Abstract
Purpose
Thymosin β4 is a highly active protein that exerts multiple biological activities such as tissue repair, anti-inflammation, and cell maturation. Thymosin β4 has also been listed as a prohibited drug by the World Anti-doping Agency (WADA). Based on its biological activities, thymosin β4 has a high potential of abuse for the performance enhancement among athletes. This study aimed to investigate and characterize the metabolism of thymosin β4 in vitro system.
Methods
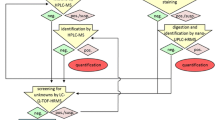
TB4 protein was metabolized in six different enzyme–buffer systems in vitro. After TB4 was metabolized with an appropriate buffer system, the resulting metabolites were detected by high resolution LC–MS/MS. The mass spectrum data of the observed metabolites were characterized in silico, and confirmed the structures based on synthesized authentic standards.
Results
Total 13 new metabolites, some of which were detected in more than one enzyme system, were found. This study characterized all of the detected metabolites according to their in silico m/z ions and compared our findings with synthesized standards. Finally, metabolites M1, M5, M7, M11, M12, and M13 were confirmed based on their synthesized authentic standards.
Conclusion
By using an approach for metabolizing a protein to detect, characterize and identify new peptide metabolites, 6 metabolites are identified among 13 expected potential metabolites. Newly detected metabolites may have the potential for biological activities after further screening compared to their parent protein.




Similar content being viewed by others
References
Bagchi S, Chhibber T, Lahooti B, Verma A, Borse V, Jayant RD (2019) In-vitro blood-brain barrier models for drug screening and permeation studies: an overview. Drug Des Dev Ther 13:3591–3605
Charman SA, Segrave AM, Edwards GA, Porter CJ (2000) Systemic availability and lymphatic transport of human growth hormone administered by subcutaneous injection. J Pharm Sci 89:168–177
Crockford D (2007) Development of thymosin beta4 for treatment of patients with ischemic heart disease. Ann N Y Acad Sci 1112:385–395
Di L (2015) Strategic approaches to optimizing peptide ADME properties. AAPS J 17:134–143
Ezan E, Becher F, Fenaille F (2014) Assessment of the metabolism of therapeutic proteins and antibodies. Expert Opin Drug Metab Toxicol 10:1079–1091
Goldstein AL, Hannappel E, Kleinman HK (2005) Thymosin beta4: actin-sequestering protein moonlights to repair injured tissues. Trends Mol Med 11:421–429
Hasan M, Min H, Rahaman KA, Muresan AR, Kim H, Han D, Kwon OS (2019) Quantitative proteome analysis of brain subregions and spinal cord from experimental autoimmune encephalomyelitis mice by TMT-based mass spectrometry. Proteomics 19:e1800355
Ho EN, Kwok WH, Lau MY, Wong AS, Wan TS, Lam KK, Schiff PJ, Stewart BD (2012) Doping control analysis of TB-500, a synthetic version of an active region of thymosin beta(4), in equine urine and plasma by liquid chromatography-mass spectrometry. J Chromatogr A 1265:57–69
Jyrkas J, Tolonen A (2021) Hepatic in vitro metabolism of peptides; Comparison of human liver S9, hepatocytes and Upcyte hepatocytes with cyclosporine A, leuprorelin, desmopressin and cetrorelix as model compounds. J Pharm Biomed Anal 196:113921
Malinda KM, Sidhu GS, Mani H, Banaudha K, Maheshwari RK, Goldstein AL, Kleinman HK (1999) Thymosin beta4 accelerates wound healing. J Investig Dermatol 113:364–368
Matthews RA (2008) Medical progress depends on animal models—doesn’t it? J R Soc Med 101:95–98
Motyan JA, Toth F, Tozser J (2013) Research applications of proteolytic enzymes in molecular biology. Biomolecules 3:923–942
Ooka M, Lynch C, Xia M (2020) Application of in vitro metabolism activation in high-throughput screening. Int J Mol Sci 21:8182
Palermo A, Alessi B, Botre F, De La Torre X, Fiacco I, Mazzarino M (2016) In vitro evaluation of the effects of anti-fungals, benzodiazepines and non-steroidal anti-inflammatory drugs on the glucuronidation of 19-norandrosterone: implications on doping control analysis. Drug Test Anal 8:930–939
Philp D, Kleinman HK (2010) Animal studies with thymosin beta, a multifunctional tissue repair and regeneration peptide. Ann N Y Acad Sci 1194:81–86
Prueksaritanont T, Tang C (2012) ADME of biologics—what have we learned from small molecules? AAPS J 14:410–419
Scarth JP, Spencer HA, Timbers SE, Hudson SC, Hillyer LL (2010) The use of in vitro technologies coupled with high resolution accurate mass LC–MS for studying drug metabolism in equine drug surveillance. Drug Test Anal 2:1–10
Sosne G, Kleinman HK (2015) Primary mechanisms of thymosin beta4 repair activity in dry eye disorders and other tissue injuries. Investig Ophthalmol vis Sci 56:5110–5117
Wrighton SA, Ring BJ, Vandenbranden M (1995) The use of in vitro metabolism techniques in the planning and interpretation of drug safety studies. Toxicol Pathol 23:199–208
Yu X, Fridman A, Bagchi A, Xu S, Kwasnjuk KA, Lu P, Cancilla MT (2020) Metabolite identification of therapeutic peptides and proteins by top-down differential mass spectrometry and metabolite database matching. Anal Chem 92:8298–8305
Zhu T, Park HC, Son KM, Kwon JH, Park JC, Yang HC (2014) Effects of thymosin beta4 on wound healing of rat palatal mucosa. Int J Mol Med 34:816–821
Zvereva I, Semenistaya E, Krotov G, Rodchenkov G (2016) Comparison of various in vitro model systems of the metabolism of synthetic doping peptides: proteolytic enzymes, human blood serum, liver and kidney microsomes and liver S9 fraction. J Proteomics 149:85–97
Acknowledgements
This research was supported, in part, by Project 2V06980 of the Korea Institute of Science and Technology (KIST).
Author information
Authors and Affiliations
Contributions
KAR conceived the study, performed experiments, analyzed data, and prepared the manuscript. ARM contributed to method development and interpreted MS/MS data. HM interpreted MS/MS data, edited and reviewed the manuscript. JS provided support, resources, and funding as a project leader and reviewed the manuscript. M-JK interpreted MS/MS data, edited and reviewed the manuscript. O-SK conceived the study, analyzed mass spectra, interpreted data, and provided support, resources, and funding as a project leader.
Corresponding author
Ethics declarations
Conflict of interest
All authors (K.A. Rahaman, A.R. Muresan, H. Min, J. Son, M.‑J. Kang, and O.-S. Kwon) declare that they have no conflicts of interest or personal relationships that could influence this work.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
About this article
Cite this article
Rahaman, K.A., Muresan, A.R., Min, H. et al. An approach for identifying in silico peptides against authentic metabolites: in vitro characterization of thymosin β4 metabolites. J. Pharm. Investig. 52, 611–621 (2022). https://doi.org/10.1007/s40005-022-00581-z
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s40005-022-00581-z