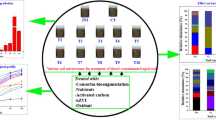
Abstract
The supernatant harvested from a mesophilic, molasses-fed, non-methanogenic bioreactor, which is rich in nitrogen, phosphorous, metals, free amino acids, Sporolactobacillus sp., Prevotella sp., and Clostridium sp., is diluted with tap water and tested as the bioagent to catalyze diesel degradation in soil. Outdoor experiments are performed under the following conditions to assess the effectiveness of the bioagent: diesel doses: 9–15 mg TPH/g soil; bioagent concentrations and dose: 1–5 % at 60 ml/day; soil sample size: 600 g; ambient temperatures: 28–32 °C, and relative humidity: 40–82 % (TPH: total petroleum hydrocarbons). Diesel degradation in soil treated with 3 % bioagent, which proceeds at the rates of 1.04–1.55 mg TPH/g soil-day, is completed in about a week with up to 83 % efficiencies. In contrast, diesel degradation in soil sprinkled with water (60 ml/day) proceeds at the rate of 0.3 mg TPH/g soil-day that achieves 15–22 % degradation efficiencies. The addition of 3 % bioagent yields desired soil moisture content (10–15 %), soil pH (6.8–8.2), and nutrition inputs. Both Sporolactobacillus sp. and Prevotella sp. grown on molasses are robust, tolerant high diesel doses, able to utilize hydrophobic hydrocarbons, and readily adaptable to the soil environment. Most notably, prior acclimatization is not required to enable these properties.




Similar content being viewed by others
References
Abioye OP (2011) Biological remediation of hydrocarbon and heavy metals contaminated soil. In: Pascucci S (ed) Soil Contamination. InTech Europe, Riejka, Croatia, pp 127–142
Abioye OP, Abdul AA, Agamuthu P (2010) Enhanced biodegradation of used engine oil in soil amended with organic wastes. Water Air Soil Pollut 209:173–179
Banat IM, Makkar RS, Cameotra SS (2000) Potential commercial application of microbial surfactants. Appl Microbiol Biotechnol 53:495–508
Bento FM, Camargo FOA, Okeke BC, Frankenberger WT (2005) Comparative bioremediation of soils contaminated with diesel oil by natural attenuation, biostimulation and bioaugmentation. Bioresour Technol 96:1049–1055
Britton L (1984) Microbial degradation of aliphatic hydrocarbons. In: Gibson TD (ed) Microbial Degradation of Organic Compounds. Marcel Dekker, New York, pp 89–129
Chang JJ, Chou CH, Ho CY, Chen WE, Lay JJ, Huang CC (2008) Syntrophic co-culture of aerobic Bacillus and anaerobic Clostridium for bio-fuels and bio-hydrogen production. Int J Hydrogen Energy 33:5137–5146
Choi SC, Kwon KK, Sohn JH, Kim SJ (2002) Evaluation of fertilizer additions to stimulate oil biodegradation in sand seashore mescocosms. J Microbiol Biotechnol 12:431–436
Chou C, Han C, Chang J, Lay J (2011) Co-culture of Clostridium beijerinckii L9, Clostridium butyricum M1 and Bacillus thermoamylovorans B5 for converting yeast waste into hydrogen. Int J Hydrogen Energy 36:13972–13983
Gentry TJ, Rensing C, Pepper IL (2004) New approaches for bioaugmentation as a remediation technology. Environ Sci Technol 34:447–494
Grishchenkov VG, Townsend RT, McDonald TJ, Autenrieth RL, Bonner JS, Boronin AM (2000) Degradation of petroleum hydrocarbons by facultative anaerobic bacteria under aerobic and anaerobic conditions. Proc Biochem 35:889–896
Han CL, Lay JJ, Shieh WK (2015) Cellulosic fermentation using the Bacillus thermoamylovorans-enabled digested sludge. J Environ Eng ASCE. doi:10.1061/(ASCE)EE.1943-7870.0000990
Kallimanis A, Frillingos S, Drainas C, Koukkou AI (2007) Taxonomic identification, phenanthrene uptake activity, and membrane lipid alterations of the PAH degrading Arthrobacter sp. strain Sphe3. Appl Microbiol Biotechnol 76(3):709–717
Kalme S, Parshetti G, Gomare S, Govindwar S (2008) Diesel and kerosene degradation by Pseudomonas desmolyticum NCIM 2112 and Nocardia hydrocarbonoxydans NCIM 2386. Curr Microbiol 56(6):581–586
Kim S, Choi DH, Sim DS, Oh Y (2005) Evaluation of bioremediation effectiveness on crude oil-contaminated sand. Chemosphere 59:845–852
Korda A, Santas P, Tenente A, Santas R (1997) Petroleum hydrocarbon bioremediation: sampling and analytical techniques, in situ treatments and commercial microorganisms currently used. Appl Microbiol Biotechnol 48:677–686
Kostka JE, Prakash O, Overholt WA, Green SJ, Freyer G, Canion A, Delgardio J, Norton N, Hazen TC, Huettel M (2011) Hydrocarbon-degrading bacteria and the bacterial Community response in Gulf of Mexico beach sands impacted by the Deepwater Horizon oil spill. Appl Environ Microbiol 77(22):7962–7974
Kvenvolden KA, Cooper CK (2003) Natural seepage of crude oil into the marine environment. Geo-Marine Lett 23:140–146
Lay JJ (2013) Restoration of rot tree roots using the supernatant from a mesophilic, molasses-fed anaerobic bioreactor without methanogenesis. Presented at the 15th conference on soil and groundwater remediation, Kaohsiung, Taiwan
Lee K, Park JW, Ahn IS (2003) Effect of additional carbon source on naphthalene biodegradation by Pseudomonas putida G7. J Hazard Mater 105:157–167
Margesin R, Schinner F (2001) Bioremediation (natural attenuation and biostimulation) of diesel-oil-contaminated soil in an alpine glacier skiing area. Appl Environ Microbiol 67:3127–3133
Masakorala K, Yao J, Cai M, Chandankere R, Yuan H, Chen H (2013) Isolation and characterization of a novel phenanthrene (PHE) degrading strain Pseudomonas sp. USTB-RU from petroleum contaminated soil. J Hazard Mater 263(2):493–500
Mnif I, Ellouze-Chaabouni S, Ayedi Y, Ghribi D (2014) Treatment of diesel- and kerosene-contaminated water by B. subtilis SPB1 biosurfactant-producing strain. Wat. Environ Res 86(8):707–716
Mulla SI, Talwar MP, Bagewadi ZK, Hoskeri RS, Ninnekar HZ (2013) Enhanced degradation of 2-nitrotoluene by immobilized cells of Micrococcus sp. strain SMN-1. Chemosphere 90(6):1920–1924
Mulligan CN (2005) Environmental applications of biosurfactants. Environ Pollut 133:183–198
Nadim F, Hoag GE, Liu S, Carley RJ, Zack P (2000) Detection and remediation of soil aquifer systems contaminated with petroleum products: an overview. J Pet Sci Eng 26:169–178
NIEA E203.56B., NIEA M165.00C., NIEA S703.62B (2013) Environmental Analysis Laboratory, Environmental Protection Administration, Executive Yuan, Republic of China
Ortega-González DK, Zaragoza D, Aguirre-Garrido J, Ramirez-Saad H, Hernández-Rodriguez C, Jan-Roblero J (2013) Degradation of benzene, toluene, and xylene isomers by a bacterial consortium obtained from rhizosphere soil of Cyperus sp. in a petroleum-contaminated area. Folia Microbiol 58:569–577
Paudyn K, Rutter A, Rowe RK, Poland JS (2008) Remediation of hydrocarbon contaminated soils in the Canadian Arctic by landfarming. Cold Reg Sci Technol 53:102–114
Pineda FG, Boll AG, Lira GC, Mesta HAM (2004) A microbial consortium isolated from a crude oil sample that uses asphaltenes as a carbon and energy source. Biodegradation 15(3):145–151
Pirnik MP, Atlas RM, Bartha R (1974) Hydrocarbon metabolism by Brevibacterium erythrogenes: normal and branched alkanes. J Bacteriol 119:868–878
Rahman KSM, Thahira-Rahman J, McClean S, Marchant R, Banat IM (2002) Rhamnolipid surfactants production by strains of Pseudomonas aeruginosa using low cost raw materials. Biotechnol Prog 18:1277–1281
Shen Y, Stehmeier LG, Voordouw G (1998) Identification of hydrocarbon-degrading bacteria in soil by reverse sample genome probing. Appl Environ Microbiol 64:637–645
Vidali M (2001) Bioremediation: an overview. Pure Appl Chem 73:1163–1172
Wasilkowski D, Swedziol Z, Mrozik A (2012) The applicability of genetically modified microorganisms in bioremediation of contaminated environment. Chemik 66:817–826
Youssef N, Simpson DR, Duncan KE, McInerney MJ, Folmsbee M, Fincher T, Knapp RM (2007) In situ biosurfactant production by Bacillus strains injected into a limestone petroleum reservoir. Appl Environ Microbial 73(4):1239–1247
Acknowledgments
This project was financially supported by the National Science Council, Republic of China (Project Number: NSC 101-2221-E-327-008-MY2). The assistance from Professor Yu Z. Huang of Chun Yuan Christian University (Chungli, Taiwan) in performing the PCR analyses was greatly appreciated.
Author information
Authors and Affiliations
Corresponding author
Appendix 1: PCR and RT-PCR procedures (Chang et al. 2008; Chou et al. 2011)
Appendix 1: PCR and RT-PCR procedures (Chang et al. 2008; Chou et al. 2011)
For PCR tests, DNA was extracted using FastDNA™ Spin Kit for Soil (MP Biomedicals, USA). The samples were processed following the procedures specified in the vendor manual. Table 3 list the primers sets used. Each reaction was performed with a mixture of 3 μl of template DNA, 2 μl (10 μM) of each primer set, 33 μl of double-distilled water and 10 μl of PCR Master Mix (BioKit, Taiwan). The thermal cycling program was: 2 min at 95 °C, 30 30-s cycles at 95 °C, 30 s at 54 °C for Prevo F/R and Sporo F/R (50 °C for the Clos L1F/R and Clos E1F/R), 30 s at 72 °C, and the program was terminated at 72 °C for 10 min. PCR results were checked with gel electrophoresis.
The fluorescent dye, SsoFast™ EvaGreen® Supermix, was used for real-time PCR (RT-PCR) reactions using the procedure specified in the vendor’s manual (Bio-Rad, Hercules, USA). Reactions were performed in a 48-well optical reaction plate and set into a 48-well MJ Mini™ Gradient Thermal-Cycler (Bio-Rad, Hercules, USA). Primers sets are listed in Table 3. Each reaction was performed in triplicate with a mixture of 2 μl of cDNA, 1 μl (3–5 μM) of each primer set, 2 μl of double-distilled water and 5 μl of enzyme Supermix. The thermal cycling program was: 60 s at 95 °C, 40 5-second cycles of at 95 °C, 5 s at 54 °C for Prevo F/R and Sporo F/R (or 50 °C for Clos L1F/R and Clos E1F/R) and followed by a dissociation stage (i.e., 15 s at 95 °C, 15 s at 65 °C, and followed by a slow temperature rise to 95 °C). Neither primer-dimer artifacts nor nonspecific PCR amplicons were observed during the melting process. The efficiency of each reaction was between 90 % and 110 %. R2 values for all standard curves were greater than 0.99.
Rights and permissions
About this article
Cite this article
Lin, Y., Lay, JJ. & Shieh, W.K. Diesel degradation in soil catalyzed by the addition of a bioagent. Int. J. Environ. Sci. Technol. 13, 551–560 (2016). https://doi.org/10.1007/s13762-015-0889-8
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s13762-015-0889-8