Abstract
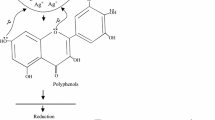
Nanobiotechnology has been emerging as an interdisciplinary act which converges materials and living organisms at nanoscale and proved to be one of the potential tools in nanotechnology to address some of the critical problems. Production of biogenic metallic nanoparticles using microorganisms and other living organisms including plants is been an attracting research activity. Herein, we report the synthesis of silver nanoparticles (AgNPs) using the bark extract of Alstonia scholaris, one of the most important medicinal plants and their promising antimicrobial activity. Stable AgNPs were formed by treating 10 % of A. scholaris bark extract with the aqueous solution of AgNO3 (1 mM). The formation of AgNPs was confirmed by UV–visible spectroscopic analysis and recorded the localized surface plasmon resonance of AgNPs at 432 nm. Fourier transform infrared spectroscopic analysis revealed that primary and secondary amine groups in combination with the proteins present in the bark extract are responsible for the reduction and stabilization of the AgNPs. X-ray diffraction micrograph indicated the face-centered cubic structure of the formed AgNPs, and morphological studies including size (average size 50 nm) were carried out using transmission electron microscopy. The hydrodynamic diameter (111.7 nm) and zeta potential (−18.9 mV) were measured using the dynamic light scattering technique. The antimicrobial activity of A. scholaris bark-extract-mediated AgNPs was evaluated (in vitro) against fungi, Gram-negative and Gram-positive bacteria using disc diffusion method.
Similar content being viewed by others
Explore related subjects
Discover the latest articles, news and stories from top researchers in related subjects.Avoid common mistakes on your manuscript.
Background
Nanoscience poses a basic scientific challenge as it requires a control over the size and shape of the aggregated atoms at nanoscale (1–100 nm). The atomic or molecular aggregates at nanoscale (at least in one dimension) are called as nanoparticles (NPs) whose properties are entirely different from their bulk counterparts. Production of nanoparticles can be achieved through different methods, for example, reduction in solutions, chemical and photochemical reactions in reverse micelles, thermal decomposition of silver compounds [1], radiation-assisted [2], electrochemical [3], sonochemical [4] and microwave-assisted methods [5]. Though there are well-established methods of synthesis of nanoparticles using chemical and physical methods, but are having inherent limitations regarding the usage of toxic chemicals, energy consumption and their biological applications. Therefore, the only left out option, i.e., using biological materials for the synthesis of metallic nanoparticles appeared as the most efficient and greener approach [6]. The present decade has witnessed the rapid shift in synthesis strategies from physicochemical methods to biological agents such as bacteria [7], fungi [8] and plants [9, 10] for the synthesis of nanoparticles. Using plants and plant material for the synthesis of metallic nanoparticles is cost effective, environmental friendly and up scalable method for the large-scale synthesis. Biosynthesis of silver nanoparticles by plants including, Desmodium triflorum [11], Cinnamomum camphora [12], Moringa oleifera [13], Eucalyptus hybrid [14], Memecylon umbellatum [15], Citrus reticulate [16] and Dioscorea batatas [17], has been reported in the literature recently.
The emergence and spread of antibiotic resistance microorganisms triggered the search of new materials through diverse sources including investigations on plants. Higher plants can serve as both potential antimicrobial crude drugs and a source of new anti-infective agents. Like many other medicinal plants, Alstonia scholaris (L.) R.Br. (Apocynaceae) is an evergreen tropical tree native to Indian subcontinent and South East Asia having grayish rough bark and milky sap rich in poisonous alkaloid. The bark also called dita bark is traditionally used by many ethnic groups of Northeast India and also other parts of the world as an antimicrobial agent against fungal infections, malarial fever, toothache, rheumatism, snake bite, dysentery, bowl disorder, etc., and the latex is used in treating coughs, through sores and fever [18, 19]. Among the several genera of Alstonia, only scholaris species has been studied for antimicrobial potency [20]. Silver has long been recognized as having an inhibitory effect toward many bacterial strains and microorganisms commonly present in medical and industrial processes [21]. Many attempts have been made to use silver nanoparticles as an anticancer agent, and they have all turned up positive [22, 23]. The role of silver nanoparticles as an anticancer agent should open new doors in the field of medicine. However, hardly reports are available on antimicrobial activity of silver nanoparticles synthesized using the aqueous extract of this plant. Therefore, in the present investigation, we have synthesized AgNPs using the aqueous bark extract of A.scholaris and evaluated the antimicrobial efficacy of as prepared AgNPs against pathogenic fungi and bacteria using disc diffusion (in vitro) method.
Materials and methods
Silver nitrate (>99 % pure) was purchased from Sigma-Aldrich, India. Nutrient broth, nutrient agar plate, was supplied by Hi-Media, India.
Collection of biofilm formed in poly vinyl chloride (PVC) pipes
The PVC biofilm samples were collected from four different regions located in and around Tirupati (Chittoor District), Andhra Pradesh, India. The samples were collected from drinking water supply PVC pipelines and stored in plastic bottles. The collected samples were in amorphous form. These samples were stored in an ice box and transported for further microbiological characterization analysis.
Collection of plant material
Healthy plant of A. scholaris was collected from Mumbai, Maharashtra, India. The identity of the plant was confirmed by Agharkar Research Institute, Pune, India. A voucher specimen (No. AHMA-23537) has been deposited for future reference. From the selected plant, bark was collected by scrapping the trunk using neat and clean knife during the month of May, 2014, and collected material was carefully washed and dried at 45 °C to constant weight. The dried bark of plant material was powdered, passed through a BSS no. 85-mesh sieve and stored in airtight container.
Preparation of aqueous extract (AE)
The collected A. scholaris bark was allowed to shade dried for 72 h and was ground to get fine powder. Then, 10 g of powder was mixed with 100 mL of distilled water and boiled for 30 min. After that, the extract was filtered by using Whatman No. 1 filter paper and collected in plastic bottle and stored at 4 °C for further characterization and experimentation.
Isolation of fungal and bacterial sp. from drinking water pipeline
Eight fungal species and ten bacterial samples were isolated from drinking water supply PVC pipelines in Tirupati, Chittoor district, AP, India. Through serial dilution pour plate technique, fungal sp. was isolated using potato dextrose agar (PDA) medium and Gram-negative and Gram-positive bacteria were isolated from nutrient agar medium. Further, it is maintained in potato dextrose agar slants (fungi) and nutrient agar slants (bacteria) for onward analysis.
Preparation of A. scholaris silver nanoparticles
To prepare the AgNPs, a 90-mL aqueous solution of 1.0 × 10−3 M silver nitrate was mixed with a 10-mL of 5 % aqueous solution of A. scholaris bark extract. The A. scholaris Ag solution was yellow in color and the solution was stirred repeatedly for an hour, and it was observed that the color of the solution has been changed to dark brown which visually confirms the formation of nanoparticles. The initial concentration of the A. scholaris silver nanoparticles was measured using inductively coupled plasma optical emission spectrophotometer (ICP-OES) and was found to be 170 ppm. Then, by diluting this solution, each sample of different concentration was used to investigate the concentration dependence of the antibacterial effect of Ag nanoparticles. These A. scholaris silver nanoparticles were characterized by using the techniques such as X-ray diffractometry (XRD), Fourier transform infrared spectrophotometry (FTIR), UV–Vis spectrophotometry, dynamic light scattering (Particle size), zeta potential and transition electron microscopy (TEM).
Measurement of concentration of AgNPs using inductively coupled plasma optical emission spectrophotometer (ICP-OES)
The concentrations of the A. scholaris bark-extract-mediated AgNPs were measured using ICP-OES (Prodigy XP, Leeman Labs, USA). The samples were prepared with 10 times dilution after centrifugation at 4,000 rpm for 15 min. Then, 20 ml of aliquot was loaded to the racks of automatic sampler and estimated the concentration of AgNPs thrice.
Assay for antimicrobial activity of A. scholaris silver nanoparticles against microorganisms (fungi and bacteria)
The antimicrobial activity of A. scholaris silver nanoparticles was examined on the basis of colony formation by in vitro Petri dish assays (disc diffusion). Each fungal and bacterial isolates was cultured on growth media that induced prolific conidia and bacterial production. The fungus isolates were grown on potato dextrose agar medium, and bacterial isolates were grown on nutrient agar medium. Conidia were collected from cultures that were incubated at 37 °C for 10 days (fungi), and bacterial cultures were collected from cultures that were incubated at 37 °C for 2 days for (bacteria) and diluted with sterile, deionized water to a concentration of 106 spores ml-1. Aliquots of the conidial suspension and bacterial suspension were mixed with serial concentrations of silver preparations to a final volume of 1 ml and were also mixed with sterile, deionized water as control. A 10 μl subsample of the conidia and A. scholaris silver mixture stock was taken at 50 ± 0.9, 100 ± 1.1 and 170 ± 1.4 ppm after silver treatments and diluted 100-fold with the deionized water. A 10 μl aliquot of the diluted spore suspension was spread on PDA (Becton, Dickson and Company, Sparks, MD) medium. Three PDA plates for fungi and three NA plates for bacteria per each combination of exposure A. scholaris silver concentration were tested. The filter paper disc dipped in different ppm and inserted on mediums (PDA), and then, the plates were incubated at 37 °C for 2–4 days for fungi and bacteria, respectively. The average number of colonies from silver-treated spore suspensions (fungi) and (bacteria) was compared with the number on the water control (percent colony formation). The zone size was determined by measuring the diameter of the zone in mm [24–26].
Characterization of silver nanoparticles
UV–visible spectrum for synthesized nanoparticles
The bioreductant nanoparticles were monitored by UV–visible (UV–Vis) spectrum at various time intervals. The UV–Vis spectra of this solution were recorded in spectra 2450, SHIMADZU Spectrophotometer, from 400 to 800 nm.
FTIR analysis for synthesized nanoparticles
The FTIR spectrum was taken in the mid-IR region of 400–4,000 cm−1. The spectrum was recorded using ATR (attenuated total reflectance) technique. The dried sample was mixed with the KBr (1:200) crystal, and the spectrum was recorded in the transmittance mode.
Particle size and zeta potential analyzer for synthesized nanoparticles
The aqueous suspension of the synthesized nanoparticles was filtered through a 0.22-μm syringe-driven filter unit, and the size and distribution of the nanoparticles were measured using dynamic light scattering technique (Nanopartica, HORIBA, SZ-100).
X-ray diffraction (XRD) analysis for synthesized nanoparticles
The phyto-reduced silver nanoparticles were characterized by XRD. The XRD pattern was recorded using computer-controlled XRD system (JEOL; Model: JPX-8030 with CuKα radiation (Ni filtered = 13418 Å)) in the range of 40 kV, 20A. The built-in software (syn master 7935) program was used for the identification of XRD peaks corresponds to the Bragg’s reflections.
Transmission electron microscopy (TEM)
The surface morphology and size of the nanoparticles were studied by transmission electron microscopy (JEOL (JEM-1010) instrument) with an accelerating voltage of 80 kV. A drop of aqueous AgNPs on the carbon-coated copper TEM grids was dried and kept under vacuum in desiccators before loading them onto a specimen holder. The particle size and surface morphology of nanoparticles were evaluated using ImageJ 1.45s software.
Results and discussion
Selection of fungi and bacteria present in drinking water pipeline and synthesis of silver nanoparticles
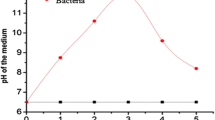
Drinking water pipeline fungi and bacterial species have unusual biological activities depending upon the metabolisms under temperature, pH and pressure. Once the bark (Fig. 1) extract (10 ml) was treated with 90 ml of silver nitrate solution, the color of the solution changed to dark brown color (Fig. 2) in <5 min. The brown color was primarily due to the surface plasmon resonance of the formed silver nanoparticles.
UV–visible spectral analysis: recording of localized surface plasmon resonance (LSPR) of AgNPs
It is well known that silver nanoparticles exhibit brown color, which arises due to excitation of surface plasmon vibrations of the silver nanoparticles. After addition of 1 mM silver nitrate solution to the bark extract, the color of the composition has been changed to dark brown color (Fig. 2). UV–Vis spectroscopy was employed to record the localized surface plasmon resonance of the silver nanoparticles. The maximum absorbance peak is observed at 432 nm (Fig. 3). The overall observations suggest that the bioreduction of (silver ions) Ag+ to Ag(0) was confirmed by UV–Vis spectroscopy.
FTIR analysis: identification of the biomolecules responsible for the reduction and stabilization of AgNPs
FT-IR spectrum is used to identify the possible chemical interactions among the silver salts and functional groups present. FT-IR spectrum of the biosynthesized silver nanoparticles using A. scholaris. Figure 4 shows the absorption peaks at 1,414.78, 1,640, 2,108.51 and 3,380 cm−1. The peak at 3,380 and 2,018.5 cm−1 reveals the presence of N–H bend, indicating the primary and secondary amine groups of protein. The band present at 565 shows the C–Br stretching, likewise the bands at 1,640 and 1,414.78 cm−1 correspond to the primary and secondary amine groups of N–H bending and carbonyl stretching vibrations of protein, respectively, indicating the involvement of proteins in reduction and stabilization of silver ions. The nanoparticles are bound to the functional organic groups (carboxyl and amine) from the A. scholaris extract, and these functional groups may act as template, reducing and capping agents of silver nanoparticles.
X-ray diffraction (XRD) analysis
X-ray diffraction pattern of A. scholaris-mediated synthesis of silver nanoparticles shows the peaks correspond to the Bragg’s reflections of (111), (200), (220) and (311) planes, which confirms the face-centered cubic (FCC) crystalline structure of silver (Fig. 5). The lattice constant calculated from this pattern was 4.0869 Å a value in agreement with the literature report (4.0855 Å) (JSPCDS file no. 89-3722). This clearly indicates that the silver nanoparticles formed by the reduction of Ag+ ions by the A. scholaris extract are crystalline in nature. The relatively higher intensity of (111) plane in FCC crystal structure supports the enhanced antimicrobial activity of the prepared AgNPs. The crystalline size was calculated from the width of the peaks present in the XRD pattern, assuming that they are free from non-uniform strains, using the Debye–Scherrer formula
where D is the average crystalline domain size perpendicular to the reflecting planes, λ (1.5406 × 10−10) is the X-ray wavelength used, β is the full width at half maximum (FWHM) and θ is the diffraction angle [27]. The calculated crystalline size of the AgNPs was 50 nm.
Dynamic light scattering analysis
Dynamic light scattering technique has been used to measure hydrodynamic diameter of the hydrosol (particle suspension). AgNO3 was found to be 111.7 nm (Fig. 6a). The recorded value of zeta potential of the silver nanoparticles was −18.9 mV (Fig. 6b), which resulted in the agglomerated state of the formed AgNPs. If the hydrosol has a large negative or positive zeta potential (≥30 mV), then the particles tend to repel with each other and show no tendency to agglomerate resulted in polydispersed particles.
TEM analysis
The surface morphology, size and shape of phyto-reduced silver nanoparticles were characterized and shown in the TEM micrograph (Fig. 7). From the TEM micrograph, it is evident that AgNPs were spherical in shape and were polydispersed. The measured average size of AgNPs was 50 nm. Occasional agglomeration of the AgNPs has been observed.
Antimicrobial efficacy of A. scholaris bark-extract-mediated silver nanoparticles
Silver nanoparticles obtained from A. scholaris have very strong inhibitory action against fungal sp, Gram-positive and Gram-negative bacteria (Plates 1 and 2). These isolates were collected from drinking water PVC pipes. Three concentrations of AgNPs (170, 100, 50 ppm) were prepared and were applied against an array of fungal species viz, Aspergillus fumigates, Aspergillus niger, Aspergillus clavatus, Cladosporium, Acremonium, Epicoccum, Trichoderma and Rhizopus oligosporus and bacterial species viz., Pseudomonas cepacia, Pseudomonas aeruginosa, Pseudomonaspickettii, Mycobacterium, Legionella, Aeromonas, Klebsiella, Campylobacter, Helicobacter pylori and Staphylococcus aureus. The higher concentration (170 ppm) of AgNPs showed significant antimicrobial effect (Tables 1 and 2) compared with other concentrations (100.50 ppm). The inhibitory action of the microbes may be attributed to the loss of replication ability of DNA upon treatment with the silver ion, besides the fact that expression of ribosomal sub-unit proteins as well as some other cellular proteins and enzymes essential to ATP production becomes inactivated [27]. The higher antimicrobial activity of smaller-sized nanoparticles than their bulk counterparts could be due to the large surface area to volume ratio and the surface activity [28–30]. Further, the fact is that nanoparticles are more abrasive in nature than bulk AgNO3, thus contributing to the greater mechanical damage to the cell membrane resulting in enhanced fungal effect [31]. But to understand the mechanisms of action of these agents, more detailed chemical structure elucidation of the bioactive components followed by therapeutic investigations and toxicological assessment are required.
Conclusions
The bark extract of A. scholaris has been proved to be one of the potential bio-sources to synthesize silver nanoparticles. The formed AgNPs are highly stable and polydispersed with the mean size of 50 nm. At higher concentrations (170 ppm), significant antimicrobial activity of AgNPs has been recorded against Fungal sp. and Gram-positive and Gram-negative bacteria and points to the commercial use of these phyto-medicine-coated AgNPs in biomedical applications and agriculture as fungicides for the effective control of disease causing pathogens. Moreover, these AgNPs could be used as coating material in drinking water PVC pipe lines to control the microbial contamination of the water.
References
Plante, I.J.L., Zeid, T.W., Yangab, P., Mokari, T.: Synthesis of metal sulfide nanomaterials via thermal decomposition of single-source precursors. J. Mater. Chem. 20, 6612–6617 (2010)
Cheng, Y., Yin, L., Lin, S., Wiesner, M., Bernhardt, E., Liu, J.: Toxicity reduction of polymer- stabilized silver nanoparticles by sunlight. J. Phys. Chem. C 115, 4425–4432 (2011)
Hirsch, T., Zharnikov, M., Shaporenko, A., Stahl, J., Weiss, D., Wolfbeis, O.S.: Size controlled electrochemical synthesis of metal nanoparticles on monomolecular templates. Angew. Chem. Int. Ed. 44, 6775–6778 (2005)
Korotchenkov, O.A., Cantarero, A., Shpak, A.P., Kunitskii, Y.A., Senkevich, A.I., Borovoy, M.O.: Doped ZnS: Mn nanoparticles obtained by sonochemical synthesis. Nanotechnology 16, 2033 (2005)
Nadagouda, M.N., Speth, T.F., Varma, R.S.: Microwave-assisted green synthesis of silver nanostructures. Acc. Chem. Res. 44, 469–478 (2011)
Bhattacharya, D., Gupta, R.K.: Nanotechnology and potential of microorganisms. Crit. Rev. Biotechnol. 25, 199 (2005)
Sunkar, S., Nachiyar, C.V.: Microbial synthesis and characterization of silver nanoparticles using the endophytic bacterium bacillus cereus: a novel source in the benign synthesis. Glob. J. Med. Res. 12(2), 43–50 (2012)
Yamac, M., Bilgili, F.: Antimicrobial activities of fruit bodies and/or mycelial cultures of some mushroom isolates. Pharm. Biol. 44, 660–667 (2006)
Cowan, M.M.: Plant products as antimicrobial agents. Clin. Microbiol. Rev. 12, 564–582 (1999)
Rios, J.L., Reico, M.C.: Medicinal plants and antimicrobial activity. J. Ethnopharmacol. 100, 80–84 (2005)
Ahmad, N., Sharma, S., Singh, V.N., Shamsi, S.F., Fatma, A., Mehta, B.R.: Biosynthesis of silver nanoparticles from Desmodium triflorum: novel approach towards weed utilization. Biotechnol. Res. Int. 1–8 (2011). doi:10.4061/2011/454090
Huang, J., Li, Q., Sun, D., Lu, Y., Su, Y., Yang, S.: Biosynthesis of silver and gold nanoparticles by novel sundried Cinnamomum camphora leaf. Nanotechnology 18, 105104 (2007)
Prasad, T.N.V.K.V., Elumalai, E.K.: Biofabrication of Ag nanoparticles using Moringa oleifera leaf extract and their antimicrobial activity. Asian Pac. J. Trop. Biomed. 1(6), 439–443 (2011)
Dubay, M., Bhadauria, S., Kushwah, B.S.: Green synthesis of nanosilver particles from extract of Eucalyptus hybrida (Safeda) Leaf. Dig. J. Nanomater. Biostruct. 4, 537–543 (2009)
Arunachalam, K.D., Annamalai, S.K., Hari, S.: One-step green synthesis and characterization of leaf extract-mediated biocompatible silver and gold nanoparticles from Memecylon umbellatum. Int. J. Nanomedicine 8, 1307 (2013)
Nagajyothi, P.C., Lee, K.D., Sreekanth, T.V.M.: Plants as green source towards synthesis of nanoparticles. J. Optoelectron. Adv. Mater. 15, 269 (2013)
Sreekanth, T.V.M., Nagajyothi, P.C., Supraja, N., Prasad, T.N.V.K.V.: Evaluation of the antimicrobial activity and cytotoxicity of phytogenic gold nanoparticles. Appl Nanosci (2014). doi:10.1007/s13204-014-0354-x
Kumar, S.: Medicinal plants of north-east region, 1st Edition. Scientific Publishers, Rajasthan, India (2002)
Khanikar, G.: Gharooa Sikitsher Nidan, 3rd edn. Puthiteertha Publication, Assam (2007)
Khan, M.R., Omoloso, A.D., Kihara, M.: Fitoterapia 74, 736–740 (2003)
Mostafa, A., Oudadesse, H., Legal, Y., Foad, E., Cathelineau, G.: Characteristics of silver- hydroxyapatite/PVP nanocomposite. Bioceram. Dev. Appl. 1, 1–3 (2011)
Murphy, C.J.: Sustainability as a design criterion in nanoparticle synthesis and applications. J. Mater. Chem. 18, 2173–2176 (2008)
Vaidyanathan, R., Kalishwaralal, K., Gopalram, S., Gurunathan, S.: Nanosilver the burgeoning therapeutic molecule and its green synthesis. Biotechnol. Adv. 27(6), 924–937 (2009)
Cynthia, H., Callaghan, O.: Assessment of a new antibiotic. In: Hugo, W.B., Russel, A.D. (eds.) Pharmaceutical microbiology, vol. 3, pp. 122–134. Blackwell Scientific Publications, Oxford (1983)
Reeves, W., Andrew, W.: Clinical antimicrobial assay, p. 25. Oxford University Press, New York (1999)
Aneja, K.R.: Experiments in microbiology plant pathology tissue culture and mushroom production technology, 3rd edn. New Age International Publisher, India (2003)
Prasad, T.N.V.K.V., Kambala, V.S.R., Naidu, R.: A critical review on biogenic silver nanoparticles and their antimicrobial activity. Curr. Nanosci. 7, 531–544 (2011)
Kouvaris, P., Delimitis, A., Zaspalis, V., Papadopoulos, D., Tsipas, S.A., Michailidis, N.: Green synthesis and characterization of silver nanoparticles produced using Arbutus unedo leaf extract. Mater. Lett. 76, 18–20 (2012)
Jae, Y.S., Beom, S.K.: Rapid biological synthesis of silver nanoparticles using plant leaf extracts. Bioprocess Biosyst. Eng. 32, 79–84 (2009)
Sivakumar, P., NethraDevi, C., Renganathan, S.: Synthesis of silver nano particles using Lantana camara fruit extract and its effect on pathogens. Asian J. Pharm. Clin. Res. 5(3), 97–101 (2012)
Padmavathy, N., Vijayaraghavan, R.: Enhanced bioactivity of AgNO3 nanoparticles—an antibacterial study. Sci. Technol. Adv. Mater. 9, 1–7 (2008)
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
This article is published under license to BioMed Central Ltd. Open Access This article is distributed under the terms of the Creative Commons Attribution License which permits any use, distribution, and reproduction in any medium, provided the original author(s) and the source are credited.
About this article
Cite this article
Shetty, P., Supraja, N., Garud, M. et al. Synthesis, characterization and antimicrobial activity of Alstonia scholaris bark-extract-mediated silver nanoparticles. J Nanostruct Chem 4, 161–170 (2014). https://doi.org/10.1007/s40097-014-0132-z
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s40097-014-0132-z