Abstract
Introduction
Discharge summaries contain valuable information about adverse drug reactions, but their unstructured nature makes them challenging to analyse and use as a signal source for pharmacovigilance. Machine learning has shown promise in identifying discharge summaries that contain related drug-adverse event pairs but has fared relatively poorer in entity extraction.
Methods
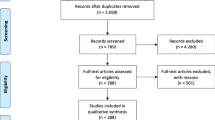
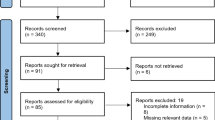
A hybrid model is developed combining rule-based and machine learning algorithms using discharge summaries with the aim of maximising capture of related drug-adverse event pairs. The rule first identifies segments containing adverse event entities within a 100-character distance from a drug term; machine learning subsequently estimates the relatedness of the drug and adverse event entities contained. The approach is validated on four independent datasets that are temporally and geographically separated from model development data. The impact of restricted drug-adverse event pair detection on recall is evaluated by using two of the four validation datasets that do not impose rule-based restrictions to annotations.
Results
The hybrid model achieves a recall of 0.80 (fivefold cross validation), 0.80 (temporal) and 0.76 (geographical) on validation using datasets containing only pre-identified target text segments that fulfil the rule-based algorithm criteria. When tested on datasets that additionally contained drug-adverse event pairs not restricted by the rule-based criteria, recall of the model declines to 0.68 and 0.62 on temporally and geographically separated datasets, respectively.
Conclusions
The proposed hybrid model demonstrates reasonable generalisability on external validation. Rule-based restriction of the detection space results in an approximately 12–14% reduction in recall but improves identification of the related drug and adverse event terms.




Similar content being viewed by others
References
Lopez-Gonzalez E, Herdeiro MT, Figueiras A. Determinants of under-reporting of adverse drug reactions: a systematic review. Drug Saf. 2009;32(1):19–31.
Hazell L, Shakir SA. Under-reporting of adverse drug reactions : a systematic review. Drug Saf. 2006;29(5):385–96.
Giardina C, Cutroneo PM, Mocciaro E, Russo GT, Mandraffino G, Basile G, et al. Adverse drug reactions in hospitalized patients: results of the FORWARD (Facilitation of Reporting in Hospital Ward) Study. Front Pharmacol. 2018;9:350.
Chan SL, Ng HY, Sung C, Chan A, Winther MD, Brunham LR, et al. Economic burden of adverse drug reactions and potential for pharmacogenomic testing in Singaporean adults. Pharmacogenom J. 2019;19(4):401–10.
Komagamine J, Kobayashi M. Prevalence of hospitalisation caused by adverse drug reactions at an internal medicine ward of a single centre in Japan: a cross-sectional study. BMJ Open. 2019;9(8): e030515.
Honigman B, Lee J, Rothschild J, Light P, Pulling RM, Yu T, et al. Using computerized data to identify adverse drug events in outpatients. J Am Med Inform Assoc. 2001;8(3):254–66.
Tang Y, Yang J, Ang PS, Dorajoo SR, Foo B, Soh S, et al. Detecting adverse drug reactions in discharge summaries of electronic medical records using Readpeer. Int J Med Inform. 2019;128:62–70.
Ang PS, Tham MY, Tan SH, Soh BLS, Foo BPQ, Loke CWP, et al. Towards human-machine collaboration in creating an evaluation corpus for adverse drug events in discharge summaries of electronic medical records. Big Data Res. 2016;4(C):37–43.
Jagannatha A, Liu F, Liu W, Yu H. Overview of the first natural language processing challenge for extracting medication, indication, and adverse drug events from electronic health record notes (MADE 1.0). Drug Saf. 2019;42(1):99–111.
Dandala B, Joopudi V, Devarakonda M. Adverse drug events detection in clinical notes by jointly modeling entities and relations using neural networks. Drug Saf. 2019;42(1):135–46.
Yang X, Bian J, Gong Y, Hogan WR, Wu Y. MADEx: a system for detecting medications, adverse drug events, and their relations from clinical notes. Drug Saf. 2019;42(1):123–33.
Chapman AB, Peterson KS, Alba PR, DuVall SL, Patterson OV. Detecting adverse drug events with rapidly trained classification models. Drug Saf. 2019;42(1):147–56.
Johnson AE, Pollard TJ, Shen L, Lehman LW, Feng M, Ghassemi M, et al. MIMIC-III, a freely accessible critical care database. Sci Data. 2016;24(3): 160035.
US Food and Drug Administration. Orange book: approved drug products with therapeutic equivalence evaluations. www.accessdata.fda.gov/scripts/cder/ob/index.cfm. Accessed Feb 2022.
WHO Adverse Reaction Terminology (WHO-ART). 2015. Collaborating Centre for International Drug Monitoring, World Health Organization. Geneva, Switzerland.
Nelson SJ, Zeng K, Kilbourne J, Powell T, Moore R. Normalized names for clinical drugs: RxNorm at 6 years. J Am Med Inform Assoc. 2011;18(4):441–8.
MedDRA®, the Medical Dictionary for Regulatory Activities. 2017. The International Council for Harmonisation of Technical Requirements for Pharmaceuticals for Human Use. Geneva, Switzerland.
Bochkarev VV, Shevlyakova AV, Solovyev V. Average word length dynamics as indicator of cultural changes in society. Soc Evol Hist. 2012;14(2):153–75.
Cocos A, Fiks AG, Masino AJ. Deep learning for pharmacovigilance: recurrent neural network architectures for labeling adverse drug reactions in Twitter posts. J Am Med Inform Assoc. 2017;24(4):813–21.
Christopoulou F, Tran TT, Sahu SK, Miwa M, Ananiadou S. Adverse drug events and medication relation extraction in electronic health records with ensemble deep learning methods. J Am Med Inform Assoc. 2020;27(1):39–46.
Lavertu A, Hamamsy T, Altman RB. Quantifying the severity of adverse drug reactions using social media: network analysis. J Med Internet Res. 2021;23(10): e27714.
Alvaro N, Miyao Y, Collier N. TwiMed: Twitter and PubMed comparable corpus of drugs, diseases, symptoms, and their relations. JMIR Public Health Surveill. 2017;3(2): e24.
Naranjo CA, Busto U, Sellers EM, Sandor P, Ruiz I, Roberts EA, et al. A method for estimating the probability of adverse drug reactions. Clin Pharmacol Ther. 1981;30(2):239–45.
Xuelan F, Graeme K. Expressing causation in written English. RELC J. 1992;23(1):62–80.
Acknowledgements
The authors thank their colleagues from the Information Management Department, Corporate Services Group, Health Sciences Authority for their assistance with the set-up required to run the machine learning models mentioned in this paper. We are grateful to the Academic Informatics Office, National University Health System for providing the data to enable the conduct of this study. We thank Prof. Cynthia Sung for her insightful comments that assisted the implementation of this study. We also thank A/Prof. Cheng Leng Chan, Group Director of the Health Products Regulation Group (HPRG), Dr. Dorothy Toh, Assistant Group Director, HPRG Vigilance, Compliance and Enforcement Cluster and Ms Jalene Poh, Director, Vigilance and Compliance Branch at the Health Sciences Authority for their support to pursue this research to advance the organisation’s pharmacovigilance mission as well as Michael Winther for his valuable programmatic coordination for the SAPhIRE (Surveillance and Pharmacogenomics Initiative for Adverse Drug Reactions) Project.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Funding
This study was conducted under the SAPhIRE Project, funded by a Strategic Positioning Fund grant from the Biomedical Research Council of the Agency for Science, Technology and Research of Singapore (SPF2014/001).
Conflict of interest
The authors have no conflicts of interest that are directly relevant to the content of this article. The view expressed in this article are the authors’ personal views and may not be understood or quoted as being made on behalf or reflect the positions of Health Sciences Authority, NUH and NUS.
Ethics approval
No ethics approval was required for this study.
Consent to participate
Not applicable.
Consent for publication
Not applicable.
Availability of data and material
The MIMIC III data used in this study are considered an open source database.
Code availability
All data were processed and analysed using Python 3.7. Codes are provided in the ESM.
Authors’ contributions
THX, TCHD and DSR designed the research, models and computational framework to analyse the data. APS, LWPC, TMY, TSH, SBLS, FPQB, TCHD and DSR provided the domain expertise for manual annotation of discharge summaries and developed the list of trigger words, short forms and acronyms. LZJ and YWLJ coordinated the provision of NUH data and participated in discussions. TYX, YJ and TKHA set up the analytic architecture to allow the package applications to run the codes for the data analysis. All authors read and approved the final version of this manuscript.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Tan, H.X., Teo, C.H.D., Ang, P.S. et al. Combining Machine Learning with a Rule-Based Algorithm to Detect and Identify Related Entities of Documented Adverse Drug Reactions on Hospital Discharge Summaries. Drug Saf 45, 853–862 (2022). https://doi.org/10.1007/s40264-022-01196-x
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s40264-022-01196-x