Abstract
Introduction
The rapid expansion of the Internet and computing power in recent years has opened up the possibility of using social media for pharmacovigilance. While this general concept has been proposed by many, central questions remain as to whether social media can provide earlier warnings for rare and serious events than traditional signal detection from spontaneous report data.
Objective
Our objective was to examine whether specific product–adverse event pairs were reported via social media before being reported to the US FDA Adverse Event Reporting System (FAERS).
Methods
A retrospective analysis of public Facebook and Twitter data was conducted for 10 recent FDA postmarketing safety signals at the drug–event pair level with six negative controls. Social media data corresponding to two years prior to signal detection of each product–event pair were compiled. Automated classifiers were used to identify each ‘post with resemblance to an adverse event’ (Proto-AE), among English language posts. A custom dictionary was used to translate Internet vernacular into Medical Dictionary for Regulatory Activities (MedDRA®) Preferred Terms. Drug safety physicians conducted a manual review to determine causality using World Health Organization-Uppsala Monitoring Centre (WHO-UMC) assessment criteria. Cases were also compared with those reported in FAERS.
Findings
A total of 935,246 posts were harvested from Facebook and Twitter, from March 2009 through October 2014. The automated classifier identified 98,252 Proto-AEs. Of these, 13 posts were selected for causality assessment of product–event pairs. Clinical assessment revealed that posts had sufficient information to warrant further investigation for two possible product–event associations: dronedarone–vasculitis and Banana Boat Sunscreen--skin burns. No product–event associations were found among the negative controls. In one of the positive cases, the first report occurred in social media prior to signal detection from FAERS, whereas the other case occurred first in FAERS.
Conclusions
An efficient semi-automated approach to social media monitoring may provide earlier insights into certain adverse events. More work is needed to elaborate additional uses for social media data in pharmacovigilance and to determine how they can be applied by regulatory agencies.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Previous work has established that social media can be a viable source for pharmacovigilance signals by drawing information directly from patients that may not have been otherwise reported to regulatory agencies or industry. |
Of 10 recent postmarketing safety signals from the US FDA, public Facebook and Twitter posts showed mentions of one product–event pair before cases were reported to the traditional spontaneous report system. |
Social media can be an adjunct to traditional safety reporting systems to possibly uncover postmarketing safety signals more rapidly, although considerations of noise and volume need to be further characterized. |
1 Introduction
Clinical trials are imperative for evaluating the safety and efficacy of new medical products; however, because of patient selection criteria, small sample sizes, and short durations, they provide limited information on rare harms. As a result, traditional clinical drug trials are not designed to detect rare adverse drug events. This is exemplified by the fact that over one-half of approved drugs have serious adverse events (AEs) identified that are not detected before approval [1]. In addition, existing postmarket AE reporting systems are significantly underutilized by clinicians and patients. It has been estimated that as much as 96% of adverse drug events go unreported to regulators [2].
The normative and burgeoning online representation of human knowledge and the corresponding growth of computing power to analyze unstructured information offer novel opportunities to access the variety of patient-generated experience, which has been referred to as “digital disease detection” when applied to health [3]. Adoption of social media tools and smartphones has surged. The USA alone has 160 million Facebook users [4] and 140 million Twitter users [5]. In the health domain specifically, 59% of US adults had looked online for health information in the past year, and 35% had gone online specifically to determine a medical condition [6]. A high volume of discussions about medical products is occurring online, with patients using social media to publicly share concerns and reactions to medications. There is increasing interest in exploring the use of social media data for pharmacovigilance, but questions remain regarding how to validate and best leverage this data source.
The general benefits and limitations of using social media for pharmacovigilance have been reviewed extensively [7–15]. One area of research has focused on creating online electronic tools that may expedite case reporting by incorporating user-friendly formats, generally referred to as ‘crowdsourcing’ [16]. Data mining or ‘social listening’ case studies have established a base of applied examples of how generally unstructured online electronic sources can be used to derive drug safety information from news and social media, commonly including Twitter, Facebook, Google+, patient forums, news reports, blogs, and others. Notable among these are studies evaluating online discussions of benfluorex [17], antibiotics [18], human papilloma virus (HPV) vaccine and infertility [19], oral antineoplastic drugs [20], duloxetine [21], anti-Parkinsonian agents [22], and opioid analgesics [23]; a study by Coloma et al. [19] used assertion analysis to demonstrate a positive association between rosiglitazone and cardiovascular events in posts acquired from Twitter. Data mining of web search logs has also been considered for pharmacovigilance and drug safety surveillance [24, 25].
Data mining approaches to social media for pharmacovigilance have been published by other academic groups and evaluated in the conduct of the research presented here [26–37]. However, an outstanding fundamental question is whether analysis of social media could lead to earlier detection of rare and serious AEs. This is the objective of traditional spontaneous reporting and represents a high-value use case by which to evaluate social media, especially for regulatory purposes.
It was hypothesized that warnings of safety signals for US FDA-regulated medications would have been generated earlier by monitoring social media in the period prior to the discovery of the safety signals via the current pharmacovigilance surveillance system, the FDA Adverse Event Reporting System (FAERS). The objective of this study was to examine whether cases of specific product–event pairs were reported in social media before they were reported to FAERS.
2 Methods
2.1 Study Overview
The main study outcome was to determine whether social media posts for selected product–event pairs occurred prior to the date of the first FAERS case report and the date when the FDA first considered the reports in FAERS a potential safety signal. The date of the potential safety signal was the time at which the FDA first determined through routine pharmacovigilance that evidence of a concern was sufficient and that a formal safety review would be required. Retrospective social media data were acquired and processed to remove spam and duplicates and to identify AEs. Causality assessment was conducted by an independent third party.
The study process was as follows:
-
1.
Product–event pairs were identified for retrospective analysis.
-
2.
Search terms were selected to identify products, including brand and generic names, and common misspellings.
-
3.
Medical Dictionary for Regulatory Activities (MedDRA®) Preferred Terms (PTs) indicative of the symptomology of the MedWatch alerts were identified.
-
4.
Queries were submitted to a third-party vendor to gather historical data.
-
5.
A vernacular-to-regulatory dictionary was applied to convert the verbatim post text to MedDRA® PTs.
-
6.
Machine learning algorithms were run on the data to automatically identify ‘posts with resemblance to adverse events’ (Proto-AEs), delete spam posts, consolidate duplicates, and remove personally identifiable information.
-
7.
Symptomatic descriptors for each product–event association were used to identify posts for manual review.
-
8.
An experienced drug safety physician manually curated auto-tagged Proto-AEs.
-
9.
An independent third-party medical reviewer conducted causality assessments on the resulting posts.
-
10.
Social media data were reviewed for concordance with the dates provided by the FDA regarding when the safety signals were first identified using the agency’s standard signal detection processes.
2.2 Data Sources and Study Period
The data source was English language public social media posts from Facebook and Twitter collected between March 2009 and October 2014 regardless of geographic location. Follow-up duration was the same for all product–event pairs, starting two calendar years before the first day of the month in which the FDA safety signal was first identified.
Social media data from public Facebook and Twitter posts were obtained from an authorized third-party data vendor, since an estimated 90% of all Twitter accounts [5] and 25% of all Facebook accounts [4] are publicly accessible. The commercially available service provided the verbatim text of the body of the original post along with metadata such as timestamp, a link to the original content, and, if available, the poster’s gender and geographical information. Posts that were truncated but still described AEs and contained links to original content (e.g., a Tweet that linked to a longer post on the discussion board MedsChats.com) were followed to retrieve the full text.
The FDA provided the month and year that the selected safety signals were first identified through the use of routine monitoring of FAERS in combination with other routine data sources for pharmacovigilance. Public data from FAERS were used to extract information on individual cases received by the FDA. Public FAERS data do not contain case narratives, so further information could not be used to determine exact date of onset or a detailed causality assessment. See the Electronic Supplementary Material (ESM) 1 for a list of data elements that were available in the public FAERS dataset and used in this analysis.
2.3 Drug–Event Selection
A total of 15 safety signals, identified between January 2011 and October 2014, retrieved from the FDA website were initially considered for use in the study [38]. The time period was selected based on the availability of historical social media data from a third-party data vendor (Datasift). The final list selected for analysis resulted in 10 product–event pairs (Table 1). The final 10 products were nine prescription medications and one over-the-counter consumer product. See the ESM 2 for a complete list of product–event pairs considered for analysis.
The following rules reduced the list of 15 initial safety signals to 10:
-
Of the 15 safety signals, three were attributed to dronedarone, with another two attributed to Tecfidera. Three of these earlier reported product–event pairs with identical products were removed after it was determined that including multiple events would complicate the selection of negative controls by restricting the number of feasible negative controls. The process for selecting negative controls is described in Sect. 2.5.
-
Acetaminophen–skin reactions was eliminated because the amount of curation required to review the high volume of posts for ‘acetaminophen’ and ‘paracetamol’ was too onerous to be feasible in this study.
Of note, the safety signal for hydroxyzine was specific to an injectable formulation, but all mentions of hydroxyzine were analyzed to reduce the possibility of missing reports by using injectable-specific search terms.
2.4 Selection of Search Terms
The brand names and misspellings in Table 2 were identified for each of the products of interest from a list of hypothesized spelling variants, using the public Twitter search. This is because drugs posted in social media can take on a variety of forms; aside from being referred to by various brand names and active ingredients, medical products can be misspelled or referred to in more general terms (synecdoche). Any search term that resulted in more than 10 posts was included. To maximize signal-to-noise ratio, an additional search criterion was added to include any posts mentioning ‘vivance’ (a common misspelling of Vyvanse) but to exclude ‘la vivance’, a common French word, because the concept is semantically unrelated to drug safety.
For the purpose of identifying cases in FAERS, which contains curated data with corrected misspellings, products were searched for using properly spelled brand and generic names.
2.5 Selection of Controls
The selection of negative controls was modeled after the approach taken by Harpaz et al. [39] while establishing a time-indexed reference standard for adverse drug reactions. This approach defines negative controls by randomly pairing each product that appeared in the set of positive controls with one event that appeared in the set of positive controls. This process is repeated until each product is paired with at least one negative control event.
In addition, we avoided assigning negative controls where the signs and symptoms indicative of the negative control medical concept may be associated with a product’s indication and therefore may trigger a false positive.
FDA safety reviewers then checked product labeling to ensure that the events were not AEs listed in the product labeling for the randomly assigned negative control product. The most recent product label in October 2014 that was available from the National Library of Medicine DailyMed site was used [40]. Of the resulting matches, four were excluded because the AE was listed as an AE in the label for that drug. The six resulting product–event pairs were selected as negative controls: dabigatran etexilate (Pradaxa) and Raynaud’s phenomenon, dimethyl fumarate (Tecfidera) and priapism, hydroxyzine (Atarax) and increased international normalized ratio (INR), ziprasidone (Geodon) and vasculitis, lisdexamfetamine (Vyvanse) and drug reaction with eosinophilia and systemic symptoms (DRESS), olmesartan (Benicar) and progressive multifocal leukoencephalopathy (PML).
2.6 Definitions
We utilized a previously created colloquial symptom dictionary in the social media listening platform to map colloquial phrases in social media posts to MedDRA®, the international medical terminology developed under the auspices of the International Council for Harmonisation of Technical Requirements for Registration of Pharmaceuticals for Human Use (ICH) [41]. The MedDRA® taxonomy comprises a five-level hierarchy of terms and codes describing, to varying degrees of specificity, a wide range of medical conditions, diagnoses, treatments, investigations, and signs and symptoms. The hierarchical levels are defined by degree of specificity, starting at the most general ‘System Organ Class’ (SOC) that describes organ- or site-specific disorders. The fourth level of specificity, ‘Preferred Term’ (PT) was chosen for this analysis. A PT represents a distinct clinical concept and is routinely used in pharmacovigilance, including in FAERS data. As described elsewhere [29], the colloquial symptom dictionary was developed over four years through a manual curation process. When curators encountered vernacular terms in social media describing medical conditions, the phrases were added to the dictionary. The dictionary encompasses euphemisms (e.g., “I look like a lobster” codes to Erythema), combined words in the form of hashtags (denoted in social media using the # symbol), variations between American and British English, misspellings, abbreviations, homonym confusion, affixes (e.g., plural -s, past tense -ed), homographs, heteronyms, metonyms or synecdoche, and other facets of semantics. At the start of this study, the colloquial symptom dictionary contained 8857 vernacular terms mapped to 1956 of the most common PTs. Phrases are mapped to the MedDRA® (version 18) PT level since PTs represent distinct clinical concepts and are routinely used in pharmacovigilance, including coding AEs in FAERS data. Each colloquial phrase identified in social media describing a symptom was mapped to a MedDRA® PT as described elsewhere [29, 42]. For our study, all PTs identified in this analysis for the 10 AEs of interest were available in the colloquial symptom dictionary, including all corresponding AE synonyms.
Another research group found low recall when using a lexicon-based approach [26]. The lexicon studied was optimized for searching biomedical literature. As such, we would not expect that specific lexicon to perform well in social media posts in Twitter and Facebook. However, the lexicon used in our analysis was specifically generated based on empirical observation of patient conversations in social media. Therefore, the results observed by Nikfarjam et al. [26] cannot be generalized to this study.
2.7 Case Detection
The unit of analysis was a single post in Facebook or Twitter that mentioned both a product name and an AE. Those AEs mentioned with a specific product are referred to as Proto-AEs, a term used to differentiate AEs described in social media posts from AE reports that have been vetted to meet the minimum regulatory requirement for reporting AEs, such as those in FAERS. Proto-AEs are conceptually modeled on the regulatory reporting definition, as we consider a network username to be similar to a patient identifier for a MedWatch report.
We identified standardized symptom lists and definitions that were used to consistently evaluate the Proto-AEs identified in social media posts. This was because patients were more likely to report signs and symptoms of an AE than a medical diagnosis.
To screen social media posts for cases for causality assessment, symptoms related to the 10 AEs of interest needed to be identified (see ESM 3) to build an event profile for automated Proto-AE detection.
The process used was as follows:
-
If available, Standardized MedDRA® Queries (SMQs) were identified, yielding definitions for two AEs: angioedema and vasculitis.
-
All terms related to medical devices, injection, infusion, implant, stents, vaccinations, and syringes or needles were removed from the angioedema list because only solid oral formulations were under review among drug products.
-
Injection site-related PTs were selected from the Extravasation events MedDRA® SMQ.
-
To give preference to regulatory precedent on definitions, the FDA website was searched (using search engine-indexed site-specific syntax) to identify additional definitions, resulting in the identification of symptom lists for renal impairment and sprue-like enteropathy.
-
If other definitions were still not available, clinical reference books (e.g., Merck Manual) and trusted medical websites (e.g., Mayo Clinic and National Institutes of Health [NIH]) were consulted and converted into MedDRA® PTs.
The number of unique PTs for each condition varied considerably; angioedema had by far the highest number of symptoms.
2.8 Indicator Score Calculation
A machine learning algorithm was employed to distinguish posts containing relevant drug safety information from non-informative posts. This was achieved using a statistical approach for automated document classification that categorized each post as either a Proto-AE or a non-Proto-AE, based on a manually coded training set using Certified MedDRA® Coders, as previously described [29, 42]. This training set consisted of data collected from Facebook and Twitter between October 2012 and December 2014 using application programming interfaces made publicly available by Facebook and Twitter. There was no overlap between the training set data and the study data. This involved feature extraction, a process by which each post was broken up into a set of discrete elements that were then tallied to determine the number of times each element (or feature) occurred in posts that had been manually coded as Proto-AEs and in posts that had been coded as non-Proto-AEs. To improve computational efficiency, the text was parsed using a single tree structure, based on the dictionary-matching algorithm originally developed for HealthMap, adapted to identify mentions of products and symptoms [29, 42]. Loading the product and symptom taxonomies into a single tree structure enabled the post to be processed a single time to identify tokens that may have represented both products and symptoms. Next, all punctuation was removed, and the post was divided into tokens comprising one- to three-word phrases. Finally, each token was processed and matched against the vernacular dictionary. Positive matches were established when a ‘leaf’ was reached in the tree and a phrase for either a product or a symptom was identified based on the taxonomy.
After the feature extraction algorithm was run on the input, the classifier, developed through statistical machine learning computation, was applied to each post. The model for the classifier was taken from Robinson’s approach to eliminating spam in email [43]. Using a Bayesian probabilistic model and drawing on a training dataset containing more than 360,000 manually classified posts, the software assigned each post an indicator score on a scale of 0 to 1. For each feature extracted from a post, we calculated a probability score representing the likelihood that the post was a Proto-AE based on the manually classified corpus. We then combined probabilities across features using the inverse chi-squared function and calculated an overall indicator score, as detailed below. A score close to 0 meant there was a low probability that the post was related to an AE discussion (usually spam), whereas a score close to 1 indicated a high probability that the post was related to an AE discussion. In general terms, each new token was compared with the priors, and two probabilities were calculated based on the number of times the token appeared in positive and negative posts in the manually classified priors. Finally, we empirically determined that if a post contained no identifiable symptom, it was less likely to contain a true AE report. To account for this, and as a performance improvement measure, we therefore deducted a penalty of 0.2 from the indicator score if the post did not contain any identifiable symptoms.
The process for computing the probabilities, combining them, and producing an indicator score was as follows. For each token ‘w’:
-
b(w) = (number of positives containing w)/(total number of positives)
g(w) = (number of negatives containing w)/(total number of negatives)
p(w) is the ‘raw’ probability that the document is a positive (AE) given that it contains word ‘w’.
To adjust for rare tokens, i.e., phrases that did not occur frequently or at all in the training set, the following statistical adjustment was undertaken:
f(w) is a scaled or ‘weighted’ transformation of p(w). The weighting is based on priors as well as the frequency of w in the training set.
s: ‘strength’ assigned to background (training) information
x: assumed probability of spam for unknown token
n: number of examples containing w
Strength (s) and probability of positive (x) are priors. 1.0 and 0.5 are considered neutral values for s and x, respectively; in our implementation, we used 1.4 for the value of s and 0.6 for the value of x. These values were determined by experimentation, namely, checking performance metrics for a range of values and selecting the values yielding the best performance. No formal optimization methodology was applied for this process.
Probabilities were derived using Fisher’s method of combining probabilities, where C −1 is shorthand for the inverse chi-squared function:
Complementary probabilities were combined in this manner:
The indicator score was then calculated by averaging the combined probabilities with the inverse probabilities.
We average the positive score (H) and the negative score (S) to produce a bimodal distribution of indicator (I) values. That is, with the resulting distribution of I in our dataset, we see large numbers of posts scoring either very low (high likelihood of spam) or very high (high likelihood of AE) and a smaller number scattered across the middle. The middle of the distribution represents content that the algorithm determined to be ambiguous. Since there is not enough human analyst time to check and reclassify all posts, we can then focus our analyst efforts on posts falling in the middle of the distribution. Second, the nature of the distribution makes the performance less sensitive to the exact indicator threshold value, since we place the threshold near the middle of the distribution.
Posts with indicator scores of 0 were generally spam, e.g., advertisements, coupons, or other posts containing text clearly generated by bots. Posts with indicator scores >0 were called ‘mentions.’
An analysis of classifier performance suggested that optimal thresholds for removing irrelevant posts without excessively sacrificing true positives were indicator scores of 0.6 to 0.7 [42]. In the previous analysis, posts with indicator scores of ≥0.65 had a positive predictive value of approximately 0.5 (i.e., 5 of every 10 posts would be relevant for drug safety), while keeping 92% of true AE posts.
2.9 Human Curation
Given the nuanced nature of social media language, previous experience has consistently suggested that review by a human curator is necessary on some posts that have been automatically identified as AEs by the classifier. This process helps to remove false positives, continuously improve the classifier, and expand the symptom and product taxonomies. This approach, called ‘active learning,’ introduces a feedback loop between the human curators and the automated classifier [29]. The high volume of data in this study meant it was not practical to manually review every auto-tagged post; therefore, we established a threshold to identify potentially ambiguous posts that may require curation based on previous analysis of social media data using the machine classifier [42]. Posts with indicator scores of at least 0.65 were reviewed by a team of human curators that included Certified MedDRA® Coders and medical professionals with a focus on removing false positives, called ‘primary curation.’ All curators received standardized training on curation methods for social media data, according to the MedDRA® coding standards and ICH-endorsed guidelines. Standard MedDRA® practices were used, including objective coding without any attempt to interpret or infer beyond the stated text. Curators receive annual refreshers, as well as training on each new release of MedDRA®. Primary curation is an ongoing process that is conducted for drugs in the general system; however, to prepare the data for causality assessment in this analysis, an additional round of curation was needed.
Accordingly, ‘secondary curation’ was then conducted to identify relevant posts of interest for the hypothesis in this paper. An experienced drug safety physician (HVL) reviewed posts that had undergone primary curation for a given drug that had any symptom (e.g., from the SMQ list) associated with the product–event pair from positive associations and negative controls. Posts that appeared to describe the product–event pairs of interest were extracted for causality assessment. For SMQs, the presence of any symptom was not sufficient to trigger a positive match; the symptoms listed for SMQs in ESM 3 are often non-specific. As such, it was left to the discretion of the physician reviewer to determine whether the constellation of symptoms described was sufficient for a plausible positive match.
2.10 Causality Review
World Health Organization-Uppsala Monitoring Centre (WHO-UMC) criteria for causality assessment [44] were assigned to each post identified during secondary curation. To accomplish this, the text of the post, along with date and data source, was sent to WHO-UMC for independent review by an experienced senior medical officer (IRE). Posts were classified as being ‘certain,’ ‘likely,’ ‘possible,’ ‘unlikely,’ or ‘unclassified.’ The differentiating factor between certain/likely and possible was the presence of time-to-event information; more granular and plausible time information elevated a case to certain as opposed to likely. It was assumed that most social media posts would likely fall in the possible category since time-to-event information is not routinely reported by patients.
2.11 Ethics Statement
All social media data used in this analysis came from public sources. Prior to causality assessment, posts were stripped of personally identifiable information (PII), such as screen names, user names, first and last names, and addresses. A hybrid automated–manual process was used to remove first and last names. Using a ‘PII dictionary’ containing a few hundred thousand names from a global database of values [42], the software removed from the posts any names that were contained in the dictionary. In addition, text-based pattern matching was used to remove structured information (e.g., phone numbers, email addresses) that could lead to individual identification, other than for the needs of manual review. ‘Verbatim’ post text presented in this manuscript has been paraphrased and/or altered in non-meaningful ways to protect patient identity and prevent unmasking using Internet search engines. AE data from FAERS were also publicly available in an anonymized format. Since all primary data sources used in the analysis were publicly available, no institutional review board approval was sought.
3 Results
3.1 Data Processing Results
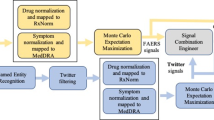
Figure 1 represents the schematic flow of posts and steps taken in data cleaning as well as the number and percent of posts resulting from each process. We started with an initial dataset of 935,246 posts containing text of selected products. These were posted on public Facebook and Twitter accounts from March 2009 to October 2014. Spam posts were identified and eliminated based on indicator scores (less than zero) implying that the posts did not contain text that met our requirements (i.e., posts with the name of a product and only an emoticon, or in non-English languages). This spam-elimination step resulted in the removal of about 25% (n = 230,963) of the remaining posts, leaving a corpus of 704,283 non-spam posts mentioning the products of interest. Next, verbatim duplicates (8.3%, n = 58,188) were consolidated. A total of 98,252 Proto-AEs were then identified based on indicator scores of ≥0.65, representing about 15% of the remaining posts. After primary and secondary manual curation, which occurred over the course of three months, 13 posts were identified as possibly relevant and were then submitted for causality assessment. This process resulted in six posts of interest that described certain, probable, and possible cases.
Data processing results. Public social media data were obtained for Facebook and Twitter, with 935,246 posts identified where each of the 10 products under review were mentioned. An automated classifier was used to remove spam, followed by exclusion of verbatim duplicate posts. Posts with an indicator score of ≥0.65 were considered a ‘post with resemblance to an adverse event’ and manually curated to remove false positives. The resulting 13 posts that mentioned product–event associations from alerts or negative controls were assessed for causality by an independent third party. Six posts contained certain, probable, and possible cases. AE adverse event, Proto-AE post with resemblance to an adverse event
3.2 Comparison of Twitter and Facebook Data
Volume of mentions varied considerably by product, with controlled substances (e.g., lisdexamfetamine, methylphenidate) showing much higher volume due in considerable part to Internet pharmacy advertisements (Fig. 2). Mentions for the products of interest varied over time, with rapid increases after new product launches (e.g., lisdexamfetamine and Tecfidera). Surges in post counts generally tended to express excitement about the new products among patient communities, but also included sales and stock price forecasts for finance audiences. Most posts came from Twitter, compared with Facebook (Fig. 3). Twitter generally yielded a higher volume of mentions overall due to the prevalence of pharmacy advertisements, references to financial reporting, and links to medication-related news articles or literature. The percent of posts from Facebook varied from 1.4% (lisdexamfetamine [Vyvanse]) to 27.3% (dimethyl fumarate [Tecfidera]). The higher proportion of posts from Facebook for dimethyl fumarate originated from patients posting within open patient groups rather than individuals posting on their personal pages. While Twitter typically generates more posts mentioning medical products, Facebook data typically consists of higher-quality posts with more detailed information. Facebook’s post format and its numerous open patient groups provide a platform for patients to share their experiences without the character limits imposed by Twitter.
3.3 Results of Causality Assessment
In total, 13 posts were submitted to WHO-UMC for causality assessment. The assessor was provided with the complete post of the text and told which product–event pair was under examination for each. The posts covered 5 of the 10 product–event pairs identified in Table 1: ziprasidone–DRESS, methylphenidate–priapism, dimethyl fumarate–PML, dronedarone–vasculitis, and Banana Boat Sunscreen Spray–skin burns. The manual curator assigned the following scores to each of the 13 product–event pair posts: ziprasidone–DRESS: three unlikely posts; dronedarone–vasculitis: three possible posts, one probable post; methylphenidate–priapism: three unlikely posts; dimethyl fumarate–PML: one unlikely post; Banana Boat Sunscreen Spray–skin burns: two certain posts. Of the posts given the classification of unlikely, the greatest limitation was that the vernacular language used was not specific enough to constitute a causal relationship with the paired condition. For example, one Twitter post stated, “I’m severely allergic to geodon—had to go to the hospital just 8 hours after I took my dose.” The phrase “severely allergic” did not adequately describe DRESS. Similarly, another post read, “my doc wants me to try a new anti-psychotic. im fuckin terrified, i dont wanna, i almost lost my life a few times from it, im the only person i know who’s allergic to abilify and geodon.” In this instance “almost lost my life” did not specifically suggest DRESS. In a third example, “So, they took me off Geodon because I had an extreme attack last week and was trying to go into cardiac arrest, that was fun [truncated].”, again “extreme attack” and “trying to go into cardiac arrest” did not provide enough detail to determine a causal relationship to DRESS. All three of these posts would be considered serious AEs and potentially useful information, but they did not meet the strict intent of this analysis. These posts were all reported in social media before the FDA signal threshold for ziprasidone–DRESS had been reached from spontaneous reports.
For dimethyl fumarate–PML, a single Facebook post was identified from secondary curation that was eventually adjudicated by WHO-UMC to be unlikely: “I tried taking Tecfidera and it made me SO sick I thought I was gonna die. Now I’m thinking of switching to Tysabri, but I tested positive for JC virus. My index value was 1.5 and the doctor told me that risk for PML was low at that range. Just wondering if anyone else is JCV positive and doing well on Tysabri?” In this instance, the link to PML could not be definitively established because no specific adverse drug reaction was being described. However, it is also worth noting that JC virus seropositivity is considered the differentiating risk factor for PML with this medication [45]. In strictly medical terms, the combination of a positive JC virus test and “SO sick I thought I was gonna die” may not be sufficient to describe PML, but nonetheless implies a potentially severe and particularly inauspicious event experienced by the patient. This post was reported in Facebook before the FDA signal threshold for dimethyl fumarate–PML had been reached.
For dronedarone, four specific posts mentioned symptoms that were consistent with vasculitis, all observed in Twitter, but two of these originated in a patient discussion board. One example stated, “Been on 400 mg of Multaq, 2× daily, for almost 6 weeks. Four days ago, noticed I had a rash on the palms of both my hands with dots that kind of look like small blood blisters.” This case was considered probable by the WHO-UMC causality reviewer because of the realistic time-to-onset mentioned in the post. The symptomology in the specific location of the extremities is a known characteristic of vasculitis, with the description of “like small blood blisters” further confirming the suspicion.
Finally, hypersexuality has long been associated with synthetic pharmaceutical stimulants, from amphetamine to methylphenidate and others [46, 47]. Identifying priapism cases specifically within a background of normative hypersexual references led to difficulty in drawing a causal link. For example, “@user says …”is one of the side effects of ritalin a hard cock?” LMFAO.” Since no pathology is described in this post, the WHO-UMC reviewer made the determination that it had unlikely causal connection. The other two posts reviewed for this product–event pair also had similar limitations.
3.4 Review of Social Media Posts
The 13 posts identified as a result of secondary curation appeared to be discussing symptoms associated with the pre-specified product–event pairs. No posts were found to contain symptoms for the negative controls after curation. The 13 posts were associated with 2 of the 10 FDA-identified safety signals, specifically dronedarone-associated vasculitis and skin burns resulting from Banana Boat Sunscreen Spray (Table 3). There did not appear to be specific symptomology discussed in social media for the other eight safety signals identified by routine surveillance methods.
For Banana Boat Sunscreen Spray, the first mention was on Facebook on 1 June 2012, followed a few hours into the next day on Twitter. Over the next week, 19 Facebook posts and 144 Twitter posts discussed this topic, fueled initially by a local television morning news story on 3 June.
3.5 Timing of Social Media Posts Compared with FAERS Reporting
For dronedarone–vasculitis, the social media case was identified from Twitter on 5 December 2010. No cases were identified in Facebook. The first FAERS primary suspect drug case was received on 7 July 2010, with signal detection reported by the FDA on 15 April 2012. Four social media reports were evaluated as possible (n = 3), probable (n = 1), or certain (n = 2), compared with 10 unique Individual Case Safety Reports (ICSRs) leading to signal detection by the FDA.
For Banana Boat Sunscreen–skin burns, the first social media case was identified from Twitter on 2 June 2012, whereas the first Facebook case was identified on 1 June 2012. The first FAERS case was received on 19 June 2012, with signal detection reported later that month by the FDA. Two social media reports were evaluated as possible or probable, compared with one unique ICSR leading to signal detection by the FDA. The FAERS data suggest that (presumably separate) events occurred on 28 May 2012 and 12 June 2012. It is possible that the case reported in social media was the one that occurred on 28 May, but this cannot be verified. Therefore, only a single post count was attributed.
4 Discussion
This analysis applied strict criteria for evaluating the usefulness of social media data in detecting serious AEs. Despite the very specific nature of this analysis, of the 10 safety signals studied, one was mentioned in social media data prior to meeting FDA signal detection thresholds, whereas another appeared in FAERS prior to social media. The events selected for this analysis were very specific diagnoses, whereas patients are more likely to discuss their medical conditions in common terms and symptoms. Therefore, it was necessary to cast a wide net using a large collection of terms consisting of any symptom that could possibly be associated with the selected medical event. This resulted in a very large number of collected posts that ultimately were deemed irrelevant to the product–events of interest, such as posts that referred to other non-medical products or individuals (e.g., musicians) that happened to have the same name as the chosen medical products. The combined lexical and machine learning approach used in this analysis had a performance of 0.88 sensitivity (recall) and 0.68 positive predictive value (precision) [29]. This approach was selected on the basis of performance after comparison with other machine learning techniques [48]. A recent analysis using Twitter and one patient forum with a training set of 6000 posts (limited to 81 drugs) returned recall of 0.68–0.78 and precision in the range of 0.76–0.86 with a semantic approach; by comparison, in that analysis, a lexical approach had 0.61–0.72 recall and 0.56–0.58 precision [26].
The WHO-UMC causation scale is a high standard when applied to social media. The drug and the AE must be well described and specific in addition to having plausible time relationships. Possible confounders will reduce the likely causality. When one considers the usefulness of trawling the Internet, WHO-UMC reviewers think of signals that describe ‘something serious happening,’ and the ‘something’ is characterized cohesively enough to suggest the effect is shared by a few unique individuals (generally more than three). Seriousness should therefore be a signal in its own right so that “extreme attack” and “trying to go into cardiac arrest” would be as important a signal as something with a suggestive causal link to a specified disease entity; WHO-UMC reviewers feel that they would always be interested in such reports when identifying and evaluating signals. Perhaps these are the most important contributions from social media—that an early serious concern is expressed about a drug harm before it is medicalized.
Why social media monitoring did not yield early safety concerns for the other eight product–event pairs bears consideration. One possible explanation is that the events used in this analysis were relatively severe and/or serious, and our assumption is that patients would have sought medical treatment for many of these outcomes. Therefore, there is a likelihood that a patient’s provider might find the event to be significant enough to report, whereas a patient might consider a more severe diagnosis to be private information, thereby increasing the chances of it being detected through formal reporting channels and not mentioned on social media. Similarly, many of the events involved in this analysis may have required formal medical diagnosis for detection or may require medical expertise to describe it to the extent that it might trigger a signal.
A previous study by the authors investigated concordance at the MedDRA® SOC level between social media reports and FAERS data [29]. General disorders (usually less serious) were most commonly reported in social media, with patients describing symptoms in more general terms, e.g. “stomach problems” (“Abdominal symptom”), implying that whereas some patients are keen to describe their side effects publicly, many do not describe symptoms using the medical terminology needed to explicitly confirm a certain condition or diagnosis.
This research contributes to the literature by providing a scientific approach to identifying potential posts containing AE information using an automated process, followed by structured manual review by drug safety physicians, including a process for assessing causality using an international standard.
As we mentioned above, the nature of the severe events selected for this analysis required us to rely on a large selection of terms for common signs and symptoms that would provide us with a large enough corpus from which to work. This methodology might have been improved upon—and search results could have been much narrower—perhaps by developing an algorithm for each separate event of interest or product that assigned weights to each sign and symptom according to the likelihood that it would indicate the event of interest.
One unresolved key question is how to operationalize social media surveillance for regulated medical products. Given the curation burden and the large volume of social media data, prospective monitoring of all regulated products may have limitations. Social media could be used to stimulate a safety assessment that the FDA was not already considering. Some pharmaceutical companies have progressed to monitoring a portfolio of products [42], but government agency monitoring of the pharmacopeia will require additional effort. Whereas other researchers have developed more directly automated screening methods to identify potential posts of interest (e.g., AEs), a critical component of our study was to elucidate a process by which machine-identified posts could be evaluated for causation within the existing global frameworks used by regulators and industry. This process is a bridge between the computer science tools and existing pharmacovigilance methods. A future can be envisioned of greater machine dependence, but adjudication of causality is likely to remain a fundamentally human function.
Beyond this, the results of this analysis elicit points for discussion. First, further research is needed to develop best practices and methods for determining what constitutes a ‘safety signal’ in social media. It can be hypothesized that higher signal thresholds than those used in traditional pharmacovigilance may be appropriate for signal detection in social media data, as these data are not specifically intended to be used for reporting adverse drug reactions. A line of interdisciplinary research has been initiated to explore alternative methods for quantitative signal detection using social media data [7] in addition to an approach using proportional reporting ratios [42]. Second, social listening for postmarketing safety surveillance is a new field, and best practices are still being established. There is some uncertainty on the appropriate use of these data from a regulatory and patient privacy point of view.
There are several technical limitations to this study that bear mentioning. First, there is inherent variability across data sources that can change rapidly over time. This may include the types of people posting on a particular website and limitations caused by website characteristics (e.g., character limitation). In 2015, Facebook enacted a policy to no longer make post-level verbatim text available for data-mining purposes, citing privacy concerns. This effectively makes the world’s largest social network unavailable for future drug safety surveillance, without special dispensation. Other social networking sites may also make their user-generated data not public, meaning it will be inaccessible using the methods described herein. Second, Internet vernacular may not accurately reflect a medical condition. To alleviate this limitation, the vernacular-to-MedDRA® dictionary used for this study was able to pick up on many such peculiarities of language, but some may have been missed. Other researchers have proposed solutions to developing an exhaustive consumer health vocabulary, which will be incorporated in future analyses [49, 50]. Third, there is the possibility that people may post the same information in multiple social media platforms. As a result, some duplication may exist. However, this was not apparent to curators while manually reviewing posts for this analysis. Potential for duplication will need to be considered in the development of quantitative signal detection methodologies, alongside the perennial problem that stems from the lack of an obvious denominator for rate calculation in spontaneously reported AE data. Finally, the manual curation steps described here were considered necessary to provide a medical understanding of the social media posts. A practical limitation to including the human curation step is in the volume of reports that can be processed. For example, acetaminophen/paracetamol products were not included in this analysis because the volume was too high for manual curation. Specific investigations may need to consider approaches that rely more on automation.
5 Conclusion
Although we do not foresee that social media monitoring will replace current spontaneous report systems, we present one example of how social media listening, even for rare and serious events, can be modeled as an early warning system for emerging drug safety concerns. In 1999, Cobert and Silvey [51] commented “we live in electronically exciting times” in reference to the then-impending explosion of Internet-based options for pharmacovigilance. A decade and a half later, we feel that their vision for the future of drug safety is being borne out and will continue to expand in coming years.
References
Deyo R. Gaps, tensions, and conflicts in the FDA approval process: implications for clinical practice. J Am Board Fam Med. 2004;17(2):142–9.
Smith CC, Bennet PM, Pearce HM, et al. Adverse drug reactions in a hospital general medical unit meriting notification to the Committee on Safety of Medicines. Br J Clin Pharmacol. 1996;42:423–9.
Brownstein JS, Freifeld CC, Madoff LC. Digital disease detection—harnessing the Web for public health surveillance. N Engl J Med. 2009;360(21):2153–7. doi:10.1056/NEJMp0900702.
Brown D. 52 Cool facts about social media--2012 edition dannybrown.me. 2012. http://dannybrown.me/2012/06/06/52-cool-facts-social-media-2012. Accessed 6 Oct 2015.
Beevolve. An exhaustive study of Twitter users across the world--social media analytics. Beevolve. 2012. http://temp.beevolve.com/twitter-statistics/-cl. Accessed 6 Oct 2015.
Fox S, Duggan M. Health Online 2013. Pew Internet: Pew Research Center, 2013. http://www.pewinternet.org/2013/01/15/health-online-2013. Accessed 15 Oct 2015.
Ghosh R, Lewis D. Aims and approaches of Web-RADR: a consortium ensuring reliable ADR reporting via mobile devices and new insights from social media. Expert Opin Drug Saf. 2015;14(12):1845–53.
Golder S, Norman G, Loke YK. Systematic review on the prevalence, frequency and comparative value of adverse events data in social media. Br J Clin Pharmacol. 2015;80(4):878–88.
Lardon J1, Abdellaoui R, Bellet F, Asfari H, Souvignet J, Texier N, Jaulent M-C, Beyens M-N, Burgun A, Bousquet C. Adverse drug reaction identification and extraction in social media: a scoping review. J Med Internet Res. 2015;17(7):e171.
Sloane R, Osanlou O, Lewis D, Bollegala D, Maskell S, Pirmohamed M. Social media and pharmacovigilance: a review of the opportunities and challenges. Br J Clin Pharmacol. 2015;80(4):910–20.
Sarker A, Ginn R, Nikfarjam A, O’Connor K, Smith K, Jayaraman S, Upadhaya T, Gonzalez G. Utilizing social media data for pharmacovigilance: a review. J Biomed Inform. 2015;54:202–12.
Norén GN. Pharmacovigilance for a revolving world: prospects of patient-generated data on the internet. Drug Saf. 2014;37(10):761–4.
White RW, Harpaz R, Shah NH, DuMouchel W, Horvitz E. Toward enhanced pharmacovigilance using patient-generated data on the internet. Clin Pharmacol Ther. 2014;96(2):239–46.
Masoni M, Guelfi MR, Conti A, Gensini GF. Pharmacovigilance and use of online health information. Trends Pharmacol Sci. 2013;34(7):357–8.
Edwards IR, Lindquist M. Social media and networks in pharmacovigilance: boon or bane? Drug Saf. 2011;34(4):267–71. doi:10.2165/11590720-000000000-00000.
Yamamoto M, Kubota K, Okazaki M, Dobashi A, Hashguchi M, Doil H, et al. Patients views and experiences in online reporting adverse drug reactions: findings of a national pilot study in Japan. Patient Prefer Adher. 2015;9:173–84. doi:10.2147/PPA.S75761.
Abou Taam M, Rossard C, Cantaloube L, Bouscaren N, Roche G, Pochard L, et al. Analysis of patients’ narratives posted on social media websites on benfluorex’s (Mediator(R)) withdrawal in France. J Clin Pharm Ther. 2014;39(1):53–5.
Dyar OJ, Castro-Sanchez E, Holmes AH. What makes people talk about antibiotics on social media? A retrospective analysis of Twitter use. J Antimicrob Chemother. 2014;69(9):2568–72.
Coloma PM, Becker B, Sturkenboom MC, van Mulligen EM, Kors JA. Evaluating social media networks in medicines safety surveillance: two case studies. Drug Saf. 2015;38(10):921–30.
Pages A, Bondon-Guitton E, Montastruc JL, Bagheri H. Undesirable effects related to oral antineoplastic drugs: comparison between patients’ internet narratives and a national pharmacovigilance database. Drug Saf. 2014;37(8):629–37. doi:10.1007/s40264-014-0203-6.
Harmark L, van Puijenbroek E, van Grootheest K. Intensive monitoring of duloxetine: results of a web-based intensive monitoring study. Eur J Clin Pharmacol. 2013;69(2):209–15.
Schroder S, Zollner YF, Schaefer M. Drug related problems with Antiparkinsonian agents: consumer Internet reports versus published data. Pharmacoepidemiol Drug Saf. 2007;16(10):1161–6.
Shutler L, Nelson LS, Portelli I, Blachford C, Perrone J. Drug use in the Twittersphere: a qualitative contextual analysis of tweets about prescription drugs. J Addict Dis. 2015;34(4):303–10.
Simmering J, Polgreen LA, Polgreen PM. Web search query volume as a measure of pharmaceutical utilization and changes in prescribing patterns. Res Soc Adm Policy. 2014;10(6):896–903.
White RW, Wang S, Pant A, Harpaz R, Shukla P, Sun W, et al. Early identification of adverse drug reactions from search log data. J Biomed Inform. 2016;59:42–8.
Nikfarjam A, Sarker A, O’Connor K, Ginn R, Gonzalez G. Pharmacovigilance from social media: mining adverse drug reaction mentions using sequence labeling with word embedding cluster features. J Am Med Inform Assoc. 2015;22(3):671–81. doi:10.1093/jamia/ocu041.
Sarker A, Gonzalez G. Portable automatic text classification for adverse drug reaction detection via multi-corpus training. J Biomed Inform. 2015;53:196–207.
Sarntivijai S, Abernethy DR. Use of internet search logs to evaluate potential drug adverse events. Clin Pharmacol Ther. 2014;96(2):149–50.
Freifeld CC, Brownstein JS, Menone CM, Bao W, Filice R, Kass-Hout T, et al. Digital drug safety surveillance: monitoring pharmaceutical products in twitter. Drug Saf. 2014;37(5):343–50.
O’Connor K, Pimpalkhute P, Nikfarjam A, Ginn R, Smith KL, Gonzalez G. Pharmacovigilance on Twitter? Mining tweets for adverse drug reactions. AMIA Annu Symp Proc. 2014;2014:924–33.
Wu H, Fang H, Stanhope SJ. Exploiting online discussions to discover unrecognized drug side effects. Methods Inf Med. 2013;52(2):152–9.
Azadeh N, Sarker A, O’Connor K, Ginn R, Gonzalez G. Pharmacovigilance from social media: mining adverse drug reaction mentions using sequence labeling with word embedding cluster features. J Am Med Inform Assoc. 2015;22:671–81. doi:10.1093/jamia/ocu041.
Yang M, Kiang M, Shang W. Filtering big data from social media—building an early warning system for adverse drug reactions. J Biomed Inform. 2015;54:230–40. doi:10.1016/j.jbi.2015.01.011.
Kunac DL, Tatley MV, Seddon ME. A new web-based Medication Error Reporting Programme (MERP) to supplement pharmacovigilance in New Zealand—findings from a pilot study in primary care. NZ Med J. 2014;127(1401):69–81.
Karimi S, Metke-Jimenez A, Kemp M, Wang C. Cadec: a corpus of adverse drug event annotations. J Biomed Inform. 2015;55:73–81.
Benton A, Ungar L, Hill S, Hennessy S, Mao J, Chung A, et al. Identifying potential adverse effects using the web: a new approach to medical hypothesis generation. J Biomed Inform. 2011;44(6):989–96.
Mao J, Chung A, Benton A, Hill S, Ungar L, Leonard CE, et al. Online discussion of drug side effects and discontinuation among breast cancer survivors. Pharmacoepidemiol Drug Saf. 2013;22(3):256–62.
US Food and Drug Administration. Medical product safety information. Silver Spring: US FDA; 2016.
Harpaz R, Odgers D, Gaskin G, DuMouchel W, Winnenburg R, Bodenreider O, et al. A time-indexed references tandard of adverse drug reactions. Scientific Data. 2014;43:1–10.
National Library of Medicine. DailyMed: National Institutes of Health; 2016. https://dailymed.nlm.nih.gov. Accessed 14 Oct 2014.
MedDRA The Medical Dictionary for Regulatory Activities. MedDRA. 2015. http://www.meddra.org. Accessed 17 Dec 2015.
Powell G, Seifert H, Reblin T, Burstein P, Blowers J, Menius J, Painter JL, Thomas M, Pierce CE, Rodriguez HW, Brownstein JS, Freifeld CC, Bell HG, Dasgupta N. Social media listening for routine post-marketing safety surveillance. Drug Saf. 2016;39(5):443–54.
Robinson G. A statistical approach to the spam problem. Linux J. 2003;107:3.
World Health Organization. The use of the WHO-UMC system for standardised case causality assessment 2011. http://www.who-umc.org/Graphics/24734.pdf. Accessed 16 Dec 2016.
Calabrese LH, Molloy E, Berger J. Sorting out the risks in progressive multifocal leukoencephalopathy. Nat Rev Rheumatol. 2015;11(2):119–23.
Jones HB. The effects of sensual drugs on behavior: clues to the function of the brain. Adv Psychobiol. 1974;2:297–312.
Greaves G. Sexual disturbances among chronic amphetamine users. J Nerv Ment Dis. 1972;155(5):363–5.
Freifeld, CC (2014). Digital pharmacovigilance: The MedWatcher system for monitoring adverse events through automated processing of Internet social media and crowdsourcing (Doctoral dissertation). Retrieved from ProQuest Dissertations and Theses database (UMI No. 3581025).
Doing-Harris KM, Zeng-Treitler Q. Computer-assisted update of a consumer health vocabulary through mining of social network data. J Med Internet Res. 2011;13(2):e37.
Zeng QT, Tse T, Divita G, Keselman A, Crowell J, Browne AC, Goryachev S, Ngo L. Term identification methods for consumer health vocabulary development. J Med Internet Res. 2007;9(1):e4.
Cobert B, Silvey J. The Internet and drug safety: what are the implications for pharmacovigilance? Drug Saf. 1999;20(2):95–107.
Acknowledgements
The authors thank the many curators who have contributed to training the classifier, including Chi Bahk, Wenjie Bao, Anne Czernek, Michael Gilbert, Melissa Jordan, Christopher Menone, and Carly Winokur. External independent review of cases was conducted by IRE; he was invited to be an author afterwards in recognition for his contributions. The authors also thank Robert Ball, Henry (Skip) Francis, and other FDA employees for their support of this project. The MedDRA® trademark is owned by the International Federation of Pharmaceutical Manufacturers and Associations (IFPMA) on behalf of ICH.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Funding
All research presented here was conducted by the authors listed. Additional general development funds for the social listening platform were provided to Epidemico through a public–private partnership but were not used to directly support the specific content of this research. This collaborative effort is provided via the WEB-RADR project (http://www.web-radr.eu), which is supported by the Innovative Medicines Initiative Joint Undertaking (IMI JU) under Grant Agreement No. 115632, resources of which are composed of financial contributions from the EU’s Seventh Framework Programme (FP7/2007-2013) and European Federation of Pharmaceutical Industries and Associations (EFPIA) companies’ in-kind contribution (http://www.imi.europa.eu). In addition, general support for the development of the social media listening platform was provided by GlaxoSmithKline (GSK), independent of the research presented herein. GSK, WEB-RADR, and IMI JU had no role in the study design, data collection and analysis, decision to publish, or preparation of the manuscript.
Conflict of interest
Carrie Pierce, Hoa Van Le, Harold Rodriguez, John Brownstein, Clark Freifeld, and Nabarun Dasgupta are employees of or contractors to Epidemico, a technology company intending to commercialize the software platform used in this research. Epidemico is a wholly owned subsidiary of Booz Allen Hamilton. I. Ralph Edwards has no conflicts of interest that are directly relevant to the content of this manuscript. Khaled Bouri, Carol Pamer, Scott Proestel, and Mark Walderhaug are employees of the United States Food and Drug Administration. Development of the social listening platform, research conducted, and manuscript development was funded by the US FDA (http://www.fda.gov) under contract with Epidemico, Inc (HHSF223201210217C).
US Government employee statement
The following authors are employees of the US Government: Khaled Bouri, Carol Pamer, Scott Proestel, and Mark Walderhaug. The opinions expressed in this manuscript are those of the authors and not intended to represent the opinions of the United States Food and Drug Administration.
Ethics statement
All human subject data used in this analysis were publicly available and used in a de-identified format whenever possible.
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
Open Access This article is distributed under the terms of the Creative Commons Attribution-NonCommercial 4.0 International License (http://creativecommons.org/licenses/by-nc/4.0/), which permits any noncommercial use, distribution, and reproduction in any medium, provided you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made.
About this article
Cite this article
Pierce, C.E., Bouri, K., Pamer, C. et al. Evaluation of Facebook and Twitter Monitoring to Detect Safety Signals for Medical Products: An Analysis of Recent FDA Safety Alerts. Drug Saf 40, 317–331 (2017). https://doi.org/10.1007/s40264-016-0491-0
Published:
Issue Date:
DOI: https://doi.org/10.1007/s40264-016-0491-0