Abstract
The decoding of the human genome, completed two decades ago, marked a revolutionary moment in biology by introducing a vast amount of data. This avalanche of information presented several computational challenges. Machine Learning has become the dominant method to address these challenges, with Natural Language Processing playing a significant role and offering promising results. In this systematic review, we will explore the application of Machine Learning and Natural Language Processing to the study of biological data. On the one hand, Machine Learning is widely used in Artificial Intelligence to improve automation, carry out tasks that require no human interaction, and perform analytical and physical activities. It helps advance our understanding of biology and improve healthcare and drug development processes in bioinformatics. On the other hand, improved machine-human language interaction is the aim of Natural Language Processing. Its three main goals are character sequence processing, pattern recognition, and algorithm development. The use of Natural Language Processing is becoming increasingly important for the analysis of omics data using both modern and conventional Machine Learning models, underscoring the necessity for a systematic review. In this work, 82 studies were included following the PRISMA guidelines, sourced from PubMed, Scopus and IEEE Xplore on April 4th, 2023. The evaluation of the publications was based on the type of the studied biological data and the employed NLP techniques. Through our in-depth exploration of NLP approaches, we highlight their significance and potential in advancing the field of bioinformatics.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
1 Introduction
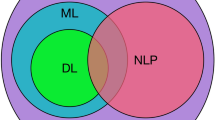
To evaluate and interpret biological activities, the interdisciplinary area of bioinformatics applies approaches from the sciences of biology, computer science, chemistry, physics, and mathematics (Sitaraman 2009). Bioinformatics has advanced quickly since the decoding of the human genome in April 2003 because of the significant amount of biological data that has been acquired and will continue to be collected. The development of high-throughput technologies, which enable the simultaneous measurement of millions of biomolecules throughout an experiment, is the reason behind the creation of Bioinformatics as we know it today. Omics data are the biological data acquired using such technologies (Allen & Cagle 2008). They are divided into distinct categories based on the kind of biomolecule being studied. The creation of effective study tools is seen to be important given the fast advancement of omics technology. To provide cutting-edge techniques and tools for the effective analysis of omics data, the discipline of artificial intelligence can provide a multitude of solutions to the challenges faced by bioinformatics. Artificial intelligence has become a cornerstone of omics data analysis as a result of the development of methodologies based on Machine Learning (ML) and Deep Learning (DL) techniques in particular, which is still evolving (Quazi 2022). The most typical forms of biological data include sequences, 3D structures, gene expression data, metabolic pathways, gene mapping, phylogenetic trees, and polymorphisms, depending on their kind, size, and complexity (Dall’alba et al. 2022). The rise of bioinformatics is a result of new, quicker, and more effective technologies taking the place of conventional molecular procedures (Manzoni et al. 2018). Omics technologies, which are high-performance analytical techniques, offer a solution to this issue. Omics scientific fields have been developed based on the types of data produced, with the most well-known being 1) genomics, 2) transcriptomics, 3) proteomics, and 4) metabolomics (Dai & Shen 2022). Since the progression of omics disciplines is intricately linked to the fundamental principles of biology, the development of omics technology greatly facilitates the comprehension and expansion of biological knowledge (Jung et al. 2020).
A subtype of AI called machine learning allows the development of programs that are not strictly programmed. Machine learning is the process of creating algorithms that can learn from experimental data to either make accurate predictions or reach insightful conclusions. The capability to utilize a vast amount of data without requiring processing by an outside user is this field’s main advantage. There are three basic approaches to machine learning: supervised, unsupervised, and reinforcement learning, depending on the subject of research, the data, and the nature of the problem (Le Glaz et al. 2021). The most used machine learning subfield is Deep Learning (Naskath et al. 2023), which enables the development of algorithms that require a substantial amount of training data to produce reliable predictions. Essentially, its reasoning is built on a neural network with three or more layers. It works based on the way the human brain functions, which takes an input, decodes it, and produces specific outputs. They are made up of many layers of synthetic neurons with a built-in hierarchical structure. The input, hidden, and the output layer are the three sub-categories that make up a Neural Network. Each layer is created by nodes that are connected along weighted edges. The term “deep” refers to the huge number of hidden layers that allow for intricate interactions between the input data. This is accomplished through the training process, which optimizes the neural network’s capability to choose the optimum course of action considering the input data. Since the model can self-adjust its parameters, this optimization is carried out without any human involvement.
The second subtype of AI we focused on was Natural Language Processing (NLP). The study of NLP is interdisciplinary, straddling the disciplines of artificial intelligence and linguistics. It entails the creation of software tools that can decode, process, and produce massive amounts of natural language data. Natural language analysis is highly challenging due to the intricate semantics of natural languages. In these languages, the overall context of the text is as crucial as the word order in determining a sentence’s meaning, as the same sentences can easily convey different meanings depending on the context. Although there isn’t clear evidence of semantics between biomolecules in biology, in this review we’ll demonstrate that such methods can produce some extremely interesting outcomes. Our main purpose was to find out how NLP algorithms have been exploited in Bioinformatics, initially in the form of abstracts in PubMed, up to nucleic acid and protein sequences. The studied methodologies we found are based on word2vec (Mikolov et al. 2013) and transformers (Vaswani et al. 2017) algorithms.
1.1 The importance of embedding methods on biological data
Natural Language Processing (NLP) stands at the intersection of linguistics and artificial intelligence, focusing on the development of algorithms capable of interpreting, analyzing, and generating extensive volumes of natural language data. The inherent complexity of natural language semantics poses a formidable challenge in NLP. It’s not merely the word order that influences meaning, but also the broader contextual framework within which the words are situated. Consequently, the same sentence can convey varying meanings based on the context.
1.2 Word2vec
Since natural language texts cannot be directly supplied to neural networks without some kind of mathematical adjustment, word2vec is built on this concept. The production of n-dimensional vectors, often known as word embedding in the literature, is one of the most widely used methods for converting text sequences into sequences of numbers. The issue that this technology has been able to solve is the proper embedding words’ semantic information. The terms king, queen, woman, and man are the most prevalent illustration of the embedding principle. Mathematically, we anticipate that the phrases “king” and “queen” will be analogous to the words “man” and “woman” in their proximity. The association was discovered by Mikolov et al. as the following formula:
vector(“King”)-vector(“Man”) + vector(“Woman”) = vector(“Queen”)
The term “embedding” refers to the process of converting data into vectors. The word2vec model is made up of three fields: a) an input field, b) an output field, and c) a hidden field (embedding field). The inclusion of the softmax activation function in the output field, which captures the probability of a word or its contexts, is a unique characteristic of these models. Depending on the study goal, the word2vec model uses two different approaches (Fig. 1).
-
the CBOW (Continuous Bag-of-Words) method, where each word is predicted based on its contextual set and
-
the reversed approach of Skip-Gram method, where the set of contexts is predicted given the word.
Depending on the chosen approach, training examples in the form of word-context pairs (CBOW) or context-word pairs (Skip-Gram) need to be generated. These examples are used to evaluate whether the neural network accurately predicts the word or the context of words through a loss function. The training process follows a standard approach for neural networks, assessing performance using a loss function, typically the cross-entropy loss function. Backpropagation is employed using the stochastic gradient descent method. The word2vec method is unsupervised, as the embeddings are derived by the model itself, without prior knowledge of the correct embeddings.
1.3 Transformers
Transformers were introduced in 2017 by Vaswani et al. as a solution to the challenges faced by traditional methodologies like Recurrent Neural Networks (RNNs) (Rezk et al. 2020) when processing natural language texts. Issues such as limited parallelization during training, difficulty handling long-word relations due to memory constraints, and fixed sentence length limitations were effectively addressed with transformers by introducing the concept of self-attention.
Self-attention revolutionized the way associations between tokens in a sequential sequence (e.g., English text) are identified, enhancing the ability to capture meaningful relationships. The architecture of a transformer is typically divided into two primary components: the encoder and the decoder (Miltiadous et al. 2023).
According to Fig. 2, we will try to outline how a transformer operates:
-
A.
Input: The initial step involves vectorizing the input data, typically text, to create the input embeddings.
-
B.
Positional Encoding: Since transformers lack recurrent or convolutional structures, positional encoding is necessary to convey the positions of units in the sequence. This encoding assigns positions to units within a vector space with dimensions similar to the input dimensions.
-
C.
Encoder: The embedded inputs, along with positional encodings, are fed into the encoder, comprising two main layers: Multi-Head Attention (MHA) and a Feed-Forward Neural Network (FNN). The MHA layer determines the relevance of each unit based on interactions with other units. The output of MHA is then passed to FNN, which introduces non-linear transformations. Crucially, residual connections (bypass connections), akin to those in RNNs, preserve the original information from the input vectorization. The outputs from the encoder layers are normalized.
-
D.
Decoder: The decoder generates predictions by utilizing its stored information and incorporating input from the encoder. The Masked Multi-Head Attention layer ensures that information from the encoder is appropriately isolated in the multi-head attention mechanism, known as encoder-decoder attention. Following this, the process is akin to the encoder, involving an input to an FNN. Similar to the encoder, the decoder incorporates bypass connections and normalizes the outputs.
-
E.
Final Prediction: The output from the decoder serves as the final prediction. It undergoes a linear transformation and is then fed into a softmax function. The role of the softmax function is to produce values ranging from 0 to 1, representing the probability of each unit being the correct prediction. These probabilities indicate the confidence levels associated with each unit’s prediction.
In summary, embedding methodologies assist bioinformaticians in processing diverse biological data types, including nucleic acids, proteins, small organic or inorganic chemical compounds, and gene ontologies. These methodologies, such as word2vec and transformers, convert the input data into vectors, enabling the extraction of relevant features in a format recognizable by machine learning algorithms (Fig. 3). The claimed superior results in training and optimization across the studied methodologies are attributed to this effective embedding process, as well as the predictive potency of machine learning models (S. Sharma & Singh 2022). Notably, even in the context of speech disorders (Anthony et al. 2022) or speech emotion recognition (Al-Dujaili & Ebrahimi-Moghadam 2023), impressive performance has been achieved.
2 Materials and methods
This work aims to explore NLP methods, specifically focusing on the application of 2vec algorithms in the field of Bioinformatics. The PRISMA protocol served as the foundation for our research because it offers precise instructions for conducting systematic reviews (Page et al. 2021). The methodology for this systematic review has been registered on PROSPERO and can be accessed using the designated PROSPERO ID, which is CRD42023459405 (Fig. 4).
Our search was constructed on a base of three highly utilized and reliable sources of scientific knowledge: MEDLINE PubMed, Elsevier Scopus, and IEEE Xplore. We utilized two separate queries since PubMed doesn’t use the “*” symbol as a wildcard in front of the requested endings, like the other two databases. In PubMed, we employed word2vec search terms based on the most widely recognized methodologies to the best of our knowledge.
Thus, the different queries are:
PubMed: (“Word2vec” OR “Doc2vec” OR “DNA2vec” OR “Protein2vec” OR “Graph2vec” OR “Pathway2vec” OR “Bio2vec” OR “Phe2vec” OR “Drug2vec” OR “Node2vec”) AND (“neural network” OR “deep learning” OR “machine learning”) AND (“DNA” OR “protein” OR “sequence” OR “GO” OR “gene ontology” OR “gene” OR “biological data” OR “health”) AND (“semantic” OR “embedding”).
Scopus και IEEE Xplore: (“*2vec”) AND (“neural network” OR “deep learning” OR “machine learning”) AND (“DNA” OR “protein” OR “sequence” OR “GO” OR “gene ontology” OR “gene” OR “biological data” OR “health”) AND (“semantic” OR “embedding”).
Our search was performed on April 4, 2023, for titles, abstracts, and keywords (Title/Abstract/Keywords). In total, we identified 591 studies in English, of which 416 belong to Scopus, 105 to Pubmed and 70 to IEEE Xplore. We used the Rayyan program (Ouzzani et al. 2016) to organize these studies.
Rayyan greatly aids in automating the systematic review process because it recognizes duplication automatically. A manual inspection resulted in the deletion of 166 articles in total, from which the 165 publications were discovered automatically. In addition, we discovered another duplicate publication with the wave2vec method that falls within the exclusion category. We attribute this to Yuan et al. conference publication (Yuan et al. 2017) before they published their work in the neurocomputing journal (Yuan et al. 2019). Then, after screening 425 studies (591−166 = 425) for their applicability to our study question, we eliminated 234 publications by looking at their abstracts only. Out of the 191 articles, seven were inaccessible without payment. We ultimately identified 82 papers out of the 184 publications we attempted to assess for eligibility.
Our excluded criteria are the following:
-
1)
Non-biological data: Refers to publications that only use text (such as article summaries), texts from social media, audio files, video files, etc. instead of biological data.
-
2)
Comparative machine learning studies: These are articles that evaluate several machine learning techniques without recommending a particular 2vec technique or learning framework.
-
3)
Additional reviews (systematic, literary, or meta-analyses).
-
4)
Publications that are unrelated to human health.
-
5)
Publications that are used as NLP tutorials.
-
6)
Studies about the best method of data pre-processing that optimizes the performance of current models.
-
7)
Studies that serve as a standard for other methodologies by using 2vec techniques.
-
8)
Publications using techniques outside of the NLP field.
As we’ll see in the following section, we divided the 82 publications into the following three subcategories, based on the year of publication and the following criteria:
-
1.
Biological inputs (e.g., DNA sequences, RNA, proteins, drug compounds, ontologies, and their interactions based on graphs) which are related to significant biological processes such as gene regulation and protein–protein interactions.
-
2.
Embedding methods used.
-
3.
Employed neural network architectures.
3 Results
Our study revealed that 82 papers in total, or 19.2% (82/425) of the initially retrieved publications, were connected to the search. We made a table (Table 1) displaying all publications and the key elements of their techniques to make the data visualization process easier. Although the word2vec methodology was published in 2013, it is evident that the year when 2vec methodologies started to be utilized in our study field was only 2018. The publications listed in these tables will be presented in detail below, organized by year.
3.1 2018
We can observe that there were only 7 studies published in 2018:
The Deep-2′-O-Me model was developed from (Mostavi et al. 2018) by creating a new 2vec methodology called rna2vec based on the dna2vec method (Ng 2017). Their objective was to predict the RNA methylation at the 2′ hydroxyl (–OH) of the ribose moiety (2′-O methylation sites). They used a Convolutional Neural Network (CNN) (Albawi et al. 2018) classifier followed by a Feed Forward Neural Network (FNN) (Sazli 2006) to get an AUC value of 0.90. The identification of such places can aid in the early diagnosis of diseases and the creation of more potent drugs, hence this model may be an effective tool for understanding gene regulation.
At the DNA stage, two different research teams investigated gene regulation via the transcription factor binding sites (TFBS) interactions on the DNA molecule. The team of (W. Zeng et al. 2018) used a type of RNN called Gated Recurrent Neural Networks (GRU) (Chung et al., 2014) to predict binding sites, and they were able to do so with predictions of AUC value as high as 0.9649. The embedding method they used was word2vec. A different approach was used by the second team of (Z. Shen et al. 2018), where the interaction of two DNA regions, the promoter and enhancer, is investigated. They created a novel model to predict promoter—enhancer interaction sites (EPI) called EP2vec, based on known EPIs. EP2vec is divided into two phases. Through the use of the paragraph vector, the region under investigation is first transformed into vectors, and then the region is divided into EPI and non-EPI regions. A type of classification tree called GBRT was employed as the classifier. Once more, the proposed model’s F1-score reached 0.933.
The research teams of (Smaili et al. 2018), (Kim et al. 2018) and (Jaeger et al. 2018) also created their 2vec methodologies for a different purpose in each case. The first team used word2vec to exploit the information of gene ontologies in protein interaction networks. To validate the capabilities of their model, they used three machine learning classifiers (Logistic Regression – LR (Peng et al. 2002), Support Vector Machines – SVM (Schölkopf 1998)and Neural Networks—NN) where SVM and NN had improved AUC values for differentiating protein interactions (reaction, activation, binding, and catalysis) from the STRING database (Szklarczyk et al. 2023). The data on cancer genetic mutations was combined by the second study team with information on protein interactions and abstracts from PubMed. Protein interactions and associated data from PubMed were used to improve the quality of their missing data (NaN). They wanted to be able to separate driver mutations from passenger mutations. The word2vec methodology is used to generate vectors. They were able to successfully discern between the two categories by utilizing Principal Component Analysis (PCA) (Jolliffe & Cadima, 2016) to visualize their data. Furthermore, they were able to Identify mutation vectors that have been identified as potential mutation leads in prior investigations. Therefore, they claim that their Mut2vec model can also identify unknown mutations, serving as a roadmap for future cancer research. Finally, a novel 2vec model that can translate the structure of tiny chemical molecules into words was created by Jaeger et al. The Morgan algorithm (Rogers & Hahn 2010), which converts each atom of a chemical compound into a numerical value called identifiers, forms the basis for applying NLP techniques to molecular structures. Small chemical compounds are seen as words by Word2vec, and complexes are seen as sentences. Consequently, this model is known as Mol2vec and is based on the word2vec approach. By combining their model with the ProtVec, they developed a new methodology called PCM2vec, through which the characteristics of protein complexes are obtained to better understand the interactions between proteins and other chemical molecules. With Random Forest as a classifier, they found a maximum AUC value equal to 0.95.
Heterogeneous graphs were employed as the input data in the work by (Zhu et al. 2018), which utilized more complicated input data and examined them in terms of drug safety. Heterogeneous Information Networks (HINs) were employed for this, and their nodes were divided into the following categories: a) Drug-drug similarity, b) gene similarity, c) drug-gene relationship, and d) drug side effect (Adverse Drug Reactions, ADRs). The node2vec technique (Grover & Leskovec 2016) which turns node relationships into vectors based on the word2vec approach, is the foundation for the metapath2vec and metapath2vec + + methodologies, which extract the graph features. Additionally, they used Kernelized Bayesian Matrix Factorization (KBMF) (Gönen et al. 2013) to enhance their model’s performance. They claim that their algorithms have an AUC of 0.8093 for metapath2vec and 0.8367 for metapath2vec + + .
3.2 2019
A total of 10 publications from the year 2019 were discovered, of which 3 were about graphs, 1 about EEG data, and the other 6 for biological sequences.
The research teams by (M. Zeng et al. 2019), (F. Shen et al. 2019), and (Z. Gao et al. 2019) used graphs. The first study involved the development of a novel technology called DeepEP, where they employed a CNN to analyze patterns in gene expression data and a node2vec model to understand the properties of protein–protein interaction graphs (PPI networks). These data were extracted as an image and the outcomes were combined with information from related graph proteins. An FNN network then receives the combined output as input, where it processes proteins that are discovered to be significant in metabolic processes. Their approach resulted in an AUC value of 0.8200. F. Shen et al. collected information from three separate animal illness databases. Their goal was to develop a technique that would produce thorough human phenotype ontologies (HPO). They built heterogeneous graphs for each base to achieve this, and then, once more using the node2vec approach, they deduced the relevant vectors for each graph. They employed 6 machine learning algorithms (Decision Tree—DT (Rokach & Maimon 2006), Logistic Regression—LR, SVM, Random Forests—RF, Naïve Bayes—NB (Rish, 2001) and Multiple Layer Perceptron—MLP) to assess the effectiveness of their methods, and they obtained AUC values of 0.81, 0.80, 0.90, and 0.92 for 4 different types of graphs, HPO-original (a graph the authors constructed in previous work), HPO-DECIPHER, HPO-OMIM, and HPO-ORPHANET. The third study group (Z. Gao et al.) assert that they developed a new technique for assessing graph features that, in contrast to node2vec, makes use of edge information as well as that of nodes. Edge2vec, their proposed approach, was used to produce an AUC value of 0.8914 using an SVM classifier.
Using a novel proprietary technique called Gene2vec that is based on word2vec, the research team of (Du et al. 2019) was able to convert co-expression data of all human genes into vectors. This approach makes it easier to classify genes and research their relationships and functions. An FNN with an AUC value of 0.720 was used to categorize the genes. With a similar approach, (Zou et al. 2019) also developed a -different- Gene2vec method to investigate mRNA sequences with m6A-type methylation sites (N6-methyladenosine sites), which are regarded as one of the most significant mRNA ribonucleotide modifications with a variety of biological activities. The produced vectors are passed through a CNN, whose output data are then supplied into a FNN. The researchers demonstrated that their model had reached AUC values equal to 0.841.
The 16 s rRNA sequences, which are frequently employed for phylogenetic purposes, are another kind of RNA that is of great importance. Researchers attempted to turn rRNA sequences into vectors in the study by (Woloszynek et al. 2019) to illustrate the various interactions between rRNAs that reflect phylogenetic relationships between bacteria. They were able to attain a maximum accuracy of 0.979 using a Multi Lasso Classifier.
Investigations on protein interactions are a crucial area of study. By creating a word2vec-based model called bio2vec, the research team of (Y. Wang et al. 2019) aimed to embed protein sequences to discover these connections. To do this, they trained their algorithm using both sequence data and information on protein interactions. The vectors obtained by bio2vec are utilized as input for the CNN network, which performs the prediction. The results are then seen using a FNN classifier. The precision of their suggested methodology is up to 0.9731.
Research with a shared interest was carried out by (Yao et al. 2019) who built Res2vec, a novel embedding technology based on word2vec. The Res2vec algorithm vectorizes proteins according to the physicochemical characteristics of each residue, and its output is then sent to a FNN network, which carries out the specified categorization of probable and improbable protein interactions. The name of their combined methodology is DeepFE-PPI.
The word2vec algorithm is also used in pharmacology. A thorough search for peptides suited for cancer treatment is conducted in the work of (C. Wu et al. 2019) using their methodology, PTPD (Prediction Therapeutic PeptiDes). They trained the word2vec algorithm to be able to extract the increased features of the peptides they are looking for, known as ACPs (AntiCancer Peptides), through a CNN network. A FNN classifier divides the output data from the CNN into ACP and non-ACP categories. They reported an impressive AUC value equal to 0.99.
In contrast with what we have observed so far, the article by (Yuan et al. 2019) addressed biosignals rather than biomolecules. The researchers developed the wave2vec approach to turn these data into vectors, and they successfully showed that it can be used to extract biosignal properties and perform automatic pathological condition detection. By correctly categorizing the data from word2vec into an autoencoder (SAE) and importing it, their declarations are verified (highest AUC value: 0.9968).
3.3 2020
Continuing our analysis, we see that 12 studies have been published in 2020. Specifically:
Word2vec is used to identify enhancers in DNA sequences in the study by (Khanal et al. 2020) and through a CNN network, they attempted to determine the intensity of each enhancer. They used DNA sequences from GenBank (Benson et al. 2013) to train the model they created, known as iEnhancer-CNN. They discovered that the FNN classifier performed much better than other existing methods at accurately detecting enhancer sequences and their strength.
In contrast, S. Yang et al. (S. Yang et al. 2020) investigated the relationships between LncRNA and miRNAs. To accomplish this, they used doc2vec (Lau & Baldwin 2016) to do feature extraction for RNA sequences (lncRNA and miRNA), as well as the methodologies of k-mer ((B. Liu et al. 2015) and CTD (Tong & Liu 2019). They created a graph using the Linear Neighborhood Similarity (LNS) (Zhou et al. 2019) approach based on these properties, with lncRNA and miRNA serving as the nodes. These graphs are embedded using role2vec in the last stage of the LncMirNet model before being imported into a CNN. Then, through a FNN (AUC: 0.9381), they determined if the model predictions were accurate or inaccurate (i.e., whether there are or are not interactions between lncRNA and miRNA). Additionally, by creating a novel methodology called kmerPR2vec, (Asim et al. 2020) investigated the topology of miRNAs. They created a novel technique because other NLP methods did not considerably improve the performance of the algorithms regarding miRNAs. Their approach involved creating random k-mers of miRNA sequences, which they subsequently encoded to identify any semantic correlations (like in transformers). The final vectors are created once this information is included in each k-mer. They found that these vectors improved the performance of the CNN. Through a FNN classifier, they confirmed the high efficiency of their model after reaching a better performance compared to the available algorithms. (C. Wang et al. 2020) conducted another investigation into RNA molecules and focused on the ITS (Internal Transcribed Spacer) portions of rRNA from fungal cells, which are unique to different fungus species and are used for phylogeny. Their model, known as Its2vec, employs word2vec to extract the crucial ITS region traits that result in the accurate classification of each species. An RF model carrying out the final classification has an accuracy of 0.9753.
Passing on gene ontologies (GO), OntoSem, a tool that identifies protein–protein interactions (PPI) based on GO, was created by (Zhao et al. 2020) as a result. To create vectors of similar length, OntoSem employs the word2vec algorithm and BERT, whose outputs are sent through two distinct RandLSTM (Randomly Long-short Term Memory) networks. The RandLSTMs outputs are fed into a CNN model, which generates the final features used by the FNN to accomplish the final classification. The model used by the researchers displayed the best performance, with a maximum AUC value of 0.993. For the same purpose, (Buchan & Jones 2020) utilized word2vec to integrate into vectors the information accessible in Pfam about the self-contained structural domains, as well as their corresponding gene ontologies. They tested their model’s performance in identifying connections between Pfam domains using kNN. The problem, however, is that the metrics of their model are low. For k = 1, the model reached mean precision and recall of 0.33 and 0.30 respectively, while for k = 10, the mean precision was 0.12 and the mean recall 0.57.
Furthermore, (Pan et al. 2020) investigated the role of toxic proteins in the development of genetically modified plants, as well as drugs that will deal with such toxic substances when introduced into the human body. To accomplish this, they developed the ToxDL model, which combines a CNN network and a word2vec model that was trained on protein domains (domain2vec) to extract features in parallel. These features are fed into a FNN network for classification, which calculates the likelihood of protein toxicity (AUC: 0.989 ± 0.002). (Y. F. Zhang et al. 2020) conducted pharmacology research focusing on identifying the characteristics that drive interactions between small chemical compounds and their target proteins. For quicker develop of novel drugs, pharmacologists essentially need to understand how proteins interact with the binding sites in cells. As a result, they created the SPVec method, based on word2vec, to achieve this goal, which turns the interactions between target proteins and their small compounds into vectors. They discovered that the vectors contain information about the physicochemical properties of the molecules, making it feasible to determine the nature of the forces that drive drug-target interactions (AUC: 0.9927).
In a similar rationale, (Thafar et al. 2020) sought to predict the interactions between drugs and their receptors to foresee potential negative effects of the drugs in question. The approach they took is based on the node2vec method, in which features of a heterogeneous network made up of known drug-receptor interactions (DTI), drug-drug interactions, and receptors are extracted. They later enriched these features with information about the optimum graph paths and fed them into an RF model. They assert that the average AUPR for their model is 0.831.
Using miRNAs, proteins, lncRNAs, drugs, and other molecules, (Z. H. Guo et al. 2020) created a heterogeneous graph (MAN). Despite having a vast quantity of information stored, a graph of this type is particularly challenging to examine using traditional graph analysis techniques. For this reason, they employed the node2vec approach to turn each node into a vector per the associations determined from MAN. With an AUC of 0.9677 ± 0.0007, they employed an autoencoder (SAE) and RF for feature analysis and classification respectively. Following the same logic, (N. Wang et al. 2020) employed protein interaction networks to discover details about proteins that are necessary for the normal development of organisms. Thus, they created a network of these proteins with 1285 nodes, which they then utilized as input for the node2vec algorithm. They succeeded in attaining an AUC value of 0.8200 using an XGBoost classifier (Chen & Guestrin, 2016).
In contrast to these approaches, X. Yang et al. (X. Yang et al. 2020) attempted to analyze protein interactions by looking at the primary structure of proteins rather than by creating graphs. They used the doc2vec approach to create r vectors with the key features identified. They argue that their model accurately predicts the known interactions of the human and viral proteins based on the sequences of the data they used. A RF classifier achieves an AUC value of up to 0.9810 when utilizing the doc2vec data.
3.4 2021
Moving on to the year 2021, we see that the number of publications increased from 11 in 2020 to 18:
So far, we have seen many attempts to study heterogeneous and homogeneous graphs by converting their nodes into vectors. In the study by (Basher & Hallam 2021) the scientists used multi-layer heterogeneous graphs that incorporate metabolic pathways (pathway layer), chemical compounds (compound layer), and enzymes (enzyme layer), all of which are documented in the MetaCyc database. They created a package called pathway2vec, which uses 5 embedding models, including the already-existing node2vec, metapath2vec, metapath2vec + + , JUST (Hussein et al., 2018), and their RUST approach, to successfully convert the graph into vectors. On multi-level graphs, they discovered that combining these models improved the accuracy of their predictions. They employed the mlLGPR approach to classify interactions, with metapath2vec having the lowest Hamming Loss value (0.0412) on test data from Leish Cyc.
Another approach for protein function analysis is the methodology of (F. Zhang et al. 2021) They employed DeepWalk to analyze protein interaction graphs and word2vec for protein sequences. The word2vec output is fed into a multi-CNN network that extracts features by individual segments of the protein sequences (local features) and a BiLSTM network that detects the necessary features along the entire sequence (global features). After the extraction of these features, a FNN is used to discover the roles of the proteins. The DeepGOA model has an AUC value of 0.9760 for proteins with the cellular component (CC) as the gene ontology term. An analogous approach was taken by (M. Zeng et al. 2021), who used three distinct types of inputs: a) protein interaction graphs from which features are extracted by node2vec, b) gene expression data investigated via a BiLSTM network, and c) topological features used as inputs to an FNN. These traits are fed into a final FNN, which will locate proteins required for cell function with an accuracy of 0.850.
The research team led by (H. Liu et al. 2021) employed the node2vec technique to extract protein characteristics associated with genes contributing to the physiological processes involved in the development of Cerebral Ischemic Stroke (ICH). They employed an SVM classifier to determine the genes associated with the disease and an SAE to further reduce the output of the embedding techniques. The lack of discrete performance values in their model is interesting, as their results are only shown as figures. The performance of the model varies depending on the values of the hyperparameters p and q. By adjusting the return parameter (p) and the in–out parameter (q), we can influence two aspects of a random walk on a graph. Firstly, we can modify the probability of revisiting nodes during the walk (p), determining whether the walk is more inclined to backtrack (bigger values) or move forward (lower values). Secondly, we can shape the exploration behavior, choosing between exploring nearby nodes (local for q > 1) or venturing to more distant ones (global for q < 1). The highest AUC value is approximately 0.73 for p = 0.1 and q = 10. In the case of L. Wang et al. (L. Wang et al. 2021), a cell–cell interaction network based on scRNA sequencing data was analyzed. They classified the node2vec vectors using the K-means algorithm to determine which genes are associated with one another to demonstrate that their methodology is effective. The researchers built six distinct networks encompassing genes implicated in the development of melanoma based on the six created clusters. Finally, (Ovens et al. 2021) tried to effectively compare Gene Co-Expression Networks (GCNs) graphs. Their model, Juxtapose, is based on the word2vec. According to the researchers, it performs better than models with comparable goals such as MAGNA + + , IsoRankN, and MUNK.
Using the dna2vec approach, (Cheng et al. 2021) tried to discover semantic connections between DNA sequence and transcription factor binding sites to examine gene expression. Their CapBind model emphasizes the utilization of dna2vec data, leveraging diverse networks such as Capsule Neural Network (CapsNet) (Sabour et al., 2017) BiGRU, and CNN to effectively extract optimal features generated by dna2ve. An FNN network (AUC: 0.8940) classifies whether a transcription factor binding region exists or not. In another research investigation on DNA sequences, (Ji et al. 2021) addressed the distinction between super-enhancer (SE) sequences and typical enhancers. They used a methodology that included the following four steps: 1) creating 4,5 and 6-mer non-overlapping DNA segments (k = 4,5,6), 2–3) transforming the entire mouse and human genomes into vectors via dna2vec, and 4) training a CNN to locate the requested motifs and classifying the results through an FNN network (maximum AUC: 0.9600).
The research team led by (L. Zhang et al. 2021) employed three different techniques to transform sequences into vectors: word2vec, RNA embedding, and one-hot encoding (OHE). The explanation behind employing so many approaches is that each method introduces unique variations to the data, thereby offering opportunities for mutual reinforcement and complementarity among them. A BiLSTM network, which is the first stage of a FNN network, is fed data from the output of each algorithm. The model they developed is called EDLm6APred and its prediction is obtained after considering the three different predictions as an ensemble vote. Their model was able to predict methylated RNA (N6-methyladenosine, m6A) with a maximum AUC value of 0.8660.
Another very important modification at the DNA sequence level is N4-methylcytosine. The 4mCNLP-Deep model was developed by researchers to locate methylation sites in the C. elegans genome (Wahab et al. 2021). The model comprises a word2vec stage for feature extraction, followed by CNN and FNN in the traditional neural network sequencing order. They claim that their model has an AUC value of 0.9798 and can distinguish between sequences in 4mC and non-4mC areas. Using a similar approach, Khanal et al. sought to uncover similar modifications in the genomes of two plants belonging to the Rosaceae family (Fragaria vesca and Rosa chinensis), in the same year. They discovered the highest AUC values for k = 3, 0.9400, and 0.9379 for F.vesca and R.chinensis species respectively.
Another study on the genome is the research of (F. Wu et al. 2021) who utilized the word2vec method to be able to identify replication origins (ORIs). The genomes of four different fungi species—Saccharomyces cerevisiae, Schizosaccharomyces pombe, Pichia pastoris, and Kluyveromyces lactis—were gathered for the study. Following a similar approach to the methodologies described earlier, the researchers utilized word2vec vectors as input for a Convolutional Neural Network (CNN) that was trained based on the features derived from word2vec embeddings. An FNN model is then used to categorize the obtained data into ORI and non-ORI sequences. Saccharomyces cerevisiae had the highest AUC value (0.9810) out of the four fungi species. Additionally, (Matougui et al. 2021) employed word2vec to categorize several microbial species according to the traits of their genomes. To accomplish this, they trained a FNN model using 14,000 microbial genomes that were transformed into vectors. Their approach, named NLP-MeTaxa, generates a tree similar to the NCBI tree viewer. On datasets with a medium number of dimensions (medium complexity datasets), their model’s highest accuracy is 0.8359.
The word2vec model was utilized by (Z. Wang & Lei 2021), to turn circRNA sequences into vectors. They were searching for patterns in the sequences of these circRNA molecules that can indicate the presence or absence of RNA binding proteins. An LSTM network is trained to recognize any motifs along the DNA that may indicate their interaction with RBPs using the word2vec data as an input. The classification for interaction sites or not is done using an FNN model, with an AUC value of 0.9570.
The (J. Gao et al. 2021) research team combined the struc2vec, node2vec, and word2vec methodologies into a new method, Protein2vec, which identifies similarities between graphs in the form of vectors. They suggest that their method is more effective than already existing techniques for the study of topological graph homogeneity such as BEAMS, SMETANA, IsoRankN, and NetCoffee. In contrast, (Ali & Patterson 2021) proposed a new 2vec method independent of the word2vec for the study of COVID-19 spike sequences. Spike2vec consists of 3 stages: a) the coronavirus spike glycoprotein (Spike glycoprotein) is generated in all possible k-mers, b) vectors are then developed depending on the frequency of each k-mer inside the sequence, and c) the low dimensional embeddings are generated. They estimated their model using three classifiers (NB, LR, and RC), with LR outperforming the others with an AUC value of 0.6900. In the publication of (Ostrovsky-Berman et al. 2021), the researchers attempted to use the word2vec approach for antibody sequence analysis (BCR). They did this by converting antibody sequences into vectors using word2vec. For the classification task, they employed an RF, which they were able to train with a maximum F1-score of 0.6589.
In the last article for 2021, (Long & Luo 2021) constructed the HNERMDA algorithm to enable them to find specific drugs for different microbial species. They developed a heterogeneous network with two types of nodes—pharmaceuticals and microbial species—and three types of interactions (microbes—microbes, microbes—drugs, drugs–drugs). After learning about each node’s features using the metapath2vec approach, they changed this network into a bipartite network. At last, to successfully find the microbe-drug interactions, they developed the Bias Network Projection Recommendation algorithm (BNPR) to achieve a more accurate conversion of the network relations into vectors, for optimal prediction of microbe-drug interactions. Compared to 5 other algorithms (HerGePred, KATZHMDA, NTSHMDA, IMCHMDA, GCMDR) they acquired the best prediction on two different datasets (MDAD: AUC = 0.9026 ± 0.0062, aBiofilm: AUC = 0.8858 ± 0.0121).
3.5 2022
In 2022, it became evident just how significant the contributions of 2vec techniques had been to bioinformatics, as indicated by a notable 76% increase in the number of publications in comparison to 2021. In particular:
Two more research teams attempted to predict N4-methylcytosine modifications. According to the amount of data used, the research of (Liang et al. 2022) focuses on the development of a model based on the dna2vec approach that is trained to vectorize DNA sequences and feed them into two separate neural network models. These subnets are Hyb_Caps, which addresses species with a smaller volume of data, and Hyb_Conv, which addresses species with more data. The second one uses a CapsNet network and an attention layer to maximize training on the critical feature, while the first uses a CNN. They tested their model on a wide range of species, with the Escherichia coli data showing the highest AUC value (0.9960). The other work (Zulfiqar et al. 2022) focuses on the creation of a Deep-4mCW2V model that is solely dedicated to identifying these alterations in E. coli. In contrast to the initial study, they used a CNN-FNN architecture. They also employed the word2vec method for feature extraction. With an AUC value of 0.962, their model was evaluated using independent data (an independent test dataset).
SplitDNA2vec, a novel embedding technique proposed by (Maruyama et al. 2022), is a modification of the dna2vec algorithm. The key difference between these two techniques lies in their approach to generating k-mers. In SplitDNA2vec, the algorithm generates k-mers that commence at random positions within the sequences under examination, in addition to varying the number of generated k-mers. This fact increases the possible DNA segments and seems to be important for the performance of their model (Accuracy = 0.949). The creation of a BiGRU network was crucial for the effective learning of the required CMIC characteristics for CpG islands, in addition to the embedding methods. For the classification process they used a FNN.
Keeping on with DNA data, the (Liao et al. 2022) research team developed an enhancer prediction algorithm in DNA sequences, iEnhancer-DCLA, using the dna2vec method. In an endeavor to discern patterns within the data, the results obtained from dna2vec were further processed using CNN and BiLSTM networks. These patterns are then enhanced by an extra layer of attention. The final classification is done by the FNN network with an accuracy of 0.7825 for the recognition of enhancer sequences and 0.7800 for the enhancer strength.
Furthermore, the dna2vec approach was applied to two additional research projects. The first is the study of (Cao et al. 2022), where they combined OHE and dna2vec embedding techniques. They also used three different types of neural networks to effectively extract the features that result in transcription factor binding due to the high dimensionality of the data. To discover the underlying DNA patterns, they combined a CNN, a BiLSTM, and an attention layer. They declare that DeepARC, their model, performs with an AUC value of 0.908 on average. The second publication was carried out by (H. Y. Liu & Du 2022), who explored the 5-hydroxymethylcytosine (5hmC) modification in RNA sequences. To achieve their goal, they trained an SVM classifier and the dna2vec model to extract the proper features from RNA sequences. They used two distinct datasets, weakRM and iRNA5hmc, and their AUC results were 0.9200 and 0.6840, respectively. It’s interesting to note that their model in the second dataset didn’t perform much better than their suggested model.
Transformers have also been used to identify DNA sequences in the case of (Helaly et al. 2022). A BERT transformer with a CNN network was used, while in the study of (Pipoli et al. 2022) they utilized word2vec, DeepLncLoc (M. Zeng et al. 2022), and a transformer. In the first case, researchers examined how well the model classified bacteria, with results up to an AUC of 1. In the second case, the researchers used a more intricate chain of neural networks, starting with an LSTM, moving on to a CNN, and ending with a FNN. Instead of classifying their data, their goal was to create a linear regression from it to uncover those patterns that affect gene expression. Depending on the number of variables (1 or 3), they were able to achieve an R2 metric equal to 0.596 and 0.760 respectively.
An equally complex architecture in their model was also followed by (Tsukiyama & Kurata 2022). In this study they tried to understand how viruses proliferate in the human body, searching for connections between viral and human proteins. For this purpose, they used the word2vec algorithm and a CNN network followed by an attention level to obtain the features of the primary structures, that guide these protein interactions. To see if their model works, they used a FNN to determine how their model performed on data with known interactions. The performance is great as they managed to achieve an AUC value of 0.9880.
In addition, (Forghani et al. 2022) sought to learn more about the evolution of novel influenza virus strains. They focused on how each viral strain is created through the interaction of hemagglutinins (H) with neuraminidases (N) on the surface of the envelope. To accomplish this task, researchers trained a word2vec model using a classified vocabulary called RAAA (reduced amino acid alphabet). They note that RAAA accelerates word2vec training without affecting the algorithm’s performance. Finally, a CNN was employed to identify the hiding patterns and as a classifier, the best results were found by the RF (Accuracy = 0.9330).
In addition to its applications in virology, the word2vec model has found utility in the field of neurobiology. In a study by (S. Chen et al. 2022) they successfully developed a model known as “NeuroPred-CLQ” that embeds peptide sequences and utilizes them as input data for a CNN-TCN-Mul model. This approach aimed to identify the features required for training the FNN model. During performance testing, FNN locates the required neuropeptides with an AUC value of 0.988. They claim that for identifying the required neuropeptides, their model surpasses all previous approaches.
In turn, word2vec was used by (Sun et al. 2022) to discover the features of miRNA and mRNA sequences. They trained word2vec twice to successfully predict how miRNA molecules interact with mRNA. In this instance, a BiLSTM network was employed to find the desired patterns, which condenses the word2vec feature set of 200 features to just 2. With a softmax function, they determine whether miRNAs and mRNA sequences would interact. Their model achieves a 0.9604 accuracy. (W. Xie et al. 2022) took a different approach for the investigation of how miRNAs control gene expression and employed 3 distinct embedding approaches. The doc2vec method to analyze the miRNA sequence, as well as the use of role2vec and GCN to analyze the miRNA interaction networks they created. While the outputs of role2vec train a BiLSTM/LSTM network, the data from doc2vec and GCN serve as inputs to various LSTM neural networks. In the end, they came up with three separate scores that represent the likelihood of miRNA-gene interaction. They determine the final measure (Sfinal) of miRNA-gene interaction by adding the weighted sum of the three scores. The model’s estimation using independent data yields the AUC value equal to 0.9000.
(Zhao et al. 2022) used an entirely different path to examine gene ontologies using two separate research approaches. One involves analyzing each GO term’s semantics using a BERT transformer that has been trained with a TSDAE (Transformer-based Sequential Denoising Auto-Encoder), which improves the quality of the BERT outputs. They also utilized a GCN network, which examines the graphs of the associated GO keywords, at the same time. The Spearman correlation is applied to evaluate the performance of the FNN network that combines the two vectors using the data from the two techniques. The correlation is calculated on already known protein interaction data (benchmark dataset) where they find a maximum AUC value equal to 0.830 in the PPI ALL3 dataset for GO term BP (Biological Process).
A different combination of embedding techniques was used by (Koca et al. 2022) who wanted to study the interactions between viral pathogens and human proteins (Pathogen-host protein–protein interaction, PHI). They created a PHI graph for this purpose, which they then trained a GCN network on. To enhance the information from the graphs for potential missing interactions, scientists applied the doc2vec method independently to viruses and humans while analyzing protein main structures. They employed an RF for this purpose, using it to extract the interactions that are most likely to occur and then finally include them in the GCN model. They employed the binary classifier GA2M, which has an AUC value of 0.9000, for the classification. A similar approach to this study was followed by (X. Wu et al. 2022). The difference in their case was that they employed 4 different embedding techniques to evaluate the sequence of Anti-Cancer Peptides (ACP). They utilized node2vec, OHE, and two physicochemical property embedding techniques, since the graph’s nodes are amino acids rather than proteins, allowing them to account for the physical properties of amino acids that node2vec and OHE might not be able to extract. The GCN receives the final vector produced by all four embedding techniques and extracts the features from the graphs. They estimated their model and classified ACP and non-ACP sequences using a ResNet-FNN network with an AUC performance of 0.8410.
In the meantime, (Chao et al. 2022) used heterogeneous graphs made up of drugs, target proteins, and diseases. They were built by combining data from 5 distinct databases, including Gene Ontology (GO), BioGrid, DrugBank, and Comparative Toxicogenomic Database (CTD). They employed node2vec, a transformer, and the GNN network to obtain the local and global aspects of the heterogeneous networks to examine the characteristics of the graphs. Their goal was to discover plant compounds that could treat the calcification of blood arteries. They used an RF classifier to evaluate their model with a F1 score of 0.7240.
(F. Xie et al. 2022) investigated the connection between lncRNAs and the emergence of different diseases. They produced three different types of graphs for this purpose: A) L-MS (lncRNA-miRNA similarity network), which has lncRNA and miRNA as its nodes; B) M-DS (miRNA-disease similarity network); and C) L-DA (lncRNA-disease association network), which has lncRNAs and disease categories as its nodes. The adjacency information matrix and topology information matrix, which serve as input to SAE and ResNet networks, respectively, were produced by node2vec. They simultaneously used SVM, RF, and XGBoost classifiers to build a weighted input for an LR model, which predicts the association of lncRNA with diseases. The AUC value of their model was 0.9750.
In addition, (Pan et al. 2022) have utilized node2vec to examine the topology of biological proteins. To find the proteins under research in 16 human cellular topologies (MCC = 0.800), they created the node2loc model, in which protein vectors (from node2vec) are initially constructed using as input STRING-based graphs.
In the field of pharmacology, (Amiri Souri et al. 2022) delved into the study of interaction networks between synthetic ligands and proteins to enhance drug development. Their approach involved the creation of two separate graphs to represent interactions between chemical molecules and proteins. Subsequently, they applied node2vec to extract pertinent features from these graphs and utilized these features to train an XGBoost classifier. According to the authors, the prediction of thermodynamically stable interactions is very successful (AUC: 0.9409). Staying in pharmacology, (Joshi et al. 2022) wanted to create a new method through which it will be possible to prevent negative side effects from the use of drugs (Adverse Drug Reactions, ADRs). To be able to implement their thinking, they built a knowledge graph consisting of medicinal substances, ADRs, target molecules, pathways, and genes. Then, they used node2vec to separate the required features for a FNN network to use in classifying harmful and benign side effects (AUC: 0.912).
As previously stated, combining 2vec techniques with transformers is a common technique for the effective embedding of data. A similar strategy was used by (Ieremie et al. 2022), who coupled the node2vec method with the transformers to investigate the interactions between Gene Ontology (GO) terms, as well as to conduct a comprehensive analysis of each GO term. Using an FNN classifier that incorporates data from the STRING database, they achieved AUC values of 0.973 for S. cerevisiae and 0.958 for H. sapiens.
At this point, it will surely have been realized that many 2vec methods deal with proteins either at the level of sequence or at the level of interactions. For this reason, the protein2vec algorithm was created (J. Zhang et al. 2022) (a different model from Protein2vec by J. Gao et al. in 2021). Three modules make up their model. In the first module, node2vec is used to extract every feature required from a graph of gene ontologies. In the second module, LSTM neural networks are utilized, which “remember” which phrases are connected and extract the information (in the form of text) of the GOs. The existence of this “memory” is the most important factor in training the FNN (the final module), which outputs whether an interaction exists or not with an average AUC value of 0.8400 (for various data and GO).
Another recommendation for the study of proteins in the light of gene ontologies is the Anc2vec model by (Edera et al. 2022). It is based on the logic of word2vec, with the difference that it creates three vectors instead of one: a) a vector concerning the ontological uniqueness of the GO term, b) a vector concerning the GO ancestors (ancestors’ hierarchy) and c) the ontology itself (CC, BP, and MF). To confirm the performance of their model they used cosine similarity between vectors as well as an FNN. The feedforward network achieved a maximum AUC value of 0.9600.
In response to the coronavirus pandemic, (Ray et al. 2022) attempted to develop a mechanism for anticipating how the human proteome will interact with the COVID-19 proteome. They created a graph with three different types of nodes: human proteins, SARS-COV2 proteins linked to humans (CoV-host), and SARS-COV2 proteins. The required features were then extracted in the form of a feature matrix using the node2vec metho d. They assessed the probability of protein interactions by employing the weighted rank aggregation (WRA) technique, as introduced by (Pihur et al. 2007). Subsequently, they compiled a catalog of human proteins along with the associated drugs that had been identified to actually interact.
Furthermore, (Ali et al. 2022) conducted a broad investigation of the coronavirus family, focusing on the sequence analysis of their spike proteins. Their goal is to find the attributes of the sequences that facilitate the targeting of particular host organisms by various coronaviruses, as well as the identification of mutations that aid in infecting other organisms. The approach is based on building every possible k-mer of the specified sequence. To calculate PWM (Position Weight Matrix) tables from k mers, a vector with the dimensions 1 × n must be created, where n is the total number of k-mer sequences. The generated vectors serve as the input for a ridge regression function, where the extracted features are fed into several machine learning techniques (7 in total). Logistic regression (LR) reached the highest mean AUC value (0.90).
Staying in protein interactions, (Qian et al. 2022) developed the SPP-CPI model, where they investigate the interaction of small compounds and proteins. Their approach consists of two steps: a) protein sequence analysis and b) feature extraction from a distance matrix they created, to depict the structures of the chemical compounds under investigation. Doc2vec is used for the protein sequence analysis, and a ResNet network along with a CNN (SPP-net) is used for the second step. The two outputs are combined into a feature matrix, which serves as the input of a FNN with an AUC value of 0.978 ± 0.004 for humans.
By using the Deep-AFPpred model, (R. Sharma et al. 2022) attempted to identify antifungal peptides from a set of 2758 peptides (4124 of which were in the training set). They employed a pre-trained seq2vec model, which was found to significantly improve the 1DCNN-BiLSTM neural network system’s performance in comparison to the OHE approach. They were able to attain an AUC value of 0.9784 ± 0.0053. The seq2vec technique is also used in the Virtifier method (Miao et al. 2022) to locate viral genetic material in metagenomics research. They argue that seq2vec and the LSTM-attention level help their model detect viral DNA more effectively because they keep in mind the potential codon combinations that result in protein expression. Their algorithm requires optimization because as the sequence length increases, its performance experiences a significant decline. More particular, it achieves the highest AUC value (AUC: 0.9354) for training and test data with a length of 500 bp. In the training and test combination of 500/10,000 and 10,000/500 bp, respectively, the minimal AUC value is less than 0.65.
Returning to nucleotide sequences, (L. X. Guo et al. 2022) developed the WSCD model to research the relationships between circRNAs and miRNAs, as well as their effects on health. In this case, two distinct embedding techniques were used to apply both the interactions between ribonucleotides and their sequences. The word2vec model was applied to the first situation, and the SDNE model to the second. A CNN-FNN neural network system was developed using a feature matrix that combined the output of the two techniques. The model’s accuracy is extremely poor, with a value of 0.8161 and a constant deviation of 0.0097.
3.6 2023
Until April 4th of 2023, the last date in our results, we only discovered 5 papers, of which only one is about graphs:
(R. Liu et al. 2023) effectively extend the node2vec technology to weighted graphs. They gave this extension the term node2vec + and demonstrated that utilizing node2vec + vectors rather than node2vec greatly improves the efficiency of LR classification (higher values of log2auPRCprior).
Regarding DNA sequences, we have two analyses. The first relates to the development of a novel computational methodology (Histone-Net) that concerns the prediction of DNA-histone interactions (Asim et al. 2023). They used two algorithms. The dna2vec that learns DNA sequence vectors in an unsupervised manner, while the second model is called SuperDNA2vec, a supervised learning algorithm that was trained on known histone-interacting DNA regions. They discovered that when predicting regulatory sequences via histones (AUC: 0.8821 on the balanced dataset), the SuperDNA2vec algorithm achieved the highest performance.
The second analysis by (Halder et al. 2023) on DNA sequences addresses the topic of epigenetic modification involving N4-methylcytosine once again. In this case, the sequences are embedded using the word2vec method, and the CNN is then trained to detect the patterns that inform the FNN whether the sequence has been changed or not. They state that compared to other 4mC modification recognition models, their GS-MLDS model has remarkable performance (maximum AUC: 0.9807).
Transitioning to protein sequences, (Tran et al. 2023) combined word2vec with an artificial feature extraction approach. The artificial approach (Feature Extraction Module) does not involve any computational algorithm, but 5 mathematical methods for feature calculation (Amino acid composition, Pseudo-amino acid composition, Amphilic Pseudo amino acid composition, Quasi-Sequencer-Order, Dipeptide composition). The two kinds of features are integrated through FNNs coupled with attention levels, which extract only the important features (Feature Combination Module). The final classification of proteins into interacting and non-interacting proteins reached an accuracy of 0.9920 ± 0.0015.
Finally, (Alves et al. 2023) extracted characteristics from EEG data and applied the word2vec algorithm to identify people with Attention Deficit Hyperactivity Disorder (ADHD). They primarily deal with EEG microstates, which serve as biomarkers for the automatic identification of ADHD patients. An FNN network is fed by the attributes for each microstate to categorize people as sick or healthy, depending on whether they classify patients into subtypes (3-class classification). They were able to reach an average accuracy of 0.9556 and 0.9906, respectively.
4 Discussion
In our analysis of the 82 papers, although different architectures are used to apply NLP techniques to bioinformatics, we can say that there is a common structure to them. Different types of biological data such as nucleic acids, proteins, small chemical compounds of organic or inorganic composition, and gene ontologies are used as input data. These data are embedded using methods such as word2vec and transformers to extract features and convert them into a format that can be recognized by ML algorithms. By training and optimizing these models, all the methods we studied claimed to produce better models precisely because of the embedding.
As we can see in the figures above, 92% of the biological data concerns the protein and nucleic acid sequences and their effects on gene regulation, as well as the interactions between them. In addition, there is one study where they used abstracts too, where they incorporated information from protein interactions and PubMed text on genes associated with mutations. Their reasoning was to remove the missing values from their data. The 2 and 5% were about biomarkers and GOs respectively (Fig. 5). Naturally, the word2vec (26%) approach is the most popular, followed closely by node2vec (18%). The dna2vec and doc2vec follow with 7 and 3% respectively. The 20% is based on the aforementioned models, and only the 7% was about novel embedding approaches. Finally, since we concentrated on 2vec approaches, the 2% that employed transformers is reasonable (Fig. 6). The main approach researchers use for the analysis of biological features is the Deep Learning methods (61%), while the traditional ML methods account for 26%. A combination of these models was used in 6% of the articles, while new embedding approaches were offered in 7% of the research (Fig. 7).
5 Conclusion
The production of data in the field of Biology is increasing rapidly, necessitating the use of robust computational methods. The progression of Machine Learning is steering research in this direction, enabling the design of more efficient algorithms. The advent of Deep Learning, in particular, has unlocked the capability to investigate intricate relationships among biomolecules within organisms’ cells. Nevertheless the efficacy of the traditional machine learning methods remains formidable (Kumari et al. 2023).This, in turn, enhances the safe production of medicines, aligning with the primary objective of Bioinformatics. In Natural Language Processing, algorithms are created to analyze vast volumes of natural language data and come to conclusions or even write on their own. The revolution in this field came in 2013, with the introduction of the word2vec model and its evolution in 2017 with transformers. The idea that biomolecules communicate using a system of principles similar to those used by humans expanded the field of bioinformatics by allowing the connection of two seemingly unrelated scientific fields, linguistics and biology. In this paper, we have shown a wide range of NLP models that have been created for the analysis of numerous biological data types, including sequences, chemical compounds, and even biosignals. This notion has given the scientific community new hope. It is now possible to identify patterns in a variety of biological data, from networks to simple structures, thanks to the development of models based on embedding methods. Omics analysis has become clearer than ever, given its newfound capabilities to discern sequence similarity, pinpoint regulatory regions, forecast the functions of biological substances, and more. The embedding approaches that we looked at are not a panacea for bioinformatics. The nature of biological data is so complex that each algorithm may require a very large amount of data to be able to decode the relationships between its data, which contributes to rising computing power requirements, training time, and unavoidably, financial costs. As a result, the training times for the models we mentioned can be time-consuming. In addition to their effect on the natural world, Deep Learning approaches have a distinct disadvantage. Since no one is aware of how these deep learning models are trained, it is frequently exceedingly difficult to study the etiology of the outputs of neural networks, making it difficult to corroborate or even to explain the results. Because of this, findings from the aforementioned approaches should be viewed with skepticism if they cannot be independently verified (lack of target data). We thus anticipate that our systematic review will help the interested reader to learn about the applications of NLP models to biological data and to be able to quickly and easily locate the methodology pertinent to their data. As more useful data are retrieved from such procedures, we are confident that the advancement of NLP methodology will henceforth be inevitably tied to the advancement of bioinformatics. We anticipate that even more algorithms for converting data into vectors will be created to improve the models we’ve discussed, judging by the ongoing increase in articles on them (Fig. 8).
Data availability
The data that support the findings of this study are available from the corresponding author, Tzallas AT, upon reasonable request.
References
Albawi S, Mohammed TA, Al-Zawi S (2018) Understanding of a convolutional neural network. Proceedings of 2017 International Conference on Engineering and Technology, ICET 2017, 2018-January, 1–6. https://doi.org/10.1109/ICENGTECHNOL.2017.8308186
Al-Dujaili MJ, Ebrahimi-Moghadam A (2023) Speech emotion recognition: a comprehensive survey. Wireless Pers Commun 129(4):2525–2561. https://doi.org/10.1007/S11277-023-10244-3/TABLES/4
Ali S, Patterson M (2021) Spike2Vec: An Efficient and Scalable Embedding Approach for COVID-19 Spike Sequences. Proceedings - 2021 IEEE International Conference on Big Data, Big Data 2021, 1533–1540. https://doi.org/10.1109/BIGDATA52589.2021.9671848
Ali S, Bello B, Chourasia P, Punathil RT, Zhou Y, Patterson M (2022) PWM2Vec: an efficient embedding approach for viral host specification from coronavirus spike sequences. Biology 11(3):418. https://doi.org/10.3390/BIOLOGY11030418
Allen TC, Cagle PT (2008) Bioinformatics and omics. In: Zander DS, Popper HH, Jagirdar J, Haque AK, Cagle PT, Barrios R (eds) Molecular Pathology of Lung Diseases. Springer, New York, pp 65–69. https://doi.org/10.1007/978-0-387-72430-0_6
Alves LM, Côco KF, de Souza ML, Ciarelli PM (2023) Contextual microstates: an approach based on word embedding of microstates sequence to identify ADHD patients. Rec Biom Eng 39(1):1–13. https://doi.org/10.1007/S42600-022-00245-9
Amiri Souri E, Laddach R, Karagiannis SN, Papageorgiou LG, Tsoka S (2022) Novel drug-target interactions via link prediction and network embedding. BMC Bioinformatics 23(1):121. https://doi.org/10.1186/S12859-022-04650-W
Anthony AA, Patil CM, Basavaiah J (2022) A Review on speech disorders and processing of disordered speech. Wireless Pers Commun 126(2):1621–1631. https://doi.org/10.1007/S11277-022-09812-W/TABLES/1
Asim MN, Ibrahim MA, Malik MI, Razzak I, Dengel A, Ahmed S (2023) Histone-net: a multi-paradigm computational framework for histone occupancy and modification prediction. Com Intel Syst 9(1):399–419. https://doi.org/10.1007/S40747-022-00802-W
Asim MN, Malik MI, Zehe C, Trygg J, Dengel A, Ahmed S (2020) Mirlocpredictor: a convnet-based multi-label microrna subcellular localization predictor by incorporating k-mer positional information. Genes 11(12):1–23. https://doi.org/10.3390/GENES11121475
Basher ARMA, Hallam SJ (2021) Leveraging heterogeneous network embedding for metabolic pathway prediction. Bioinformatics 37(6):822–829. https://doi.org/10.1093/BIOINFORMATICS/BTAA906
Benson DA, Cavanaugh M, Clark K, Karsch-Mizrachi I, Lipman DJ, Ostell J, Sayers EW (2013) GenBank. Nucleic Acids Res 41(D1):S36-D42. https://doi.org/10.1093/NAR/GKS1195
Buchan DWA, Jones DT (2020) Learning a functional grammar of protein domains using natural language word embedding techniques. Proteins: Struct, Funct, Bioinf 88(4):616–624. https://doi.org/10.1002/PROT.25842
Cao L, Liu P, Chen J, Deng L (2022) Prediction of transcription factor binding sites using a combined deep learning approach. Front Oncol 12:893520. https://doi.org/10.3389/FONC.2022.893520/FULL
Chao CT, Tsai YT, Lee WT, Yeh HY, Chiang CK (2022) Deep learning-assisted repurposing of plant compounds for treating vascular calcification: an in silico study with experimental validation. Oxid Med Cell Longev 202:4378413. https://doi.org/10.1155/2022/4378413
Chen T, Guestrin C (n.d.). XGBoost: A Scalable Tree Boosting System. https://doi.org/10.1145/2939672.2939785
Chen S, Li Q, Zhao J, Bin Y, Zheng C (2022) NeuroPred-CLQ: incorporating deep temporal convolutional networks and multi-head attention mechanism to predict neuropeptides. Brief Bioinform 23(5):1–12. https://doi.org/10.1093/BIB/BBAC319
Cheng J, Wang Z, Liu Y, Huang W (2021) CapBind: Prediction of transcription factor binding sites based on capsule network. Proceedings - 2021 6th International Conference on Computational Intelligence and Applications, ICCIA 2021, 31–35. https://doi.org/10.1109/ICCIA52886.2021.00014
Chung J, Gulcehre C, Cho K (n.d.). Empirical Evaluation of Gated Recurrent Neural Networks on Sequence Modeling.
Dai X, Shen L (2022) Advances and trends in omics technology development. Front Med 9:1546. https://doi.org/10.3389/FMED.2022.911861/BIBTEX
Dallalba G, Casa PL, De Abreu FP, Notari DL, De Avila E, Silva S (2022) A survey of biological data in a big data perspective. Big Data 10(4):279–297. https://doi.org/10.1089/BIG.2020.0383/ASSET/IMAGES/LARGE/BIG.2020.0383_FIGURE1.JPEG
Du J, Jia P, Dai Y, Tao C, Zhao Z, Zhi D (2019) Gene2vec: distributed representation of genes based on co-expression. BMC Genomics 20:82. https://doi.org/10.1186/S12864-018-5370-X
Edera AA, Milone DH, Stegmayer G (2022) Anc2vec: embedding gene ontology terms by preserving ancestors relationships. Brief Bioinform 23(2):bbac003. https://doi.org/10.1093/BIB/BBAC003
Forghani M, Khachay M, Firstkov A, Ramsay E (2022) An artificial neural network based ensemble model for predicting antigenic variants: application of reduced amino acid alphabets and word2Vec. Proceedings - 2022 8th International Iranian Conference on Signal Processing and Intelligent Systems, ICSPIS 2022. https://doi.org/10.1109/ICSPIS56952.2022.10044061
Gao J, Tian L, Lv T, Wang J, Song B, Hu X (2021) Protein2Vec: aligning multiple ppi networks with representation learning. IEEE/ACM Trans Comput Biol Bioinf 18(1):240–249. https://doi.org/10.1109/TCBB.2019.2937771
Gao Z, Fu G, Ouyang C, Tsutsui S, Liu X, Yang J, Gessner C, Foote B, Wild D, Ding Y, Yu Q (2019) Edge2vec: Representation learning using edge semantics for biomedical knowledge discovery. BMC Bioinformatics 20(1):306. https://doi.org/10.1186/S12859-019-2914-2
Gönen M, Suleiman Khan, aaltofi A, Kaski S (2013) Kernelized Bayesian Matrix Factorization (Vol. 28, pp. 864–872). PMLR. https://proceedings.mlr.press/v28/gonen13a.html
Grover A, Leskovec J (2018) node2vec: scalable feature learning for networks. KDD : Proceedings. International Conference on Knowledge Discovery & Data Mining, 2016, 855. https://doi.org/10.1145/2939672.2939754
Guo LX, You ZH, Wang L, Yu CQ, Zhao BW, Ren ZH, Pan J (2022) A novel circRNA-miRNA association prediction model based on structural deep neural network embedding. Brief Bioinform 23(5):bbac391. https://doi.org/10.1093/BIB/BBAC391
Guo ZH, You ZH, Yi HC (2020) Integrative construction and analysis of molecular association network in human cells by fusing node attribute and behavior information. Mol Ther Nucleic Acids 19:498–506. https://doi.org/10.1016/J.OMTN.2019.10.046
Halder RK, Uddin MN, Uddin MdA, Aryal S, Islam MdA, Hossain F, Jahan N, Khraisat A, Alazab A (2023) A grid search-based multilayer dynamic ensemble system to identify dna n4—methylcytosine using deep learning approach. Genes 14(3):582. https://doi.org/10.3390/GENES14030582
Helaly MA, Rady S, Aref MM (2022) BERT contextual embeddings for taxonomic classification of bacterial DNA sequences. Expert Syst Appl 208:117972. https://doi.org/10.1016/J.ESWA.2022.117972
Hussein R, Yang D, Cudré-Mauroux P, Cudré P (n.d.). Are Meta-Paths Necessary? Revisiting Heterogeneous Graph Embeddings. 10. https://doi.org/10.1145/3269206.3271777
Ieremie I, Ewing RM, Niranjan M (2022) TransformerGO: predicting protein-protein interactions by modelling the attention between sets of gene ontology terms. Bioinformatics 38(8):2269–2277. https://doi.org/10.1093/BIOINFORMATICS/BTAC104
Jaeger S, Fulle S, Turk S (2018) Mol2vec: unsupervised machine learning approach with chemical intuition. J Chem Inf Model 58(1):27–35. https://doi.org/10.1021/ACS.JCIM.7B00616
Ji QY, Gong XJ, Li HM, Du PF (2021) DeepSE: detecting super-enhancers among typical enhancers using only sequence feature embeddings. Genomics 113(6):4052–4060. https://doi.org/10.1016/J.YGENO.2021.10.007
Jolliffe IT, Cadima J (2016) Principal component analysis: a review and recent developments. Philos Trans A Math Phys Eng Sci 374(2065):20150202. https://doi.org/10.1098/rsta.2015.0202
Joshi PVM, Mukherjee A (2022) A knowledge graph embedding based approach to predict the adverse drug reactions using a deep neural network. J Biomed Inform 132:104122. https://doi.org/10.1016/J.JBI.2022.104122
Jung GT, Kim KP, Kim K (2020) How to interpret and integrate multi-omics data at systems level. Anim Cells Syst 24(1):1. https://doi.org/10.1080/19768354.2020.1721321
Khanal J, Tayara H, Chong KT (2020) Identifying enhancers and their strength by the integration of word embedding and convolution neural network. IEEE Access 8:58369–58376. https://doi.org/10.1109/ACCESS.2020.2982666
Khanal J, Tayara H, Zou Q, Chong KT (2021) Identifying DNA N4-methylcytosine sites in the rosaceae genome with a deep learning model relying on distributed feature representation. Comput Struct Biotechnol J 19:1612–1619. https://doi.org/10.1016/j.csbj.2021.03.015
Kim S, Lee H, Kim K, Kang J (2018) Mut2Vec: distributed representation of cancerous mutations. BMC Med Genomics 11:33. https://doi.org/10.1186/S12920-018-0349-7
Koca MB, Nourani E, Abbasoğlu F, Karadeniz İ, Sevilgen FE (2022) Graph convolutional network based virus-human protein-protein interaction prediction for novel viruses. Comput Biol Chem 101:107755. https://doi.org/10.1016/J.COMPBIOLCHEM.2022.107755
Kumari N, Anwar S, Bhattacharjee V (2023) A comparative analysis of machine and deep learning techniques for EEG evoked emotion classification. Wireless Pers Commun 128(4):2869–2890. https://doi.org/10.1007/S11277-022-10076-7/TABLES/3
Lau J H, Baldwin T (2016) An Empirical Evaluation of doc2vec with Practical Insights into Document Embedding Generation. 78–86. https://groups.google.com/forum/#!topic/
Le Glaz A, Haralambous Y, Kim-Dufor DH, Lenca P, Billot R, Ryan TC, Marsh J, DeVylder J, Walter M, Berrouiguet S, Lemey C (2021) Machine learning and natural language processing in mental health: systematic review. J Med Internet Res 23(5):e15708. https://doi.org/10.2196/15708
Liang Y, Wu Y, Zhang Z, Liu N, Peng J, Tang J (2022) Hyb4mC: a hybrid DNA2vec-based model for DNA N4-methylcytosine sites prediction. BMC Bioinformatics 23(1):258. https://doi.org/10.1186/S12859-022-04789-6
Liao M, Jian-ping Z, Tian J, Zheng CH (2022) iEnhancer-DCLA: using the original sequence to identify enhancers and their strength based on a deep learning framework. BMC Bioinformatics 23(1):480. https://doi.org/10.1186/S12859-022-05033-X
Liu B, Fang L, Wang S, Wang X, Li H, Chou KC (2015) Identification of microRNA precursor with the degenerate K-tuple or Kmer strategy. J Theor Biol 385:153–159. https://doi.org/10.1016/J.JTBI.2015.08.025
Liu HY, Du PF (2022) i5hmCVec: Identifying 5-hydroxymethylcytosine sites of drosophila RNA using sequence feature embeddings. Front Genet 13:896925. https://doi.org/10.3389/FGENE.2022.896925/FULL
Liu H, Hou L, Xu S, Li H, Chen X, Gao J, Wang Z, Han B, Liu X, Wan S (2021) Discovering cerebral ischemic stroke associated genes based on network representation learning. Front Genet 12:728333. https://doi.org/10.3389/FGENE.2021.728333/FULL
Liu R, Hirn M, Krishnan A (2023) Accurately modeling biased random walks on weighted networks using node2vec. Bioinformatics 39(1):btad047. https://doi.org/10.1093/BIOINFORMATICS/BTAD047
Long Y, Luo J (2021) Association mining to identify microbe drug interactions based on heterogeneous network embedding representation. IEEE J Biomed Health Inform 25(1):266–275. https://doi.org/10.1109/JBHI.2020.2998906
Manzoni C, Kia DA, Vandrovcova J, Hardy J, Wood NW, Lewis PA, Ferrari R (2018) Genome, transcriptome and proteome: the rise of omics data and their integration in biomedical sciences. Brief Bioinform 19(2):286. https://doi.org/10.1093/BIB/BBW114
Maruyama O, Li Y, Narita H, Toh H, Au Yeung WK, Sasaki H (2022) CMIC: predicting DNA methylation inheritance of CpG islands with embedding vectors of variable-length k-mers. BMC Bioinformatics 23(1):371. https://doi.org/10.1186/S12859-022-04916-3
Matougui B, Boukelia A, Belhadef H, Galiez C, Batouche M (2021) NLP-metaxa: a natural language processing approach for metagenomic taxonomic binning based on deep learning. Curr Bioinform 16(7):992–1003. https://doi.org/10.2174/1574893616666210621101150
Miao Y, Liu F, Hou T, Liu Y (2022) Virtifier: a deep learning-based identifier for viral sequences from metagenomes. Bioinformatics 38(5):1216–1222. https://doi.org/10.1093/BIOINFORMATICS/BTAB845
Mikolov T, Chen K, Corrado G, Dean J (2013) Efficient Estimation of Word Representations in Vector Space. http://ronan.collobert.com/senna/
Miltiadous A, Gionanidis E, Tzimourta KD, Giannakeas N, Tzallas AT (2023) DICE-Net: a novel convolution-transformer architecture for alzheimer detection in eeg signals. IEEE Access 11:71840–71858. https://doi.org/10.1109/ACCESS.2023.3294618
Mostavi M, Salekin S, Huang Y (2018) Deep-2’-O-Me: predicting 2′-O-methylation sites by convolutional neural networks. Proceedings of the Annual International Conference of the IEEE Engineering in Medicine and Biology Society, EMBS, 2018-July, 2394–2397. https://doi.org/10.1109/EMBC.2018.8512780
Naskath J, Sivakamasundari G, Alif A, Begum S (2023) A study on different deep learning algorithms used in deep neural nets: MLP SOM and DBN. Wireless Pers Commun 128:2913–2936. https://doi.org/10.1007/s11277-022-10079-4
Ng P (2017) dna2vec: Consistent vector representations of variable-length k-mers. https://arxiv.org/abs/1701.06279v1
Ostrovsky-Berman M, Frankel B, Polak P, Yaari G (2021) Immune2vec: embedding B/T cell receptor sequences in ℝ N using natural language processing. Front Immunol 12:680687. https://doi.org/10.3389/FIMMU.2021.680687/FULL
Ouzzani M, Hammady H, Fedorowicz Z, Elmagarmid A (2016) Rayyan-a web and mobile app for systematic reviews. Syst Rev 5(1):1–10. https://doi.org/10.1186/S13643-016-0384-4/FIGURES/6
Ovens K, Maleki F, Eames BF, McQuillan I (2021) Juxtapose: a gene-embedding approach for comparing co-expression networks. BMC Bioinformatics 22(1):125. https://doi.org/10.1186/S12859-021-04055-1
Page MJ, McKenzie JE, Bossuyt PM, Boutron I, Hoffmann TC, Mulrow CD, Shamseer L, Tetzlaff JM, Akl EA, Brennan SE, Chou R, Glanville J, Grimshaw JM, Hróbjartsson A, Lalu MM, Li T, Loder EW, Mayo-Wilson E, McDonald S, Moher D (2021) The PRISMA 2020 statement: an updated guideline for reporting systematic reviews. BMJ 372:n71. https://doi.org/10.1136/BMJ.N71
Pan X, Chen L, Liu M, Niu Z, Huang T, Cai YD (2022) Identifying protein subcellular locations with embeddings-based node2loc. IEEE/ACM Trans Comput Biol Bioinf 19(2):666–675. https://doi.org/10.1109/TCBB.2021.3080386
Pan X, Zuallaert J, Wang X, Shen HB, Campos EP, Marushchak DO, De Neve W (2020) ToxDL: deep learning using primary structure and domain embeddings for assessing protein toxicity. Bioinformatics 36(21):5159–5168. https://doi.org/10.1093/BIOINFORMATICS/BTAA656
Peng CYJ, Lee KL, Ingersoll GM (2002) An introduction to logistic regression analysis and reporting. J Educ Res 96(1):3–14. https://doi.org/10.1080/00220670209598786
Pihur V, Datta S, Datta S (2007) Weighted rank aggregation of cluster validation measures: a monte carlo cross-entropy approach. Bioinformatics 23(13):1607–1615. https://doi.org/10.1093/BIOINFORMATICS/BTM158
Pipoli V, Cappelli M, Palladini A, Peluso C, Lovino M, Ficarra E (2022) Predicting gene expression levels from DNA sequences and post-transcriptional information with transformers: predicting gene expression levels from DNA sequences. Comput Methods Programs Biomed 225:107035. https://doi.org/10.1016/J.CMPB.2022.107035
Qian Y, Li X, Zhang Q, Zhang J (2022) SPP-CPI: predicting compound-protein interactions based on neural networks. IEEE/ACM Trans Comput Biol Bioinf 19(1):40–47. https://doi.org/10.1109/TCBB.2021.3084397
Quazi S (2022) Artificial intelligence and machine learning in precision and genomic medicine. Med Oncol 39(8):1–18. https://doi.org/10.1007/S12032-022-01711-1
Ray S, Lall S, Bandyopadhyay S (2022) A deep integrated framework for predicting SARS-CoV2-human protein-protein interaction. IEEE Trans Emerg Topics Comput Intell 6(6):1463–1472. https://doi.org/10.1109/TETCI.2022.3182354
Rezk NM, Purnaprajna M, Nordstrom T, Ul-Abdin Z (2020) Recurrent neural networks: an embedded computing perspective. IEEE Access 8:57967–57996. https://doi.org/10.1109/ACCESS.2020.2982416
Rish I (n.d.). An empirical study of the naive Bayes classifier.
Rogers D, Hahn M (2010). Extended-Connectivity Fingerprints. https://doi.org/10.1021/ci100050t
Rokach L, Maimon O (2006) Decision Trees. Data Mining and Knowledge Discovery Handbook, 165–192. https://doi.org/10.1007/0-387-25465-X_9
Sabour S, Frosst N, Hinton GE (n.d.). Dynamic Routing Between Capsules.
Sazli MH (2006) A brief review of feed-forward neural networks. Commun Fac Sci Univ Ank Series 1:11–17
Schölkopf B (1998) SVMs - a practical consequence of learning theory. IEEE Intel Syst Their Appl 13(4):18–21. https://doi.org/10.1109/5254.708428
Sharma R, Shrivastava S, Singh SK, Kumar A, Saxena S, Singh RK (2022) Deep-AFPpred: Identifying novel antifungal peptides using pretrained embeddings from seq2vec with 1DCNN-BiLSTM. Brief Bioinform 23(1):bbab422. https://doi.org/10.1093/BIB/BBAB422
Sharma S, Singh S (2022) Recognition of indian sign language (ISL) using deep learning model. Wireless Pers Commun 123(1):671–692. https://doi.org/10.1007/S11277-021-09152-1/TABLES/8
Shen F, Peng S, Fan Y, Wen A, Liu S, Wang Y, Wang L, Liu H (2019) HPO2Vec+: leveraging heterogeneous knowledge resources to enrich node embeddings for the human phenotype ontology. J Biomed Inform 96:103246. https://doi.org/10.1016/J.JBI.2019.103246
Shen Z, Bao W, Huang DS (2018) Recurrent neural network for predicting transcription factor binding sites. Sci Rep 8(1):15270. https://doi.org/10.1038/S41598-018-33321-1
Sitaraman R (2009) The first paper in bioinformatics? Microbe (washington, D.c.) 4:485–486. https://doi.org/10.1128/microbe.4.485.2
Smaili FZ, Gao X, Hoehndorf R (2018) Onto2Vec: joint vector-based representation of biological entities and their ontology-based annotations. Bioinformatics 34(13):i52–i60. https://doi.org/10.1093/BIOINFORMATICS/BTY259
Sun Y, Xiong F, Sun Y, Zhao Y, Cao Y (2022) A miRNA target prediction model based on distributed representation learning and deep learning. Comput Math Methods Med 2022:4490154. https://doi.org/10.1155/2022/4490154
Szklarczyk D, Kirsch R, Koutrouli M, Nastou K, Mehryary F, Hachilif R, Gable AL, Fang T, Doncheva NT, Pyysalo S, Jensen LJ, Von Mering C (2023) The STRING database in 2023: protein-protein association networks and functional enrichment analyses for any sequenced genome of interest. Database Issue Published Online 51:D638–D646. https://doi.org/10.1093/nar/gkac1000
Thafar MA, Albaradie S, Olayan RS, AshoorH, Essack M, Bajic VB (2020) Computational drug-target interaction prediction based on graph embedding and graph mining. ACM International Conference Proceeding Series, 14–21. https://doi.org/10.1145/3386052.3386062
Tong X, Liu S (2019) CPPred: coding potential prediction based on the global description of RNA sequence. Nucleic Acids Res 47(8):e43. https://doi.org/10.1093/NAR/GKZ087
Tran HN, Xuan QNP, Nguyen TT (2023) DeepCF-PPI: improved prediction of protein-protein interactions by combining learned and handcrafted features based on attention mechanisms. Appl Intell. https://doi.org/10.1007/S10489-022-04387-2
Tsukiyama S, Kurata H (2022) Cross-attention PHV: prediction of human and virus protein-protein interactions using cross-attention–based neural networks. Comput Struct Biotechnol J 20:5564–5573. https://doi.org/10.1016/J.CSBJ.2022.10.012
Vaswani A, Shazeer N, Parmar N, Uszkoreit J, Jones L, Gomez AN, Kaiser Ł, Polosukhin I (2017) Attention Is All You Need. Advances in Neural Information Processing Systems, 2017-December, 5999–6009. https://arxiv.org/abs/1706.03762v5
Wahab A, Tayara H, Xuan Z, Chong KT (2021) DNA sequences performs as natural language processing by exploiting deep learning algorithm for the identification of N4-methylcytosine. Sci Rep 11(1):212. https://doi.org/10.1038/S41598-020-80430-X
Wang C, Zhang Y, Han S (2020) Its2vec: fungal species identification using sequence embedding and random forest classification. Biomed Res Int 2020:2468789. https://doi.org/10.1155/2020/2468789
Wang L, Liu F, Du L, Qin G (2021) Single-cell transcriptome analysis in melanoma using network embedding. Front Genet 12:700036. https://doi.org/10.3389/FGENE.2021.700036/FULL
Wang N, Zeng M, Zhang J, Li Y, Li M (2020) Ess-NEXG: predict essential proteins by constructing a weighted protein interaction network based on node embedding and XGBoost. Lecture Notes in Computer Science (Including Subseries Lecture Notes in Artificial Intelligence and Lecture Notes in Bioinformatics) 12304 LNBI:95–104. https://doi.org/10.1007/978-3-030-57821-3_9/FIGURES/3
Wang Y, You ZH, Yang S, Li X, Jiang TH, Zhou X (2019) A high efficient biological language model for predicting protein-protein interactions. Cells 8(2):122. https://doi.org/10.3390/CELLS8020122
Wang Z, Lei X (2021) Prediction of RBP binding sites on circRNAs using an LSTM-based deep sequence learning architecture. Brief Bioinform 22(6):bbab342. https://doi.org/10.1093/BIB/BBAB342
Woloszynek S, Zhao Z, Chen J, Rosen GL (2019) 16S rRNA sequence embeddings: meaningful numeric feature representations of nucleotide sequences that are convenient for downstream analyses. PLoS Comput Biol 15(2):e1006721. https://doi.org/10.1371/JOURNAL.PCBI.1006721
Wu C, Gao R, Zhang Y, De Marinis Y (2019) PTPD: predicting therapeutic peptides by deep learning and word2vec. BMC Bioinformatics 20(1):456. https://doi.org/10.1186/S12859-019-3006-Z
Wu F, Yang R, Zhang C, Zhang L (2021) A deep learning framework combined with word embedding to identify DNA replication origins. Sci Rep 11(1):844. https://doi.org/10.1038/S41598-020-80670-X
Wu X, Zeng W, Lin F (2022) GCNCPR-ACPs: a novel graph convolution network method for ACPs prediction. BMC Bioinformatics 23:560. https://doi.org/10.1186/S12859-022-04771-2
Xie F, Yang Z, Song J, Dai Q, Duan X (2022) DHNLDA: a novel deep hierarchical network based method for predicting lncRNA-disease associations. IEEE/ACM Trans Comput Biol Bioinf 19(6):3395–3403. https://doi.org/10.1109/TCBB.2021.3113326
Xie W, Zheng Z, Zhang W, Huang L, Lin Q, Wong KC (2022) SRG-vote: predicting mirna-gene relationships via embedding and LSTM ensemble. IEEE J Biomed Health Inform 26(8):4335–4344. https://doi.org/10.1109/JBHI.2022.3169542
Yang S, Wang Y, Lin Y, Shao D, He K, Huang L (2020) LncMirNet: predicting LncRNA–miRNA interaction based on deep learning of ribonucleic acid sequences. Molecules 25(19):4372. https://doi.org/10.3390/MOLECULES25194372
Yang X, Yang S, Li Q, Wuchty S, Zhang Z (2020) Prediction of human-virus protein-protein interactions through a sequence embedding-based machine learning method. Comput Struct Biotechnol J 18:153–161. https://doi.org/10.1016/J.CSBJ.2019.12.005
Yao Y, Du X, Diao Y, Zhu H (2019) An integration of deep learning with feature embedding for protein–protein interaction prediction. PeerJ 7(6):e7126. https://doi.org/10.7717/PEERJ.7126
Yuan Y, Xun G, Suo Q, Jia K, Zhang A (2017) Wave2Vec: learning deep representations for biosignals. Proceedings - IEEE International Conference on Data Mining, ICDM, 2017-November, 1159–1164. https://doi.org/10.1109/ICDM.2017.155
Yuan Y, Xun G, Suo Q, Jia K, Zhang A (2019) Wave2Vec: deep representation learning for clinical temporal data. Neurocomputing 324:31–42. https://doi.org/10.1016/J.NEUCOM.2018.03.074
Zeng M, Li M, Fei Z, Wu FX, Li Y, Pan Y, Wang J (2021) A deep learning framework for identifying essential proteins by integrating multiple types of biological information. IEEE/ACM Trans Comput Biol Bioinf 18(1):296–305. https://doi.org/10.1109/TCBB.2019.2897679
Zeng M, Li M, Wu FX, Li Y, Pan Y (2019) DeepEP: a deep learning framework for identifying essential proteins. BMC Bioinformatics 20:506. https://doi.org/10.1186/S12859-019-3076-Y
Zeng M, Wu Y, Lu C, Zhang F, Wu FX, Li M (2022) DeepLncLoc: a deep learning framework for long non-coding RNA subcellular localization prediction based on subsequence embedding. Brief Bioinform 23(1):bbab360. https://doi.org/10.1093/BIB/BBAB360
Zeng W, Wu M, Jiang R (2018) Prediction of enhancer-promoter interactions via natural language processing. BMC Genomics 19:84. https://doi.org/10.1186/S12864-018-4459-6
Zhang F, Song H, Zeng M, Wu FX, Li Y, Pan Y, Li M (2021) A deep learning framework for gene ontology annotations with sequence- and network-based information. IEEE/ACM Trans Comput Biol Bioinf 18(6):2208–2217. https://doi.org/10.1109/TCBB.2020.2968882
Zhang J, Zhu M, Qian Y (2022) Protein2vec: predicting protein-protein interactions based on LSTM. IEEE/ACM Trans Comput Biol Bioinf 19(3):1257–1266. https://doi.org/10.1109/TCBB.2020.3003941
Zhang L, Li G, Li X, Wang H, Chen S, Liu H (2021) EDLm6APred: ensemble deep learning approach for mRNA m6A site prediction. BMC Bioinformatics 22(1):288. https://doi.org/10.1186/S12859-021-04206-4
Zhang YF, Wang X, Kaushik AC, Chu Y, Shan X, Zhao MZ, Xu Q, Wei DQ (2020) SPVec: a word2vec-inspired feature representation method for drug-target interaction prediction. Front Chem 7:895. https://doi.org/10.3389/FCHEM.2019.00895/FULL
Zhao L, Wang J, Cheng L, Wang C (2020) Ontosem: an ontology semantic representation methodology for biomedical domain. Proceedings - 2020 IEEE International Conference on Bioinformatics and Biomedicine, BIBM 2020, 523–527. https://doi.org/10.1109/BIBM49941.2020.9313128
Zhao L, Sun H, Cao X, Wen N, Wang J, Wang C (2022) Learning representations for gene ontology terms by jointly encoding graph structure and textual node descriptors. Brief Bioinform 23(5):bbac124. https://doi.org/10.1093/BIB/BBAC318
Zhou S, Yue X, Xu X, Liu S, Zhang W, Niu Y (2019) LncRNA-miRNA interaction prediction from the heterogeneous network through graph embedding ensemble learning. Proceedings - 2019 IEEE International Conference on Bioinformatics and Biomedicine, BIBM 2019, 622–627. https://doi.org/10.1109/BIBM47256.2019.8983044
Zhu S, Bing J, Min X, Lin C, Zeng X (2018) Prediction of drug-gene interaction by using metapath2vec. Front Genet 9:248. https://doi.org/10.3389/FGENE.2018.00248/FULL
Zou Q, Xing P, Wei L, Liu B (2019) Gene2vec: gene subsequence embedding for prediction of mammalian N 6 -methyladenosine sites from mRNA. RNA 25(2):205–218. https://doi.org/10.1261/RNA.069112.118
Zulfiqar H, Sun ZJ, Huang QL, Yuan SS, Lv H, Dao FY, Lin H, Li YW (2022) Deep-4mCW2V: a sequence-based predictor to identify N4-methylcytosine sites in escherichia coli. Methods 203:558–563. https://doi.org/10.1016/J.YMETH.2021.07.011
Acknowledgements
The publication of the article in OA mode was financially supported by HEAL-Link. In addition, we acknowledge support of this work by the project “Dioni: Computing Infrastructure for Big-Data Processing and Analysis.” (MIS No. 5047222) which is implemented under the Action “Reinforcement of the Research and Innovation Infrastructure”, funded by the Operational Programme “Competitiveness, Entrepreneurship and Innovation” (NSRF 2014-2020) and co-financed by Greece and the European Union (European Regional Development Fund.
Funding
Open access funding provided by HEAL-Link Greece. Hellenic Academic Libraries Link,Dioni: Computing Infrastructure for Big-Data Processing and Analysis.
Author information
Authors and Affiliations
Contributions
Emmanouil Oikonomou performed the literature search as well as the data analysis. The idea for the article was by Emmanouil Oikonomou, Petros Karvelis, Nikolaos Giannakeas and Alexandros Tzallas. Petros Karvelis, Nikolaos Giannakeas and Alexandros Tzallas critically revised the work. Aristidis Vrachatis and Evripidis Glavas helped to draft the manuscript. All authors read and approved the final manuscript.
Corresponding authors
Ethics declarations
Conflict of interests
The authors have declared no competing interests.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Oikonomou, E.D., Karvelis, P., Giannakeas, N. et al. How natural language processing derived techniques are used on biological data: a systematic review. Netw Model Anal Health Inform Bioinforma 13, 23 (2024). https://doi.org/10.1007/s13721-024-00458-1
Received:
Revised:
Accepted:
Published:
DOI: https://doi.org/10.1007/s13721-024-00458-1