Abstract
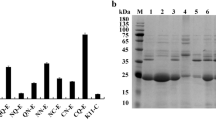
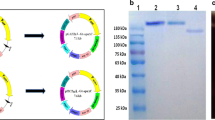
The native and the N-terminal signal peptide sequence deleted gene encoding for α-amylase from Lactobacillus plantarum S21 were cloned into the inducible lactobacilli expression vectors pSIP409 and pSIP609 and expressed in L. plantarum WCFS1 and food-grade L. plantarum TGL02, respectively. Only the native amylase gene was expressed and secreted extracellular amylase at a level of approximately 2000 U/L with 90 % secretion efficiency from both hosts. The purified extracellular amylase from the L. plantarum TGL02 retained unique properties of the wild-type enzyme, particularly the broad pH stability (4.0–8.0) and maltose-forming activity. The results indicate high compatibility of L. plantarum S21 signal peptide sequence to both recombinant lactobacilli hosts. The recombinant lactobacilli exhibited high efficiency for direct lactic acid production from starch as found with L. plantarum S21. The efficient compatible signal peptide is also expected to be applied in secretory expression for production of valuable proteins in food-grade lactobacilli host.






Similar content being viewed by others
References
Abdel-Rahman MA, Tashiro Y, Sonomoto K (2010) Lactic acid production from lignocellulose-derived sugars using lactic acid bacteria: overview and limits. J Biotechnol 156:286–301
Aukrust T, Blom H (1992) Transformation of Lactobacillus strains used in meat and vegetable fermentations. Food Res Int 25:253–261
Cho MH, Park SE, Lee MH, Ha SJ, Kim HY, Kim MJ, Lee SJ, Madsen SM, Park CS (2007) Extracellular secretion of a maltogenic amylase from Lactobacillus gasseri ATCC33323 in Lactococcus lactis MG1363 and its application on the production of branched maltooligosaccharides. J Microbiol Biotechnol 17:1521–1526
Huang LP, Jin B, Lant P, Zhou J (2005) Simultaneous saccharification and fermentation of potato starch wastewater to lactic acid by Rhizopus oryzae and Rhizopus arrhizus. Biochem Eng J 23:265–276
Inoue H, Nojima H, Okayama H (1990) High efficiency transformation of Escherichia coli with plasmids. Gene 96:23–28
Kanpiengjai A, Lumyong S, Pathom-aree W, Khanongnuch C (2014a) Starchy effluent from rice noodle manufacturing process as feasible substrate for direct lactic acid production by Lactobacillus plantarum S21. J Korean Soc Appl Biol Chem 57:217–220
Kanpiengjai A, Rieantrakoonchai W, Pratanaphon R, Pathom-aree W, Lumyong S, Khanongnuch C (2014b) High efficacy bioconversion of starch to lactic acid using an amylolytic lactic acid bacterium isolated from Thai indigenous fermented rice noodles. Food Sci Biotechnol 23:1541–1550
Kanpiengjai A, Haltrich D, Nguyen T-H, Pathom-aree W, Lumyong S, Khanongnuch C (2015) Characterization of a maltose-forming α-amylase from an amylolytic lactic acid bacterium, Lactobacillus plantarum S21. J Mol Catal B Enzym 120:1–8
Karlskås IL, Maudal K, Axelsson L, Rud I, Eijsink VGH, Mathiesen G (2014) Heterologous protein secretion in lactobacilli with modified pSIP vectors. PLoS One 9:e91125
Kleerebezem M, Boekhorst J, van Kranenburg R, Molenaar D, Kuipers OP, Leer R, Tarchini R, Peters SA, Sandbrink HM, Fiers MW (2003) Complete genome sequence of Lactobacillus plantarum WCFS1. Proc Natl Acad Sci USA 100:1990–1995
Mathiesen G, Sveen A, Piard JC, Axelsson L, Eijsink VGH (2008) Heterologous protein secretion by Lactobacillus plantarum using homologous signal peptides. J Appl Microbiol 105:215–226
Muhialdin BJ, Hassan Z, Saari N (2013) Lactic acid bacteria in biopreservation and the enhancement of the functional quality of bread. In: Marcelino K (ed) Lactic acid bacteria R & D for food, health and livestock purposes. InTech, Rijeka, pp 155–172
Nguyen T-T, Mathiesen G, Fredriksen L, Kittl R, Nguyen T-H, Eijsink VG, Haltrich D, Peterbauer CK (2011) A food-grade system for inducible gene expression in Lactobacillus plantarum using an alanine racemase-encoding selection marker. J Agric Food Chem 59:5617–5624
Nguyen HA, Nguyen T-H, Nguyen T-T, Peterbauer CK, Mathiesen G, Haltrich D (2012) Chitinase from Bacillus licheniformis DSM13: expression in Lactobacillus plantarum WCFS1 and biochemical characterisation. Protein Expr Purif 81:166–174
Sasikumar P, Gomathi S, Anbazhagan K, Selvam GS (2013) Secretion of biologically active heterologous oxalate decarboxylase (OxdC) in Lactobacillus plantarum WCFS1 using homologous signal peptides. Biomed Res Int 2013:280432
Sharma A, Satyanarayana T (2013) Microbial acid-stable α-amylases: characteristics, genetic engineering and applications. Process Biochem 48:201–211
Sorvig E, Mathiesen G, Naterstad K, Eijsink VG, Axelsson L (2005) High-level, inducible gene expression in Lactobacillus sakei and Lactobacillus plantarum using versatile expression vectors. Microbiology (Reading, England) 151:2439–2449
Vishnu C, Naveena BJ, Altaf M, Venkateshwar M, Reddy G (2006) Amylopullulanase—a novel enzyme of Lactobacillus amylophilus GV6 in direct fermentation of starch to L(+) lactic acid. Enzym Microb Technol 38:545–550
Acknowledgments
The authors are grateful to the ASEAN-European Academic University Network (ASEA Uninet) funded by the Austrian Federal Ministry of Science, Research and Economy (BMWFW). This work was also supported by Postdoctoral fellowship granted by Chiang Mai University. We also acknowledge the Thailand Research Fund (RTA 5880006).
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
No conflict of interest declared.
Rights and permissions
About this article
Cite this article
Kanpiengjai, A., Lumyong, S., Wongputtisin, P. et al. Efficient secretory expression of gene encoding a broad pH-stable maltose-forming amylase from Lactobacillus plantarum S21 in food-grade lactobacilli host. J Korean Soc Appl Biol Chem 58, 901–908 (2015). https://doi.org/10.1007/s13765-015-0121-z
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s13765-015-0121-z