Abstract
A healthy gut microbiota is essential in maintaining the human body in a homeostatic state by its functions in digestion and immune tolerance. Under states of aberrant microbial composition or function (dysbiosis), the gut microbiota induces systemic inflammation that can lead to the onset of many diseases. In this review, we describe some evidence, largely from rodent studies, that supports the possible role of a dysbiotic gut microbiota in the onset and exacerbation of ocular diseases, primarily diabetic retinopathy, age-related macular degeneration, choroidal neovascularization, and uveitis. Furthermore, we examine several potential therapeutic measures that show promise in restoring the gut microbiota to a eubiotic state, preventing the aforementioned disease pathologies.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
The gut microbiota is integral in health maintenance. |
Aberrations in the gut microbiota have been shown to be associated with ocular diseases in both human and animal studies. |
Cellular and antibiotic therapies, intermittent fasting, and altered diet have been shown to restructure the gut microbiota and decrease ocular pathologies. |
Future studies of the gut microbiota and its relationship with ocular disease will increase our understanding of the gut–eye axis and lead to innovative therapeutic approaches to treating ocular diseases. |
Introduction
The trillions of viruses, protists, and bacteria in the human small and large intestines that comprise the gut microbiome are integral to maintaining health. The primary roles of these microbes include assisting the body in digestion and facilitating the function of the immune system. Without this complex ecosystem, the human body would fail to function. Thus, preserving appropriate composition of these microbes, specifically the bacteria, is essential in promoting health and maintaining homeostasis. Following shifts in gut bacterial composition, the immune system surpasses microbial tolerance and progresses to an inflammatory state, which can induce tissue damage throughout the body and promote disease pathogenesis. The focus of this brief review is to describe the alterations in the gut microbiota that may exacerbate ocular phenotypes, including diabetic retinopathy, age-related macular edema, choroidal neovascularization, uveitis, and Sjögren’s disease. The following search terms were used in PubMed and Google Scholar: [(“gut microbiota” OR “gut microbiome) AND (“diabetic retinopathy” OR “DR” OR “age related macular degeneration” OR “AMD” OR “choroidal neovascularization” OR “CNV” OR “uveitis” OR “Sjögren’s disease”)]. The resultant articles were filtered by publication date, with all publications prior to the year 2000 being omitted.
Here, we investigate interactions between the gut microbiota and ocular pathology and their implications for progression of disease, and propose several potential therapeutic approaches aimed at promoting gut microbial homeostasis and reducing adverse ocular outcomes. This article is based on previously conducted studies and does not contain any studies with human participants or animals performed by any of the authors.
The Gut Microbiota
The human intestine is home to approximately 1014 microbes, comprising over 4000 species. The concentration of microbes increases from the stomach to the distal colon [1, 2]. The normal human gut microbiota consists primarily of organisms from two bacterial phyla, Firmicutes and Bacteroidetes, representing 65% and 23% of organisms, respectively [3]. (Fig. 1) Interestingly, it has been well documented that the human gut microbiota is highly variable between individuals [3,4,5]. Under homeostatic conditions, these bacteria function during development and in digestion, metabolism, and immune defense/tolerance [5, 6]. The massive complexity of this system is necessary to properly maintain the aforementioned functions. Thus, alterations of the gut microbiota have been implicated in the pathogenesis of many acute and chronic diseases, such as diabetes, obesity, inflammatory bowel disease, Crohn’s disease, Alzheimer’s disease, anxiety, and depression [5, 7,8,9,10,11,12,13,14,15]. It is necessary to further delineate aberrations in gut microbial composition and function in order to understand how these enhance or inhibit disease states.
The effects of gut dysbiosis. A dysbiotic gut microbiota, typically associated with an increased Firmicutes/Bacteroidetes (F/B) ratio, results in thinning of the intestinal mucus layer and disruption of the gut–vascular barrier. Together, loss of these physiological barriers allows for translocation of gut commensal bacteria and bacterial products into the circulation, where they travel to tissues and induce pathological inflammation
Analysis of microbial populations and communities, such as those within the gut, is performed via various well-described methods. The most common and cost-effective method for taxonomic identification of bacteria within desired fecal or gut mucosal samples is sequencing of bacterial 16S rRNA. Bacteria 16S rRNA is composed of nine hypervariable regions (V1–V9) separated by nine conserved regions [16]. The identification of taxa relies on the hypervariable regions selected, with hypervariable regions 3 and 4 (V3–V4) enabling the greatest taxa resolution and identification to the genus level [17,18,19]. 16S rRNA sequences are then grouped into operational taxonomic units (OTUs), which are clusters of sequences that differ by less than a fixed diminished threshold (typically 3%) [20]. Other identification techniques include shotgun metagenomics, offering species- and strain-level classification. Shotgun metagenomics also allows for inference of functional relationships between bacteria and host by determining the functional content of samples directly. This method is more costly and requires more advanced bioinformatics [17]. Methods for defining the functional profiles of bacterial communities involve metatranscriptomic analysis of sample RNA sequences [13, 18, 21, 22]. Extracted RNA is sequenced and mapped to protein-coding sequences from the KEGG Orthology database. The resulting data are informative of the functional gene families within a sample. Therefore, one is able to infer the functional pathways that are abundant within a sample. Taken together, these methods allow for the identification and characterization of taxonomic genera and respective metabolic functions within a given sample [13, 18, 21, 22]. These results are important for understanding gut microbial changes that occur under various disease states and how the gut microbiota promotes or protects against said conditions.
Intestinal Immunity
The diversity of immune cells involved in maintaining intestinal homeostasis is vast. These cells, specifically macrophages, neutrophils, plasma cells, B cells, and a variety of T cells, work together to ensure resolution of inflammation and prevention of infections. Any aberration in the intestinal immune response may lead to systemic disease, sepsis, and under the most extreme conditions even death. Within the intestinal lamina propria, resident macrophages exist in a state of inflammatory anergy in which they reside in close proximity to potentially pathogenic bacteria that would, in any other part of the body, cause macrophages to engage a number of proinflammatory signaling cascades with the purpose of destroying said bacteria. However, intestine resident macrophages do not produce proinflammatory cytokines such as IL-1, IL-6, IL-12, RANTES, TGF-β, or TNF-α, but remain poised to combat invading pathogens via their phagocytic and bactericidal repertoires [23]. The purpose of inflammatory anergy is to prevent constant immune activity in the intestine against the commensal, physiologically protective gut microbiota, a state which, if allowed to proceed uninhibited, would destroy the very tissue the macrophages were intended to protect.
The sole structural barrier between commensal and potentially pathogenic bacteria in the intestinal lumen is the intestinal single-cell layer of epithelial cells and underlying vascular endothelial cells, the so-called gut–vascular barrier (GVB). The gut epithelium allows the selective entry of substances present in the lumen and prevents the entry of harmful elements, including bacteria and their bio-products, into the circulation [24]. However, not all secreted microbial metabolites are harmful to humans. For example, bacterial short-chain fatty acids exert beneficial effects [25]. The luminal surface of the intestinal epithelium is covered with a sticky layer of mucus, secreted by goblet cells, which traps luminal bacteria attempting to traverse the GVB. Further, Paneth cells, epithelial cells, and plasma cells secrete antimicrobial peptides, cytokines, and antibodies, respectively. These molecules act to directly eliminate potentially pathogenic bacteria, recruit a number of immune cells, such as neutrophils and T helper cells, to the site of potential infection, or mark bacteria for destruction by intestinal lymphocytes. However, these responses often result in damage to the GVB, allowing for the translocation of gut microbes or their secreted products across the GVB and into the extraluminal system.
Reduced integrity of the GVB allows the translocation of gut microbial peptides, such as peptidoglycan (PGN), a component of bacterial cell walls, into the circulation [13]. Both Gram-positive and Gram-negative bacteria disassemble their PGN lattices during replication and growth. Gram-negative bacteria, which constitute the majority of the healthy gut microbiota, import much of the shed PGN back into the cell via the action of AmpG, a muropeptide permease [26]. Though the majority of this PGN is recycled back into the cell to be used again, some is lost into the extracellular space, where it acts as a pathogen-associated molecular pattern (PAMP) to activate pro-inflammatory pathways of the host [27,28,29,30,31]. Specifically, PGN has been implicated in the pathogenesis of diabetic retinopathy [13].
The Gut Microbiota in Models of Ocular Diseases
Diabetic Retinopathy
Diabetic retinopathy (DR) is a leading cause of blindness in individuals between the ages of 20 and 74 years, and in 2010 was estimated to affect more than one-third of diabetics worldwide, or approximately 100 million persons [32]. It is known that prediabetes and type 1 and type 2 diabetes are associated with gut microbial dysbiosis [33,34,35,36,37,38]. However, there has been much confusion in the field as to whether this association is inherent in the pathology of diabetes, or whether it is a secondary effect of medications used to treat the disease. Several studies have found that diabetic medications, particularly metformin, influence the gut microbiota, and medication has been identified as a confounding variable in microbiota studies of diabetic individuals [39, 40]. Therefore, studies of diabetic subjects that aim to assess gut microbial dysbiosis must be stratified to remove confounding variables such as medication.
Type 2 diabetes (T2D), unlike T1D, is not the consequence of autoimmunity, but the result of progressive insulin intolerance. The gut microbiota has been shown to exacerbate the progression of insulin resistance [41]. Recent studies in a murine model of T2D (db/db) have demonstrated alterations of gut microbial composition with increased generation of beneficial secondary bile acids, namely tauroursodeoxycholate (TUDCA) [12]. TUDCA was shown to provide a neuroprotective effect in the retina via activation of its receptor, TGR5, preventing exacerbation of DR [12]. T2D mice were found to contain larger concentrations of bacteria in the Verrucomicrobia and Tenericutes phyla, and decreased concentrations of bacteria in the Bacteroidetes and Firmicutes phyla, compared to control mice (db/m) [12]. However, there was no difference in the Firmicutes/Bacteroidetes (F/B) ratio of the gut microbiota, a measure which has been used as an indicator of changes in the microbiota with obesity [42]. T2D was also associated with decreased abundance of goblet cells, producers of intestinal mucus, and increased concentrations of plasma PGN. Furthermore, an altered gut microbiota in T2D was associated with exacerbation of DR, assessed by abundance of retinal acellular capillaries [12] (Figs. 2 and 3).
The effects of cellular and nutritional therapies on the prevention of diabetic retinopathy. a In a model of type 1 diabetes with ACE2 deficiency (ACE2−/y-Akita), intraperitoneal injection of FACS-sorted wild-type (WT) bone marrow myeloid angiogenic cells (MACs) was shown to restructure the gut microbiota and repair the gut–vascular barrier (GVB). The intact GVB resulted in a decrease in the levels of systemic peptidoglycan (PGN), inhibiting a noncanonical TLR2-mediated signaling cascade in the retina, reducing retinal barrier dysfunction. b In a model of type 2 diabetes (db/db), intermittent fasting on an alternate-day regimen was shown to restructure the gut microbiota and activate production of beneficial secondary bile acids, specifically tauroursodeoxycholate (TUDCA), in the liver. TUDCA prevented progression of DR via activation of its receptor, TGR5, in the neural retina
The gut–eye axis in ocular diseases. Alterations in the gut microbiota have been shown to be associated with multiple ocular diseases, including diabetic retinopathy, age-related macular degeneration, choroidal neovascularization, and uveitis. Various groups have shown that restructuring the gut microbiota by cellular and antibiotic therapies, intermittent fasting, and dietary intervention have potential therapeutic benefits in preventing/treating ocular diseases
Dysfunction of the renin–angiotensin system (RAS) is associated with diabetes and vascular dysfunction, particularly diabetic retinopathy [43,44,45,46,47,48,49,50,51]. This effect is largely mediated by the loss of angiotensin-converting enzyme 2 (ACE2), the primary protein responsible for shifting the effects of RAS between its vasoprotective and vasodeleterious arms by its action on the MAS receptor and ATR1/ATR2, respectively [52,53,54]. Duan et al. demonstrated that ACE2 deficiency in a murine model of T1D (ACE2−/y-Akita) was associated with a restructuring of the gut microbiota. Of particular interest, ACE2−/y-Akita mice were shown to possess a gut microbiota with increased PGN biosynthesis. Via GVB disruption, initiated by the effect of hyperglycemia on intestinal epithelial cells, PGN from the intestinal lumen is able to translocate into the circulation and travel to the retina [13]. In retinal endothelial cells, PGN was shown to activate a noncanonical TLR2-mediated MyD88/ARNO/ARF6 signaling cascade, exacerbating DR by compromising retinal endothelial cell adherens junctional integrity [13] (Figs. 2 and 3). It is, however, important to note that murine models of diabetic retinopathy are limited in their translatability to human disease, as mice only develop mild disease pathology and do not have a macula. Thus, the pathology that is observed cannot be directly translated to human disease in particular vision-threatening diabetic retinopathy such as diabetic macular edema and proliferative diabetic retinopathy.
Age-Related Macular Degeneration
Age-related macular degeneration (AMD), the leading cause of irreversible blindness in industrialized countries, is a disease in which the central area of the retina, the macula, is damaged, leading to progressive central vision loss, particularly in people over the age of 55 [55,56,57]. Obesity is a risk factor for AMD [58, 59]. In fact, Adams et al. showed that male subjects with an increase of 0.1 in waist/hip ratio had a 13% and 75% increase in the probability of developing early and late AMD, respectively [59]. As diet is known to be a primary contributor to the development of obesity, Rowan et al. examined the effect of high- and low-glycemic diets on the gut microbiota and development of AMD. It was determined that aged mice fed a high-glycemic (HG) diet developed retinal pathology similar to AMD (AMDf), whereas mice on a low-glycemic (LG) diet did not [60, 61] (Fig. 3). Interestingly, in mice raised on an HG diet and switched to an LG diet late in life, this AMDf phenotype was arrested or reversed. Furthermore, alterations in the gut microbiota were determined to be associated with the AMDf phenotype. Specifically, risk for AMDf is associated with increases in gut abundance of bacteria within the Clostridiales order, while protection from AMDf is associated with the Bacteroidetes order [60] (Fig. 3). The above studies yield important associations between alterations of the gut microbiota and AMD; however, these studies are limited in their translatability and applicability to the human condition, as no single murine model exists which exhibits all features of human AMD.
Choroidal Neovascularization
Choroidal neovascularization (CNV), which is seen as the vascular pathology associated with wet AMD [62], is classified into multiple types based on vessel growth pattern: type 1—between the retinal pigment epithelium (RPE) and Bruch’s membrane, type 2—between the retina and RPE, or a combination of both (combined pattern) [62]. Using a murine model of CNV via rupture of Bruch’s membrane with an argon laser, Andriessen et al. investigated the interaction between diet, gut microbiota, and CNV pathophysiology. In concordance with previous findings, a high-fat diet (HFD, 60% kcal fat, 26% kcal carbohydrate, 14% kcal protein) induced gut dysbiosis compared to mice fed standard chow (RD, 16% kcal fat, 63% kcal carbohydrate, 21% kcal protein) (Fig. 3). Specifically, HFD induced decreased abundance of Bacteroidetes and increased abundance of Firmicutes, increasing the F/B ratio. Interestingly, HFD-fed mice experienced exacerbated CNV [63]. HFD-fed mice showed increased gut–vascular permeability. HFD-fed mice also had increased systemic inflammation by elevated pattern recognition receptor (PRR) activation [63,64,65,66,67]. Furthermore, choroidal inflammation was exacerbated via increased quantities of mononuclear phagocytes, microglia, Il6 mRNA, Tnf mRNA, and Vegfa mRNA [63]. These data suggest that a high-fat diet plays a role in exacerbating the pathogenesis of CNV by increasing inflammation as a result of an increase in the F/B ratio in the gut microbiota. Studies of the gut microbiota of individuals with neovascular AMD revealed an enrichment in Anaerotruncus, Oscillibacter, Ruminococcus torques, and Eubacterium ventriosum compared to control subjects [68]. This study also found that the gut microbiota of said subjects was enriched in genes related to l-alanine fermentation, glutamate degradation, and arginine biosynthesis, and reduced in genes related to fatty acid elongation [68]. Additional studies are needed to understand the impact of the gut microbiota on the complex pathogenesis of human neovascular AMD. It is important to take into consideration the limitations of the rodent laser-induced CNV model. In particular, this model has features of wound healing and does not take into account aging. However, despite these limitations, the laser-induced CNV model is one of the most widely used animal models which mimics the angiogenic aspects of neovascular AMD [69].
Uveitis
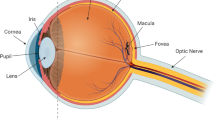
Uveitis (UVT) is an inflammatory condition of the eye affecting the iris, ciliary body, and choroid, but may involve other ocular tissues such as the retina, optic nerve, and vitreous humor [70]. UVT is a common ocular disease primarily affecting middle-aged individuals and accounting for 10% of legal blindness in the United States and approximately 25% in the developing world [70]. UVT is generally associated with either a systemic infection or immunological disease; however, approximately 50% of uveitis cases affect only the eye and do not have a known etiology, so-called idiopathic uveitis [71].
The gut microbiota has been implicated in the severity of UVT. A recent study by Chakravarthy et al. found alterations in the gut microbiota of a cohort of Chinese patients with UVT compared to healthy controls. In particular, those with UVT demonstrated decreases in operational taxonomic units (OTUs) of several anti-inflammatory bacteria, including Faecalibacterium, Bacteroides, Lachnospira, and Ruminococcus, and increases in colonization by Prevotella and Streptococcus organisms [72]. Furthermore, oral administration of combinatorial broad-spectrum antibiotics [metronidazole (1 g/L) + vancomycin (500 mg/L) + ampicillin (1 g/L) + neomycin (1 g/L)] and single oral administration of metronidazole (1 g/L) or vancomycin (500 mg/L) resulted in a reduction in uveitis clinical scores in an interphotoreceptor retinoid-binding protein (IRBP) peptide-induced model of experimental autoimmune uveitis (EAU) in B10.RIII mice [73] (Fig. 3). Broad-spectrum antibiotic administration in this model altered gut microbial composition by a reduction in Bacteroidetes spp., Firmicutes spp., Alphaproteobacteria, and Gammaproteobacteria. The IRBP-induced EAU model in B10.RIII mice accurately reproduces the histopathological features of human uveitis; however, this model is limited in that it is triggered by introduction of a retinal antigen which is not believed to be present in naturally occurring forms of the disease. The gut microbiota has been shown to influence uveitis via activation of retinal T cells before the onset of clinical uveitis. With this in mind, Horai et al. developed a spontaneous uveitis model in which transgenic mice express a T-cell receptor for IRBP, the R161H model of uveitis, which is pathologically similar to the IRBP-induced EAU model [74]. The onset of uveitis was shown to be reduced when R161H mice were raised under germ-free conditions, supporting the notion that the gut microbiota negatively regulates uveitis onset [74]. A recent study by Janowitz et al. demonstrated that the gut of EAU mice exhibit reduced alpha diversity compared to non-immunized mice prior to onset of ocular inflammation. At peak uveitis pathology in EAU mice, the gut microbiota contained increased abundance of Prevotella, Lactobacillus, Anaeroplasma, Parabacteroides, Firmicutes, and Clostridium species [75]. Furthermore, EAU mice experienced increased production of ileal antimicrobial peptides, such as Reg3γ, S100A8, and lipocalin, compared with non-immunized mice [75]. The aforementioned studies support a role for the gut microbiota in the onset and severity of autoimmune uveitis. However, additional studies using models which more closely represent human disease are still warranted [74, 76].
Glaucoma
Primary open-angle glaucoma (POAG) is an ocular neurodegenerative disease characterized by loss of retinal ganglion cells which causes irreversible blindness and affects approximately 79.6 million people worldwide [77]. A recent study by Gong et al. found that the gut microbiota of subjects with POAG, compared to healthy subjects, had increased abundance of Prevotellaceae, Enterobacteriaceae, and Escherichia coli and decreased abundance of Megamonas and Bacteroides plebeius [78]. Furthermore, they found that mean visual acuity was negatively correlated with bacteria of the Megamonas genus, visual field mean defect (VF-MD) was negatively correlated with members of the Faecalibacterium genus, and average retinal nerve fiber layer (RNFL) thickness was positively correlated with members of the Streptococcus genus. [78] In a study of 1999 African American subjects (1217 with POAG and 782 controls), Collins et al. found that variants in two mitochondrial DNA (mtDNA) haplogroups, L2a1 (m.15784T>C) and L2 haplogroups (m.16390G>A), were enriched in DNA pools of POAG subjects compared to controls [79]. These variants were associated with the composition of the gut microbiota. Specifically, the variant m.15784T>C was associated with members of the Firmicutes phylum, whereas the variant m.16390G>A was associated with members of the Proteobacteria phylum [80].
Additionally, murine models have shown that the gut microbiota may play a role in the development of glaucoma. Specifically, Chen et al. demonstrated that DBA/2J mice, which spontaneously develop increased intraocular pressure (IOP) and glaucoma by 6–8 months of age, exhibit an absence of glaucomatous neural degeneration at 12 months of age when raised under germ-free conditions. DBA/2J mice raised under specific-pathogen-free (SPF) conditions, on the other hand, exhibit progressive retinal ganglion cell (RGC) and axon loss by ~ 25% and ~ 50% at 8–10 months and 12 months of age, respectively [81] (Fig. 3). Together, the aforementioned studies provide support for the involvement of an altered gut microbiota in glaucoma; however, additional work is required to further elucidate this association and to determine whether there is a definitive role for the gut microbiota in glaucoma progression.
Sjögren’s Syndrome
Sjögren’s syndrome (SS) is an autoimmune condition characterized by lymphocytic infiltration in exocrine glands, particularly the lacrimal glands, causing dry eyes [82, 83]. Moon et al. recently published a study in which changes in the gut microbiota were found to be associated with SS in human subjects compared to healthy volunteers and subjects with environmental dry eye syndrome (DES) [82]. SS was associated with a decreased F/B ratio, decreased Actinobacteria, and decreased abundance of members of the Bifidobacterium, Blautia, Dorea, and Agathobacter genera, with increased abundance of members of the Prevotella, Odoribacter, and Alistipes genera [82]. Furthermore, severe dysbiosis, as assessed by the Dysbiosis Index score (DIS), was prevalent in subjects with primary SS (pSS). pSS subjects with severe dysbiosis (DIS = 5) had higher disease activity as assessed by the European League Against Rheumatism (EULAR) Sjögren’s Syndrome Disease Activity Index (ESSDAI) and Clinical ESSDAI (ClinESSDA) [84]. However, dysbiosis severity was not associated with disease duration or EULAR Sjögren’s Syndrome Patient Reported Index (ESSPRI) total score [84].
Potential Therapeutics
Intermittent Fasting
Intermittent fasting (IF), the restriction of food intake to specific days or times of day, has shown many beneficial effects in human and animal studies, including facilitating regeneration and tissue repair [85]. There are currently three intermittent fasting regimens reported to have beneficial health outcomes: alternate-day fasting—alternation of fasting and feeding days; modified fasting—consumption of 20–25% of recommended energy needs on fasting days, usually 2 nonconsecutive days per week, with 5 days of ad libitum feeding; and time-restricted fasting—allowance of ad libitum feeding during specific time frames, typically 8 h per day [86]. Studies on the IF in both rodent and human models have proposed the following mechanisms for its beneficial effects including regulation of inflammation, insulin levels, lipid levels, and satiety hormones via modification of the gut microbiota, circadian rhythms, and lifestyle factors such as diet, activity, and sleep [86]. In rodent models, within 1 week of starting an alternate-day fasting regimen, IF has been shown to reduce blood pressure and heart rate, both of which are known risk factors for ocular vascular diseases such as DR [87]. This effect is suggested to be due to increased brain-derived neurotrophic factor signaling by increasing the activity of brainstem cholinergic neurons [88,89,90]. Furthermore, Sutton et al. performed the first supervised controlled feeding trial in which early time-restricted feeding (eTRF, ad libitum feeding from 8 a.m. to 2 p.m. and fasting for the remaining 18 h) was shown to decrease postprandial insulin, blood pressure, oxidative stress, and evening appetite while increasing insulin sensitivity and β-cell function in human subjects [91]. This study and others further support the beneficial effects of intermittent fasting and suggest it may have therapeutic potential in ocular vascular diseases by decreasing blood pressure and insulin sensitivity [91,92,93].
It was recently shown that alternate-day fasting for 7 months increased the quantity of intestinal mucus-producing goblet cells and decreased concentrations of plasma PGN, indicating improved GVB integrity. IF was shown to restructure the gut microbiota by an increase in the F/B ratio in db/db-IF compared to db/m-IF [12]. Furthermore, IF altered bile acid metabolism in db/db–IF by increased generation of beneficial secondary metabolites, such as tauroursodeoxycholate (TUDCA), which was shown to cross the blood–retinal barrier and activate its receptor, TGR5, reducing retinal inflammation, compared to db/db mice on an ad libitum diet (db/db-AL) [12]. Finally, IF improved DR pathology by reducing the number of retinal acellular capillaries in db/db-IF mice compared to controls (db/db-AL) [12] (Fig. 3). It is important to note that the beneficial effects observed in this study were glucose-independent, as no difference in glycated hemoglobin was found between db/db-AL and db/db-IF mice. Thus, the beneficial effects of IF on DR progression in the db/db mouse model of T2D was attributed to changes in the gut microbiota that modulated the production of secondary bile acids, and not improvement in glucose control.
Fecal Microbial Transplant
Fecal microbial transplant (FMT) describes the method of transferring the gut microbiota of control or healthy donors into that of unhealthy recipients. The goal of such a transplant is to colonize the gut of the recipient with a microbiota which is considered “normal”. Several murine studies have been performed in this regard in which the gut microbiota of disease-state mice are transplanted into germ-free (GF) mice, those lacking a gut microbiota [94,95,96]. Bäckhed et al. showed that GF mice colonized with the microbiota of their counterparts experienced weight gain and increased insulin resistance [41]. Furthermore, weight gain was observed in GF mice with reconstituted gut microbiota of obese (ob/ob) donor mice [42]. These studies demonstrate the ability of the gut microbiota to transfer disease states. A number of human studies have been conducted using FMT as a method for altering the microbiota in patients with various diseases. Of note, FMT increased the alpha diversity, i.e., species richness, in patients with Crohn’s disease and pediatric ulcerative colitis [97, 98]. To our knowledge, no studies have been performed in which FMT is used as treatment for ocular disease. However, this technique may prove useful in restructuring the gut microbiota and improving the pathophysiology of individuals with the aforementioned ocular conditions.
Pre- and Probiotics
Prebiotics are nondigestible short-chain carbohydrates which function as metabolic substrates for gut bacteria such as Lactobacillus and Bifidobacterium species. These compounds are naturally occurring and exert their positive health benefits by improving gut immunity and gut barrier function [99]. Prebiotics have been shown to restructure the microbiota; however, increased efficacy has been observed when combined with a probiotic, a live microbial supplement, in human fecal sample studies [99]. Probiotics have been shown to improve intestinal health via inhibition of pathogenic species colonization, improvement of gut barrier function, and modulation of gut immunity [99]. As mentioned previously, diabetic mice exhibit decreased intestinal levels of ACE2 [13, 51]. It is hypothesized that probiotic replenishment of ACE2 via oral administration of live bacteria which produce ACE2 will improve gut barrier function, restructure the gut microbiota to homeostatic levels, and prevent pathological translocation of PGN into the circulation [100]. Together, this treatment has the potential to reduce DR pathology, as these have all been shown to play a role in the progression of DR in mouse models of type 1 and type 2 diabetes [13, 51]. Pre- and probiotic interventions show promise in that they are a less invasive therapeutic option with minimal risk for complications.
Antibiotics
Antibiotics have been used for modulation of the microbiome experimentally and as a therapy. Orally administered combination broad-spectrum antibiotics [ampicillin (1 g/L), neomycin (1 g/L), metronidazole (1 g/L), and vancomycin (500 mg/L)] were shown to reduce mean uveitis clinical scores in EAU mice [73] (Fig. 3). Administered alone, metronidazole or vancomycin were shown to decrease mean uveitis clinical scores. Combinatorial broad-spectrum antibiotics induced phyla-level and class-level alterations in gut microbial composition, assessed by quantitative polymerase chain reaction (qPCR), as follows: decreased quantities of Bacteroidetes, Firmicutes, and Alphaproteobacteria, and increased quantities of Gammaproteobacteria [73]. Antibiotics have also been found to influence the gut microbiota in models of neurodegeneration, autism, anxiety, Alzheimer’s disease, hepatic encephalopathy, and obesity [101,102,103,104].
Conclusion
A eubiotic gut is essential to the maintenance of human health. Under aberrant composition, the gut microbiota induces pathological inflammation via stimulation of both the innate and adaptive arms of the immune system. This inflammation may proceed to destroy tissues throughout the body. Of particular interest, pathological inflammation induces disruption of the ocular tissues, exacerbating the progression of diabetic retinopathy, age-related macular degeneration, choroidal neovascularization, uveitis, glaucoma, and Sjögren’s syndrome. Much work remains to be done to understand the mechanisms involved in regulating the gut–eye axis. This review highlights key studies which link various models of ocular diseases with alterations in the gut microbiota; however, further studies are required to determine whether direct mechanistic links exist between an aberrant gut microbiota and initiation, progression, or exacerbation of, or protection from, ocular diseases.
References
Ley RE, Peterson DA, Gordon JI. Ecological and evolutionary forces shaping microbial diversity in the human intestine. Cell. 2006;124(4):837–48.
Atarashi K, Honda K. Microbiota in autoimmunity and tolerance. Curr Opin Immunol. 2011;23(6):761–8.
Frank DN, Pace NR. Gastrointestinal microbiology enters the metagenomics era. Curr Opin Gastroenterol. 2008;24(1):4–10.
Eckburg PB, Bik EM, Bernstein CN, Purdom E, Dethlefsen L, Sargent M, et al. Diversity of the human intestinal microbial flora. Science. 2005;308(5728):1635–8.
Frank DN, St Amand AL, Feldman RA, Boedeker EC, Harpaz N, Pace NR. Molecular-phylogenetic characterization of microbial community imbalances in human inflammatory bowel diseases. Proc Natl Acad Sci USA. 2007;104(34):13780–5.
Hooper LV, Gordon JI. Commensal host-bacterial relationships in the gut. Science. 2001;292(5519):1115–8.
Bibbò S, Dore MP, Pes GM, Delitala G, Delitala AP. Is there a role for gut microbiota in type 1 diabetes pathogenesis? Ann Med. 2017;49(1):11–22.
Kasselman LJ, Vernice NA, DeLeon J, Reiss AB. The gut microbiome and elevated cardiovascular risk in obesity and autoimmunity. Atherosclerosis. 2018;2(271):203–13.
Lin L, Zhang J. Role of intestinal microbiota and metabolites on gut homeostasis and human diseases. BMC Immunol. 2017;18(1):2.
Sohail MU, Althani A, Anwar H, Rizzi R, Marei HE. Role of the gastrointestinal tract microbiome in the pathophysiology of diabetes mellitus. J Diabetes Res. 2017;26(2017):9631435.
Gavin PG, Mullaney JA, Loo D, Cao K-AL, Gottlieb PA, Hill MM, et al. Intestinal metaproteomics reveals host-microbiota interactions in subjects at risk for type 1 diabetes. Diabetes Care. 2018;41(10):2178–86.
Beli E, Yan Y, Moldovan L, Vieira CP, Gao R, Duan Y, et al. Restructuring of the gut microbiome by intermittent fasting prevents retinopathy and prolongs survival in db/db mice. Diabetes. 2018;67(9):1867–79.
Duan Y, Prasad R, Feng D, Beli E, Li Calzi S, Longhini ALF, et al. Bone marrow-derived cells restore functional integrity of the gut epithelial and vascular barriers in a model of diabetes and ACE2 deficiency. Circ Res. 2019;125(11):969–88.
Dinan TG, Cryan JF. Brain-Gut-Microbiota Axis and Mental Health. Psychosom Med. 2017;79(8):920–6.
Jiang C, Li G, Huang P, Liu Z, Zhao B. The gut microbiota and Alzheimer’s disease. J Alzheimers Dis. 2017;58(1):1–15.
Yang B, Wang Y, Qian P-Y. Sensitivity and correlation of hypervariable regions in 16S rRNA genes in phylogenetic analysis. BMC Bioinformatics. 2016;22(17):135.
Rausch P, Rühlemann M, Hermes BM, Doms S, Dagan T, Dierking K, et al. Comparative analysis of amplicon and metagenomic sequencing methods reveals key features in the evolution of animal metaorganisms. Microbiome. 2019;7(1):133.
Human Microbiome Project Consortium. Structure, function and diversity of the healthy human microbiome. Nature. 2012;486(7402):207–14.
Graspeuntner S, Loeper N, Künzel S, Baines JF, Rupp J. Selection of validated hypervariable regions is crucial in 16S-based microbiota studies of the female genital tract. Sci Rep. 2018;8(1):9678.
Callahan BJ, McMurdie PJ, Rosen MJ, Han AW, Johnson AJA, Holmes SP. DADA2: high-resolution sample inference from Illumina amplicon data. Nat Methods. 2016;13(7):581–3.
Franzosa EA, Morgan XC, Segata N, Waldron L, Reyes J, Earl AM, et al. Relating the metatranscriptome and metagenome of the human gut. Proc Natl Acad Sci USA. 2014;111(22):E2329–38.
Chan JP, Wright JR, Wong HT, Ardasheva A, Brumbaugh J, McLimans C, et al. Using bacterial transcriptomics to investigate targets of host-bacterial interactions in Caenorhabditis elegans. Sci Rep. 2019;9(1):5545.
Smythies LE, Sellers M, Clements RH, Mosteller-Barnum M, Meng G, Benjamin WH, et al. Human intestinal macrophages display profound inflammatory anergy despite avid phagocytic and bacteriocidal activity. J Clin Invest. 2005;115(1):66–75.
Barreau F, Hugot JP. Intestinal barrier dysfunction triggered by invasive bacteria. Curr Opin Microbiol. 2014;17:91–8.
Den Besten G, van Eunen K, Groen AK, Venema K, Reijngoud D-J, Bakker BM. The role of short-chain fatty acids in the interplay between diet, gut microbiota, and host energy metabolism. J Lipid Res. 2013;54(9):2325–40.
Goodell EW. Recycling of murein by Escherichia coli. J Bacteriol. 1985;163(1):305–10.
Hergott CB, Roche AM, Tamashiro E, Clarke TB, Bailey AG, Laughlin A, et al. Peptidoglycan from the gut microbiota governs the lifespan of circulating phagocytes at homeostasis. Blood. 2016;127(20):2460–71.
Sabroe I, Prince LR, Jones EC, Horsburgh MJ, Foster SJ, Vogel SN, et al. Selective roles for Toll-like receptor (TLR)2 and TLR4 in the regulation of neutrophil activation and life span. J Immunol. 2003;170(10):5268–75.
Prince LR, Whyte MK, Sabroe I, Parker LC. The role of TLRs in neutrophil activation. Curr Opin Pharmacol. 2011;11(4):397–403.
Sabroe I, Jones EC, Usher LR, Whyte MKB, Dower SK. Toll-like receptor (TLR)2 and TLR4 in human peripheral blood granulocytes: a critical role for monocytes in leukocyte lipopolysaccharide responses. J Immunol. 2002;168(9):4701–10.
Hayashi F, Means TK, Luster AD. Toll-like receptors stimulate human neutrophil function. Blood. 2003;102(7):2660–9.
Lee R, Wong TY, Sabanayagam C. Epidemiology of diabetic retinopathy, diabetic macular edema and related vision loss. Eye Vis (Lond). 2015;30(2):17.
Qin J, Li Y, Cai Z, Li S, Zhu J, Zhang F, et al. A metagenome-wide association study of gut microbiota in type 2 diabetes. Nature. 2012;490(7418):55–60.
Larsen N, Vogensen FK, van den Berg FWJ, Nielsen DS, Andreasen AS, Pedersen BK, et al. Gut microbiota in human adults with type 2 diabetes differs from non-diabetic adults. PLoS One. 2010;5(2):e9085.
Zhang Y, Zhang H. Microbiota associated with type 2 diabetes and its related complications. Food Science and Human Wellness. 2013;2(3–4):167–72.
Yassour M, Lim MY, Yun HS, Tickle TL, Sung J, Song Y-M, et al. Sub-clinical detection of gut microbial biomarkers of obesity and type 2 diabetes. Genome Med. 2016;8(1):17.
Karlsson FH, Tremaroli V, Nookaew I, Bergström G, Behre CJ, Fagerberg B, et al. Gut metagenome in European women with normal, impaired and diabetic glucose control. Nature. 2013;498(7452):99–103.
Allin KH, Tremaroli V, Caesar R, Jensen BAH, Damgaard MTF, Bahl MI, et al. Aberrant intestinal microbiota in individuals with prediabetes. Diabetologia. 2018;61(4):810–20.
Forslund K, Hildebrand F, Nielsen T, Falony G, Le Chatelier E, Sunagawa S, et al. Disentangling type 2 diabetes and metformin treatment signatures in the human gut microbiota. Nature. 2015;528(7581):262–6.
Sun L, Xie C, Wang G, Wu Y, Wu Q, Wang X, et al. Gut microbiota and intestinal FXR mediate the clinical benefits of metformin. Nat Med. 2018;24(12):1919–29.
Bäckhed F, Ding H, Wang T, Hooper LV, Koh GY, Nagy A, et al. The gut microbiota as an environmental factor that regulates fat storage. Proc Natl Acad Sci USA. 2004;101(44):15718–23.
Turnbaugh PJ, Ley RE, Mahowald MA, Magrini V, Mardis ER, Gordon JI. An obesity-associated gut microbiome with increased capacity for energy harvest. Nature. 2006;444(7122):1027–31.
Clermont A, Bursell S-E, Feener EP. Role of the angiotensin II type 1 receptor in the pathogenesis of diabetic retinopathy: effects of blood pressure control and beyond. J Hypertens Suppl. 2006;24(1):S73–80.
Hayden MR, Sowers KM, Pulakat L, Joginpally T, Krueger B, Whaley-Connell A, et al. Possible mechanisms of local tissue renin-angiotensin system activation in the cardiorenal metabolic syndrome and type 2 diabetes mellitus. Cardiorenal Med. 2011;1(3):193–210.
Miller AG, Tan G, Binger KJ, Pickering RJ, Thomas MC, Nagaraj RH, et al. Candesartan attenuates diabetic retinal vascular pathology by restoring glyoxalase-I function. Diabetes. 2010;59(12):3208–15.
Rojas M, Zhang W, Lee DL, Romero MJ, Nguyen DT, Al-Shabrawey M, et al. Role of IL-6 in angiotensin II-induced retinal vascular inflammation. Invest Ophthalmol Vis Sci. 2010;51(3):1709–18.
Saiki A, Ohira M, Endo K, Koide N, Oyama T, Murano T, et al. Circulating angiotensin II is associated with body fat accumulation and insulin resistance in obese subjects with type 2 diabetes mellitus. Metab Clin Exp. 2009;58(5):708–13.
Silva KC, Rosales MAB, Biswas SK, de Faria LJB, de Faria LJM. Diabetic retinal neurodegeneration is associated with mitochondrial oxidative stress and is improved by an angiotensin receptor blocker in a model combining hypertension and diabetes. Diabetes. 2009;58(6):1382–90.
Wiecek A, Chudek J, Kokot F. Role of angiotensin II in the progression of diabetic nephropathy-therapeutic implications. Nephrol Dial Transplant. 2003;18(Suppl 5):v16–20.
Wilkinson-Berka JL, Rana I, Armani R, Agrotis A. Reactive oxygen species, Nox and angiotensin II in angiogenesis: implications for retinopathy. Clin Sci. 2013;124(10):597–615.
Duan Y, Beli E, Li Calzi S, Quigley JL, Miller RC, Moldovan L, et al. Loss of angiotensin-converting enzyme 2 exacerbates diabetic retinopathy by promoting bone marrow dysfunction. Stem Cells. 2018;36(9):1430–40.
Dominguez JM, Hu P, Caballero S, Moldovan L, Verma A, Oudit GY, et al. Adeno-associated virus overexpression of angiotensin-converting enzyme-2 reverses diabetic retinopathy in type 1 diabetes in mice. Am J Pathol. 2016;186(6):1688–700.
Jarajapu YPR, Bhatwadekar AD, Caballero S, Hazra S, Shenoy V, Medina R, et al. Activation of the ACE2/angiotensin-(1-7)/Mas receptor axis enhances the reparative function of dysfunctional diabetic endothelial progenitors. Diabetes. 2013;62(4):1258–69.
Verma A, Shan Z, Lei B, Yuan L, Liu X, Nakagawa T, et al. ACE2 and Ang-(1-7) confer protection against development of diabetic retinopathy. Mol Ther. 2012;20(1):28–36.
García-Layana A, Cabrera-López F, García-Arumí J, Arias-Barquet L, Ruiz-Moreno JM. Early and intermediate age-related macular degeneration: update and clinical review. Clin Interv Aging. 2017;3(12):1579–87.
Cook HL, Patel PJ, Tufail A. Age-related macular degeneration: diagnosis and management. Br Med Bull. 2008;85:127–49.
Maberley DAL, Hollands H, Chuo J, Tam G, Konkal J, Roesch M, et al. The prevalence of low vision and blindness in Canada. Eye (Lond). 2006;20(3):341–6.
Zhang Q-Y, Tie L-J, Wu S-S, Lv P-L, Huang H-W, Wang W-Q, et al. Overweight, obesity, and risk of age-related macular degeneration. Invest Ophthalmol Vis Sci. 2016;57(3):1276–83.
Adams MKM, Simpson JA, Aung KZ, Makeyeva GA, Giles GG, English DR, et al. Abdominal obesity and age-related macular degeneration. Am J Epidemiol. 2011;173(11):1246–55.
Rowan S, Jiang S, Korem T, Szymanski J, Chang M-L, Szelog J, et al. Involvement of a gut-retina axis in protection against dietary glycemia-induced age-related macular degeneration. Proc Natl Acad Sci USA. 2017;114(22):E4472–81.
Rinninella E, Mele MC, Merendino N, Cintoni M, Anselmi G, Caporossi A, et al. The role of diet, micronutrients and the gut microbiota in age-related macular degeneration: new perspectives from the gut-retina axis. Nutrients. 2018;10(11):1677.
Grossniklaus HE, Green WR. Choroidal neovascularization. Am J Ophthalmol. 2004;137(3):496–503.
Andriessen EM, Wilson AM, Mawambo G, Dejda A, Miloudi K, Sennlaub F, et al. Gut microbiota influences pathological angiogenesis in obesity-driven choroidal neovascularization. EMBO Mol Med. 2016;8(12):1366–79.
Radu M, Chernoff J. An in vivo assay to test blood vessel permeability. J Vis Exp. 2013;73:e50062.
Cani PD, Possemiers S, Van de Wiele T, Guiot Y, Everard A, Rottier O, et al. Changes in gut microbiota control inflammation in obese mice through a mechanism involving GLP-2-driven improvement of gut permeability. Gut. 2009;58(8):1091–103.
Membrez M, Blancher F, Jaquet M, Bibiloni R, Cani PD, Burcelin RG, et al. Gut microbiota modulation with norfloxacin and ampicillin enhances glucose tolerance in mice. FASEB J. 2008;22(7):2416–26.
Neves AL, Coelho J, Couto L, Leite-Moreira A, Roncon-Albuquerque R. Metabolic endotoxemia: a molecular link between obesity and cardiovascular risk. J Mol Endocrinol. 2013;51(2):R51–64.
Zinkernagel MS, Zysset-Burri DC, Keller I, Berger LE, Leichtle AB, Largiadèr CR, et al. Association of the intestinal microbiome with the development of neovascular age-related macular degeneration. Sci Rep. 2017;17(7):40826.
Liu C-H, Wang Z, Sun Y, Chen J. Animal models of ocular angiogenesis: from development to pathologies. FASEB J. 2017;31(11):4665–81.
Abdulaal MR, Abiad BH, Hamam RN. Uveitis in the aging eye: incidence, patterns, and differential diagnosis. J Ophthalmol. 2015;18(2015):509456.
Horai R, Sen HN, Caspi RR. Commensal microbiota as a potential trigger of autoimmune uveitis. Expert Rev Clin Immunol. 2017;13(4):291–3.
Kalyana Chakravarthy S, Jayasudha R, Sai Prashanthi G, Ali MH, Sharma S, Tyagi M, et al. Dysbiosis in the gut bacterial microbiome of patients with uveitis, an inflammatory disease of the eye. Indian J Microbiol. 2018;58(4):457–69.
Nakamura YK, Metea C, Karstens L, Asquith M, Gruner H, Moscibrocki C, et al. Gut microbial alterations associated with protection from autoimmune uveitis. Invest Ophthalmol Vis Sci. 2016;57(8):3747–58.
Horai R, Caspi RR. Microbiome and autoimmune uveitis. Front Immunol. 2019;19(10):232.
Janowitz C, Nakamura YK, Metea C, Gligor A, Yu W, Karstens L, et al. Disruption of intestinal homeostasis and intestinal microbiota during experimental autoimmune uveitis. Invest Ophthalmol Vis Sci. 2019;60(1):420–9.
Chen J, Caspi RR. Clinical and functional evaluation of ocular inflammatory disease using the model of experimental autoimmune uveitis. Methods Mol Biol. 2019;1899:211–27.
Chaiwiang N, Poyomtip T. Microbial dysbiosis and microbiota-gut-retina axis: the lesson from brain neurodegenerative diseases to primary open-angle glaucoma pathogenesis of autoimmunity. Acta Microbiol Immunol Hung. 2019;66(4):541–58.
Gong H, Zhang S, Li Q, Zuo C, Gao X, Zheng B, et al. Gut microbiota compositional profile and serum metabolic phenotype in patients with primary open-angle glaucoma. Exp Eye Res. 2020;191:107921.
Collins DW, Gudiseva HV, Trachtman B, Bowman AS, Sagaser A, Sankar P, et al. Association of primary open-angle glaucoma with mitochondrial variants and haplogroups common in African Americans. Mol Vis. 2016;16(22):454–71.
Ma J, Coarfa C, Qin X, Bonnen PE, Milosavljevic A, Versalovic J, et al. mtDNA haplogroup and single nucleotide polymorphisms structure human microbiome communities. BMC Genom. 2014;3(15):257.
Chen H, Cho K-S, Vu THK, Shen C-H, Kaur M, Chen G, et al. Commensal microflora-induced T cell responses mediate progressive neurodegeneration in glaucoma. Nat Commun. 2018;9(1):3209.
Moon J, Choi SH, Yoon CH, Kim MK. Gut dysbiosis is prevailing in Sjögren’s syndrome and is related to dry eye severity. PLoS One. 2020;15(2):e0229029.
Kuklinski E, Asbell PA. Sjogren’s syndrome from the perspective of ophthalmology. Clin Immunol. 2017;2(182):55–61.
Mandl T, Marsal J, Olsson P, Ohlsson B, Andréasson K. Severe intestinal dysbiosis is prevalent in primary Sjögren’s syndrome and is associated with systemic disease activity. Arthritis Res Ther. 2017;19(1):237.
Cheng C-W, Villani V, Buono R, Wei M, Kumar S, Yilmaz OH, et al. Fasting-mimicking diet promotes Ngn3-driven β-cell regeneration to reverse diabetes. Cell. 2017;168(5):775–788.e12.
Patterson RE, Sears DD. Metabolic effects of intermittent fasting. Annu Rev Nutr. 2017;21(37):371–93.
Wong TY, Cheung CMG, Larsen M, Sharma S, Simó R. Diabetic retinopathy. Nat Rev Dis Primers. 2016;17(2):16012.
Wan R, Weigand LA, Bateman R, Griffioen K, Mendelowitz D, Mattson MP. Evidence that BDNF regulates heart rate by a mechanism involving increased brainstem parasympathetic neuron excitability. J Neurochem. 2014;129(4):573–80.
Wan R, Camandola S, Mattson MP. Intermittent food deprivation improves cardiovascular and neuroendocrine responses to stress in rats. J Nutr. 2003;133(6):1921–9.
Mattson MP, Longo VD, Harvie M. Impact of intermittent fasting on health and disease processes. Ageing Res Rev. 2017;39:46–58.
Sutton EF, Beyl R, Early KS, Cefalu WT, Ravussin E, Peterson CM. Early time-restricted feeding improves insulin sensitivity, blood pressure, and oxidative stress even without weight loss in men with prediabetes. Cell Metab. 2018;27(6):1212–1221.e3.
Jamshed H, Beyl RA, Della Manna DL, Yang ES, Ravussin E, Peterson CM. early time-restricted feeding improves 24-hour glucose levels and affects markers of the circadian clock, aging, and autophagy in humans. Nutrients. 2019;11(6):1234.
Peterson CM. Intermittent fasting induces weight loss, but the effects on cardiometabolic health are modulated by energy balance. Obesity (Silver Spring). 2019;27(1):11.
de Groot PF, Frissen MN, de Clercq NC, Nieuwdorp M. Fecal microbiota transplantation in metabolic syndrome: history, present and future. Gut Microbes. 2017;8(3):253–67.
Sommer F, Anderson JM, Bharti R, Raes J, Rosenstiel P. The resilience of the intestinal microbiota influences health and disease. Nat Rev Microbiol. 2017;15(10):630–8.
Brunkwall L, Orho-Melander M. The gut microbiome as a target for prevention and treatment of hyperglycaemia in type 2 diabetes: from current human evidence to future possibilities. Diabetologia. 2017;60(6):943–51.
Nusbaum DJ, Sun F, Ren J, Zhu Z, Ramsy N, Pervolarakis N, et al. Gut microbial and metabolomic profiles after fecal microbiota transplantation in pediatric ulcerative colitis patients. FEMS Microbiol Ecol. 2018;94(9):fiy133.
Vaughn BP, Vatanen T, Allegretti JR, Bai A, Xavier RJ, Korzenik J, et al. Increased intestinal microbial diversity following fecal microbiota transplant for active crohn’s disease. Inflamm Bowel Dis. 2016;22(9):2182–90.
Murphy AB, Dinan TG, Cryan JF, Stanton C, Ross RP. Probiotics as curators of a healthy gut microbiota. In: Hyland N, Stanton C (eds) The Gut-Brain Axis. Elsevier; 2016. p. 61–88.
Verma A, Xu K, Du T, Zhu P, Liang Z, Liao S, et al. Expression of human ACE2 in lactobacillus and beneficial effects in diabetic retinopathy in mice. Mol Ther Methods Clin Dev. 2019;13(14):161–70.
Gérard P. Gut microbiota and obesity. Cell Mol Life Sci. 2016;73(1):147–62.
Mangiola F, Ianiro G, Franceschi F, Fagiuoli S, Gasbarrini G, Gasbarrini A. Gut microbiota in autism and mood disorders. World J Gastroenterol. 2016;22(1):361–8.
Schmidtner AK, Slattery DA, Gläsner J, Hiergeist A, Gryksa K, Malik VA, et al. Minocycline alters behavior, microglia and the gut microbiome in a trait-anxiety-dependent manner. Transl Psychiatry. 2019;9(1):223.
Quigley EMM. Microbiota-brain-gut axis and neurodegenerative diseases. Curr Neurol Neurosci Rep. 2017;17(12):94.
Acknowledgements
All figures were created with BioRender.com.
Funding
This manuscript was supported by the National Institutes of Health grants R01EY025383, R01EY012601, R01EY028858, R01EY028037 to M.B. Grant and T32HL105349 to J.L. Floyd. This manuscript was supported by unrestricted grant from Research to Prevent Blindness to UAB Department of Ophthalmology and Visual Sciences. No funding or sponsorship was received for the publication of this article.
Authorship
All named authors meet the International Committee of Medical Journal Editors (ICMJE) criteria for authorship for this article, take responsibility for the integrity of the work as a whole, and have given their approval for this version to be published.
Disclosures
Jason L. Floyd and Maria B. Grant have no financial disclosures.
Compliance with Ethics Guidelines
This article is based on previously conducted studies and does not contain any studies with human participants or animals performed by any of the authors.
Data Availability
Data sharing is not applicable to this article as no datasets were generated or analyzed during the current study.
Author information
Authors and Affiliations
Corresponding author
Additional information
Digital Features
To view digital features for this article go to https://doi.org/10.6084/m9.figshare.12497978.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial 4.0 International License, which permits any non-commercial use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc/4.0/.
About this article
Cite this article
Floyd, J.L., Grant, M.B. The Gut–Eye Axis: Lessons Learned from Murine Models. Ophthalmol Ther 9, 499–513 (2020). https://doi.org/10.1007/s40123-020-00278-2
Received:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s40123-020-00278-2