Abstract
After the World War II, every country throughout the world is experiencing the biggest crisis induced by the devastating Coronavirus disease (COVID-19), which initially arose in the city of Wuhan in December 2019. This global pandemic has severely affected not only the health of billions of people but also the economy of countries all over the world. It has been evident that novel virus has infected a total of 20,674,903 lives as on 12 August 2020. The dissemination of the virus can be regulated by detecting the positive COVID cases as soon as possible. The reverse-transcriptase polymerase chain reaction (RT-PCR) is the basic approach used in the identification of the COVID-19. As RT-PCR is less sensitive to determine the novel virus at the beginning stage, it is worthwhile to develop more robust and other diagnosis approaches for the detection of the novel coronavirus. Due to the accessibility of medical datasets comprising of radiography images publicly, more robust diagnosis approaches are contributed by the researchers and technocrats for the identification of COVID-19 images using the techniques of deep leaning. In this paper, we proposed VGG16 and MobileNet-V2, which makes use of ADAM and RMSprop optimizers for the automatic identification of the COVID-19 images from other pneumonia chest X-ray images. Then, the efficiency of the proposed methodology has been enhanced by the application of data augmentation and transfer learning approach which is used to overcome the overfitting problem. From the experimental outcomes, it can be deduced that the proposed MobileNet-V2 model using ADAM and RMSprop optimizer achieves better accomplishment in terms of accuracy, sensitivity and specificity when contrasted with the VGG 16 using ADAM and RMSprop optimizers.
Similar content being viewed by others
Introduction
Since the beginning of twenty-first century, several viruses belonging to the family of coronaviruses that cause serious impact on the health of human being have been emerging. Severe acute respiratory syndrome (SARS) and Middle East respiratory syndrome (MERS) belonging to the family of coronaviruses have been reported in 2003 and 2015, respectively. In December 2019, the Wuhan city in South china have been reported with a series of unidentified pneumonia cases. Most of these cases are associated with the Wholesale wet market which is famous for selling large variety of fresh meat and sea food. On 7 January 2020, Centers for Disease Control declared the virus as novel coronavirus pneumonia after making an examination of the samples gathered from the throat swab of the infected patients [1]. As the genome structure of novel coronavirus is identical to the structure of SARS-CoV, it was renamed as Severe Acute Respiratory Syndrome Coronavirus 2 (SARS-CoV-2) by the International Committee on Taxonomy of Viruses [2, 3], and the disease was renamed on 11 February 2020 as COVID-19 by the World Health organization (WHO) [4]. On 11 March 2020, when the number of cases has raised thirteen time all over the world apart from China then the WHO declared the COVID-19 disease as pandemic. The clinical symptoms of COVID-19 infected patient are identical to the signs of illness caused by Influenza, bacterial pneumonia and respiratory syncytial virus (RSV). The symptoms like fever, dry cough, sore throat, shortness of breath, diarrhea and lung infiltrates are the most common signs of illness observed in coronavirus infected patients. In the beginning, the spread of novel COVID-19 is related with recent travel history to china. Later on, the increase in number of cases occurred because of person-to-person contact through respiratory droplets [5]. Therefore, the necessary measure in preventing the transmission of disease is the immediate diagnosis of COVID-19 infected patients. Presently, RT-PCR is the standard screening technique utilized for the detection of COVID-19 infection [6, 7]. Due to less sensitivity of RT-PCR in the identification of COVID-19 at the initial stage and unavailability of RT-PCR kits, computed radiography images are used as an alternative approach for the diagnosis of COVID-19 infection. Several clinical analyses reveal that most of the COVID-19 infected patients experience the symptom of lung infection [8]. Even though computed tomography (CT) images are more accurate in the diagnosis of diseases associated with lungs, chest X-ray images are widely used. This is mainly due to the availability of more number of X-ray devices and also because of its low cost and less radiation when compared to CT scans.
From the past few years, the usage of computer vision and deep learning algorithms for the automatic identification of different diseases and lesions and for the analysis of biomedical images has produced sufficient results [9]. These techniques have been effectively used in the determination of types of tumor [10, 11], classification of electrocardiogram (ECG) [12], detection of skin cancer [13], etc. In recent years, due to the substantial increase in the number of pneumonia cases it becomes one of the hot topic areas in the research. As deep learning techniques can be utilized with low-cost imaging techniques, the usage of deep learning methods on chest X-ray images is gaining popularity. Togaçar et al. [14] have used convolutional neural network (CNN) as feature extractor for the detection of pneumonia in chest X-ray images. Initially, they performed reduction in number of deep features in each deep model, such as AlexNet, VGG-16 and VGG-19. Later, they performed classification by using different approaches, such as decision tree, K-nearest neighbors, linear discriminant analysis, linear regression, and support vector machine (SVM). The acquired outcomes reveal that the deep features generate coherent features for the identification of pneumonia in X-ray images. A unique deep learning architecture for the categorization of child pneumonia images was proposed by Liang and Zeng [15]. The proposed model utilizes residual structure and dilated convolution to avoid the problem of depth model, such as degradation and overfitting. The development of prediction models for the detection of post-stroke pneumonia using deep learning techniques has been suggested by Ge et al. [16]. The temporal series of information available in electronic health records have been utilized by implementing classical methods and methods depending on the concepts of multiple layer perceptron (MLP) and recurrent neural networks (RNN). From the results, it can be deduced that the suggested model achieves enhanced performance when analyzed with the performance of traditional machine learning approaches.
The main intention of this paper is to improve the novel COVID-19 detection by using Deep learning architectures, such as VGG16 and MobileNet-V2 with ADAM and RMSprop optimizers. Further, we have applied techniques, such as image preprocessing, augmentation of data and transfer learning techniques to avoid the overfitting problem. Finally, to assess the performance of the models, the proposed models have been evaluated on chest X-ray image dataset.
Related Work
From the last few months, COVID-19 is rapidly spreading across different regions of the world. To inhibit the spread of the novel virus, it is required to identify the COVID-19 positive cases as soon as possible. As the signs of coronavirus infection are related to pneumonia, it can be revealed through the diagnosis of medical imaging datasets. Chest X-ray images are extensively used by the medical practitioners in the identification of COVID-19 disease because of the availability of sufficient number of X-ray devices. Several studies have been conducted on the diagnosis of pneumonia and other viral diseases using chest X-ray images [17,18,19,20]. As deep learning techniques are efficiently utilized in the screening, segmentation and classification of medical images [21], these techniques are used as alternative to the standard RT-PCR for the identification of COVID-19 infected images.
To assist clinical experts in effective identification of COVID-19 disease, an efficient detection method known as Cascaded-SEMEnet was developed by Lv et al. [22]. The proposed model consists of SEME-ResNet50 and SEME-DenseNet169. Vast input sizes and SE-structure are adopted by both cascaded network of the proposed model. Further, for the enhancement of data it utilizes MOEx and Histogram equalization techniques. Initially, SEME-ResNet50 was used for the screening of chest X-ray images and for identification of chest X-ray images as normal, viral pneumonia a bacteria class. Afterwards, SEME-DenseNet169 was utilized for further fine categorization of viral pneumonia to determine whether it has been generated by COVID-19. While training the SEME-DenseNet169, U-Net is used for preprocessing the data to remove the impact of non-pathological characteristics on the network. Moreover, from the outcomes it can be noticed that the suggested model produces an accuracy of 85.6% in deciding the category of pneumonia infection and accuracy of 97.1% in the fine categorization of COVID-19. Further, authors have utilized Grad-CAM for anticipating the judgment which provides assistance to the physicians in understanding the chest X-ray image for the efficient diagnosis of COVID-19 disease. Rahimzadeh et al. [23] have trained various deep convolutional paradigms for the efficient categorization of chest X-ray images as normal, pneumonia and COVID-19 infected classes using two publicly available datasets. These datasets consist of 180 X-ray images of COVID-19 infected patients. They also used some training approaches that assist the network models to perform better even if it consists of unbalanced datasets. Further, a neural network framework that combines both Xception and ResNet50 models has been proposed. The multiple features obtained from the two network models helps in achieving better accuracy results in the categorization of chest X-ray images. To reveal the original accuracy attainable in real situations, the proposed model was tested on 11,302 images. The results specify that the proposed model attains an average accuracy of 99.50% for identifying the COVID-19 infected images and an overall accuracy of 91.4% for the identification of all classes. To reduce the limitation of inadequacy of trained practitioners in remote villages, a novel approach for the automatic identification of COVID-19 infection from the chest X-ray images was proposed by Ozturk et al. [24]. The proposed model was mainly designed for the precise categorization of X-ray images as two class (COVID-19 and normal images) and multiple class (COVID-19, Pneumonia and Normal) images. The proposed model was implemented using 17 convolutional layers and by using various filtering techniques in every layer of the model. The experimental outcomes reveal that the suggested approach achieves 98.08% and 87.02% for the categorization of X-ray images into two class and multi-class images. The YOLO (you only look once) real-time detection system was developed by using Darknet approach as classifier. Moreover, the proposed model can be implemented to provide help to the radiologist in ascertaining the initial screening. It also provides prompt screening of the patients by implementing the proposed model via cloud. To overcome the challenges of unbiased learning, various random oversampling and weighted class loss function techniques have been suggested by Punn et al. [25]. These approaches provide fine-tuned transfer learning in different pre-trained deep learning models to execute classification of posteroanterior X-ray image into two classes, i.e., normal and COVID-19 infected images and multi-class images, i.e., normal, pneumonia and COVID-19 infected images. The effectiveness of the model is assessed by using evaluation metrics, such as accuracy, loss, recall, precision and area under curve (AUC). From the experimental results, it can be noticed that the effectiveness of the models mainly relies on the scenario. Among the available models, NASNetLarge achieves better score. Moreover, the authors also compared the performance of NASNetLarge with other newly proposed applications to show its efficiency in classification of X-ray radiography images. The following Table 1 represents the other research works carried on the diagnosis of coronavirus infection from chest X-ray radiographs using deep learning approaches.
Methodology
From the past few years, deep learning frameworks have been extensively utilized for the diagnosis of pneumonia [32, 33]. In this paper, we have chosen VGG16 and MobileNetV2 for the automatic diagnosis of the COVID infected images from the other pneumonia X-ray images using ADAM and RMSprop optimizer.
VGG16 Architecture
The architecture of VGG (Visual Geometry Group) was proposed by Simonyan and Zisserman in 2014 to win the competition of ILVSR (ImageNet) [34]. The model VGG16 belongs to CNN framework. As it consists of 16 layers, this model was named as VGG 16. The VGG16 model comprises of layers, such as Convolutional, Max Pooling, Activation and Fully connected layers. This framework has been trained on ImageNet dataset that consists of 14 million images and 1000 classes and attains top-5 accuracy of 92.7%.
It outperforms AlexNet network by substituting the large filters of size 11 × 5 used in the first and second convolution layers with small filters of size 3 × 3. Therefore, VGG16 is the modified version of AlexNet. This network is large even by modern standards. It has around 138 million parameters. Most of the parameters are used in the fully connected layers. It has a total memory of 96 MB per image for only forward propagation. The block diagram of VGG-16 architecture is mentioned in Fig. 1.
MobileNetV2 Architecture
MobileNetV2 is issued as a component of TensorFlow-Slim Image Classification Library. MobileNetV2 is a convolutional neural network builds upon the ideas from MobileNetV1 [35]. The architecture of MobileNetV2 consists of 54 layers that takes as input an image of size 224 × 224. The main feature of this architecture is that it utilizes depth wise separable convolution as the effective building blocks of the model. Due to this feature, MobileNet-V2 makes use of two 1D convolutions with two kernels rather than operating on 2D convolution with one kernel. Moreover, it also adds two new features to the framework, such as usage of linear bottlenecks between the layers and shortcut connections between the bottlenecks.
As depicted in Fig. 2, the MobileNet-V2 consists of two types of blocks. One block k with stride1is the residual block and the other block with stride 2 is used for downsizing. Both the blocks consist of three layers. The first layer consists of 1 × 1 convolution with ReLU. Depth wise convolution of 3 × 3 is used in the second layer. The third layer again makes use of 1 × 1 convolution that is linear. The third layer does not make use of ReLU. If the third layer uses ReLU again, then the deep networks can be used only as linear classifier on the nonzero part of the output.
Each layer in the MobileNet-V2 is represented as series of one or more identical layers that are repeated for n number of times. The number of output channels is same in all the layers that have the same series. Stride s is used only in the first layer of each series, while all other layers make use of stride 1. The MobileNet-V2 makes use of 3 × 3 kernels in all the spatial convolutions. MobileNetV1 requires 1600 k memory space but MobileNetV2 requires only 400 k memory space (1/4 of MobileNetV1). So, the model has taken the MobileNet-V2 for data training. The overall architecture of the MobileNet-V2 is depicted in Fig. 3.
The directed acyclic hypergraph ‘G’ generated by TensorFlow comprises of edges and nodes which are used to describe the operations and tensors in between computations. The total number of tensors that needs to be stored in memory can be reduced by the computations. In general, it explores all computational orders Σ(G) and selects the one that minimizes.
In Eq. 1, the list of intermediate tensors linked to any of πi... πn nodes is denoted by R(i, π, G), size of the tensor A is denoted by |A| and the total amount of memory required for internal storage during each operation i is denoted by size (πi).
The amount of memory required is denoted by the maximum total size of the integrated inputs and outputs over each operation as shown in Eq. 2
Bottleneck Residual Block
Equation 3 shows a bottleneck block operator \({\mathcal{F}}(x)\) expresses as a composition of three operator \({\mathcal{F}}(x) = [A \circ {\mathcal{N}} \circ B]x\), where A if linear transform A: \({\mathcal{R}}^{s \times s \times k} \to {\mathcal{R}}^{s \times s \times n} ,{\mathcal{N}}\) is nonlinear transform \({\mathcal{N}}:{\mathcal{R}}^{s \times s \times k} \to {\mathcal{R}}^{{s^{\prime } \times s\prime \times n}}\), B is also linear transform(o/p) B: \({\mathcal{R}}^{{s^{\prime } \times s^{\prime } \times n}} \to {\mathcal{R}}^{{s^{\prime } \times s^{\prime } \times k^{\prime } }}\). For this network \({\mathcal{N}} = ({\text{Relu}} \circ {\text{dwise}} \circ {\text{Relu}})\). The memory space needed to calculate F(X) can be as small as \(\left| {s^{2} k} \right| + \left| {s^{\prime 2} k^{\prime } } \right| + {\mathcal{O}}(\max (s^{2} ,s^{\prime 2} )).\)
The basic phenomenon of the algorithm mainly depends on the factor that the inner tensor denoted by I can be expressed as sequence of t tensors each of size n/t. The function used in bottle residual block can be represented as shown in Eq. 3
Optimization Algorithm
Both VGG16 and MobileNet-V2 makes use of ADAM and RMSprop as optimizers for the identification of the COVID infected X-ray images from other pneumonia images. The algorithms for ADAM [36] and RMSprop [37] are given below.


Experimental Set up
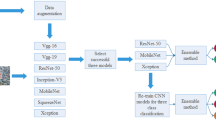
The experiment has carried out on: Acer Predator PH315-51 Helios 300; CPU: Intel® Core™ i7 + 8th Gen -8750H @ 2.20 GHz 2.21 GHz; RAM: 8.00 GB; GPU: NVIDIA® GEFORCE® GTX 1060; Optane Memory: 16 GB; OS: Windows 10 Home, Anaconda Distribution package jupyter notebook and Python programming language along with tensorflow, keras, Numpy, matplotlib and OpenCV. Fig. 4 represents the in-detail steps in the proposed methodology.
Description of Dataset
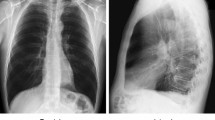
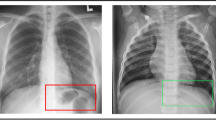
In this study, we considered the image dataset from the Kaggle COVID-19 X-ray dataset [38], organized into dataset folder and contains subfolders for each image category (COVID-19/Normal). This dataset comprises of 624 X-ray images of which 390 are COVID-19 X-Ray images and 234 are Normal X-ray images. Figure 5 depicts the examples of X-ray images of patients with COVID-19 infection and normal X-ray images. Figure 5a displays COVID-19 infected X-ray images with opacities in the lobes of the lungs, while Fig. 5b displays normal X-ray images with no abnormal opacification zones.
Data Preprocessing
To enhance the training process, the input images are pre-processed utilizing various preprocessing approaches. The main reason for preprocessing the images is to enhance the visual capacity of the input image by increasing contrast, by deleting low or high frequencies in the image and by reducing the noise present in the original image. In this paper, we applied image resizing and image normalization preprocessing techniques for enhancing the quality of image.
Image Resizing
The dimensions of the images in this dataset are of different ranges. Therefore, to achieve fixed dimensions of all images, the original images are resized to the size of 224 × 224 pixels to fit to the input image size in VGG16 and MobileNetV2 pretrained models.
Image Normalization
In this paper, we applied intensity normalization as preprocessing technique which is widely used in image processing applications [39]. The intensity values of all the images from 0 to 255 are normalized to the basic normal distribution by utilizing min–max normalization as shown in Eq. 15. As this technique removes bias and achieves uniform distribution, it assists in accelerating the convergence of the model.
where x is the pixel intensity. xmin and xmax are minimum and maximum intensity values of the input image in Eq. 15.
Approach of Data Augmentation
Data augmentation is a technique used by the practitioners to significantly enhance the diversity of available data available for training models, without obtaining any new data. To train the large neural networks, techniques of data augmentation, such as cropping, padding and horizontal flipping, are used. However, most of the approaches use only primitive type of augmentation for training the large neural networks. As the main focus of the neural network architecture is on analyzing the depth, less attention has been given in developing strong type of data augmentation techniques. In our research, we have used the following parameter values as shown in Table 2 in data augmentation technique to avoid the data overfitting.
Data Splitting
In this paper, further image dataset is divided into 80:20 ratio where 80% of the image dataset is used for training the model and 20% of the image dataset is used for validating the model.
Transfer Learning
The process of replacing random initialization of weights with the existing convolutional neural network models that were trained on huge image datasets is known as transfer learning. Transfer learning can be performed in two ways. The first way is concerned with removing the fixed features and the whole classifier is trained by considering data from the intermediate layer. The second way is concerned with fine-tuning of the image dataset. In this, new classes are used to replace the last fully connected layer and then the entire convolutional neural network model is retrained with the new alterations. Further, the weights in the deeper layer are updated using these alterations [2, 13]. In this paper, we performed transfer learning by using the net weights of the VGG16 and MobileNetV2 models to replace the last fully connected layer. Further, we added the global average pooling and required final two class labels.
Result and Discussions
We validated our proposed models on 390 COVID and 234 normal Chest X-ray images using ADAM and RMSprop optimizers. It is also significant that transfer learning techniques have been used in the training process improve the learning ability of models. We performed transfer learning by using the net weights of the VGG16 and MobileNetV2 models to substitute the weights of the last fully connected layer. Further, we added the global average pooling and required final two class labels.
Performance Metrics Used for the Interpretation of Image Data
The efficiency of any machine learning model can be determined using performance metrics, such as F1_score, Recall, Precision, Specificity, Confusion Matrix, Mean and Standard Deviation. In this study, the efficiency of the proposed approach is also evaluated using these methods based on confusion matrix. The equations for Recall, Fall_out, Specificity, F1_score and Accuracy are specified in Eqs. 16 to 20
where True Positive, False Positive, True Negative and False Negative of dataset are denoted by TP, FP, TN and FN, respectively.
Classification Report
Table 3 highlights the detailed comparative analysis and best scores of the VGG16 and MobileNetV2 models using ADAM and RMSprop optimizers for the identification of COVID-19 infected images by means of discussed evaluation metrics. From Table 3, it can be observed that MobileNet-V2 using ADAM and RMSprop optimizer attains better performance in terms of precision, recall, and F1_score when compared with VGG 16 precision, recall, and F1_score for identification of COVID-19 infected chest X-ray images. Furthermore, MobileNet-V2 using ADAM and RMSprop achieves an overall accuracy of 92.13% and 92.91%, respectively, when compared with accuracy of VGG16 using ADAM and RMSprop optimizer. It can be also observed that MobileNetV2 using RMSprop and ADAM achieves better accuracy in identification of COVID and normal chest X-ray images when compared with the accuracy of VGG16 using ADAM and RMSprop optimizer.
Confusion Matrix with Heat Map
Figures 6,7,8,9 represent the confusion matrix of VGG16 and MobileNet-V2 with ADAM and RMSprop optimizers that illustrates the detailed description of what happens with images after image classification.
From Fig. 6, it can be observed that the VGG16 model using ADAM optimizer was capable of recognizing 74 images as COVID infected images, while 6 images belonging to COVID class are wrongly identified as normal images. In normal class, 41 images were correctly identified as normal images, but 6 images were wrongly identified as COVID images. In Fig. 7, VGG 16 using RMSprop optimizer in COVID class was able to recognize 78 images correctly as COVID infected images, but 2 images were wrongly identified as normal images. While 37 images were identified as normal images and 10 were wrongly determined as COVID infected in Normal class. From Fig. 8, it can be observed that MobileNet-V2 with ADAM optimizer can diagnose 73 X-ray images correctly as COVID infected images in COVID class, but 7 images were wrongly diagnosed as normal images. While MobileNet-V2 with ADAM optimizer in normal class was able to recognize 44 images as normal images and 3 images as COVID-19 infected images. It can be observed from Fig. 9 that MobileNet-V2 with RMSprop optimizer was able to analyze 72 X-ray images correctly as COVID class images, while 8 images were wrongly identified as normal images. In normal class, MobileNet-V2 was able to diagnose 46 images as normal images and only one image was wrongly identified as COVID infected image. The accuracy, sensitivity and specificity of the VGG16 and MobileNet-V2 using ADAM and RMSprop optimizer are represented in Table 4.
It can be also observed from Table 4 that MobileNet-V2 with ADAM optimizer and RMSprop optimizers achieves accuracy of 92.13% and 92.91%, sensitivity of 91.25% and 90% and specificity of 93.62% and 97.87%, respectively, when compared with VGG16 using ADAM (accuracy is 90.55%, sensitivity is 92.50%, specificity is 87.23%) and VGG16 using RMSprop optimizer (accuracy is 90.05%, sensitivity is 97.50%, specificity is 78.72%). Moreover, the results achieved in this study are analyzed with the outcomes obtained using these models in other studies to identify COVID-19 from X-ray images as shown in Table 5.
It can be noticed from Table 5 that our proposed approach achieves better accuracy and precision when compared with the same models in other recent studies. In the proposed work, the visual capacity of the image dataset has been enhanced by applying image resizing and normalization as data preprocessing techniques. Then, data augmentation technique is applied to avoid the overfitting of the model. Further, transfer learning results in saving training time of the module. Because of these techniques, the proposed work achieves better accuracy when compared with other previous studies.
Performance Testing of Four Different Method
This section is used to represent and discuss the classification results of chest X-ray images. The classification results are represented using train curve and validation curve. From the training dataset, the train curve is calculated which represents how appropriately the prototype is learning, while the validation curve which is determined from the hold-out validation dataset is used for representing how properly the model is generalizing. Figures 10,11,12,13 represent the training and validation curve of VGG16 with ADAM optimizer, VGG16 with RMSprop optimizer, MobileNetV2 with ADAM optimizer, MobilenetV2 with RMSprop optimizer.
Training and Validation Curve of VGG16 with ADAM Optimizer
Fig. 10 represents the accuracy and loss curve of VGG16 using ADAM optimizer. From the figure, it can be noticed that the train data accuracy curve is gradually raising from epoch 1 to epoch 22 and then it converges to a value around 90.98%. Similarly, the validation accuracy curve is also gradually raising from epoch 1 to epoch 25 and reaches a value of 90.55% for the epoch 25. It is also noticed that in the train data loss curve is gradually decreasing from epoch 1 to last epoch and reaches and the loss is equal to 0.26 for the epoch 25. The same is noticed for the validation data loss curve and the loss value for the last 25 epoch is 0.23.
Training and Validation Curve of VGG16 with RMSprop Optimizer
From Fig. 11, it can be observed that the train and test data accuracy curve of VGG 16 using RMSprop increases from epoch 1 to last epoch and reaches a value of 90.3% and 90.5% for the last epoch 25 of train and test data. In the same way, for the loss curve that represents train and validation data, the loss value gradually decreases from epoch 1 to last epoch and reaches a value of 0.25 and 0.24 for the last epoch of train and test data, respectively.
Training and Validation Curve of MobileNet-V2 with ADAM Optimizer
As shown in Fig. 12, the training and validation accuracy curve of the MobileNet-V2 increases from epoch 1 to 23 and then converge to a value around 95.5% and 92.1%, respectively, for the train and validation data. For the loss curve, the loss value gradually decreases from epoch 1 to epoch 25 and reaches a value of 0.11 for the train data and 0.18 for the test data.
Training and Validation curve of MobileNet-V2 with RMSprop optimizer
In Fig. 13, the accuracy curve increases gradually from epoch 1 to epoch 19 and then for the last epoch it reaches a value of 94.39% and 92.91% for the train and test data of MobileNet-V2 using RMSprop. In the case of loss curve, the value decreases from epoch 1 to epoch 25 and for the lost epoch the loss value is 0.16 for the train data and 0.20 for the test data.
Conclusion
The COVID-19 infection induced by SARS-CoV2 is one of the recent viruses that loomed in late December 2019 in the Wuhan city of South China. This virus causes pneumonia that causes infection to the lung air sacs. RT-PCR is the basic approach used in the identification of COVID-19 infection. The alternative approach that is used in the efficient detection of infection is computed radiography images as RT-PCR is less sensitive for the determination of novel coronavirus at the initial stage. Therefore, in this paper, we proposed VGG16 and Mobilenet-V2, which makes use of ADAM and RMSPROP optimizers for the automatic identification of the COVID-19 X-ray images from other X-ray images consisting of pneumonia. Then, the efficiency of the proposed methodology has been enhanced by the application of data preprocessing, augmentation of data and transfer learning approaches which are utilized to enhance the visual quality of the image as well as to overcome the overfitting problem. Precision, recall, f1score, and accuracy are applied to estimate the efficiency of the models. The performance of the proposed models has been validated on a publicly available COVID-19 dataset consisting of 390 COVID-19 and 234 normal X-ray images. From the experimental outcomes, it can be deduced that proposed MobileNet-V2 prototype using ADAM and RMSPROP optimizers achieved accuracy of 92.13% and 92.91%, sensitivity of 91.25% and 90% and specificity of 93.62% and 97.87%, respectively, when compared with the VGG16 using ADAM and RMSPROP optimizers. Though this model had performed incredibly well for image classification, there is no concept of image segmentation and localization. In future, we will look forward to apply Image segmentation using deep learning approaches.
References
H. Lu, C.W. Stratton, Y.-W. Tang, Outbreak of pneumonia of unknown etiology in Wuhan, China: the mystery and the miracle. J. Med. Virol. 92(4), 401–402 (2020)
C. Huang et al., Clinical features of patients infected with 2019 novel coronavirus in Wuhan, China. Lancet 395(10223), 497–506 (2020)
C.-C. Lai, et al. Severe acute respiratory syndrome coronavirus 2 (SARS-CoV-2) and corona virus disease-2019 (COVID-19): the epidemic and the challenges. Int. J. Antimicrob. Aagent 55(3), 105924 (2020). https://doi.org/10.1016/j.ijantimicag.2020.105924
World Health Organization. Laboratory testing for coronavirus disease 2019 (COVID-19) in suspected human cases: interim guidance, 2 March 2020. No. WHO/COVID-19/laboratory/2020.4. World Health Organization (2020)
S. Chavez, et al. Coronavirus Disease (COVID-19): A primer for emergency physicians. Am. J. Eemergency Med. (2020). https://doi.org/10.1016/j.ajem.2020.03.036
W. Wang et al., Detection of SARS-CoV-2 in different types of clinical specimens. JAMA 323(18), 1843–1844 (2020)
V.M. Corman et al., Detection of 2019 novel coronavirus (2019-nCoV) by real-time RT-PCR. Eurosurveillance 25(3), 2000045 (2020)
Y.-R. Guo et al., The origin, transmission and clinical therapies on coronavirus disease 2019 (COVID-19) outbreak–an update on the status. Military Med Res 7(1), 1–10 (2020)
G. Litjens et al., A survey on deep learning in medical image analysis. Med Image Anal 42, 60–88 (2017)
J.-Z. Cheng et al., Computer-aided diagnosis with deep learning architecture: applications to breast lesions in US images and pulmonary nodules in CT scans. Sci Rep 6(1), 1–13 (2016)
S.K. Lakshmanaprabu et al., Optimal deep learning model for classification of lung cancer on CT images. Futur. Gener. Comput. Syst. 92, 374–382 (2019)
A. M. Alqudah, Q. Shoroq, A. Amin. Automated systems for detection of COVID-19 using chest x-ray images and lightweight convolutional neural networks. (2020).
A.M. Alqudah, H. Alquraan, I.A. Qasmieh. Segmented and non-segmented skin lesions classification using transfer learning and adaptive moment learning rate technique using pretrained convolutional neural network. J. Biomimet. Biomater. Biomed. Eng. 42 (2019). https://doi.org/10.4028/www.scientific.net/JBBBE.42.67
M. Toğaçar, et al. A deep feature learning model for pneumonia detection applying a combination of mRMR feature selection and machine learning models. IRBM (2019)
G. Liang, L. Zheng, A transfer learning method with deep residual network for pediatric pneumonia diagnosis. Comput. Methods Programs Biomed. 187, 104964 (2020)
Y. Ge et al., Predicting post-stroke pneumonia using deep neural network approaches. Int. J. Med. Informatics 132, 103986 (2019)
J.P. Cohen, et al. "COVID-19 image data collection: prospective predictions are the future. arXiv preprint http://arxiv.org/abs/arXiv:2006.11988 (2020).
J. Irvin, et al. Chexpert: A large chest radiograph dataset with uncertainty labels and expert comparison, in Proceedings of the AAAI Conference on Artificial Intelligence. Vol. 33 (2019).
L. Oakden-Rayner. CheXNet: an in-depth review. Luke Oakden-Rayner (PhD Candidate/Radiologist) Blog (2018)
A. Stein. Pneumonia dataset annotation methods. rsna pneumonia detection challenge discussion, 2018. https://www.kaggle.com/c/rsna-pneumonia-detection-challenge/discussion (2020)
G. Litjens et al., A survey on deep learning in medical image analysis. Med. Image Anal. 42, 60–88 (2017)
D. Lv, et al. A cascade network for Detecting COVID-19 using chest x-rays. arXiv preprint http://arxiv.org/abs/arXiv:2005.01468 (2020).
M. Rahimzadeh, A. Attar. A modified deep convolutional neural network for detecting COVID-19 and pneumonia from chest X-ray images based on the concatenation of Xception and ResNet50V2. Informat. Med. Unlock 100360 (2020). https://doi.org/10.1016/j.imu.2020.100360
T. Ozturk, et al. Automated detection of COVID-19 cases using deep neural networks with X-ray images. Comput. Biol. Med. 103792 (2020). https://doi.org/10.1016/j.compbiomed.2020.103792
N.S. Punn, S. Agarwal. Automated diagnosis of COVID-19 with limited posteroanterior chest X-ray images using fine-tuned deep neural networks. arXiv preprint http://arxiv.org/abs/arXiv:2004.11676 (2020).
L.O. Hall, et al. Finding covid-19 from chest x-rays using deep learning on a small dataset. arXiv preprint http://arxiv.org/abs/arXiv:2004.02060 (2020).
A.I. Khan, J.L. Shah, M.M. Bhat. Coronet: a deep neural network for detection and diagnosis of COVID-19 from chest x-ray images. Comput. Methods Program Biomed. 105581 (2020). https://doi.org/10.1016/j.cmpb.2020.105581
E.E.-D. Hemdan, M.A. Shouman, M.0E. Karar. Covidx-net: a framework of deep learning classifiers to diagnose COVID-19 in x-ray images. arXiv preprint http://arxiv.org/abs/arXiv:2003.11055 (2020).
L. Wang, A. Wong. COVID-Net: a tailored deep convolutional neural network design for detection of COVID-19 cases from chest x-ray images. arXiv preprint http://arxiv.org/abs/arXiv:2003.09871 (2020).
E.J.daS. Luz, et al. Towards an effective and efficient deep learning model for COVID-19 patterns detection in x-ray Images. CoRR (2020).
X. Li, D. Zhu. Covid-xpert: an ai powered population screening of covid-19 cases using chest radiography images. arXiv preprint http://arxiv.org/abs/arXiv:2004.03042 (2020).
A. Bhandary et al., Deep-learning framework to detect lung abnormality–A study with chest X-Ray and lung CT scan images. Pattern Recogn Lett 129, 271–278 (2020)
D.S. Kermany et al., Identifying medical diagnoses and treatable diseases by image-based deep learning. Cell 172(5), 1122–1131 (2018)
K. Simonyan, A. Zisserman. Very deep convolutional networks for large-scale image recognition. arXiv preprint http://arxiv.org/abs/arXiv:1409.1556 (2014).
M. Sandler, et al. Mobilenetv2: Inverted residuals and linear bottlenecks, in Proceedings of the IEEE conference on computer vision and pattern recognition. 2018.
D.P. Kingma, J. Ba. Adam: A method for stochastic optimization. arXiv preprint http://arxiv.org/abs/arXiv:1412.6980 (2014).
G. Somua-Gyimah, et al. A computer vision system for terrain recognition and object detection tasks in mining and construction environments, in SME Annual Conference (2019)
https://www.kaggle.com. Accessed Sep 4 2020.
S.H. Kassani, et al. Classification of histopathological biopsy images using ensemble of deep learning networks. arXiv preprint http://arxiv.org/abs/arXiv:1909.11870 (2019).
M.R. Karim, et al. DeepCOVIDExplainer: Explainable COVID-19 diagnosis based on chest x-ray images.
X. Li, C. Li, D. Zhu. COVID-MobileXpert: On-Device COVID-19 screening using snapshots of chest x-ray. arXiv preprint http://arxiv.org/abs/arXiv:2004.03042 (2020).
Author information
Authors and Affiliations
Corresponding author
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
About this article
Cite this article
Swapnarekha, H., Behera, H.S., Roy, D. et al. Competitive Deep Learning Methods for COVID-19 Detection using X-ray Images. J. Inst. Eng. India Ser. B 102, 1177–1190 (2021). https://doi.org/10.1007/s40031-021-00589-3
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s40031-021-00589-3