Abstract
Purpose of Review
We discuss the current state of genomic testing for cancer in the UK, how this has been impacted by whole genome sequencing (WGS) and the 100,000 Genomes Project, along with approaches to reviewing whole genome analyses.
Recent Findings
The 100,000 Genomes Project has led to the development of new pathways for tissue handling and processing, variant interpretation and clinical reporting of cancer genomic testing. To our knowledge, this is the first paper discussing the recommended review process for WGS reports by the Genomics Tumour Advisory Board.
Summary
Through wider use of WGS and next-generation sequencing technologies, the new NHS Genomic Medicine Service aims to expand precision oncology research and personalised cancer care. As research in cancer genomics progresses, the standards and guidelines for interpretation of WGS reports will continue to evolve.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
In 15 years since the human genome was first mapped, genomic technologies have begun to transform our understanding of the pathology of solid tumours, providing insights into how they are best classified, what drives them and how these drivers might be targeted. This has been enabled by a precipitous decline in the cost of sequencing and prior knowledge from the field of molecular biology. Following the completion of 100,000 Genomes Project, the UK is beginning a transformation in cancer genomics as it moves from the more limited targeted approach for genomic testing to wider use of panels and whole genome sequencing (WGS) through the NHS Genomic Medicine Service. Here, we explore some of the groundwork laid by 100,000 Genomes and how this builds on previous genomic testing strategies. We discuss a step-by-step approach to systematic analysis and interpretation of the cancer WGS report and the potential next steps for somatic cancer genomics in the UK.
Next-Generation Sequencing and the Cancer Genome
The genome is the entire DNA content of a single cell. Cancer is a disease of the genome, caused by mutations in DNA, which may occasionally be germline (passed on via parental gametes and present in every cell in the body) or, more commonly, sporadic (occurring spontaneously and present only in the patient’s tumour). The sea change in genomic technologies that has occurred in the last two decades has therefore revolutionised cancer diagnostics and research, and the field continues to evolve.
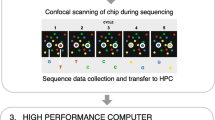
The Human Genome Project utilised cloning technologies and Sanger sequencing but since its completion in 2003, next-generation sequencing (NGS) has enabled low-cost sequencing of human genomes that can be completed in days [1, 2]. The technology involves massively parallel sequencing of short DNA fragments (‘reads’) by imaging luminescent signals given off during DNA base incorporation, and then processing the vast amount of data generated to produce a consensus sequence that is able to differentiate true mutations from errors that may occur during the sequencing process.
Following the advent of NGS, efforts such as The Cancer Genome Atlas (TGCA) have provided insights into the common somatic mutations across various tumour types but linked clinical data regarding patient’s outcomes and response to therapy according to molecular subtype is scarce [3]. Meanwhile, genomic testing for clinical diagnostics has lagged behind, utilising more primitive technologies such as targeted polymerase chain reaction (PCR) or immunohistochemistry to look for mutations that have prognostic or therapeutic impact. More recently, NGS has begun to be utilised by health services for sequencing of limited panels of genes commonly mutated in cancer [4].
Announced in 2012 and launched in 2014, the cancer arm of the Genomics England 100,000 Genomes Project set out to bridge this gap by performing paired germline and somatic tumour whole genome sequencing with the aims of (I) building a research database of cancer genomes with high-quality clinical and outcomes data, (II) providing NHS patients with clinically relevant results that may enable use of targeted drugs or recruitment to trials and (III) developing an NHS Genomic Medicine Service with established protocols and expertise to continue to deliver standardised genomic testing to cancer patients beyond the life of the original project [5,6,7]. It recruited across 13 Genomic Medicine Centres (GMCs) and their affiliated hospitals across the UK and the first 100,000 genomes across rare disease and cancer completed in December 2018.
Changes to NHS Tissue Handling for Genomic Testing
Genomic testing of cancer tissue requires relatively large amounts of high-quality DNA, requiring changes to current standard tissue handling in the NHS to facilitate this. Cancer panel analysis can be performed on as little as 10 ng of DNA, but work from the 100,000 Genomes Project and TRACERx study (Tracking non-small cell lung cancer evolution through therapy) has shown that whole genome sequencing (WGS) or whole exome sequencing (WES), respectively, requires 1–2 μg of DNA [8,9,10]. Initial pilot work performed as part of the 100,000 Genomes Project confirmed what was suspected in molecular testing laboratories; DNA extraction from FFPE often leads to insufficient yields of DNA as well as introducing artefactual mutations [11] making interpretation difficult.
Cancer tissue is procured by one of two routes: via surgical resections or via diagnostic biopsies. To obtain fresh frozen tumour tissue, this requires changes to the standard approach for handling of tissue, where the specimen is placed into formalin, and closer co-ordination between the clinical teams sampling or resecting the tumour tissue and the histopathology team, as shown in Fig. 1. For resections, it is essential to produce a formalin-fixed mirror image of the fresh sampling site for WGS or NGS for the pathologist to assess tumour content, cellularity and necrosis. For biopsy samples, these must be snap frozen and a second biopsy sample that has been fixed is used to assess these parameters. Work from the 100,000 Genomes Project demonstrated ability to sample tumours greater than 15 mm for WGS without interfering with the routine histopathological analysis [8]. There are projects looking at alternative methods of sampling for tumours less than 15 mm, but these are not yet validated. Despite advances in establishing formalin-free pathways, there are still considerable challenges to overcome before it is adopted widely as standard of care.
WGS Variant Calling and Interpretation
A human genome consists of roughly 3 billion base pairs or 1.5 gigabytes worth of data [2]. In the case of whole genome sequencing, bioinformatic algorithms are required to create reports that contain the relevant clinically actionable information.
Quality control and identification of mutations (variant calling) of WGS within the cancer arm of 100,000 Genomes is enabled through a standardised bioinformatics pipeline [5, 8, 9]. For each patient, two genomes are sequenced, the germline, from the patient’s blood, and the somatic cancer genome, from the patient’s tumour sample. The germline genome must be subtracted to differentiate somatic from germline mutations.
Analysis of the somatic genome identifies single nucleotide variants (SNVs) and small insertion/deletions (indels), as well as larger structural variants (SVs) including translocations and copy number variants (CNVs).
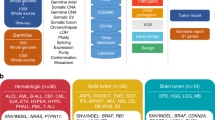
In the 100,000 Genomes WGS report, somatic SNVs and indels are tiered according to pathogenicity into 3 domains [12]. Domain 1 encompasses all variants in ‘actionable or potentially actionable genes’, e.g. those related to subtype, prognosis or a targeted drug. This information is currently taken from GenomOncology knowledge management system [13]. Domain 2 is SNVs and indels not in domain 1 but in genes associated with cancer according to the Cancer Gene Census [14]. Finally, domain 3 contains the remaining SNVs and indels [12].
Given these broad domains, it is crucial that further appraisal of somatic variants is undertaken within the Genomic Tumour Advisory Board (GTAB). To guide interpretation of somatic variants in cancer, several bodies have produced guidance—the Association for Molecular Pathology, American Society of Clinical Oncology and College of American Pathologists have produced consensus guidance that groups variants into 4 tiers according the evidence of their pathogenicity and clinical relevance: variants of strong clinical significance (tier 1), variants of potential clinical significance (tier 2), variants of unknown clinical significance (tier 3) and benign or likely benign variants (tier 4) [15]. Variants reported within domain 1 of 100,000 Genomes may fall into tier I or II under this guidance. The European Society of Medical Oncology (ESMO) has also produced the ESMO Scale for Clinical Actionability of Molecular Targets (ESCAT), a tiered system by which to evaluate therapeutic actionability of variants specifically (Table 1) [17]. It is anticipated that these and other guidelines will be incorporated into the WGS analysis and this is likely to influence the evaluation of which genes are incorporated into targeted cancer panels and funded as part of the new Genomic Test Directory for Cancer [6, 9, 16].
Germline SNVs and indels are also reported by a tiering system within 100,000 Genomes to determine the likelihood of a causal inherited cancer predisposition. All tier 1 germline variants are reported, which are those with a ‘high burden of evidence’ (e.g. 3 star rating on ClinVar) and part of the ‘pertinent cancer susceptibility gene panel’ (PCSGP) for the specific cancer [12]. If there is a strong family history or a priori suspicion of inherited cancer, tier 3 germline susceptibility and therapeutic variants are also provided, which may not be matched for tumour type and have a smaller evidence base. There would require ACGS classification in line with the latest guidelines [18].
Interpretation of a whole genome is not without its complexities. As mentioned, calling errors are possible, especially if the input tissue was of low cellularity or sequencing was of poor coverage. These errors also mean that actionable mutations found on whole genome sequencing must be validated ‘using an orthogonal method’ appropriate to the type of mutation detected, as detailed in the Guidance for the Validation and Reporting [12].
The Genomics Tumour Advisory Board
A multidisciplinary approach is required when analysing cancer genome results. As testing moves from single gene to cancer panels and whole genomes, specialist knowledge is required from clinical geneticists and scientists who sit outside of the tumour site–specific multidisciplinary team meeting. The Guidance for the Validation and Reporting of Whole Genome Sequencing Results for the 100,000 Genomes Project Cancer Programme details the recommended structure for the GTAB and how WGS reports should be reviewed, but it would be beneficial to bring all cancer panel test results to a similar meeting [12]. The GTAB core members should comprise an oncologist or haemato-oncologist, a pathologist, a clinical geneticist and a clinical scientist, all with specialist knowledge of somatic or inherited cancer genomics. Ideally, additional members including, a tumour site–specific oncologist or haemato-oncologist, clinical bioinformatician, GTAB coordinator and medical trainees (as part of their training in genomic healthcare) should be invited to help facilitate a thorough review of results and an easy-to-interpret report for clinicians from the genomic data.
Analysing the WGS Report
For each patient whose WGS report discussed at the GTAB, there should be a structured step-by-step review of results as follows.
Clinical information
The first step is to discuss the clinical context of the patient. This includes information regarding whether the sample submitted for WGS was from a resection or biopsy sample. Was the pathological diagnosis the same as that clinically suspected when the tissue was sent for WGS? Does the patient have localised disease and if so, have they undergone surgical resection? Does the patient have locally advanced or metastatic disease, and if so, what line of treatment are they receiving? Has the clinical context changed from when the biopsy was taken? Are they clinically fit enough for further treatment?
Quality control
When reviewing the WGS report, firstly, all demographic information should be confirmed, followed by a review of the quality metrics of the sample and the sequencing undertaken, including cellularity, tumour content and sequencing coverage.
Somatic variants
Somatic domain 1 and 2 variants should be reviewed as these may have a clinical impact with regard to prognosis or therapeutics. The report contains relevant information to appraise the variant, including the gene it is identified in, the cDNA and protein change, and the predicted consequence (i.e. frameshift variant or missense variant). Certain metrics are critical for variant interpretation, such as supporting reads (depth of coverage) and variant allele frequency (VAF), and should be reviewed when evaluating the called variant, along with the COSMIC ID (if present for this variant) and links to clinical trials at the gene, and for some tumour types, at the variant level. If a variant is not well characterised, further investigation or evidence may be required by a member of the GTAB, before or after the meeting to aid interpretation and advise on whether validation is appropriate. Structural variants, including translocations and CNVs, should also be reviewed, particularly if there is a known standard-of-care testing for a particular tumour type, such as EML4-ALK in non-small cell lung cancer or ERBB2 amplification in breast cancer.
Germline variants
As discussed above, only known pathogenic mutations in cancer genes with a 3-star ClinVar rating are flagged to the clinical team. A senior clinical scientist should review and interpret any report tier 3 variants. This ensures that only highly penetrant mutations as reviewed by an expert panel are reported, alleviating concerns about identifying variants of unknown significance (VUS). Any findings are highlighted to the clinical genetics team to contact the patient and arrange an appointment. Importantly, a discrepancy in suspected and eventual tumour type may mean some germline variants are not reported because of lack in evidence in one tumour type compared with another. This would necessitate a repeat analysis. Secondly, if a GTAB is reviewing a cancer panel test rather than WGS, the germline is unlikely to be sequenced. In this case, it is important that the clinical scientists and clinical geneticists identify common germline variants if found in the panel and organise appropriate germline testing. In the case of deceased patients, the guidance recommends that only pertinent germline variants need to be reviewed due to the potential impact for relatives.
Tumour mutational burden
The whole genome report provides a wealth of information, including pan-genomic markers detailed in the supplementary information of the genome report [9]. The first of these is tumour mutational burden (TMB) which is quantified in coding SNVs per megabase. Recent work has shown that TMB may serve as a predictive or prognostic biomarker of response to immunotherapy, due to its correlation with neoantigens, which are expressed on the tumour cell surface and recognised by the immune system. If TMB is greater than 10 coding SNVs per megabase, this may prompt consideration of immunotherapy, within a trial setting. It may also support the finding of a mismatch repair deficiency/microsatellite instability on standard-of-care testing. The 100,000 Genomes WGS report includes a rainfall plot which visually displays mutational burden across the genome and highlights areas of hypermutation or ‘kataegis’ [19], but the clinical relevance of these areas of hypermutation is not yet fully understood.
Mutational signatures
A second pan-genomic marker includes mutational signatures. These are patterns of mutational combinations associated with particular mutational processes that may be intrinsic or extrinsic [20]. There are over 30 mutational signatures with the vast majority of unknown aetiology. However, some are clinically relevant, including those associated with tobacco exposure, UV light and failures of DNA repair mechanisms including mismatch repair and double-strand break repair. They therefore may simply act to support the known pathological diagnosis, e.g. tobacco exposure in a small cell lung cancer, or may provide insights into the possible aetiology of a cancer such as failure of DNA repair mechanisms [21]. In the 100,000 WGS report, this is displayed by both a bar graph showing the percentage of each mutational signature within the tumour (Fig. 2a) and a histogram of the percentage of each type of base substitution (in a trinucleotide context) (Fig. 2b).
Figures from the 100,000 Genomes WGS Report. a Bar graph showing the percentage of each mutational signature within the tumour. This patient has > 20% signature 6, representing mismatch repair (MMR) deficiency. b Histogram of the percentage of each type of base substitution in a trinucleotide context, also for a patient with MMR deficiency. c Circos plot: the multicoloured bars of the innermost ring represent the chromosomes and the lines traversing the circle represent aberrant structural rearrangements between the chromosomes. The red and green rings represent the number of single nucleotide variants (SNVs) and small insertions and deletions (indels) across the chromosomes. The outermost blue ring represents the sequencing coverage across the chromosomes, and the second outermost ring the copy number changes
Circos plot
In the case of the 100,000 Genomes WGS report, somatic information, including sequencing coverage, small variants and structural variants, is visually represented via a circos plot (Fig. 2c).
Outcome
Following the review of a 100,000 Genomes WGS report, the GTAB may issue one of five outcomes in their clinical report (Table 2) [12]. They will also provide additional information to the treating clinician including advice on validation method (see above) and relevant clinical trials. The 100,000 Genomes report provides links to clinical trials with information about inclusion criteria and recruiting status, but this should always be verified by the GTAB and treating clinician.
Genomic Testing and Clinical Trials
WGS and panel testing are both useful in enhancing recruitment to existing stratified medicine trials within oncology, particularly umbrella trials recruiting patients with a single tumour type and matching them with an appropriate arm corresponding to their genomic variants, such as the Lung MATRIX trial in NSCLC [22]. WGS can also help recruit to basket trials which recruit patients with particular (often rare) mutations across multiple tumour types [23]. Given that these trials recruit patients with metastatic cancer, it is crucial that there is a clinically meaningful turnaround time of the genomic test. The 100,000 Genomes Project was able to operate Fast Track testing for patients with this type of clinical need, with a median time from the sample being sent to genomic report being returned to the GMC of 10–20 days. The benefit of WGS over panel testing is the broader range of variants it is able to pick up, as well as pan-genomic markers, which are increasingly featuring in clinical trials.
WGS can also be used to help design future clinical trials. Current estimates suggest that clinical trial success rates are less than 5% but the success rate is higher in trials that use biomarkers compared with trials that do not [24]. WGS may also provide a better understanding of the disease process, which may in turn help identify novel biomarkers or targets for subsequent clinical trials.
Tumour Heterogeneity and Liquid Biopsy
Recent work has shown that tumour heterogeneity contributes significantly to the difficulties in treating cancer [25,26,27]. A tumour consists of different sub-clonal populations which under selection pressures develop treatment resistance. In individuals who have a ‘mixed response’ to therapeutic regimen, e.g. growth of some sites of disease and shrinkage in others, there are different sub-clonal populations accounting for this. It is not yet clear how many samples should be taken from a tumour sample to accurately capture tumour heterogeneity, or, in the context of widespread metastatic disease, how many different sites should be biopsied. WGS from a single sample may not be representative of the whole tumour, and performing additional biopsies may not be clinically safe or feasible. This may be overcome by using ‘liquid biopsy’ of cell-free DNA (cfDNA) or circulating tumour cells (CTCs), but more work is underway on utilising this technology for disease monitoring as it is not yet clear whether sampling of peripheral blood for cfDNA and CTCs can be used to accurately assay tumour heterogeneity [28]. cfDNA from tumours allow molecular targets to be identified without the need for biopsy, and this is already being utilised for detection of targeted resistance mutations in lung cancer [29].
The NHS Genomic Medicine Service
With around 50% of tumours sequenced as part of 100,000 Genomes containing actionable or potentially actionable variants, the clinical utility of genomics and more specifically WGS for cancer has become more evident [6]. On 1 January 2019, the NHS Genomic Medicine Service was launched. It builds on the work done as part of 100,000 Genomes Project and will deliver genomic testing for rare disease and cancer across the UK, with a standardised Genomic Testing Directory which includes single-gene tests, panels and WGS and continued opportunity for patients to participate in research [5, 6, 16]. It is anticipated that testing will be delivered predominantly by seven Genomic Lab Hubs (GLHs) complying with national standards. Initially, WGS will be available in a small number of cancer types of clinical need, e.g. sarcomas and paediatric tumours, with development of targeted panels across other cancer types. As the cost of WGS continues to fall, and with further research completed on 100,000 Genomes and other WGS efforts in cancer, the breadth of the genome tested for other tumour types is likely to increase as it becomes more economical and offers greater clinical utility.
Multi-omics—The Next Step in Cancer Genomics
Mutations within the germline and somatic genomes only reveal part of the aetiology of cancer. The epigenome, including levels of CpG island methylation and histone modification, are also well known to be implicated in cancer development and progression. There are already several clinically relevant biomarkers including methylation of MLH1 in sporadic MSI colorectal cancer and MGMT promoter methylation in glioma which help define the molecular subtype of a tumour and appropriate therapy [30, 31]. As well as these specific targets, there are methylation array–based classification systems under development including EPICUP for cancer of unknown primary and MolecularNeuropathology.org for primary brain tumours [32, 33]. As they are validated, it is likely that more and more of these will be incorporated into the Genomic Test Directory to allow better classification and prognostication of disease. More challenging will be the measurement of the transcriptome, although this has proven an excellent research tool for profiling tumours; preservation of RNA of sufficient amount and quality is challenging for NHS Pathology departments, as it requires more rapid processing of frozen tissue or use of bespoke fixatives [34]. As ‘genome-friendly’ pathways become more standard practice, this may form part of future strategies within the Genomic Medicine Service.
Conclusions
Cancer genomics aims to transform cancer care in the UK through the NHS Genomic Medicine Service. The 100,000 Genomes Project has laid the groundwork for tissue handling, data analysis and interpretation, but further research by GeCIPs and the wider cancer research community will enable the Genomic Medicine Service to evolve. It appears that we are moving towards wider adoption of WGS, pan-genomic markers, multi-omics and liquid biopsy, ushering in a new era of personalised medicine and precision oncology trials.
References
Goodwin S, McPherson JD, McCombie WR. Coming of age: ten years of next-generation sequencing technologies. Nat Rev Genet. 2016;17:333–51.
Collins FS, Lander ES, Rogers J, Waterson RH. Finishing the euchromatic sequence of the human genome. Nature. 2004;431:931–45.
Weinstein JN, Collisson EA, Mills GB, Shaw KM, Brad A, Ellrott K, Shmulevich I, Sander C, Stuart JM. NIH public access. 2014;45:1113–1120.
AW K, JM F. Multigene panel testing in oncology practice: how should we respond? JAMA Oncol. 2015;1:277–8.
Turnbull C, Scott RH, Thomas E, et al. The 100 000 genomes project: bringing whole genome sequencing to the NHS. BMJ. 2018;361:1–7.
Hill S. Introducing genomics into cancer care. Br J Surg. 2018;105:e14–5.
Cancer N, Programme T. NATIONAL CANCER TRANSFORMATION PROGRAMME Publications Gateway Reference: 07318. 2016.
Caulfield M, Davies J, Dennys M, et al. The 100,000 Genomes Project Protocol. Genomics Engl Protoc. 2017;1–112.
Turnbull C. Introducing whole-genome sequencing into routine cancer care: the Genomics England 100 000 Genomes Project. Ann Oncol. 2018;29:784–7.
ThermoFisher Scientific. Oncomine focus assay. 2019.
Do H, Dobrovic A. Sequence artifacts in DNA from formalin-fixed tissues: causes and strategies for minimization. Clin Chem. 2015;61:64–71.
Alikian M, Deans S, Diaz-Cano S, et al. Guidance for the Validation and Reporting of Whole Genome Sequencing Results for the 100,000 Genomes Project Cancer Programme. 2018;1–32.
Vanderbilt Ingram Cancer Centre. My cancer genome. 2019.
Futreal PA, Coin L, Marshall M, Down T, Hubbard T, Wooster R, et al. A census of human cancer genes. Nat Rev Cancer. 2004;4:177–83.
Li MM, Datto M, Duncavage EJ, et al. Standards and guidelines for the interpretation and reporting of sequence variants in cancer: A Joint Consensus Recommendation of the Association for Molecular Pathology, American Society of Clinical Oncology, and College of American Pathologists. J Mol Diagn. 2017;19:4–23.
Keogh SB, Medical N. NHS ENGLAND – BOARD PAPER Creating a genomic medicine service to lay the foundations to deliver personalised interventions and treatments introduction potential benefits of genomic technologies. 2017;1–8.
Mateo J, Chakravarty D, Dienstmann R, Jezdic S, Gonzalez-Perez A, Lopez-Bigas N, et al. A framework to rank genomic alterations as targets for cancer precision medicine: the ESMO Scale for Clinical Actionability of Molecular Targets (ESCAT). Ann Oncol. 2018;29:1895–902.
Ellard S, Baple E, Owens M, Eccles D, Abbs S, Deans Z, McMullan D. ACGS best practice guidelines for variant classification 2017. 2017;1–16.
Nik-Zainal S, Alexandrov LB, Wedge DC, van Loo P, Greenman CD, Raine K, et al. Mutational processes molding the genomes of 21 breast cancers. Cell. 2012;149:979–93.
Sanger Institute. Signatures of mutational processes in human cancer. 2019.
Alexandrov LB, Nik-Zainal S, Wedge DC, et al. Signatures of mutational processes in human cancer. Nature. 2013;500:415–21.
Middleton G, Crack LR, Popat S, Swanton C, Hollingsworth SJ, Buller R, et al. The National Lung Matrix Trial: translating the biology of stratification in advanced non-small-cell lung cancer. Ann Oncol. 2015;26:2464–9.
West HJ. Novel precisionmedicine trial designs umbrellas and baskets. JAMA Oncol. 2017;3:423.
Wong CH, Siah KW, Lo AW. Estimation of clinical trial success rates and related parameters. Biostatistics kxx069. 2018.
Jamal-Hanjani M, Hackshaw A, Ngai Y, et al. Tracking genomic cancer evolution for precision medicine: the lung TRACERx study. PLoS Biol. 2014;12:1–7.
McGranahan N, Swanton C. Clonal heterogeneity and tumor evolution: past, present, and the future. Cell. 2017;168:613–28.
Sottoriva A, Kang H, Ma Z, Graham TA, Matthew P, Zhao J, Marjoram P, Siegmund K, Press MF, Curtis C. HHS public access. 2015;47:209–216.
Domínguez-Vigil IG, Moreno-Martínez AK, Wang JY, Roehrl MHA, Barrera-Saldaña HA. The dawn of the liquid biopsy in the fight against cancer. Oncotarget. 2018;9:2912–22.
Remon J, Caramella C, Jovelet C, Lacroix L, Lawson A, Smalley S, et al. Osimertinib benefit in EGFR-mutant NSCLC patients with T790M-mutation detected by circulating tumour DNA. Ann Oncol. 2017;28:784–90.
Bettstetter M, Dechant S, Ruemmele P, Grabowski M, Keller G, Holinski-Feder E, et al. Distinction of hereditary nonpolyposis colorectal cancer and sporadic microsatellite-unstable colorectal cancer through quantification of MLH1 methylation by real-time PCR. Clin Cancer Res. 2007;13:3221–8.
Esteller M, Garcia-Foncillas J, Andion E, Goodman SN, Hidalgo OF, Vanaclocha V, et al. Inactivation of the DNA-repair gene MGMT and the clinical response of gliomas to alkylating agents. N Engl J Med. 2000;343:1350–4.
Capper D, Jones DTW, Sill M, Hovestadt V, Schrimpf D, Sturm D, et al. DNA methylation-based classification of central nervous system tumours. Nature. 2018;555:469–74.
Moran S, Martínez-Cardús A, Sayols S, Musulén E, Balañá C, Estival-Gonzalez A, et al. Epigenetic profiling to classify cancer of unknown primary: a multicentre, retrospective analysis. Lancet Oncol. 2016;17:1386–95.
Posorski N, Kaemmerer D, Ernst G, Grabowski P, Hoersch D, Hommann M, et al. Localization of sporadic neuroendocrine tumors by gene expression analysis of their metastases. Clin Exp Metastasis. 2011;28:637–47.
Acknowledgments
We thank the participants of the 100,000 Genomes Project, the NHS England Genomics team that has led the NHS implementation and service transformation and the future genomic medicine service developments, and the NHS staff undertaking recruitment and sample processing, as well as all additional people who have supported the project in multitudes of other ways. We would like to thank Alona Sosinsky, Head of Cancer Analysis, and John Ambrose, Bioinformatics Team at Genomics England.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of Interest
Alison May Berner and George J Morrissey each declare no potential conflicts of interest.
Nirupa Murugaesu declares an Advisory Board relationship with Pfizer and Roche, and a Speaker’s Bureau relationship with Merck, Pfizer, and Roche.
Human and Animal Rights and Informed Consent
This article does not contain any studies with human or animal subjects performed by any of the authors.
Additional information
Publisher’s Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
This article is part of the Topical Collection on Cancer Genomics
Rights and permissions
Open Access This article is distributed under the terms of the Creative Commons Attribution 4.0 International License (http://creativecommons.org/licenses/by/4.0/), which permits unrestricted use, distribution, and reproduction in any medium, provided you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made.
About this article
Cite this article
Berner, A.M., Morrissey, G.J. & Murugaesu, N. Clinical Analysis of Whole Genome Sequencing in Cancer Patients. Curr Genet Med Rep 7, 136–143 (2019). https://doi.org/10.1007/s40142-019-00169-4
Published:
Issue Date:
DOI: https://doi.org/10.1007/s40142-019-00169-4