriptional gene silencing in controlling viruses of the Tomato yellow leaf curl virus complex. Arch Virol 151(12):2349–2363. https://doi.org/10.1007/s00705-006-0819-7
Agong SG (2001) Genetic variation of Kenyan tomato (Lycopersicon esculentum L.) germplasm. J Food Technol Afric 6:13–17. https://doi.org/10.4314/jfta.v6i1.19277
Anfoka G, Abhary M, Haj Ahmed F, Hussein AF, Rezk A, Akad F, Abou-Jadah Y, Lapidot M, Vidavaski F, Nakhla MK, Sobh H, Atamian H, Cohen L, Sobol I, Mazyad H, Maxwell DP, Czosnek H (2008) Survey of tomato yellow leaf curl disease-associated viruses in the eastern Mediterranean Basin. J Plant Pathol 90:311–320
AOAC, Association of Official Agricultural Chemists (1990) Official methods of analysis, 15th ed. Washington, DC
Bartleet MS (1937) Properties of sufficiency and statistical tests. Proc R Soc Lond 160A:268–282. https://doi.org/10.1098/rspa.1937.0109
Bello O, Ige S, Azeez M, Afolabi M, Abdulmalig S, Mahamood J (2012) Heritability and genetic advance for grain yield and its component characters in maize (Zea mays L.). Int J Plant Res 2:138–145. https://doi.org/10.5923/j.plant.20120205.01
Bernardo R (2010) Breeding for quantitative traits in plants. Stemma Press, Woodbury
Bhattarai K, Louws FJ, Williamson JD, Panthee DR (2016) Diversity analysis of tomato genotypes based on morphological traits with commercial breeding significance for fresh market production in eastern USA. Aust J Crop Sci 10:1098–1103. https://doi.org/10.21475/ajcs.2016.10.08.p7391
Brejda JJ, Moorman TB, Karlen DL, Dao TH (2000) Identification of regional soil quality factors and indicators. I. Central and southern high-plains. Soil Sci Soc Amer J 64:2115–2124. https://doi.org/10.2136/sssaj2000.6462115x
Brown JK, Idris AM, Torres JI, Banks GK, Wyatt SD (2001) The core region of the coat protein gene is highly useful for establishing the provisional identification and classification of begomoviruses. Arch Virol 146:1581–1598. https://doi.org/10.1007/s007050170080
Burton GW, De Vane EH (1953) Estimating heritability in tall fescue (Festuca arundiancea) from replicated clonal material. Agron J 45:478–481. https://doi.org/10.2134/agronj1953.00021962004500100005x
Burton GW (1952) Quantitative inheritance in grasses. In: Proceedings of the 6th international grassland congress, Pennsylvania, PA, USA, August 1952, vol 1, pp 277–283
Chávez-Servia JL, Vera-Guzmán AM, Linares-Menèndez LR, Carrillo-Rodríguez JC, Aquino-Bolaños EN (2018) Agromorophological traits and mineral content in tomato accessions from El Salvador. Central America Agronomy 8:32. https://doi.org/10.3390/agronomy80830032
Chernet S, Belew D, Abay F (2014) Genetic diversity studies for quantitative traits of tomato (Solanum lycopersicon L.) genotypes in Western Tigray, Northern Ethiopia. J Plant Breed Crop Sci 6:105–113. https://doi.org/10.5897/JPBCS2014.0470
Dhaliwal MS, Jindal SK, Sharma A, Prasanna HC (2020) Tomato yellow leaf curl virus disease of tomato and its management through resistance breeding: a review. J Hort Sci Biotechnol 95:425–444. https://doi.org/10.1080/146203.2019.1691060
Duncan DB (1955) Multiple range and multiple F-test. Biometrics 11:1–5
Ene CO, Abtew WG, Oselebe HO, Ozi FU, Ikeogu UN (2022) Genetic characterization and quatitative trait relationship using multivariate techniques reveal diversity among tomato germplasms. Food Sci Nutr 10:2426–2442. https://doi.org/10.1002/fsn3-2850
Falconer DS, Mackay TFC (1996) Introduction to quantitative genetics, 4th edn. Pearson Education Limited, Edinburgh Gate, Harlow
Falconer DS (1952) The problem of environment and selection. Am Nat 86:293–298. https://doi.org/10.1086/281736
Figás MR, Prohens J, Casanova C, Fernández-de-Córdova P, Soler S (2018) Variation of morphological descriptors for the evaluation of tomato germplasm and their stability across different growing conditions. Sci Hort 238:107–115. https://doi.org/10.1016/j.scienta.2018.04.039
Freed R, Einensmith SP, Gutez S, Reicosky D, Smail VW, Wolberg P (1989) Guide to MSTAT-C analysis of agronomic research experiments. Michigan State University, East Lansing
Glogovac S, Takač TA, Šumic Z, Gvozdanović-Varga J, Červenski J, Vasić M, Popović V (2012) Principal component analysis of tomato genotypes based on some morphological and biochemical quality indicators. Ratar Povrt 49:296–301. https://doi.org/10.5937/ratpov49.2452
Gronenborn B (2007) The tomato yellow leaf curl virus: Genome and function of its proteins. In: Czosnek H (ed) Tomato yellow leaf curl virus disease: management, molecular biology, and breeding for resistance. Springer, Dordrecht, pp 67–84. https://doi.org/10.1007/978-1-4020-4769-5_5
Grozeva S, Nankar AN, Ganeva D, Tringovska I, Pasev G, Kostova D (2021) Characterization of tomato accessions for morphological, agronomic, fruit quality, and virus resistance traits. Can J Plant Sci 101:476–489. https://doi.org/10.1139/cjps-2020-0030
Hallauer AR, Carena MJ, Miranda Filho JB (2010) Quantitative genetics in maize breeding. Springer, Dordrecht
Hassan AA, Abdel-Ati KEA, Mahmoud AMA (2009) Tomato germplasm evaluation and selection for tomato yellow leaf curl virus resistance. Ann Agric Sci Moshtohor 47:261–274
Hassan AA, Abdel-Ati KEA, Mahmoud AMA, Mohamed HAA (2022) Potential adoption of the F2 generation of some TYLCV-resistant tomato hybrids for commercial production under natural field infection. Egypt J Bot 62:59–72. https://doi.org/10.21608/EJBO.2021.67705.1649
Hussain I, Khan SA, Ali S, Farid A, Ali N, Ali S, Masaud S, Hussain I, Azzem K, Raza H (2018) Genetic diversity among tomato accessions based on agro-morphological traits. Sains Malaysiana 47:2637–3645. https://doi.org/10.27576/jsm-2018-4711-06
Iqbal Q, Saleem MY, Hameed A, Asghar M (2014) Assessment of genetic divergence in tomato through agglomerative hierarchical clustering and principal component analysis. Pak J Bot 46:1865–1870
Islam S, Hassan L, Hossain MA (2022) Breeding potential of some exotic tomato lines: a combined study of morphological variability, genetic divergence, and association of traits. Phyton 91:97–114. https://doi.org/10.32604/phyton.2022.017251
Ji Y, Scott JW, Hanson P, Graham E, Maxwell DP (2007) Sources of resistance, inheritance, and location of genetic loci conferring resistance to members of the tomato infecting begomoviruses. In: Czosnek H (ed) Tomato yellow leaf curl virus disease: management, molecular biology, breeding for resistance. Springer Netherlands, Drodrecht, pp 343–362. https://doi.org/10.1007/978-1-4020-4769-5_20
Jin L, Zhao L, Wang Y, Zhou R, Song L, Xu L, Cui X, Li R, Yu W, Zhao T (2019) Genetic diversity of 324 cultivated tomato germplasm resources using agronomic traits and InDel markers. Euphytica 215:69. https://doi.org/10.1007/s10681-019-22391-8
Johnson HW, Robinson HF, Comstock RE (1955) Genotypic and phenotypic correlations in soybeans and their implications in selection. Agron J 47:477–482. https://doi.org/10.2134/agronj1955.00021962004700100008x
Johnson RA, Wichern DW (1988) Applied multivariate statistical analysis, 2nd edn. Prentice-Hall, Englewood Cliffs
Kumar D, Kumar R, Kumar S, Bhardwaj ML, Thakur MC, Kumar R, Thakur KS, Dogra BS, Vikram A, Thakur A, Kumar P (2013) Genetic variability, correlation and path coefficient analysis in tomato. Int J Veget Sci 19:313–323. https://doi.org/10.1080/19315260.2012.726701
Lapidot M, Legg JP, Wintermantel WM, Polston JE (2014) Management of whitefly-transmitted viruses in open-field production systems. Adv Virus Res 90:147–206. https://doi.org/10.1016/B978-0-12-801246-8.00003-2
Lefeuvre P, Martin DP, Harkins G, Lemey P, Gray AJ, Meredith S, Lakay F, Monjane A, Lett JM, Varsani A (2010) The spread of tomato yellow leaf curl virus from the Middle East to the world. PloS Pathog 6:e1001164. https://doi.org/10.1371/journal.ppat.1001164
Lynch M, Walsh B (1998) Genetics and analysis of quantitative traits. Sinauer Associates Inc, Sunderland
Mabvakure B, Martin DP, Kraberger S, Cloete L, van Brunschot S, Geering DW, Thomas JE, Bananej K, Lett JM, Lefeuvre P, Varasani A, Harkins GW (2016) Ongoing geographical spread of tomato yellow leaf curl virus. Virology 498:257–264. https://doi.org/10.1016/j.virol.2016.08.033
Mahmoud AMA (2015) Genetic analysis to select good combiners for TYLCV-tolerance and yield components in tomato. Ann Agric Sci Moshtohor 53:221–232
Mazyed HM, Khalil EM, Rezk AA, Abdel-Hakem MA, Aboul-Ata AE (2007) Genetic studies on tomato yellow leaf curl begomovirus (TYLCV) resistance in Egypt: Six-population analysis. Int J Virol 3:88–95
Merk HL, Yarnes SC, Van Deynze A, Tong N, Menda N, Mueller LA, Mustchler MA, Loewen SA, Myers JR, Francis DM (2012) Trait diversity and potential for selection indices based on variation among regionally adapted processing tomato. J Am Soc Hort Sci 137:427–437. https://doi.org/10.21273/JASHS.137.6.427
Mohammadi SA, Prasanna BM (2003) Analysis of genetic diversity in crop plants-salient statistical tools and considerations. Crop Sci 43:1235–1248. https://doi.org/10.2135/cropsci2003.1235
Moriones E, Navas-Castillo J (2000) Tomato yellow leaf curl virus, an emerging virus complex causing epidemics worldwide. Virus Res 71:123–134
Murray HG, Thompson WF (1980) Rapid isolation of high molecular weight DNA. Nucl Acids Res 8:4321–4325. https://doi.org/10.1093/nar/8.19.4321
Navez S, Letard M, Graselly D, Jost M (1999) The criteria of tomato quality. Infos-Ctifl 115:41–47
Óconnor BP (2000) SPSS and BAS programs for determining the number of components using parallel analysis and velicer’s MAP test. Behav Res Methods Instrum Comput 32:396–402
Palumbo JC, Hotowitz AR, Prabhaker N (2001) Insecticidal control and resistance management for Bemisia tabaci. Crop Protec 20:739–765. https://doi.org/10.1016/S0261-2194(01)00117-X
Pandey SK, Singh H (2011) A simple, cost-effective method for leaf area estimation. J Bot 2011:658240–658241. https://doi.org/10.1155/2011/658240
Ramos RS, Kumar L, Shabani F, Picanço MC (2018) Mapping global risk levels of Bemisia tabaci in areas of suitability for open field tomato cultivation under current and future climates. PLoS ONE 14:e0198925. https://doi.org/10.1371/journal.pone.0198925
Reich D, Price AL, Patterson N (2008) Principal component analysis of genetic data. Nat Genet 40:491–492. https://doi.org/10.1038/ng0508-491
Robinson HF, Comstock RE, Harvey PH (1949) Estimates of heritability and degree of dominance in corn. Agron J 41:353–359. https://doi.org/10.2134/agronj1949.00021962004100080005x
Schouten HJ, Tikunov Y, Verkerke W, Finkers R, Bovy A, Bai Y, Visser RGF (2019) Breeding has increased the diversity of cultivated tomato in the Netherlands. Frontiers 10:1606. https://doi.org/10.3389/fpls.2019.01606
Sehgal N, Chadha S, Kumar S, Ravita (2021) Variability and traits association analyses in bacterial wilt resistant F4 progenies of tomato, Solanum lycopersicum L. for yield and biochemical traits. Indian J Exper Biol 59:617–625
Shapiro SS, Wilk MB (1965) An analysis of variance test for normality (complete samples). Biometrika 52:591–611. https://doi.org/10.2307/2333709
Sharma S (1996) Applied multivariate techniques. Wiley, New York
Shteinberg M, Mishra R, Anfoka G, Altaleb M, Brotman Y, Moshelion M, Gorovits R, Czosnek H (2021) Tomato yellow leaf curl cirus (TYLCV) promotes plant tolerance to drought. Cells 10:2875. https://doi.org/10.3390/cells10112875
Singh BD (2001) Plant breeding: principles and methods. Kalyani Publishers, New Delhi
Singh RK, Chaudhary BD (1985) Biometrical methods in quantitative analysis. Kalayani Publishers, New Delhi
Singh RK, Rai N, Singh AK, Kumar P, Singh B (2019) A critical review on tomato leaf curl virus resistance in tomato. Int J Veget Sci 25:373–393. https://doi.org/10.1080/19315260.2018.1520379
Tembe KO, Chemining’wa G, Ambuko J, Owino W (2018) Evaluation of African tomato landraces (Solanum lycopersicum) based on morphological and horticultural traits. Agric Nat Resour 52:536–542. https://doi.org/10.1016/j.anres.2018.11.014
Tessema GL, Mohammed AW, Abebe DT (2022) Genetic variability studies for tuber yield and yield attributes in Ethiopian released potato (Solanum tuberosum L.) varieties. Peer J 10:e12860. https://doi.org/10.7717/peerj.12860
Tripodi P, Soler S, Campanelli G, Díez MJ, Esposito S, Sestili S, Figás MR, Leteo F, Casanova C, Platani C, Soler E, Bertone A, Pereire-Dias L, Palma D, Burguet R, Pepe A, Rosa-Martínez PJ, Cardi T (2021) Genome wide association mapping for agronomic, fruit quality, and root architectural traits in tomato under organic farming conditions. BMC Plant Biol 21:481. https://doi.org/10.1186/s12870-03271-4
Vidavski F, Czosnek H, Gazit S, Levy D, Lapidot M (2008) Pyramiding of genes conferring resistance to tomato yellow leaf curl virus from different wild tomato species. Plant Breed 127:625–631. https://doi.org/10.1111/j.1439-0523.2008.01556.x
Vidavski FS (2007) Exploitation of resistance genes found in wild tomato species to produce resistant cultivars; pile up of resistant genes. In: Czosnek H (ed) Tomato yellow leaf curl virus disease: management, molecular biology, breeding for resistance. Springer, Dordrecht, pp 363–372. https://doi.org/10.1007/978-1-4020-4769-521
Wickens TD, Keppek G (2004) Design and analysis: a researcher’s handbook. Pearson Prentice-Hall, Upper Saddle River
Yan W, Kang MS (2003) GCE-biplot analysis: a graphical tool for breeders, geneticists, and agronomists. CRC Press, Boca Raton
Yan W, Rajcan I (2002) Biplot analysis of test sites and trait relations of soybean in Ontario. Crop Sci 42:11–20. https://doi.org/10.2135/cropsci2002.1100
Yan Z, Pérez-de-Castro A, Díez MJ, Hutton SF, Visser RGF, Wolters A-MA, Bai Y, Li J (2018) Resistance to tomato yellow leaf curl virus in tomato germplasm. Front Plant Sci 9:1198. https://doi.org/10.3389/fpls.2018.01198
Yassin TE (1973) Genotypic and phenotypic variances and correlations in field beans (Vicia faba L.). J Agric Sci 81:445–448. https://doi.org/10.1017/S0021859600086494
Yeager AF (1937) Studies on the inheritance and development of fruit size and shape in the tomato. J Agric Res 55:141–152
Zengin S, Kabas A, Ilbi H (2020) Relationship between some morphological traits of the tomato lines and resistance to tomato yellow leaf curl virus disease. Notulae Botanicae Horti Agrobotanici Cluj-Napoca 48:388–397. https://doi.org/10.15835/nbha48111732
Funding
Open access funding provided by The Science, Technology & Innovation Funding Authority (STDF) in cooperation with The Egyptian Knowledge Bank (EKB). No funding.
Author information
Authors and Affiliations
Contributions
The study's conception and design, material preparation, data collection and analysis, and the writing of the first draft of the manuscript were all done with participate from both authors.
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no known competing financial interests or personal relationships that could have appeared to influence the work reported in this paper.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Mahmoud, A.M.A., Osman, N.H. Utilizing genetic diversity to select tomato lines tolerant of tomato yellow leaf curl virus based on genotypic coefficient of variation, heritability, genotypic correlation, and multivariate analyses. Braz. J. Bot 46, 609–624 (2023). https://doi.org/10.1007/s40415-023-00908-6
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s40415-023-00908-6