Abstract
Background and Objective
Amikacin is preferred in treating Gram-negative infections in neonates and it has a narrow therapeutic window. The population pharmacokinetic modeling approach can aid in designing optimal dosage regimens for amikacin in neonates. In this study, we attempted to identify the suitable population pharmacokinetic model from the published reports for the study population from an Indian setting.
Methods
Published population pharmacokinetic studies for amikacin in neonates were identified. Data on structural models and typical pharmacokinetic parameters were extracted from the studies. For the clinical study, neonates who met the inclusion criteria were enrolled in the study from the NICU, Kasturba Medical College, Manipal, during Jan 2020 to March 2022. Drug concentrations were estimated, and demographic and clinical data were collected. Identified population pharmacokinetic models were used to predict the amikacin concentrations in neonates. Predicted concentrations were compared against the observed concentrations. Differences between predicted and observed concentrations were quantified using statistical measures. The population pharmacokinetic model, which was able to predict the data well, is considered a suitable model for the study population. Dosing regimens were suggested for neonates using the pharmacometric simulation approach generated by the selected model.
Results
A total of 43 plasma samples were collected from 31 neonates. Twelve population pharmacokinetic models were found for amikacin in neonates. The predictive performance of the 12 studies was performed using clinical data. A two-compartment model reported by Illamola et al. predicted the amikacin concentrations better than other models. Illamola et al. reported creatinine clearance and body weight as the significant covariates impacting the pharmacokinetic parameters of amikacin. This model was able to predict the clinical data with 29.97% and 0.686 of relative median absolute prediction error and relative root mean square error, respectively, which is the best among the published models. The Illamola et al. model was selected as the final model to perform pharmacometric simulations for the subjects with different combinations of creatinine clearance and body weight. Dosage regimens were designed to attain target therapeutic concentrations for the virtual subjects and a nomogram was developed.
Conclusions
The population pharmacokinetic model reported by the Illamola et al. model was selected as the final model to explain the clinical data with the lowest relative median absolute prediction error and relative root mean square error when compared with other models. An amikacin nomogram was developed for the neonates whose creatinine clearance and body weight ranged between 10 and 90 mL/min and between 2 and 4 kg, respectively. A developed nomogram can assist clinicians to design an optimal dosage regimen of amikacin for term neonates.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
We have assessed all published population pharmacokinetic models for amikacin in neonates using the clinical data collected from the Indian clinical setting. |
The model that predicted the data better was considered for designing the dosage regimen. |
Nomograms proposed in the study can potentially aid clinicians to design optimal dosage regimens of amikacin in term neonates. |
1 Introduction
Amikacin is a semi-synthetic aminoglycoside, which is used alone or in combination to treat Gram-negative bacterial infections in neonates [1]. Amikacin exhibits concentration-dependent bactericidal activity [2]. Optimal therapeutic effects will be obtained when the maximum concentration is eight to ten times higher than the minimum inhibitory concentration [3]. The recommended therapeutic range for peak and trough concentrations of amikacin was reported as 24–35 mg/L and less than 5 mg/L, respectively [4]. Concentrations above the therapeutic range could lead to nephrotoxicity and ototoxicity and the concentrations below the range results in ineffective therapy and could potentially lead to anti-microbial resistance to amikacin [1]. In the current scenario, the potential minimum inhibitory concentration for any bacterial activity is 2–4 mg/L for amikacin, implying that the minimum needed peak concentration is around 30 mg/L [5, 6]. As there is a minimal difference between effective and toxic concentrations; amikacin is categorized as a narrow therapeutic index drug [7]. Understanding the drug pharmacokinetics in the target patient population could help in enhancing the therapeutic efficacy of narrow therapeutic index drugs.
Neonates are a vulnerable population and there is a lack of detailed pharmacokinetic (PK) information for many drugs in this group compared with the adult population [8]. It is difficult to get multiple blood samples from neonates; therefore, traditional PK studies are not available in this population for amikacin. Population PK (PopPK) studies will help to understand the PK behavior of a drug using sparse samples in the neonatal population [9]. Population pharmacokinetics is a branch of clinical pharmacology that focuses on quantifying the PK characteristics and identifying the sources and correlates of variability [10, 11].
Several PopPK studies were reported on amikacin in neonates [4, 12,13,14,15,16,17,18,19,20,21,22]. These studies identified variability in PK parameters in neonates as per their demographic and clinical conditions. The PopPK models reported by these studies were developed and validated in their respective study populations. There were no reports on PopPK of amikacin from an Indian setting. Therefore, it was necessary to evaluate these models using data collected from neonates in Indian settings. Validation of PopPK models with external clinical data will give confidence to the clinician to implement model-informed dosing decisions. In this study, we aimed to analyze the predictive performance of reported PopPK models for amikacin in neonates using clinical data collected from an Indian neonatal setting. It was also aimed to develop a dosing nomogram for amikacin from a suitable model.
2 Methodology
2.1 Literature Search and Data Extraction
A literature search was conducted in PubMed to retrieve articles published from January 2000 to June 2022. Published models of PopPK for amikacin were identified. The search was conducted using the terminologies “amikacin,” “aminoglycosides,” “pharmacokinetics,” “population pharmacokinetics,” “nonlinear mixed effect modeling,” “NONMEM,” “therapeutic drug monitoring,” “neonates,” and “pediatrics.” A manual search was also conducted by inspecting the bibliography of relevant articles. Studies that reported a PopPK model from neonatal data were included. Models were excluded if the information provided was not sufficient to reproduce the model. Demographic details and PopPK model information including the model structure and typical values of parameters were extracted from the selected studies.
2.2 Patients and Data Collection
This prospective longitudinal observational study was approved by the institutional ethics committee (ECR/146/Inst/KA/2013/RR-19; IEC:558/2019). Study was conducted at neonatal ICU, department of pediatrics, Kasturba Medical College, Manipal Academy of Higher Education. Data was collected from the period of January 2020 to March 2022 was included in this study. Clinical and demographic data were collected from medical records, and study participants were enrolled considering inclusion and exclusion criteria. Informed consent was obtained from the parents/caretakers of neonates. Full-term neonates, who were admitted to the neonatal intensive care unit and were treated with amikacin, were included in the study. Neonates termed clinically unstable by neonatologists were excluded from the study. Creatinine clearance (CrCl) was calculated using the Schwartz formula [23].
Serum creatinine (SCr) levels were estimated by the Jaffe Rate Blanked method in our hospital.
A convenience sampling approach was used to collect blood samples. Blood samples were collected only when blood was withdrawn for routine laboratory investigations, no sample was collected only for the study purpose. Dosing information and sampling time were recorded in the predesigned data collection form. Blood samples were collected into EDTA-coated tubes and centrifuged at 7000 RPM for 5 min to separate plasma. Plasma was stored at − 80 °C until quantification by the liquid chromatography with tandem mass spectrometry method. The quantification of amikacin in plasma was performed using an in-house developed and validated liquid chromatography with tandem mass spectrometry bioanalytical method, which has a working range of 0.5–100 µg/mL. The chromatographic separation was obtained in a HILICPAK VC-50 2D 150 mm × 2.0 mm column with a mobile phase having a mixture of 1.5% v/v ammonia solution and acetonitrile [24].
2.3 Predictive Performance of Published Models
Population PK models identified from the literature search were used for the predictive performance assessment. The model structure and parameter values were used from the literature. Demographic data and clinical and dosing information were used from the study conducted in the current study set-up. All these details were used to predict the concentrations at the same timepoints of the current clinical study. Predicted concentrations were compared against observed concentrations from the present clinical study. Bias and precision were calculated using relative prediction error percentage (rPE%), relative mean prediction error percentage (rMPE%), relative median absolute prediction error percentage (rMAPE%), and relative root mean squared error (rRMSE) [25]. All the simulations and data analysis were carried out using the Julia computing language PUMAS package version 1.6 [26].
Cpred and Cobs in the above equations represent predicted and observed concentrations, respectively.
2.3.1 Selection of Models
The performance of models was considered satisfactory when their rMPE and rMAPE values were within − 20 to 20%, and less than 30% respectively. A model with the lowest rMAPE and rRMSE was selected if more than one model satisfied these criteria. Visual predictive check predicted versus observed concentrations plots were used to visualize the predictive performance of each model.
2.4 Bayesian Forecasting
Bayesian forecasting has been performed to evaluate the applicability of the final model in the neonatal setting to implement therapeutic drug monitoring. Individual parameters were estimated using the initially observed concentrations of the subject. Later concentrations were predicted using the estimated parameters of the individual. Individual predictions were compared against observed concentrations to evaluate model performance. Relative mean individual prediction error percentage (rMIPE%) and relative median absolute individual prediction error percentage (rMAIPE%) were calculated. The percentage of total predictions falling within 20 and 30% absolute prediction errors were considered satisfactory.
2.5 Developing a Dosing Nomogram
Pharmacometric simulations were performed using the selected final model for the range of covariates included in the models, and what-if scenarios for the neonates with different CrCl and weight (WT) were explored using the virtual subjects. A dosing nomogram was proposed to keep the peak concentrations within 25–35 mg/L and trough concentrations within 2–5 mg/L. Dosage regimens were selected based on the attainment of target peak and trough concentrations in the recommended therapeutic window.
3 Results
Data from 31 neonates recruited into the study were included in the final dataset. All the subjects were full-term neonates and the post-natal age (PNA) was below 28 days. The median gestational age and PNA of neonates/subjects during the time of recruitment were 39 weeks and 1.6 days, respectively. A total of 43 plasma samples were collected from 31 neonates at random timepoints. The spread of plasma samples collected against the time after the recent dose of amikacin was described in Fig. 1. The median PNA when the sample was collected was 5.6 days. Most of the neonates were prescribed a once-daily dosage regimen. Table 1 shows the demographic details of the subjects recruited into the study. All the clinical information was collected from the patient case records.
Time after dose versus concentration profile of amikacin in neonates. Amikacin dosing regimens were intended to produce peak and trough concentrations of 25–35 and 2–5 mg/L, respectively. Neonates with high serum creatinine levels and lower body weight had concentrations of more than 5 mg/L of amikacin after 23 hours (hrs) of administration. MIC minimum inhibitory concentration, MSC maximum safe concentration
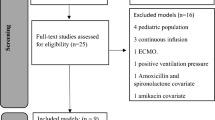
A total of 11 PopPK studies were identified. Twelve PopPK models were extracted from 11 studies. Illamola et al. reported two PopPK models using a different set of covariates. The median PNA for all the studies was less than 30 days. All the reported models had both preterm and term neonates in their study. The demographic details of all studies are described in Table I of the Electronic Supplementary Material (ESM). The PK model parameters and compartmental structures are presented in Table II of the ESM.
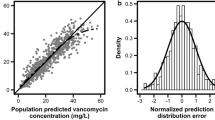
Population predictions were used to analyze the prediction-based diagnostics. rMPE%, rMAPE%, and rRMSE were used to quantify the predictive performance for bias and precision. Models M5, M3, M7, and M11 had the lowest bias when compared with the other models. Models M7, M5, M1, and M11 were found to be more precise than other models. Models M7, M10, and M11 had the lowest error in analyzing the overall performance. Model M7 performed better than the other reported models for the present data, followed by model M11. The predictive performance of all models is presented in Table 2. The overall predictive performance of the models is described in Fig. 2. Observed concentrations versus population-predicted concentrations of models are presented in Fig. 3. The percentage of predicted concentrations with less than 30% rMAPE for each model is shown in Fig. 4.
Model M7 was selected as the final model that predicted the concentrations better than other models.
Bayesian forecasting was performed using model M7. A total of nine subjects had more than one concentration. Three out of nine subjects had SCr and WT values close to the day of sample collection. In these subjects, predictions of concentrations were better with 0.6 and 13.04 of rMIPE% and rMAIPE%, respectively. Predicted and observed concentrations of these three subjects are shown in Fig. 5.
Bayesian forecasting using posterior knowledge derived from the first concentration of the subject to predict the second and third concentrations for the same subject. The solid line shows the predicted concentration profile using empirical Bayesian forecasting, overlapped scatters are the observed concentration from the individual
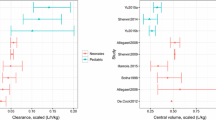
Weight and CrCl were the covariates reported in this model, which influenced the PK parameters. Virtual subjects were developed with different combinations of WT and CrCl. Weight ranged from 2 to 4 kg and CrCl ranged from 10 to 90 mL/min. Appropriate dosing regimens for the combination of these covariates are presented in Table 3. These dosing regimens were expected to produce peak and trough concentrations around 30 and 3 mg/L, respectively, after the first dose.
4 Discussion
This is the first study to evaluate the predictive performance of published PopPK models for amikacin using clinical data obtained from an external neonatal setting in India. This analysis can assist in identifying the models that are suitable for the Indian population and explore potential what-if scenarios for amikacin dosing in neonates. The PopPK model that performs best can be considered for designing the optimal dosing regimen for amikacin in neonates. Most of the models were developed using the data collected from term and preterm neonates, some models have included disease states as covariates. These models can help to understand the pharmacokinetics of amikacin in term neonates, as they were part of all the published models. Published models differed considerably in their predictive performance. Most of the models underpredicted the amikacin concentrations for the present data.
Amikacin is eliminated renally and overdosing or under-dosing can result in toxicity or treatment failure, respectively [27]. Amikacin is mainly dosed based on WT and it is important to maintain amikacin peak and trough concentrations within respective therapeutic ranges. Renal function also needs to be considered while amikacin dosing is planned, as per label recommendations by the US Food and Drug Administration [27]. However, guidelines for neonatal dosing based on renal function are not well established. Eight out of 11 reviewed studies did not report SCr levels in their demographic details. Only two studies have reported SCr/CrCl as a covariate on clearance [18, 22]. Model M7, which reported CrCl as a covariate on clearance, predicted amikacin concentrations better than other models. The predictive performance of Model M6 with SCr identified as a covariate on clearance was not better than models that did not include SCr. It is known that the clearance of amikacin is similar to the CrCl of the individual [28]. This established relationship between clearance and CrCl might be the key to better performance of the Illamola et al. model compared with other models.
Maternal creatinine levels affect neonatal SCr during the first few days of neonatal life. It is difficult to measure true neonatal SCr levels during this time [29]. This could have been one of the reasons for most of the studies to not include SCr/CrCl as a covariate in their models. Clearance of aminoglycosides was reported to be similar [28]; therefore, we attempted to analyze the SCr clouding effect reported in another PopPK model [30], but it did not perform better than the model M7. This could be the result of maternal SCr clouding, which lasts for 48 hours in term newborns [31]. The majority of the SCr levels in the clinical data/study may be a true reflection of neonatal SCr levels, as there were only four amikacin concentrations, which were collected before 48 hours of neonatal life.
Models M5 and M11 reported birth WT and PNA as covariates on the clearance of amikacin. Birth WT and PNA are directly proportional to renal maturation in neonates [17, 19, 32]. These two models could be used as alternative models to M7 in the absence of SCr levels in individuals. Model M11 was updated from the information collected from model M5. Model M11 was able to predict the data with better rRMSE and similar MAPE to M5, hence model M11 can be considered the second-best model, especially when SCr levels are not available or during the first 2 days of neonatal life. Smits et al. have provided the dosing recommendations for amikacin in neonates using model M11 [19]. Missing SCr can be predicted for the subjects using the model developed by Krzyzanski et al. to implement model M7 [33].
Bayesian forecasting has been performed on the subjects who had more than one concentration using model M7. We were successfully able to forecast the second concentrations in the subjects who had covariates collected close to the date of sampling. Bayesian forecasting did not predict concentrations satisfactorily in subjects, who did not have covariate information like WT and CRCL close to the day of sampling. Weight could not be measured in intubated neonates and SCr could not be measured frequently to get values close to the day of blood sampling.
The use of dosing recommendations informed by model M7 can assist the clinician to design a precise dosage regimen for the neonates [18]. Illamola et al. provided the dosing guidelines informed by model M7, but these guidelines were provided at wide intervals for covariates. In our study, we have provided the dosing regimens with narrow intervals for a range of covariates. The CrCl interval was defined as 10 mL/min and the WT interval was defined as 250 g. These dosing regimens can potentially result in a therapeutic range of peak and trough concentrations for the individual. Quantification of amikacin concentration after the first dose can give the actual information of PK parameters of an individual when the Bayesian approach is used for estimating parameters. These parameters in turn can be used for designing individualized dosage regimens.
The study had a few limitations. Serum creatinine and WT values were filled from previous values of the same subject, wherever they were not available at the time of amikacin sampling. Clinically unstable patients were not included in the study, thus the proposed dosing recommendations may not result in target outcomes for them. Suggested dosing regimens are yet to be evaluated in the patient population and there is a need to prospectively evaluate the suggested nomogram in various clinical settings. We will try to prospectively validate the suggested dosing nomogram and such studies from other settings are highly encouraged.
5 Conclusions
A two-compartment model developed by Illamola et al. performed better to predict the amikacin concentration data collected from a neonatal setting in India. Creatinine clearance and WT were the significant covariates that influence amikacin clearance. Dosing regimens were proposed using the final model to result in peak concentrations of around 30 mg/L and trough concentrations of around 3 mg/L. These dosing recommendations can be used for determining the first dose of amikacin in term neonates. Dosing recommendations from Smits et al. can be considered for neonates, whose SCr levels are not available. Drug concentration monitoring is advisable after the first dose and further dosage individualization is possible using Bayesian forecasting approaches.
References
Roberts JA, Norris R, Paterson DL, Martin JH. Therapeutic drug monitoring of antimicrobials. Br J Clin Pharmacol. 2012;73(1):27–36.
Garraffo R, Drugeon HB, Dellamonica P, Bernard E, Lapalus P. Determination of optimal dosage regimen for amikacin in healthy volunteers by study of pharmacokinetics and bactericidal activity. Antimicrob Agents Chemother. 1990;34(4):614–21.
Radigan EA, Gilchrist NA, Miller MA. Management of aminoglycosides in the intensive care unit. J Intensive Care Med. 2010;25(6):327–42.
Sherwin CM, Svahn S, Van der Linden A, Broadbent RS, Medlicott NJ, Reith DM. Individualised dosing of amikacin in neonates: a pharmacokinetic/pharmacodynamic analysis. Eur J Clin Pharmacol. 2009;65(7):705–13.
Kato H, Hagihara M, Hirai J, Sakanashi D, Suematsu H, Nishiyama N, et al. Evaluation of amikacin pharmacokinetics and pharmacodynamics for optimal initial dosing regimen. Drugs R D. 2017;17(1):177–87.
Alqahtani S, Abouelkheir M, Alsultan A, Elsharawy Y, Alkoraishi A, Osman R, et al. Optimizing amikacin dosage in pediatrics based on population pharmacokinetic/pharmacodynamic modeling. Paediatr Drugs. 2018;20(3):265–72.
Germovsek E, Barker CI, Sharland M. What do I need to know about aminoglycoside antibiotics? Arch Dis Child Educ Pract Ed. 2017;102(2):89–93.
Coppini R, Simons SHP, Mugelli A, Allegaert K. Clinical research in neonates and infants: challenges and perspectives. Pharmacol Res. 2016;108:80–7.
Aarons L. Population pharmacokinetics: theory and practice. Br J Clin Pharmacol. 1991;32(6):669–70.
Mould DR, Upton RN. Basic concepts in population modeling, simulation, and model-based drug development. CPT Pharmacometr Syst Pharmacol. 2012;1(9): e6.
Ette EI, Williams PJ. Population pharmacokinetics I: background, concepts, and models. Ann Pharmacother. 2004;38(10):1702–6.
Allegaert K, Anderson BJ, Cossey V, Holford NH. Limited predictability of amikacin clearance in extreme premature neonates at birth. Br J Clin Pharmacol. 2006;61(1):39–48.
Allegaert K, Scheers I, Cossey V, Anderson BJ. Covariates of amikacin clearance in neonates: the impact of postnatal age on predictability. Drug Metab Lett. 2008;2(4):286–9.
Bleyzac N, Varnier V, Labaune JM, Corvaisier S, Maire P, Jelliffe RW, et al. Population pharmacokinetics of amikacin at birth and interindividual variability in renal maturation. Eur J Clin Pharmacol. 2001;57(6–7):499–504.
Botha JH, du Preez MJ, Miller R, Adhikari M. Determination of population pharmacokinetic parameters for amikacin in neonates using mixed-effect models. Eur J Clin Pharmacol. 1998;53(5):337–41.
Cristea S, Smits A, Kulo A, Knibbe CAJ, van Weissenbruch M, Krekels EHJ, et al. Amikacin pharmacokinetics to optimize dosing in neonates with perinatal asphyxia treated with hypothermia. Antimicrob Agents Chemother. 2017;61(12):e01282-e1317.
De Cock RF, Allegaert K, Schreuder MF, Sherwin CM, de Hoog M, van den Anker JN, et al. Maturation of the glomerular filtration rate in neonates, as reflected by amikacin clearance. Clin Pharmacokinet. 2012;51(2):105–17.
Illamola SM, Colom H, van Hasselt JG. Evaluating renal function and age as predictors of amikacin clearance in neonates: model-based analysis and optimal dosing strategies. Br J Clin Pharmacol. 2016;82(3):793–805.
Smits A, De Cock RF, Allegaert K, Vanhaesebrouck S, Danhof M, Knibbe CA. Prospective evaluation of a model-based dosing regimen for amikacin in preterm and term neonates in clinical practice. Antimicrob Agents Chemother. 2015;59(10):6344–51.
Wang J, Liang WQ, Wu JJ, Pan CM. Population pharmacokinetic analysis of amikacin and validation on neonates using Monte Carlo method. Acta Pharmacol Sin. 2000;21(10):954–60.
Amponsah SK, Adjei GO, Enweronu-Laryea C, Bugyei KA, Hadji-Popovski K, Kurtzhals JAL, et al. Population pharmacokinetic characteristics of amikacin in suspected cases of neonatal sepsis in a low-resource African setting: a prospective nonrandomized single-site study. Curr Ther Res Clin Exp. 2017;84:e1-6.
Caceres Guido P, Travaglianti M, Castro G, Licciardone N, Ferreyra O, Bramuglia G, et al. Population pharmacokinetics of amikacin in neonatal intensive care unit patients. Aust Med J. 2017;10(02):140–4
Schwartz GJ, Feld LG, Langford DJ. A simple estimate of glomerular filtration rate in full-term infants during the first year of life. J Pediatr. 1984;104(6):849–54.
Matcha S, Chaudhari BB, Mallayasamy S, Lewis LE, Moorkoth S. Ion-pairing reagent-free hydrophilic interaction LC-MS/MS method for therapeutic drug monitoring of amikacin in neonates. J Appl Pharm Sci. 2023;13(2):029–38.
Sheiner LB, Beal SL. Some suggestions for measuring predictive performance. J Pharmacokinet Biopharm. 1981;9(4):503–12.
Rackauckas C, Ma Y, Noack A, Dixit V, Mogensen PK, Elrod C, et al. Accelerated predictive healthcare analytics with pumas, a high performance pharmaceutical modelling and simulation platform. BioRxiv. 2022;1:1–23.
Micromedex. Amikacin: in depth answers. Greenwood Village: IBM Corporation. 2021. www.micromedexsolutions.com. Accessed 15 Oct 2021.
Winter ME. Basic clinical pharmacokinetics. 5th ed. Philadelphia: Wolters Kluwer/Lippincott Williams & Wilkins Health; 2010.
Go H, Momoi N, Kashiwabara N, Haneda K, Chishiki M, Imamura T, et al. Neonatal and maternal serum creatinine levels during the early postnatal period in preterm and term infants. PLoS ONE. 2018;13(5): e0196721.
Wang H, Sherwin C, Gobburu JVS, Ivaturi V. Population pharmacokinetic modeling of gentamicin in pediatrics. J Clin Pharmacol. 2019;59(12):1584–96.
Falcao MC, Okay Y, Ramos JL. Relationship between plasma creatinine concentration and glomerular filtration in preterm newborn infants. Rev Hosp Clin Fac Med Sao Paulo. 1999;54(4):121–6.
Sulemanji M, Vakili K. Neonatal renal physiology. Semin Pediatr Surg. 2013;22(4):195–8.
Krzyzanski W, Smits A, Van Den Anker J, Allegaert K. Population model of serum creatinine as time-dependent covariate in neonates. AAPS J. 2021;23(4):86.
Acknowledgments
We gratefully acknowledge the Indian Council of Medical Research (Extramural Ad-hoc) for providing a grant to carry out the research.
Funding
Open access funding provided by Manipal Academy of Higher Education, Manipal.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Funding
This work was supported by the Indian Council of Medical Research, grant number 5/7/1694/CH/Adhoc/RBMCH-2020. Surulivelrajan Mallayasamy is the principal investigator.
Conflict of interest
Saikumar Matcha, Jayashree Dillibatcha, Arun Prasath Raju, Bhim Bahadur Chaudhari, Sudheer Moorkoth, Leslie E. Lewis, and Surulivelrajan Mallayasamy have no conflicts of interest that are directly relevant to the content of this article.
Ethics approval
The prospective longitudinal observational study was approved by Kasturba Medical College and Kasturba Hospital Institutional Ethics Committee, Manipal, India. (IEC-558/2019). We confirm that the study was performed in accordance with the ethical standards as laid down in the 1964 Declaration of Helsinki and its later amendments or comparable ethical standards.
Consent to participate
Neonates were recruited into the study after obtaining consent from their parents/guardians.
Consent for publication
Not applicable.
Availability of data and material
Data are available from the corresponding author upon reasonable request.
Code availability
This work has used the PUMAS-AI software, the code is available from the corresponding author on request.
Author contributions
Surulivelrajan Mallayasamy (SRM) and Lesli E. Lewis (LL) designed the protocol for the project. SRM took the responsibility to acquire funds for the research. Saikumar Matcha (SM) and Bhim Bahadur Chaudhari (BC) initiated the work. SM lead the patient recruitment along with BC. BC initiated the bioanalytical method development using the liquid chromatography with tandem mass spectrometry method under the supervision of Sudheer Moorkoth (SUM). BC and SM carried out the quantification of amikacin from plasma samples, SUM provided critical inputs for smooth conduction of the work. Jayashree Dillibatcha (JD) and Arun Prasath Raju (AR) along with SM contributed to analyze the predictive performance of published models of amikacin under the guidance of SRM. SM, JD, and AR developed the dosing nomogram under the supervision of SRM and LL. SM initiated the manuscript writing along with JD, AR, and BC. SRM, LL, and SUM provided the intellectual inputs. All the authors read and approved the final version of the manuscript.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial 4.0 International License, which permits any non-commercial use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc/4.0/.
About this article
Cite this article
Matcha, S., Dillibatcha, J., Raju, A.P. et al. Predictive Performance of Population Pharmacokinetic Models for Amikacin in Term Neonates. Pediatr Drugs 25, 365–375 (2023). https://doi.org/10.1007/s40272-023-00564-z
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s40272-023-00564-z