Abstract
The design, synthesis and characterization of a novel nucleoamino acid derivative based on an l-tryptophanamide functionalised with a thymine nucleobase (named TrpT) is here described. The novel construct’s tendency to self-assemble into supramolecular networks in aqueous solution was demonstrated by dynamic light scattering (DLS), circular dichroism (CD), fluorescence and UV spectroscopic measurements. TrpT nanoaggregates showed good stability (up to 5 h) at 140 µM and proved to comprise species of mean hydrodynamic diameter 330 nm and a homogeneous size distribution; scanning electron microscopy (SEM) analysis further revealed these to be spherical-shaped assemblies. The ability of TrpT nanoaggregates to bind curcumin, selected as a model anticancer drug, was also evaluated and its release was monitored over time by confocal microscopy. Molecular docking studies were performed on both TrpT self-assembly and curcumin-loaded nanoaggregates suggesting that the phytomolecule can be accommodated in the interior of the supramolecular network via hydrophobic (π−π and π-alkyl) interactions; the formation of TrpT-curcumin adducts may improve the polarity of the highly-hydrophobic curcumin with a resulting logP closer to the optimal values expected for a good drug bioavailability, as estimated by the ADMETlab software. Finally, the high stability of TrpT nanoassembly in human serum, and the absence of significant toxic effects on human model cells in a cell viability assay, were also demonstrated. Despite its thymine-based scaffold, TrpT was shown not to bind adenine-bearing nucleic acids, suggesting that this interaction is hindered by its intrinsic propensity to self-assemble in preference to forming A-T base pairings. Instead, TrpT was able to interact with a serum protein such as bovine serum albumin (BSA), known to improve the bloodstream transportation and bioavailability of its cargo. Collectively, our findings support the potential use of TrpT for the development of new drug delivery systems.
Graphical abstract

Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Synthetic conjugates composed of nucleobases inserted into a peptide backbone, mainly of diamino-acidic nature, may show many advantageous properties including nucleic acid-binding ability [1, 2]. Nonetheless, these molecules, also classed as nucleopeptides, are able to form supramolecular networks, cross cellular membranes and exhibit useful bioactivities [3,4,5,6,7]. Since the overall nucleopeptide properties arise from both the amino acid and nucleobase moieties, it is worth underlining that short peptides are able to self-assemble into different nanostructures [8,9,10,11], while nucleobases in synthetic molecules and especially in modified nucleosides [12,13,14,15,16,16] may lead to several biological properties. For example, when single nucleobases were inserted into polyamino acid chains, certain protein and peptide structures were stabilised, improving their function [17, 18], while nucleobase-conjugated diphenylalanines were found to self-assemble into nanostructures useful in various biomedical applications [19,20,21].
The ability of nucleopeptides to form highly ordered architectures [22] has been recently employed to develop controlled supramolecular systems such as hydrogels, nanospheres, nanotubes, nanofibers, nanovesicles, or micelles (of diverse shapes, including cylindrical, spherical, and worm-like). These are amenable to various bio-nanotechnological applications, such as drug/gene delivery, and take advantage of the well-documented biocompatibility of the nucleopeptides [23].
Nucleopeptide building blocks, i.e. nucleoamino acids, are biologically-relevant compounds isolated from natural sources (e.g. l-willardiine extracted from Fagus sylvatica) or routinely synthesised monomers for nucleopeptide synthesis [24,25,26,27,28,29,30]. The interest in these molecules stems not only from the crucial roles that the related peptidyl nucleosides [31] play in biology and therapy but also from their utility to build up new supramolecular networks. In fact, individual nucleoamino acids have been observed to self-assemble in supramolecular networks, as in the case of l-spinacine (Scheme 1A) and nucleobase-bearing phenylalanines [32, 33]. Also short nucleopeptides such as nucleobase-containing diserine, diphenylalanine, and dityrosine have also been shown to efficiently self-assemble [34,35,36].
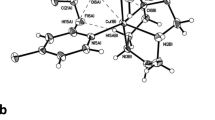
A Molecular structures of TrpT and SpinT [32] and their natural precursors. B Self-stacked 3D structure of TrpT predicted by ICM 2D to 3D (www.molsoft.com/2dto3d.html, accessed on 7th April 2022) software. C Energy-minimised 3D structure of TrpT realised by MOLVIEW (http://molview.org, accessed on 7th April 2022) software
Herein we report on the synthesis, characterization and potential self-assembling properties of a novel nucleoamino acid derivative based on an l-tryptophanamide functionalised with a thymine nucleobase (TrpT, Scheme 1A). As for the thyminyl-l-spinacine conjugate [32] (SpinT, Scheme 1A), our derivative, carrying an l-Trp in place of l-spinacine, was similarly expected to form superstructures held by intermolecular H-bonding, aromatic and hydrophobic interactions. Structurally, the substitution of l-spinacine by the l-Trp moiety (Scheme 1A) was intended to improve (i) the flexibility of the self-assembling monomer (compared to a more rigid l-spinacine scaffold), (ii) the H-bonding propensity (secondary vs. tertiary amide), (iii) the aromatic interactions.
Famously, the tryptophan residue, intrinsically able to aggregate forming nanotubes [37], when conjugated to nucleobases leads to self-assembling derivatives forming supramolecular polymers, as previously ascertained in the solid state by X-ray studies [38]. Interestingly, l-tryptophan derivatives N-functionalised with aromatic functionalities can also form hydrogels, and efficient drug nanocarriers [39, 40].
On the other hand, the nucleobase moiety not only contributes to reinforce the overall self-assembly capability but expectedly provides interactions with bioactive compounds for possible drug delivery applications.
For our TrpT structure, the conformations that lead to the intramolecular stacking (Scheme 1B) of the two tethered aromatic moieties are predicted in silico to be partially disfavoured by the tendency of the rings to lie almost perpendicular to one another (Figure S1, Scheme 1C), despite the previously reported self-stacking ability for adenine/thymine [41] and indole/thymine containing molecules [42]. Overall, we expected that both the self-stacked (Scheme 1B) and the open conformations (Scheme 1C, featuring the l-Trp and thymine moieties free from any self-stacking and exposed to intermolecular interactions with surrounding molecules) could favour the self-assembly, as this study sought confirmation by exploiting different techniques suggesting the potential use of TrpT as a self-assembling nanomaterial with potential nano-based biomedical applications.
Materials and methods
General methods
All the used reagents and solvents were purchased with the highest commercially available quality and were used as such.
1H- and 13C-NMR experiments were acquired at 25 °C on Varian Unity 400 MHz or 600 MHz spectrometers, as specified. Chemical shifts (δ) and coupling constants (J) are expressed in parts per million (ppm) and in Hertz (Hz), respectively. Proton and carbon chemical shifts were referenced to residual CHD2OD (δ = 3.30, quin) and CD3OD (δ = 49.3, sept) solvent signals, respectively. To explain signal multiplicity the following abbreviations were used: s = singlet; d = doublet; t = triplet; q = quartet; m = multiplet; b = broad.
All the samples were chromatographically analysed and characterised by LC–MS using a MSQ mass spectrometer (ThermoElectron, Milan, Italy) equipped with an ESI source—operating at 3 kV needle voltage and 320 °C—and a complete Surveyor HPLC system, comprising an MS pump, an autosampler, and a PDA detector. For sample injections, Phenomenex Jupiter (4u, 90A) Proteo (4 µm, 4.6 × 150 mm) columns were used. A gradient elution at a flow rate of 0.8 mL/min was applied starting with buffer A (0.05% TFA in water) and applying buffer B (0.05% TFA in acetonitrile) monitoring the signal at 260 nm.
MALDI-TOF MS measurements were performed on a TOF/TOF™ 5800 system using 2,5-dihydroxybenzoic acid (DHB) as a matrix and following previously optimized procedures [43, 44].
Analytical chromatograms were obtained on a Hewlett Packard/Agilent 1200 series HPLC, equipped with a diode array detector, by using a Phenomenex Jupiter C18 300 Å (5 µm, 4.6 × 250 mm) column. A gradient elution starting with buffer A' (0.1% TFA in water) and then ramping up buffer B' (0.1% TFA in acetonitrile) was performed. The elution was monitored at 25 °C and at a wavelength of 260 nm with a flow rate of 1 mL/min. Samples were lyophilised (FD4 Freeze Dryer, Heto Lab Equipment) for 16 h.
Synthesis of (2S)-3-(1H-Indol-3-yl)-2-[2-(5-methyl-2,4-dioxo-3,4-dihydro-2H-pyrimidin-1-yl)-acetylamino]-propionamide (TrpT)
TrpT was synthesized on a Rink Amide MBHA resin (0.65 mmol/g, 77 mg, 50 µmol) previously treated with 40% piperidine in DMF (Scheme 2) to remove the Fmoc protection. Subsequently, Fmoc-l-Trp-OH (250 µmol, 5 equiv.), DIEA (500 µmol, 10 equiv.) and HATU (250 µmol, 5 equiv.) were dissolved in anhydrous DMF (0.8 mL), added to the resin and left under stirring for 30 min. The resin was then drained and washed with DMF (3 × 1 ml). We repeated this procedure twice before the Fmoc deprotection, which was accomplished by treating the resin with 40% piperidine in DMF for 20 min. The l-Trp-functionalised resin was then reacted with thymine-1-acetic acid (250 µmol, 5 equiv.), in presence of DIEA (500 µmol, 10 equiv.) and HATU (250 µmol, 5 equiv.) in DMF (0.8 mL) for a half an hour. Following the washes, this coupling was repeated once again. Finally, TrpT—detached from the solid support with TFA/TIS/H2O (95/2.5/2.5, v/v/v) over 2 h—was obtained pure as a white solid (91% overall yield) after partial solvent evaporation, followed by precipitation from cold diethyl ether, centrifugation and final lyophilization. Before its use in our experiments, the solid TrpT sample was dissolved in ethanol/H2O = 1/1 (v/v) at a 14.3 mM concentration. This stock solution was then diluted to the desired concentrations with suitable amounts of sodium phosphate buffer (25 mM, pH 7.4).
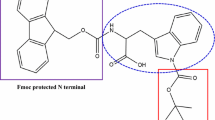
Schematic representation of the synthesis of TrpT realized in the solid phase. Abbreviations: Boc (tert-butyloxycarbonyl); DIEA (N,N-diisopropylethylamine); DMF (N,N-dimethylformamide); Fmoc (9-fluorenylmethoxycarbonyl); HATU (O-(7-azabenzotriazole-1-yl)-1,1,3,3-tetramethyluronium hexafluorophosphate); TFA (trifluoroacetic acid); TIS (triisopropylsilane); Trp (tryptophan)
1H-NMR (400 MHz, CD3OD, Figure S2): δ 7.62–6.99 (6H, m, Ar-Trp, C6-H), 4.67 (1H, dd, J = 5.6 and 8.4, CHα), 4.38 (1H, d, J = 16.8, NCHaH), 4.26 (1H, d, J = 16.8, NCHHb), 3.33 (1H, dd, J = 5.6 and 14.8, CHβH), 3.10 (1H, dd, J = 8.4 and 14.8, CHHβ'), 1.80 (3H, s, CH3);
13C-NMR (150 MHz, CD3OD, Figure S3): δ 176.8, 169.7, 167.3, 153.5, 143.7, 138.4, 129.2 124.9, 122.8, 120.2, 119.7, 112.7, 111.5, 111.4, 55.9, 51.3, 29.2, 12.5.
LC–ESI–MS (Figure S4A): m/z 369.77 (found), 370.39 (expected for [C18H19N5O4 + H]+); 391.86 (found), 392.37 (expected for [C18H19N5O4 + Na]+).
MALDI-TOF (Figure S4B): m/z 392.33 (found), 392.37 (expected for [C18H19N5O4 + Na]+); 408.31 (found), 408.38 (expected for [C18H19N5O4 + K]+).
Molecular modelling and in silico pharmacokinetic properties prediction
We realized the two models for the energy-minimised 3D structures (obtained as random low energy conformers)—reported in Scheme 1B, C, as well as Figure S1—using the online software MOLVIEW (http://molview.org) and DataWarrior (see http://openmolecules.org/).
Self-stacked TrpT 3D structure was obtained with the ICM 2D to 3D software (www.molsoft.com/2dto3d.html) using MMFF atom type assignment and force-field optimization. We computed the molecular volume of TrpT (Figure S6A) with WebLab ViewerPro 3.7 (Molecular Simulations, San Diego, California).
The Simplified Molecular Input Line Entry System (SMILES) code of TrpT was computed with MOLVIEW and applied to estimate—by using the SwissADME web service (http://www.swissadme.ch/index.php)—the physico-chemical properties of the molecule, including solubility in water, tPSA (topological polar surface area), molecular weight (MW), logP value (octanol–water partition coefficient) in five variants and the consensus LogP (cLogP) which was an average of the five mentioned predictions. All these data are reported in Figure S6B.
Moreover, the HDOCK software [45, 46] was exploited for the molecular docking studies described in this work. HDOCK is a docking software employed in macromolecule-macromolecule [45] and macromolecule-small molecule [47] docking simulations, including those involving RNA and DNA G-quadruplex structures [48, 49]. This server uses ITScore-PP, an iterative knowledge-based scoring function, ranking the Top 1–10 poses provided after the docking runs. The software provides dimensionless HDOCK scores that are correlated to binding affinities [50]. Examining HDOCK scores one can compare the binding affinities of different ligands for the same biomolecular target, with the most negative scores being associated with the highest binding affinities [50]. Blind molecular dockings for the (TrpT)n complexes (n = 2,4,8…0.256) and the curcumin/(TrpT)256 system were carried out using default parameters for all HDOCK dockings and the PDB files of curcumin and TrpT obtained after structure editing in Discovery studio software [51] that was used also to analyse the intermolecular interactions between the TrpT units in the self-assembly and between the self-assembly and the curcumin. The 3D structure, including H-atoms, for curcumin was retrieved by us from the PubChem database (https://pubchem.ncbi.nlm.nih.gov/). More details on HDOCK docking server and the procedures for the dockings are available at http://hdock.phys.hust.edu.cn/. Herein, the top-ranked poses (Top 1) for the (TrpT)n (n = 2,4,8…256) and curcumin/(TrpT)n complexes predicted by HDOCK according to the energy scores provided by the program were analysed and described.
In turn, the ADMETlab software (https://admet.scbdd.com/calcpre/calc_cf_single_mol/) was employed to predict the LogP values of the adduct TrpT-curcumin and curcumin alone, using for TrpT and TrpT-curcumin the following SMILES identifiers:
-
COc1cc(C=CC(=O)CC(=O)C=Cc2ccc(O)c(OC)c2)ccc1O (curcumin);
-
COc5cc(C=CC(=O)CC(=CC=c1ccc(=O)c(OC)c1)N(C(=O)Cn2cc(C)c(=O)[nH]c2=O)C(Cc4cc3ccccc3[nH]4)C(N)=O)ccc5O (TrpT-curcumin).
Circular dichroism (CD) and UV studies
CD and UV spectra were recorded on a JASCO J-815 CD spectropolarimeter (Jasco Inc., Easton, MD, USA). For these experiments, TrpT solutions were prepared in 25 mM sodium phosphate (pH 7.4) in the 5–140 µM concentration range. Spectra were acquired by averaging three scans in the 190–320 nm range with a 2 nm bandwidth and using a 100 nm/min scan rate. All the spectra were corrected for background by subtracting the proper blank.
CD and UV experiments were performed in triplicate: representative results were reported in Fig. 1 (for each technique the observed behaviour among the three repetitions was consistent within ± 2%).
Optical characterizations of TrpT at varying concentrations (from 5 to 140 µM) in sodium phosphate buffer (pH 7.4, 25 mM) at 25 °C. A Overlayed CD spectra of TrpT (recorded with a 2 nm bandwidth and using a 100 nm/min scan rate). Plots of the CD values recorded at λ = 214 nm (B) and λ = 271 nm (C) vs. TrpT concentration. D Selected UV spectra of TrpT. E Plot of UV absorbance (Abs) at 270 nm vs. TrpT concentration. F Selected fluorescence spectra of TrpT. G Plot of fluorescence intensity (F.I.) at 365 nm vs. TrpT concentration. The red lines in panels E and G represent the linear best-fitting of low concentration (≤ 20 µM) data points. Experiments were performed in triplicate. The observed behaviour among the three repetitions was consistent within ± 2%
Kinetic and temperature-dependent (heating–cooling) UV–vis measurements were carried out on a JASCO V-770 spectrophotometer using a quartz cell with a 10 mm optical path length (Hellma). TrpT was prepared in 25 mM sodium phosphate buffer (pH 7.4) at 50 and 140 µM concentrations and, immediately, the UV–vis spectra of each solution were recorded over time (0–24 h) with a scan rate of 100 nm/min, a 1 nm bandwidth and corrected for background by subtracting the spectrum of the buffer. After 24 h, in parallel experiments, each solution was subjected to heating/cooling cycles by monitoring the absorbance at 271 nm with a scan rate of 1 °C/min.
Fluorescence
Fluorescence experiments were performed on a spectrofluorometer (FluoroMax-4 Horiba Scientific) at 25, 50 and 80 °C. For these measurements, TrpT samples were prepared in 25 mM sodium phosphate (pH 7.4) in the 5–140 µM concentration range. All the fluorescence spectra were recorded in the 310–500 nm (λem) range upon excitation at 280 nm (λex) using 5 nm slit bandwidths for both excitation and emission and finally corrected for background fluorescence by subtracting the proper buffer blank. All the fluorescence experiments were performed in triplicate; representative experiments were reported in Fig. 1 (for each technique the observed behaviour among the three repetitions was consistent within ± 2%).
Dynamic light scattering (DLS)
DLS measurements were carried out on a Zetasizer Nano ZS (Malvern Instrument) by using 12 mm square polystyrene cuvettes (DTS0012, Malvern Instrument) according to previously optimized protocols [52, 53]. A TrpT solution was prepared at a 140 μM concentration in 25 mM sodium phosphate (pH 7.4) and then filtered using a 0.2 μm membrane filter (Merck Millipore). All the analyses were performed in triplicate at r.t. with a scattering angle of 173° and an equilibration time of 60 s. Samples were also monitored up to 5 h with the same instrument settings.
Scanning electron microscopy (SEM)
The morphological characterization of our nanosystems was carried out by SEM analysis through the collection of images using an FE-SEM Ultra Plus (Zeiss) microscope at 5 kV. For sample preparation, 20 µL (diluted in phosphate buffer pH 7.4, 25 mM) of TrpT solution (1 mM) were deposited on a thin glass slide, air-dried for 16 h at room temperature and then sputtered with a 10 nm thick gold layer. After these steps, different images of dried nanoassemblies were acquired.
Curcumin binding/release study
UV–visible spectroscopy was used to assess the entrapment of curcumin in TrpT assemblies. For this purpose, a curcumin solution (10 µM in 25 mM sodium phosphate buffer, pH 7.4) was prepared and then sonicated for several minutes. Then the solution was analysed using UV–Vis spectroscopy to observe a characteristic peak at 430 nm. Subsequently, a concentrated TrpT solution was gradually added to the curcumin solution and any changes in the intensity of the peak located at 430 nm were recorded after 10 min of incubation.
For confocal microscope imaging acquisition, a 250 µM curcumin solution (dissolved in Milli-Q water and sonicated for several minutes) was diluted twenty-five-fold into a TrpT solution (140 µM in 25 mM sodium phosphate buffer, pH 7.4), so to obtain a final 10 µM drug concentration, and then kept overnight at 25 °C. After dialysis (using a membrane pore cut-off = 12 kDa), to remove the non-entrapped drug molecules, 10 µL of this solution was spotted onto a glass slide and observed under a fluorescence microscope. In detail, the curcumin-TrpT nanoassemblies were detected with a confocal microscope (CLSM Leica SP5, Objective 63 × oil, scan speed of 400 Hz, excitation wavelength (λex) 458 nm, emission wavelength (λem) 478–520 nm) to evaluate the fluorescence emission of curcumin from the self-assemblies at zero time, which was fixed as 100%. The fluorescence intensity was then monitored over time to follow the curcumin release. The fluorescence values were normalised and subtracted from the background intensity. The results from two different experiments were averaged and shown in the figure (error bars associated with Fig. 7C histograms).
Human serum stability assay
1 µL of a 150 µM solution of TrpT was added to 99 µL of fresh human serum (Sigma Aldrich) and incubated at 37 °C. Samples withdrawn from the reaction mixture at intervals (0, 1, 2, 3, 4, 5, 24 and 168 h) were treated with 7 M urea at 95 °C for 2 min and subjected to HPLC analysis (detection at 260 nm) on a Hewlett Packard/Agilent 1100 series instrument, equipped with a diode array detector, using a Phenomenex Jupiter C18 300 Å column (5 μm, 4.6 × 250 mm).
Cell culture and cell-based assay
Human prostate cancer PC-3 cells were maintained in RPMI 1640 medium (GIBCO, USA) supplemented with 10% fetal bovine serum (GIBCO, USA), 2 mM l-glutamine (LONZA, Belgium) at 37 °C in a 5% CO2 humidified atmosphere.
Cell cytotoxicity was assessed by the crystal violet assay. Briefly, 8 × 104/ml cancer cells, after adhesion, were incubated with different concentrations of TrpT diluted with ethanol (Sigma-Aldrich) for 24 or 48 h. Subsequently, the cells were washed with Phosphate-Buffered Saline (PBS), fixed and stained with 0.1% (w/v) crystal violet in 25% methanol (Sigma-Aldrich); after 30 min, the cells were washed twice with double distilled water and let dry. Then, crystal violet was dissolved in a 10% acetic acid solution and absorbance was measured at 595 nm using a microplate reader.
For determining the effect of TrpT solutions on cell viability, the trypan-blue exclusion method was used. Cells, seeded at a density of 5 × 105/ml, were treated with TrpT (5–100 μM range) and after 24 and 48 h were counted in a Burker haemocytometer using a microscope (Leica, Germany). Cell viability was calculated as the ratio of live to dead cells, expressed as a percentage.
All data were expressed as mean ± SD of experiments performed in quadruplicate and unpaired.
CD binding studies
The CD binding assays with dA12, poly rA and BSA were registered with a JASCO J-715 spectropolarimeter equipped with a PTC-4235/15 Peltier thermostated cell holder, using a two-chamber quartz cell (Hellma, 238-QS, 2 × 0.4375 cm optical path length).
Sum and mixing CD spectra with nucleic acids were obtained with TrpT (16 nmol in T) and dA12 or poly rA (16 nmol in A each) in 25 mM sodium phosphate buffer (pH 7.5) at 10 °C.
Sum and mixing CD spectra with BSA were recorded using a BSA concentration of 1 µM (2 nmol) and a 1:10 BSA/TrpT molar ratio in the same conditions.
Results and discussion
Synthesis and characterization of TrpT
In analogy to the synthesis of other nucleoaminoacids [33, 54], the thyminyl l-tryptophanamide TrpT was obtained by a solid phase synthesis through the couplings on the resin of, first, the Fmoc/Boc protected amino acid l-tryptophan, and then − after removal of the Fmoc group with a piperidine treatment − of the thymine-1-acetic acid, using in both cases HATU/DIEA as condensing agents (Scheme 2). After cleavage from the solid support by a strong acid (95% TFA), precipitation, centrifugation, resuspension in H2O/CH3CN and lyophilisation, the TrpT product, as amidate (-CONH2), was obtained pure in 91% overall yield without need for chromatographic purification. Both the molecule identity and purity (> 98%) were confirmed by HPLC, LC–ESI–MS, NMR and MALDI-TOF (section 4.2 and supplementary Figures S2–S4). In particular, the mass spectrometry measurements showed molecular ions corresponding to the protonated form of TrpT and its adducts with either sodium and potassium ions (Figure S4).
Subsequently, a detailed spectroscopic analysis of the novel nucleoamino acid derivative was undertaken to assess its behaviour in pseudo-physiological solutions. All the performed experiments pointed to the self-assembling tendency of TrpT, in analogy to aromatic amino acids [55, 56] and aromatic di(oligo)peptides [9, 19], as described in the following sections.
CD, UV, and fluorescence analysis of TrpT solutions
CD measurements were performed exploring different TrpT concentrations (from 5 to 140 µM in phosphate buffer). At the lower concentration (5 µM), the spectrum showed a positive band at 195 nm with a small shoulder at 210 nm (red line, Fig. 1A). Some significant CD changes were detected upon increasing TrpT concentration. Specifically, the 195 nm band gradually decreased, finally disappearing at 30 µM with the concomitant emergence of a minimum at 205 nm and a wide band centred at about 220 nm. The latter band, going from 30 to 140 µM, increased in intensity with a significant shift towards shorter wavelengths (214 nm) and the appearance of a pronounced shoulder (228 nm). Simultaneously, a new, less-intense positive band (271 nm) emerged (Fig. 1A–C). These changes in the CD spectra indicated different structural organizations of TrpT, each favoured at different molecule concentrations. The appearance of new positive bands could be attributed to conformational changes following self-assembly processes and therefore to probable new morphologies.
The UV–visible and fluorescence spectra were also recorded exploring different concentrations of the nucleoamino acid derivative (Fig. 1D–G). The UV–vis spectra evidenced a wide band centred at 270 nm, which increased proportionally with the TrpT concentration (Fig. 1D). However, by plotting the UV absorption at 270 nm (Abs270) as a function of concentration, a linear behaviour in the range 5–40 µM and a negative deviation at higher concentrations (prior to signal saturation; Fig. 1E), were observed, which is a behaviour typical for aggregate formation [57, 58].
The fluorescence analysis of TrpT solutions showed concentration-dependent emission spectra following a 280 nm excitation, the characteristic λex for tryptophan (Fig. 1F). Moreover, the fluorescence emission peak resulted at a longer wavelength (λem∼365 nm) than the corresponding value usually observed for tryptophan in proteins or peptides (300–345 nm). Initially, the dependence between concentration and emission intensity was linear, but a non-linear trend was seen beyond 40 µM (Fig. 1G). These results confirmed the ability of TrpT molecules to mutually associate in a concentration-dependent manner, forming higher-order nanostructures, similarly to self-assembling aromatic-containing peptides [59]. Moreover, the tryptophan units in the formed supramolecular structures are likely solvent-exposed as hypothesised by virtue of: i) their red-shifted fluorescence emission, similar to that reported for denatured proteins or for the most red-shifted polypeptides such as glucagon and melittin [60] (λem∼352 and 346 nm, respectively), and ii) the constant λem of the fluorescence maximum upon increasing TrpT concentration, i.e. upon the growth of the self-assembling architecture. Indeed, superstructures in which tryptophan units are buried in a hydrophobic cavity often exhibit a marked blue-shifting of λem with intensity values higher than predicted [60, 61], which are effects opposite to those observed for TrpT.
In short, this spectroscopic analysis demonstrated that the novel amidate nucleoamino acid has the capability to self-assemble in solution leaving the tryptophan moieties exposed to the aqueous environment.
To further investigate the TrpT self-association, we performed i) UV-kinetic experiments at 50 µM and 140 µM, concentrations close to, respectively, the minimal expected value for the onset of nanoaggregation, and the maximum concentration tested; and ii) temperature-dependent UV–vis and fluorescence experiments.
The UV–vis spectra of the freshly prepared 50 and 140 µM solutions of TrpT were monitored over 24 h. A gradual increase of the absorbance of the 270 nm band, more evidently for the lower concentration (ΔAbs271 = + 22%, Fig. 2A) than for the more concentrated solution (ΔAbs271 = + 5%, Fig. 2B), was observed. For the two solutions a signal stabilization was observed after 6 h (50 µM) and after 4 h (140 µM).
UV-kinetic experiments. Overlayed UV spectra of TrpT measured at 50 µM (A) and 140 µM (B) in sodium phosphate buffer (NaPi, 25 mM, pH 7.4) at 60-min intervals. The scatter plots depict the time dependence of the same data: UV absorbance at λmax (271 nm) for TrpT at 50 µM (C) and 140 µM (D). Experiments were performed in triplicate. The observed behaviour among the three repetitions was consistent within ± 2%
After 24 h, the two TrpT solutions were subjected to a heating–cooling cycle (20 → 95 → 20 °C), whilst monitoring in parallel UV absorbance at λmax (271 nm) (Fig. 3). This treatment resulted in a decrease of Abs271 up to 50 °C (for 50 µM, Fig. 3A) and 65 °C (140 µM, Fig. 3B), followed by a subsequent absorbance increase up to 95 °C for both solutions. The decreases in Abs271 observed during the first stage of heating likely indicate a reorganization of the TrpT assembly with a concomitant increase in the π−π stacking interactions of the TrpT aromatic rings (Fig. 3E). Heating beyond the absorbance minimum of the Abs vs. T plot causes intermolecular binding, including stacking, to gradually disintegrate for both the systems (see Fig. 3E for the 140 μM system).
Absorbance at 271 nm vs. temperature during heating and subsequent cooling, measured at 50 µM (A, C) and 140 µM (B, D) in sodium phosphate buffer (NaPi, 25 mM, pH 7.4). E Hypothesised prevalence of various aggregate structures (TrpT 140 μM) and their likely kinetic and thermodynamically controlled formation/destruction during the heating–cooling cycle; a similar mechanism can be hypothesised also for the 50 μM concentration. Experiments were performed in triplicate. The observed behaviour among the three repetitions was consistent within ± 2%
After UV signal stabilization at 95 °C, both systems were cooled down to 20 °C, resulting in a progressive further increase in Abs271 following a sigmoidal behaviour with inflection points at about 60 °C in both cases (Fig. 3C, D). This trend suggests a difference in the pathway between aggregate disruption (during heating) of the kinetically formed superstructures and refolding, which occurs under thermodynamic control during cooling. The cooling process in both cases is accompanied by a gradual and cooperative reorganization of the system with disruption of self-stacking and probable concomitant formation of other binding modes, including H-bonding and NH-π interactions. Thus, during the thermodynamic self-aggregation stage, self-stacking interactions of the isolated molecules, dominant at high temperatures, become reduced in favour of an overall less self-stacked aggregated system (see Fig. 3E describing the hypothesised mechanism for the 140 μM concentration; a similar mechanism can be assumed also for the 50 μM concentration) [42]. In this regard, it is noteworthy that the six-membered aromatic Phe rings have been shown to interact almost perpendicularly with the indole moieties of tryptophan in proteins [62], perhaps explaining the less stacked nature of the nanoaggregated systems formed by TrpT at 20 °C.
Following the return to ambient conditions, the second heating cycle displayed almost the same behaviour as the first (Figure S5) but with somewhat different starting points of the absorbance values (especially for 50 µM). This similarity points to a significant degree of reversibility of the self-assembly. In other words, the supramolecular structures obtained under kinetic control at room temperature exhibit essentially the same form as those obtained via thermodynamic control, apart from a slightly greater extent of intermolecular stacking seen in post-cooled samples.
Fluorescence experiments on the TrpT solution were also performed at different temperatures. Most noticeably, heating was marked by a significant drop in fluorescence intensity (Fig. 4A). Such fluorescence decreases usually indicate a change in the chromophore environment towards lower hydrophobicity, in this case probably due to the disruption of supramolecular interactions [63].
A Fluorescence spectra of TrpT measured at 140 µM concentration in phosphate buffer (pH = 7.4, 25 mM) at 25, 50, and 80 °C during initial heating. B Overlay of two spectra recorded at 25 °C, before and after keeping the solution at 80 °C. Experiments were performed in triplicate. The observed behaviour among the three repetitions was consistent within ± 2%
Such a scenario is again consistent with the tryptophan units becoming progressively more exposed to the aqueous environment during the disruption of the TrpT nanostructures. Interestingly, the fluorescence spectrum of TrpT after heating the solution to 80 °C and subsequently cooling it to 25 °C differed from that initially recorded at the same temperature, consistent with our UV evidence for the non-reversibility of the denaturation (Fig. 4B). Collectively, our findings indicate that TrpT can form aggregates whose structures are affected by both temperature and concentration.
Morphological characterization of TrpT self-assembly: DLS and SEM analyses
The tendency of TrpT to form nanostructures in solution was further confirmed by DLS and SEM. In particular, DLS—performed on TrpT (140 µM in the selected buffer solution) at 25 °C—allowed the determination of the average particle size, homogeneity and stability over 5 h.
Immediately after dissolution, TrpT displayed a DLS profile relative to a nanoassembly with a single population of narrow distribution and a mean hydrodynamic diameter of ca. 330 nm (Fig. 5). Particle size homogeneity was further evidenced by the relatively low polydispersity index (PdI) values (Table 1).
The analysis of the sample in pseudo-physiological conditions over time suggested a good stability of the TrpT aggregates for up to 5 h (with no significant changes observed in the overall size and PdI values, Fig. 5 and Table 1).
A volume of 238.4 Å3 was estimated for a single TrpT molecule by WebLab ViewerPro 3.7 (Fig. S6A Molecular Simulations, Inc., San Diego-CA, 2000), equivalent to a spherical radius (r = [3 V/4π]1/3) of 3.85 Å. Considering the TrpT cluster radius of ~ 1500 Å (dH ~ 300 nm) suggested by our DLS data, this implies that many millions of molecules contribute to each self-assembled vesicle.
The morphological features of the molecular self-assemblies of TrpT were examined by SEM following a standard sample preparation involving drop-casting on glass slides (see the Experimental section). Our images clearly show that TrpT forms well-defined structures that are predominantly spherical in shape (light gray structures in Fig. 6A, B). This evidence corroborates our CD measurements, in which bands at 200 nm, (typically attributed to fibrillar/nanotubular morphologies [64, 65]) remained absent, even at high compound concentrations (Fig. 1A, B). The observed morphology is more likely due to the Trp − Trp interaction. The consequent exclusion of water around the Trp units is expected to facilitate nanoaggregate bending, leading to vesicle formation [66], in agreement with our variable temperature fluorescence data. This behaviour can also find support in the predicted lipophilicity of TrpT (single-molecule cLogP = 0.57 by SwissADME software, Figure S6B), corresponding to a slight partition preference for hydrophobic phases over polar media. This estimated cLogP value falls in the range typical of compounds characterised by good cell membrane permeability [67].
Furthermore, by measuring the diameter of 100 randomly selected particles on SEM images, an analysis of the size distribution was obtained (Figure S6C) and was found to be consistent with a Gaussian distribution (R2 = 0.9799), revealing the presence of a main population of TrpT-based nanostructures with a mean diameter of ~ 750 nm (950 ± 190 nm, PDI = 0.04), which is in line with vesicles from a single tryptophan-bearing small molecule assembly (average diameter of ~ 790 nm) [42, 66]. The found PDI value is indicative of a homogeneous population of structures.
Entrapment and release of curcumin by TrpT self-assembling nanostructure
One of the most appealing applications of nanostructures is their potential to transport bioactive molecules/drugs for therapeutic delivery, as required in the case of several marketed drugs delivered by vesicles [68]. To this end, we sought to monitor the binding and release capabilities of TrpT nanostructures by loading them with curcumin, used to model a hydrophobic drug. Despite some potential therapeutic applications being somewhat compromised by its poor bioavailability, and rapid metabolism [69, 70], curcumin has been shown to exhibit a range of anti-inflammatory, antibacterial, anticancer and anti-Alzheimer's properties [71].
Curcumin loading by TrpT aggregates was investigated by UV–visible spectrophotometric titrations. A progressive intensity decrease of the curcumin 426 nm (n → π* transition) band with increasing the concentration of TrpT added to a fixed concentration curcumin solution (Fig. 7A), was observed. This behaviour suggested that interactions of various nature were established between the drug and the nanoaggregates as already reported for similar systems [72].
A Visible absorption spectra of curcumin (10 µM) in the presence of varying TrpT concentrations. B Confocal microscopy image of curcumin molecule entrapped in self-assembled TrpT nanostructures. C Normalised fluorescence intensity (λem = 478–520 nm) of curcumin (10 µM)—TrpT (140 µM) assemblies measured by confocal fluorescence microscope. The reported fluorescence intensities were averaged from two independent experiments and reported as mean values ± SD (error bars). The p-value was calculated using the unpaired two-tailed Student’s t-test to determine whether the differences between the reference sample at time zero and the sample recorded over time were significant (*p < 0.1; **p < 0.05; and NS, not significant)
Confocal microscopy imaging studies of curcumin-treated TrpT self-assemblies confirmed the presence of the fluorescent drug bound to the nucleoamino acid-based vesicles (Fig. 7B), indicating that curcumin is loaded throughout the nanostructures probably by hydrophobic/aromatic stacking interactions. The fluorescence emission intensity of selected particles was found to gradually diminish with a 90% reduction after 1 h, whilst the shape of the vesicle remained unchanged (Fig. 7C). Therefore, no loss of the self-assembly structure morphology of the selected particles was detected during this monitoring period, consistent with good stability of the TrpT assemblies. Our findings suggest that curcumin-loaded TrpT complexes could have potential in anti-inflammatory and antitumor therapy in analogy to other similar systems [71].
These experiments pointed out the good efficiency of curcumin entrapment in the self-assembling TrpT nanosystem. Indeed, in our experimental conditions, almost all the curcumin, at a 10 µM concentration, was efficiently entrapped by the 140 μM solution of TrpT, as evidenced by fluorescence measurements of the solution dialyzed for the confocal microscopy experiment (fluorescence was negligible), and also by the UV-titration experiments in which a continuum decrease of the absorbance of the curcumin was observed still at 140 μM concentration of the nucleopeptide.
Seeking an additional computational description of TrpT self-assembly, molecular docking studies were performed using HDOCK software. Iteratively docking TrpT units (self-docking; (TrpT)n n = 2, 4, 8…256) afforded a model of supramolecular aggregation (Fig. 8A) driven mainly by thymine-thymine π−π stacking (Figure S9). Introducing curcumin into an n = 256 scenario, our model predicted the incorporation of this ligand throughout the interior of the supramolecular assembly, consistent with our previously formulated hypothesis. Interestingly, the docking study illustrated the drug-vesicle binding to be mainly hydrophobic in nature; specific interactions of note included a π−π (T-shaped) contact and two π-alkyl interactions, involving one curcumin phenol moiety and the indole ring of TrpT, as well as one curcumin methyl and the TrpT indole moiety, respectively (Fig. 8B).
Furthermore, the ADMETlab software allowed us to predict the LogP of the adduct between curcumin and TrpT. Considering that a LogP < 0 corresponds generally to poor lipid bilayer permeability, whereas a LogP > 3 is associated with poor aqueous solubility, the optimal range is 0 < LogP < 3 (https://admet.scbdd.com/home/interpretation/#part6). A LogP value of 3.1 for the TrpT-curcumin adduct with respect to the predicted value of free curcumin (LogP = 3.4) was found, suggesting that the bioavailability of curcumin could be improved by TrpT. Indeed, TrpT makes the polarity and, thus, the LogP value of TrpT-curcumin closer to the optimal range expected for good drug bioavailability.
TrpT is highly stable in human serum and does not show toxicity to human cells
To establish the feasibility of its potential applications to nanomaterials and drug delivery systems, the stability of TrpT in human serum and its effects on human cells using PC-3 as a model cell line were evaluated.
The enzymatic stability of TrpT was evaluated by incubating a sample of the compound (150 µM) in 99% fresh human serum at T = 37 °C and through HPLC analysis of the samples withdrawn from the mixture at different intervals up to 168 h. As deducible from Fig. 9A, the covalent/molecular structure of TrpT is highly stable, remaining essentially intact even after about 170 h incubation.
A Overlayed representative RP-HPLC chromatograms; HPLC peaks for samples withdrawn at 0 and 168 h from incubation of TrpT in 99% human serum at T = 37 °C. Chromatography was performed on an analytical C-18 column employing a linear gradient of 2% (5 min) up to 80% of solution B′ in A′ over 20 min (flow rate of 1 mL/min; solution A′ = 0.1% TFA in water; solution B′ = 0.1% TFA in acetonitrile. Detection was at 260 nm). B Cytotoxicity assay featuring TrpT against PC-3 cells. PC-3 were treated with different concentrations of TrpT, then the medium was removed, and crystal violet assay was performed. Cells treated with ethanol (0.01%) were included as a control. The relative number of viable cells was expressed as optical density (O.D.). Each value is the mean ± SD of two experiments performed in quadruplicate. C Images relative to the morphology of PC-3 cells treated for 24 h with ethanol 0.01% (left) and 50 μM TrpT (right). Optical microscope Axivert 200 M Zeiss; 400×; bright field
Finally, PC-3 prostate cancer cells were treated with TrpT solutions (5–100 µM range). From our experiments, TrpT was non-cytotoxic across the concentration range explored (Figs. 9B, C and S10B).
In addition, the cell viability was assayed in all samples also with the trypan blue dye exclusion method and it resulted higher than 90% (Figure S10A).
Investigation of the TrpT interaction with nucleic acids
To further confirm the non-toxicity of TrpT in view of potential drug delivery applications, its possible off-target interaction with nucleic acids was explored. In particular, the capability of TrpT to bind homoadenylic oligonucleotide sequences, specifically a dA12 DNA oligonucleotide and poly(A) RNA, was evaluated. To this end, CD binding experiments were carried out between TrpT and nucleic acids using a dual-chamber cell (inset of Figure S7). The reservoirs of this cuvette were filled with two solutions, the DNA or RNA on one side and the nucleoamino acid derivative on the other, set up with TrpT/DNA(RNA) in a T:A ratio of 1:1. No significant differences were observed between the CD spectra corresponding to the two separated components, for either TrpT/DNA or TrpT/RNA mixtures (Figure S7). This finding reveals a lack of interaction with either homoadenylic DNA or RNA sequences under our experimental conditions and renders unlikely any side effects (toxicity) due to such aspecific and undesirable nucleic acid binding.
Investigation of TrpT interaction with bovine serum albumin (BSA)
To further prove the potential of TrpT as a drug delivery system, its possible interaction with BSA, a model of a serum protein able to transport many classes of ligands, including amino acids present in the bloodstream to their target organs [73], was explored. Interestingly, this interaction has also driven the pharmaceutical use of serum albumins as drug carriers as they can improve the bioavailability of their cargoes [74].
Thus, CD binding experiments were carried out between TrpT and BSA in a dual-chamber quartz cell similarly to the experiments with nucleic acids described in the previous subsection. The reservoirs of the cuvette were filled with the solution of BSA, on one side, and the nucleoamino acid derivative, on the other. Subsequently, the ‘sum’ spectrum corresponding to the two separated solutions as well as the spectrum obtained after mixing them were recorded. The evident differences observed between the two spectra proved the interaction between BSA and TrpT (Figure S8). This preliminary finding is indicative of the potential role of serum albumins in the transportation of TrpT in the bloodstream and in sustaining the bioavailability of TrpT-drug nanosystems for biomedical applications.
Conclusions
TrpT, a novel nucleobase-containing α-amino acid derivative, obtained in the nucleoamino amide form, was synthesised in the solid phase, and characterised by NMR and MS. The analyses of the behaviour in a pseudo-physiological solution of this hybrid compound—performed by CD, UV and fluorescence spectroscopies at different sample concentrations and variable temperatures—demonstrated its ability to self-assemble into nanoaggregates which likely expose the tryptophan moieties of the outer layers to the aqueous environment, while creating hydrophobic pockets that include other tryptophan units within the particle interior. The capability of TrpT to form nanostructures in solution was also confirmed by DLS and SEM analyses, which demonstrated the formation of species that are largely spherical in shape and showed good stability up to 5 h (with a hydrodynamic diameter of about 330 nm at 140 µM TrpT and narrow distribution and homogenous size). Remarkably, the confocal fluorescence microscopy analysis showed that the supramolecular TrpT network can host the hydrophobic drug-like molecule curcumin and release it over time, suggesting its potential as a drug delivery system. Molecular docking indicates predominantly hydrophobic binding, specifically π−π (T-shaped) and π-alkyl bonding, involving one curcumin phenol moiety and the TrpT indole ring, as well as one ligand methyl and the nucleoamino amide indole moiety, respectively. Moreover, a predicted logP value of the TrpT-curcumin adduct closer to the optimal range expected for good drug lipophilicity with respect to curcumin alone, suggested an improved bioavailability of curcumin combined with the TrpT nanosystem.
The potential drug delivery application of the herein-described molecular systems was also supported by the high stability of TrpT in human serum, its non-cytotoxicity towards human cells, and its inability to perturb the secondary structure of adenine-containing DNA and RNA that excludes possible side effects due to the undesired, possible interactions with nucleic acids. Finally, TrpT interacted with a serum albumin-like BSA, which is known to improve the bloodstream transportation and bioavailability of its cargo.
Data availability
All data obtained or analysed in the frame of this work are displayed in the published article and its supporting information file.
References
Van der Laan, A.C., Van Amsterdam, I., Tesser, G.I., Van Boom, J.H., Kuyl-Yeheskiely, E.: Synthesis of chirally pure ornithine based PNA analogues. Nucleosides Nucleotides 17, 219–231 (1998)
Roviello, G.N., Musumeci, D., Pedone, C., Bucci, E.M.: Synthesis, characterization and hybridization studies of an alternate nucleo-ε/γ-peptide: complexes formation with natural nucleic acids. Amino Acids 38, 103–111 (2010)
Roviello, G.N., Musumeci, D., Castiglione, M., Bucci, E.M., Pedone, C., Benedetti, E.: Solid phase synthesis and RNA-binding studies of a serum-resistant nucleo-ε-peptide. J. Pept. Sci. 15, 155–160 (2009)
Musumeci, D., Mokhir, A., Roviello, G.N.: Synthesis and nucleic acid binding evaluation of a thyminyl l-diaminobutanoic acid-based nucleopeptide. Bioorg. Chem. 100, 103862 (2020). https://doi.org/10.1016/j.bioorg.2020.103862
Contini, A., Erba, E., Bondavalli, V., Barbiroli, A., Gelmi, M.L., Romanelli, A.: Morpholino-based peptide oligomers: synthesis and DNA binding properties. Biochem. Biophys. Res. Commun. 549, 8–13 (2021)
Giraud, T., Hoschtettler, P., Pickaert, G., Averlant-Petit, M.C., Stefan, L.: Emerging low-molecular weight nucleopeptide-based hydrogels: state of the art, applications, challenges and perspectives. Nanoscale 14, 4908–4921 (2022)
Scognamiglio, P.L., Platella, C., Napolitano, E., Musumeci, D., Roviello, G.N.: From prebiotic chemistry to supramolecular biomedical materials: exploring the properties of self-assembling nucleobase-containing peptides. Molecules. 26(12), 3558 (2021). https://doi.org/10.3390/molecules26123558
Wang, H., Yang, Z.: Short-peptide-based molecular hydrogels: novel gelation strategies and applications for tissue engineering and drug delivery. Nanoscale 4, 5259–5267 (2012)
Guo, C., Luo, Y., Zhou, R., Wei, G.: Triphenylalanine peptides self-assemble into nanospheres and nanorods that are different from the nanovesicles and nanotubes formed by diphenylalanine peptides. Nanoscale 6, 2800–2811 (2014)
Bhandaru, N., Kaur, G., Panjla, A., Verma, S.: Spin coating mediated morphology modulation in self assembly of peptides. Nanoscale 13, 8884–8892 (2021)
Gray, V.P., Amelung, C.D., Duti, I.J., Laudermilch, E.G., Letteri, R.A., Lampe, K.J.: Biomaterials via peptide assembly: design, characterization, and application in tissue engineering. Acta Biomater. 140, 43–75 (2022)
D’Errico, S., Oliviero, G., Borbone, N., Amato, J., D’Alonzo, D., Piccialli, V., Mayol, L., Piccialli, G.: A facile synthesis of 5′-Fluoro-5′-deoxyacadesine (5′-F-AICAR): a novel non-phosphorylable AICAR Analogue. Molecules 17, 13036–13044 (2012)
Musumeci, D., Irace, C., Santamaria, R., Montesarchio, D.: Trifluoromethyl derivatives of canonical nucleosides: synthesis and bioactivity studies. MedChemComm 4, 1405–1410 (2013)
D’Errico, S., Oliviero, G., Borbone, N., Di Gennaro, E., Zotti, A.I., Budillon, A., Cerullo, V., Nici, F., Mayol, L., Piccialli, V., Piccialli, G.: Synthesis and evaluation of the antiproliferative properties of a tethered tubercidin-platinum(II) complex. Eur. J. Org. Chem. 2015, 7550–7556 (2015)
D’Errico, S., Borbone, N., Piccialli, V., Di Gennaro, E., Zotti, A., Budillon, A., Vitagliano, C., Piccialli, I., Oliviero, G.: Synthesis and evaluation of the antitumor properties of a small collection of PtII complexes with 7-deazaadenosine as scaffold. Eur. J. Org. Chem. 17, 4935–4947 (2017)
D’Errico, S., Borbone, N., Catalanotti, B., Secondo, A., Petrozziello, T., Piccialli, I., Pannaccione, A., Costantino, V., Mayol, L., Piccialli, G., Oliviero, G.: Synthesis and biological evaluation of a new structural simplified analogue of cADPR, a calcium-mobilizing secondary messenger firstly isolated from sea urchin eggs. Mar. Drugs 16, 89 (2018)
Watanabe, S., Tomizaki, K.Y., Takahashi, T., Usui, K., Kajikawa, K., Mihara, H.: Interactions between peptides containing nucleobase amino acids and T7 phages displaying S. cerevisiae proteins. Biopolym. Pept. Sci. Sect. 88, 131–140 (2007)
Uozumi, R., Takahashi, T., Yamazaki, T., Granholm, V., Mihara, H.: Design and conformational analysis of natively folded β-hairpin peptides stabilized by nucleobase interactions. Biopolymers 94, 830–842 (2010)
Datta, D., Tiwari, O., Ganesh, K.N.: New archetypes in self-assembled Phe-Phe motif induced nanostructures from nucleoside conjugated-diphenylalanines. Nanoscale 10, 3212–3224 (2018)
Avitabile, C., Diaferia, C., Roviello, V., Altamura, D., Giannini, C., Vitagliano, L., Accardo, A., Romanelli, A.: Fluorescence and morphology of self-assembled nucleobases and theirdiphenylalanine hybrid aggregates. Chem. A Eur. J. 25, 14850–14857 (2019)
Tiwari, O.S.: Synthesis, characterization, and investigation of capacitance and redox properties of self-assembled Phe-Phe with ferrocene conjugates. Mol. Syst. Des. Eng. 7, 171–181 (2022)
Avitabile, C., Diaferia, C., Della Ventura, B., Mercurio, F.A., Leone, M., Roviello, V., Saviano, M., Velotta, R., Morelli, G., Accardo, A., Romanelli, A.: Self-assembling of Fmoc-GC peptide nucleic acid dimers into highly fluorescent aggregates. Chem. A Eur. J. 24, 4729–4735 (2018)
Giraud, T., Bouguet-Bonnet, S., Marchal, P., Pickaert, G., Averlant-Petit, M.C., Stefan, L.: Improving and fine-tuning the properties of peptide-based hydrogels: via incorporation of peptide nucleic acids. Nanoscale 12, 19905–19917 (2020)
Filippov, D., Kuyl-Yeheskiely, E., Van Der Marel, G.A., Tesser, G.I., Van Boom, J.H.: Synthesis of a nucleopeptide fragment from poliovirus genome. Tetrahedron Lett. 39, 3597–3600 (1998)
Geotti-Bianchini, P., Crisma, M., Peggion, C., Bianco, A., Formaggio, F.: Conformationally controlled, thymine-based α-nucleopeptides. Chem. Commun. 2009, 3178–3180 (2009)
Geotti-Bianchini, P., Moretto, A., Peggion, C., Beyrath, J., Bianco, A., Formaggio, F.: Replacement of Ala by Aib improves structuration and biological stability in thymine-based α-nucleopeptides. Org. Biomol. Chem. 8, 1315–1321 (2010)
Kramer, R.A., Bleicher, K.H., Wennemers, H.: Design and synthesis of nucleoproline amino acids for the straightforward preparation of chiral and conformationally constrained nucleopeptides. Helv. Chim. Acta 95, 2621–2634 (2012)
Simeone, L., Irace, C., Di Pascale, A., Ciccarelli, D., D’Errico, G., Montesarchio, D.: Synthesis, self-aggregation and bioactivity properties of a cationic aminoacyl surfactant, based on a new class of highly functionalized nucleolipids. Eur. J. Med. Chem. 57, 429–440 (2012)
Noel, O., Xie, J.: Synthesis of nucleo aminooxy acid derivatives. Synthesis 45, 134–140 (2013)
Pomplun, S., Gates, Z.P., Zhang, G., Quartararo, A.J., Pentelute, B.L.: Discovery of nucleic acid binding molecules from combinatorial biohybrid nucleobase peptide libraries. J. Am. Chem. Soc. 142, 19642–19651 (2020)
Walsh, C.T., Zhang, W.: Chemical logic and enzymatic machinery for biological assembly of peptidyl nucleoside antibiotics. ACS Chem. Biol. 6, 1000–1007 (2011)
Roviello, G.N., Mottola, A., Musumeci, D., Bucci, E.M., Pedone, C.: Synthesis and aggregation properties of a novel enzymatically resistant nucleoamino acid. Amino Acids 43, 1465–1470 (2012)
Roviello, G.N.: Novel insights into nucleoamino acids: biomolecular recognition and aggregation studies of a thymine-conjugated l-phenyl alanine. Amino Acids 50, 933–941 (2018)
Roviello, G.N., Ricci, A., Bucci, E.M., Pedone, C.: Synthesis, biological evaluation and supramolecular assembly of novel analogues of peptidyl nucleosides. Mol. Biosyst. 7, 1773–1778 (2011)
Roviello, G.N., Musumeci, D., Bucci, E.M., Pedone, C.: Evidences for supramolecular organization of nucleopeptides: synthesis, spectroscopic and biological studies of a novel dithymine l-serine tetrapeptide. Mol. Biosyst. 7, 1073–1080 (2011)
Roviello, G.N., Oliviero, G., Di Napoli, A., Borbone, N., Piccialli, G.: Synthesis, self-assembly-behavior and biomolecular recognition properties of thyminyl dipeptides. Arab. J. Chem. 13, 1966–1974 (2020)
Babar, D.G., Sarkar, S.: Self-assembled nanotubes from single fluorescent amino acid. Appl. Nanosci. 7, 101–107 (2017)
Ishida, T., Tokura, Y., Shimamoto, M., Doi, M., Inoue, M.: Crystal structure of (uracil-1-ylethyl)(adenin-9-ylethyl)tryptophan dipeptide: an interaction model between nucleic acid base and aromatic amino acid. Chem. Pharm. Bull. 35, 1691–1701 (1987)
Haridas, V., Kumar, P.P.P., Bhardwaj, I., Venugopalan, P.: Spatially placed tryptophan residues: a strategy for generating molecules with unique self-assembly and molecular recognition properties. Chem. Sel. 2, 130–135 (2017)
Dube, T., Mandal, S., Panda, J.J.: Nanoparticles generated from a tryptophan derivative: physical characterization and anti-cancer drug delivery. Amino Acids 49, 975–993 (2017)
Itahara, T.: Stacking conformation of 9-[ω-(thymin-1-yl)alkyl]adenine in aqueous solution. Nucleosides Nucleotides Nucleic Acids 22, 309–317 (2003)
Mutai, K., Gruber, B.A., Leonard, N.J.: Synthetic spectroscopic models. XIV. Intramolecular stacking interactions between indole and connected nucleic acid bases. Hypochromism and fluorescence. J. Am. Chem. Soc. 97, 4095–4104 (1975)
Riccardi, C., Musumeci, D., Capuozzo, A., Irace, C., King, S., Russo Krauss, I., Paduano, L., Montesarchio, D.: “Dressing up” an old drug: an aminoacyl lipid for the functionalization of Ru(III)-based anticancer agents. ACS Biomater. Sci. Eng. 4, 163–174 (2018)
Riccardi, C., Capasso, D., Coppola, A., Platella, C., Montesarchio, D., Di Gaetano, S., Roviello, G.N., Musumeci, D.: Synthesis, antiproliferative activity and DNA binding studies of nucleoamino acid-containing Pt(II) complexes. Pharmaceuticals 13, 284 (2020)
Yan, Y., Zhang, D., Zhou, P., Li, B., Huang, S.Y.: HDOCK: a web server for protein-protein and protein-DNA/RNA docking based on a hybrid strategy. Nucleic Acids Res. 45, W365–W373 (2017)
Yan, Y., Tao, H., He, J., Huang, S.Y.: The HDOCK server for integrated protein–protein docking. Nat. Protoc. 15, 1829–1852 (2020)
Majumder, A., Kanti Mondal, S., Mukhoty, S., Bag, S., Mondal, A., Begum, Y., Sharma, K., Banik, A.: Virtual screening and docking analysis of novel ligands for selective enhancement of tea (Camellia sinensis) flavonoids. Food Chem. X. 13, 100212 (2022)
Roy, A., Chatterjee, O., Banerjee, N., Roychowdhury, T., Dhar, G., Mukherjee, G., Chatterjee, S.: Curcumin arrests G-quadruplex in the nuclear hyper-sensitive III1 element of c-MYC oncogene leading to apoptosis in metastatic breast cancer cells. J. Biomol. Struct. Dyn. 40, 10203–10219 (2021)
Ji, D., Juhas, M., Tsang, C.M., Kwok, C.K., Li, Y., Zhang, Y.: Discovery of G-quadruplex-forming sequences in SARS-CoV-2. Brief. Bioinform. 22, 1150–1160 (2021)
Stoddard, S.V., Wallace, F.E., Stoddard, S.D., Cheng, Q., Acosta, D., Barzani, S., Bobay, M., Briant, J., Cisneros, C., Feinstein, S., Glasper, K., Hussain, M., Lidoski, A., Lingareddy, P., Lovett, G., Matherne, L., McIntosh, J., Moosani, N., Nagge, L., Nyamkondiwa, K., Pratt, I., Root, E., Rutledge, M.R., Sawyer, M., Singh, Y., Smith, K., Tanveer, U., Vaghela, S.: In silico design of peptide-based SARS-CoV-2 fusion pnhibitors that target WT and mutant versions of SARS-CoV-2 HR1 domains. Biophysica 1, 311–327 (2021)
Pawar, S.S., Rohane, S.H.: Review on discovery studio: an important tool for molecular docking. Asian J. Res. Chem. 14, 86–88 (2021)
Riccardi, C., Fàbrega, C., Grijalvo, S., Vitiello, G., D’Errico, G., Eritja, R., Montesarchio, D.: AS1411-decorated niosomes as effective nanocarriers for Ru(III)-based drugs in anticancer strategies. J. Mater. Chem. B. 6, 5368–5384 (2018)
Riccardi, C., Piccolo, M., Ferraro, M.G., Graziano, R., Musumeci, D., Trifuoggi, M., Irace, C., Montesarchio, D.: Bioengineered lipophilic Ru (III) complexes as potential anticancer agents. Biomater. Adv. 139, 213016 (2022)
Roviello, G.N., Roviello, V., Autiero, I., Saviano, M.: Solid phase synthesis of TyrT, a thymine-tyrosine conjugate with poly(A) RNA-binding ability. RSC Adv. 6, 27607–27613 (2016)
Uyaver, S., Hernandez, H.W., Gokhan Habiboglu, M.: Self-assembly of aromatic amino acids: a molecular dynamics study. Phys. Chem. Chem. Phys. 20, 30525–30536 (2018)
Bera, S., Xue, B., Rehak, P., Jacoby, G., Ji, W., Shimon, L.J.W., Beck, R., Král, P., Cao, Y., Gazit, E.: Self-assembly of aromatic amino acid enantiomers into supramolecular materials of high rigidity. ACS Nano 14, 1694–1706 (2020)
Musumeci, D., Roviello, G.N., Rigione, G., Capasso, D., Di Gaetano, S., Riccardi, C., Roviello, V., Montesarchio, D.: Benzodifuran derivatives as potential antiproliferative agents: possible correlation between their bioactivity and aggregation properties. ChemPlusChem 82, 251–260 (2017)
Vicidomini, C., Cioffi, F., Broersen, K., Roviello, V., Riccardi, C., Montesarchio, D., Capasso, D., Gaetano, S.D., Roviello, G.N.: Benzodifurans for biomedical applications: BZ4, a selective anti-proliferative and anti-amyloid lead compound. Future Med. Chem. 11, 285–302 (2019)
Martin, A.D., Wojciechowski, J.P., Robinson, A.B., Heu, C., Garvey, C.J., Ratcliffe, J., Waddington, L.J., Gardiner, J., Thordarson, P.: Controlling self-assembly of diphenylalanine peptides at high pH using heterocyclic capping groups. Sci. Rep. 7, 43947 (2017)
Ghisaidoobe, A.B.T., Chung, S.J.: Intrinsic tryptophan fluorescence in the detection and analysis of proteins: a focus on förster resonance energy transfer techniques. Int. J. Mol. Sci. 15, 22518–22538 (2014)
Vivian, J.T., Callis, P.R.: Mechanisms of tryptophan fluorescence shifts in proteins. Biophys. J. 80, 2093–2109 (2001)
Samanta, U., Pal, D., Chakrabarti, P.: Packing of aromatic rings against tryptophan residues in proteins. Acta Crystallogr. Sect. D Biol. Crystallogr. 55, 1421–1427 (1999)
Stepanenko, O.V., Stepanenko, O.V., Kuznetsova, I.M., Verkhusha, V.V., Turoverov, K.K.: Beta-barrel scaffold of fluorescent proteins. Int. Rev. Cell Mol. Biol. 302, 221–278 (2013)
Mishra, A., Chauhan, V.S.: Probing the role of aromaticity in the design of dipeptide based nanostructures. Nanoscale 3, 945–949 (2011)
Adler-Abramovich, L., Reches, M., Sedman, V.L., Allen, S., Tendler, S.J.B., Gazit, E.: Thermal and chemical stability of diphenylalanine peptide nanotubes: implications for nanotechnological applications. Langmuir 22, 1313–1320 (2006)
Haridas, V.: Tailoring of peptide vesicles: a bottom-up chemical approach. Acc. Chem. Res. 54, 1934–1949 (2021)
Benet, L.Z., Hosey, C.M., Ursu, O., Oprea, T.I.: BDDCS, the rule of 5 and drugability. Adv. Drug Deliv. Rev. 101, 89–98 (2016)
Halwani, A.A.: Development of pharmaceutical nanomedicines: from the bench to the market. Pharmaceutics 14, 106 (2022)
Yallapu, M.M., Nagesh, P.K.B., Jaggi, M., Chauhan, S.C.: Therapeutic applications of curcumin nanoformulations. AAPS J. 17, 1341–1356 (2015)
Riccardi, C., Napolitano, F., Montesarchio, D., Sampaolo, S., Melone, M.A.B.: Nanoparticle-guided brain drug delivery: expanding the therapeutic approach to neurodegenerative diseases. Pharmaceutics 13, 1897 (2021)
Ðoković, J.B., Savić, S.M., Mitrović, J.R., Nikolic, I., Marković, B.D., Randjelović, D.V., Antic-Stankovic, J., Božić, D., Cekić, N.D., Stevanović, V., Batinić, B., Arandelović, J., Savić, M.M., Savić, S.D.: Curcumin loaded pegylated nanoemulsions designed for maintained antioxidant effects and improved bioavailability: a pilot study on rats. Int. J. Mol. Sci. 22, 7991 (2021)
Deka, S.R., Yadav, S., Kumar, D., Garg, S., Mahato, M., Sharma, A.K.: Self-assembled dehydropeptide nano carriers for delivery of ornidazole and curcumin. Coll. Surf. B Biointerfaces 155, 332–340 (2017)
Szymaszek, P., Fiedor, P., Chachaj-Brekiesz, A., Tyszka-Czochara, M., Świergosz, T., Ortyl, J.: Molecular interactions of bovine serum albumin (BSA) with pyridine derivatives as candidates for non-covalent protein probes: a spectroscopic investigation. J. Mol. Liq. 347, 118262 (2022)
Spada, A., Emami, J., Tuszynski, J.A., Lavasanifar, A.: The uniqueness of albumin as a carrier in nanodrug delivery. Mol. Pharm. 18, 1862–1894 (2021)
Acknowledgements
We thank Mr. Leopoldo Zona and Dr. Antonella Gargiulo for their invaluable technical assistance. C.R. was supported by Fondazione Umberto Veronesi.
Funding
Open access funding provided by Consiglio Nazionale Delle Ricerche (CNR) within the CRUI-CARE Agreement.
Author information
Authors and Affiliations
Contributions
All authors contributed to the conceptualization, literature collection, experimental analysis, sample analysis, writing, data analysis, and editing and reviewing of the article. All authors have read and agreed to the published version of the manuscript.
Corresponding authors
Ethics declarations
Conflict of interest
The authors have no conflict of interests to declare.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Scognamiglio, P.L., Riccardi, C., Palumbo, R. et al. Self-assembly of thyminyl l-tryptophanamide (TrpT) building blocks for the potential development of drug delivery nanosystems. J Nanostruct Chem (2023). https://doi.org/10.1007/s40097-023-00523-7
Received:
Accepted:
Published:
DOI: https://doi.org/10.1007/s40097-023-00523-7