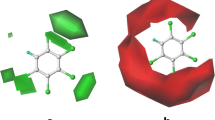
Abstract
Molecular docking was used to calculate the affinity energy between biphenyl dioxygenases(BphA), i ncluding 1ULJ, 1WQL, 2YFJ, 2YFL, 2GBX, 2XSH, 2E4P, 3GZX, and 3GZY(selected from the Protein Data Bank) and 209 polychlorinated biphenyl(PCB) congeners. The relationships between the calculated affinity energy and the persistent organic pollutant characteristics(migration, octanol-air partition coefficients, lgKOA; persistence, half-life, lgt1/2; toxicity, half-maximal inhibitory concentration, lgIC50; bioaccumulation, bioconcentration factor, lgBCF) of the PCBs were studied to understand the BphA mediated degradation of PCBs. The effect of substituent characteristics on the affinity energy was explored through full factorial experimental design. The affinities of nine kinds of BphA pr oteins on PCBs ranked as follows: 2GBX>2YFJ>2YFL>3GZX>2XSH>3GZY>2E4P>1WQL>1ULJ. The relationships between the calculated affinity energy and the molecular weight, lgKOA, lgBCF, and lgt1/2 of the PCBs were statistically significant(p<0.01), whereas the relationship with the lgIC50 of PCBs was not statistically significant(p>0.05). PCBs were more difficult to degrade following an increase in the free energy of binding. Correlation analysis showed that the average affinity energy values of PCBs gradually increased as the number of chlorine atoms increased, r egardless of the substituent position. The substituents at the ortho-positions interacted mainly through a second-order interaction, whereas those at the para-positions did not participate via a second-order interaction.
Similar content being viewed by others
References
Beyer A., Biziuk M., Rev. Environ. Contam. T., 2009, 201, 137
O’Sullivan G., Sandau C., Environmental Forensics for Persistent Organic Pollutants, Elsevier, Amsterdam, 2013
Kjellerup B. V., Paul P., Ghosh U., May H. D., Sowers K. R., App. Environ. Soil Sci., 2012, 2012, 1
Kjellerup B. V., Sun X., Ghosh U., May H. D., Sowers K. R., Environ. Microbiology., 2008, 10, 1296
Park J. S., Petreas M., Cohn B. A., Cirillo P. M., Factor-Litvak P., Environ. Int., 2009, 35, 937
Weijs L., Das K., Siebert U., van Elk N., Jauniaux T., Neels H., Blust R., Covaci A., Environ. Int., 2009, 35, 842
Zhang P., Song J. M., Liu Z. G., Zheng G. X., Zhang N. X., He Z. P., Mar. Pollut. Bull., 2007, 54, 1105
Alkhatib E., Weigand C., Environ. Monit. Assess., 2002, 78, 1
Barakat A. O., Mostafa A., Wade T. L., Sweet S. T., EI Sayed N. B., Chemosphere, 2013, 93, 545
Saba T., Su S., J. Hazard. Mater., 2013, 260, 634
Frederiksen M., Meyer H. W., Ebbehøj N. E., Gunnarsen L., Chemosphere, 2012, 89, 473
DellaValle C. T., Wheeler D. C., Deziel N. C., De Roos A. J., Cerhan J. R., Cozen W., Severson R. K., Flory A. R., Locke S. J., Colt J. S., Hartge P., Ward M. H., Environ. Sci. Technol., 2013, 47, 10405
Rawn D. F. K., Sadler A. R., Quade S. C., Sun W. F., Kosarac I., Hayward S., Ryan J. J., Chemosphere, 2012, 89, 929
Su G. Y., Liu X. H., Gao Z. S., Xian Q. M., Feng J. F., Zhang X. W., Giesy J. P., Wei S., Liu H. L., Yu H. X., Environ. Int., 2012, 42, 138
Hassine S. B., Ameur W. B., Gandoura N., Driss M. R., Chemosphere, 2012, 89, 369
Shen H. T., Ding G. Q., Wu Y. N., Pan G. S., Zhou X. P., Han J. L., Li J. G., Wen S., Environ. Int., 2012, 42, 84
Jotaki T., Fukata H., Mori C., Chemosphere, 2011, 82, 107
Arrebola J. P., Fernandez M. F., Porta M., Rosell J., de la Ossa R. M., Olea N., Martinolmedo P., Environ. Int., 2010, 36, 705
Field J. A., Sierra-Alvarez R., Environ. Pollut., 2008, 155, 1
Furukawa K., Fujihara H., J. Biosci. Bioeng., 2008, 105, 433
Monika C., Zdena K., Alena F., Stefano C., Tomáš C., Chemosphere, 2012, 88, 1317
Erickson B. D., Mondello F. J., J. Bacteriol., 1992, 174, 2903
Kitagawa W., Miyauchi K., Masai E., Fukuda M., J. Bacteriol., 2001, 183, 6598
Bedard D. L., Haberl M. L., May R. J., Brennan M. J., Appl. Environ. Microb., 1987, 53, 1103
Jia L. Y., Jia L. Y., Zheng A. P., Xu L., Huang X. D., Zhang Q., Yang F. L., J. Microbiol. Biotechn., 2008, 18, 952
Bulter C. S., Mason J. R., Adv. Microb. Physiol., 1997, 38, 47
Broadus R. M., Haddock J. D., Arch. Microbiol., 1998, 170, 106
Furusawa Y., Nagarajan V., Tanokura M., Masai E., Fukuda F., Senda T., J. Microbiol. Biotechn., 2004, 342, 1041
Kumamaru T., Suenaga H., Mitsuoka M., Watanabe T., Furukawa K., Nat. Biotechnol., 1998, 16, 663
Yang W. H., Mu Y. S., John P. G., Zhang A. Q., Yu H. X., Chemosphere, 2009, 75, 1159
Shoichet B. K., Bodian D. L., Kuntz I. D., J. Comput. Chem., 1992, 13, 380
Yutaka F., Venugopalan N., Masaru T., EijiMasai M. F., Toshiya S., J. Mol. Biol., 2004, 342, 1041
Dong X. S., Shinya F., Eriko F., Tohru T., Shugo N., Kentaro S., Hideaki N., Toshio O., Hirofumi S., Takayoshi W., J. Bacteriol., 2005, 187, 2483
Mohammadi M., Viger J. F., Kumar P., Barriault D., Bolin J. T., Sylvestre M., J. Biol. Chem., 2011, 286, 27612
Kumar P., Mohammadi M., Dhindwal S., My Pham T. T., Jeffrey T. B., Sylvestre M., Biochem. Bioph. Res. Co., 2012, 421, 757
Daniel J. F., Eric N. B., Yu C. L., Rebecca E. P., David T. G., Ramaswamy S., BMC. Struct. Biol., 2007, 7, 1
Kumar P., Mohammadi M., Viger J. F., Barriault D., Leticia G. G., Lindsay D. E., Jeffrey T. B., Michel S., J. Mol. Biol., 2011, 405, 531
Senda M., Kishigami S., Kimura S., Fukuda M., Ishida T., Senda T., J. Mol. Biol., 2007, 373, 382
Christopher L., Colbert N. Y. R. A., Pravindra K., Mathew N. C., Sangita C. S., Justin B. P., Lindsay D. E., Jeffrey T. B., Plos One, 2013, 8, e52550
Qu Q. J., Liu H. X., Feng M. B., Yang X., Wang Z. Y., J. Chem. Eng. Data., 2012, 57, 2442
Halgren T. A., J. Comput. Chem., 1996, 17, 490
Wang Z. Y., Chang Y. Q., Han Y. S., Liu K. J., Hou J. S., Dai C. L., Zhai Y. H., Guo J. L., Sun P. H., Lin J., Chen W. M., J. Mol. Struct., 2016, 1123, 335
Jain A. N., J. Comput. Aid. Mol. Des., 2007, 21, 281
Holt P. A., Chaires J. B., Trent J. O., J. Chem. Inf. Model., 2008, 48, 1602
Li X. L., Ye L., Wang X. X., Shi W., Liu H. L., Qian X. P., Zhu Y. L., Yu H. X., Chemosphere, 2013, 92, 795
Chen Y., Cai X. Y., Jiang L., Li Y., Ecotox. Environ. Safe, 2016, 124, 202
Melo E. B. D., Ecotox. Environ. Safe, 2012, 75, 213
Xu Z., Chen Y., Qiu Y. L., Gu W. W., Li Y., Chem. Res. Chinese Universities, 2016, 32(3), 348
Pender J. L., Kerr J. M., Agr. Econ., 1999, 21, 279
Wu B., Zhang Y., Kong J., Zhang X. X., Cheng S. P., Toxicol. Lett., 2009, 191, 69
Cao Y. M., Xu L., Jia L. Y., New Biotechnol., 2011, 29, 90
Shi J. Q., Qu R. J., Feng M. B., Wang X. H., Wang L. S., Yang S. G., Wang Z. Y., Environ. Sci. Technol., 2015, 49, 4209
Zeng X. L., Qu R. J., Feng M. B., Chen J., Wang L. S., Wang Z. Y., Environ. Sci. Technol., 2016, 50, 8128
Liu Z. Q., Expression of Biphenyl Dioxygense and the Binding Properties with Substrates, Dalian University of Technology, Dalian, 2012
Author information
Authors and Affiliations
Corresponding author
Additional information
Supported by the Fundamental Research Funds for the Central Universities in 2017, China(No.2017XS058) and the Key Projects in the National Science & Technology Pillar Program in the Eleventh Five-year Plan Period, China (No.2008BAC43B01).
Electronic supplementary material
Rights and permissions
About this article
Cite this article
Zhao, X., Qiu, Y., Jiang, L. et al. Analysis of Affinity Energy Between Biphenyl Dioxygenase and Polychlorinated Biphenyls Using Molecular Docking. Chem. Res. Chin. Univ. 35, 325–332 (2019). https://doi.org/10.1007/s40242-019-8340-1
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s40242-019-8340-1