Abstract
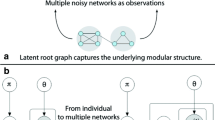
As a general approach to study interactions among small biological molecules such as genes and proteins, network analysis has aroused great interest of people from various research disciplines. However, the construction of network is usually quite sensitive to noise which is unavoidable in real data. Besides, the parameter selections for network construction can also affect the result significantly. These two factors largely decrease the consistency of results generated in network analysis. In particular, we consider detecting closely connected subgraphs named module structure. As an important common property of biological networks, this module structure is often destroyed corrupted by both noise and poor parameter selections in network construction. To conquer these two disadvantages to improve the consistency of module structure identified, we propose to process multiple networks for same set of biological molecules simultaneously for common module structure. More specifically, we combine multiple networks together by building an order 3 tensor data with each layer as one of the multiple networks. Then given any molecule(s) as prior information, a novel tensor-based Markov chain algorithm is proposed to iteratively detect the module that includes the prior node. Moreover, the proposed algorithm is capable of evaluating the contribution scores of each network to the detected module structure. The contribution scores from multiple networks can be not only useful criteria to measure the consistency of module structure, but also valid indicator of corruption in networks. To demonstrate the effectiveness and efficiency of the proposed tensor-based Markov chain algorithm, experimental results on synthetic data set as well as two real gene co-expression data sets of human beings are reported. We also validate that the identified common modules are biologically meaningful.









Similar content being viewed by others
References
Altaf-Ul-Amin M, Shinbo Y, Mihara K, Kurokawa K, Kanaya S (2006) Development and implementation of an algorithm for detection of protein complexes in large interaction networks. BMC Bioinform 7:207
Bernatsky S, Ramsey-Goldman R, Clarke A (2005) Exploring the links between systemic lupus erythematosus and cancer. Rheum Dis Clin N Am 31(2):387–402
Brun C, Chevenet F, Martin D, Wojcik J, Guenoche A, Jacq B (2003) Functional classification of proteins for the prediction of cellular function from a protein–protein interaction network. Genome Biol 5:R6
Chen J, Yuan B (2006) Detecting functional modules in the yeast protein protein interaction network. Bioinformatics 22:2283–2290
Chikina M, Huttenhower C, Murphy C, Troyanskaya O (2009) Global prediction of tissue-specific gene expression and context-dependent gene networks in caenorhabditis elegans. PLoS Comput Biol 5:e1000417
Chua H, Sung W, Wong L (2006) Exploiting indirect neighbours and topological weight to predict protein function from protein–protein interactions. Bioinformatics 22:1623–1630
Danon L et al (2005) Comparing community structure identification. J Stat Mech Theory Exp 2005:P09008
Dong J, Horvath S (2007) Understanding network concepts in modules. BMC Syst Biol 1
Eisen M, Spellman P, Brown P, Botstein D (1998) Cluster analysis and display of genome-wide expression patterns. Proc Natl Acad Sci USA 95:14863–14868
Estrada E, Hatano N (2008) Communicability in complex networks. Phys Rev E 77:036111
Fortunato S (2010) Community detection in graphs. Phys Rep 486:75–174
Girvan M, Newman MEJ (2002) Community structure in social and biological networks. Proc Natl Acad Sci USA 99:7821–7826
Hu H, Yan X, Huang Y, Han J, Zhou XJ (2005) Mining coherent dense subgraphs across massive biological networks for functional discovery. Bioinformatics 21:i213–i221
Huang Y, Li H, Hu H, Yan X, Waterman MS et al (2007) Systematic discovery of functional modules and context-specific functional annotation of human genome. Bioinformatics 23:i222–i229
Huang D, Sherman BT, Lempicki RA (2009a) Systematic and integrative analysis of large gene lists using david bioinformatics resources. Nat Protoc 4(1):44–57
Huang D, Sherman BT, Lempicki RA (2009b) Bioinformatics enrichment tools: paths toward the comprehensive functional analysis of large gene lists. Nuclear Acids Res 37(1):1–13
Koyuturk M, Grama A, Szpankowski W (2004) An efficient algorithm for detecting frequent subgraphs in biological networks. Bioinformatics 20:i200–i207
Li S, Armstrong CM, Bertin N, Ge H, Milstein S et al (2004) A map of the interactome network of the metazoan C. elegans. Science 303:540–543
Li W, Liu CC, Zhang T, Li H, Waterman MS et al (2012) Integrative analysis of many weighted co-expression networks using tensor computation, p e1001106
Mucha PJ, Richardson T, Macon K, Porter MA, Onnela J-P (2010) Community structure in time-dependent, multiscale, and multiplex networks. Science 328:876–878
Newman MEJ (2006a) Modularity and community structure in networks. Proc Natl Acad Sci USA 103:8577–8582
Newman MEJ (2006b) Finding community structure in networks using the eigenvectors of matrices. Phys Rev E 74:036104
Parikh-Patel A, White RH, Allen M, Cress R (2008) Cancer risk in a cohort of patients with systemic lupus erythematosus (SLE) in California. Cancer Causes Control 19(8):887–894
Porter MA et al (2010) Communities in networks. Not AMS 56:1082–1102
Radicchi F et al (2004) Defining and identifying communities in networks. Proc Natl Acad Sci USA 101:2658–2663
Ross S (2003) Introduction to probability models. Academic Press, New York
Segal E, Friedman N, Koller D, Regev A (2004) A module map showing conditional activity of expression modules in cancer. Nat Genet 36:1090–1098
Sharan R, Ulitsky I, Shamir R (2007) Network-based prediction of protein function. Mol Syst Biol 3:88
Shen C, Ng M, Jing L (2013) Sparse-MIML: a sparsity-based multi-instance multi-learning algorithm. Energy minimization methods in computer vision and pattern recognition. Lect Notes Comput Sci 8081:204–306
Shen C, Zhang S, Ng M (2014) A tensor-based Markov chain method for module identification from multiple networks. In: IEEE proceeding of 2014 8th international conference on systems biology, pp 49–58
Tang W, Lu Z, Dhillon IS (2009) Clustering with multiple graphs. In: IEEE international conference on data mining (ICDM), pp 1016–1021
Vazquez A, Flammini A, Maritan A, Vespignani A (2003) Global protein function prediction from protein–protein interaction networks. Nat Biotechnol 21:697–700
Wang X, Chen G (2003) Complex networks: small-world, scale-free and beyond. In: IEEE circuits and systems magazine, pp 6–20
Wu Q, Ng MK, Ye Y (2013) Markov-MIML: a markov chain-based multi-instance multi-label learning algorithm. Knowl Inf Syst 37:83–104
Yu S, Liu X, Tranchevent L-C, Glanzel W, Suykens JAK, Moor BD, Moreau Y (2010) Optimized data fusion for \(K\)-means Laplacian clustering. Bioinformatics 27(1):118–126
Zhang S, Zhao H (2012) Community identification in networks with unbalanced structure. Phys Rev E 85:066114
Zhang S, Zhao H (2013) Normalized modularity optimization method for community identification with degree adjustment. Phys Rev E 88:052802
Acknowledgments
S. Zhang’s research is supported in part by NSFC Grants 10901042, 91130032,11471082 and Shanghai Natural Science Foundation 13ZR1403600. M. Ng’s research is supported in part by Hong Kong Research Grant Council GRF Grant No. 12302715.
Author information
Authors and Affiliations
Corresponding author
Appendix
Appendix
Let us first consider the following lemma:
Lemma 1
Proof
we also easy get \(\sum ^{n_2}_{i=1}[\mathbf {M_2}]_{i, k}=1\) via the similar way. \(\square\)
This Lemma 1 guarantees that both \(\mathbf {M}_1\) and \(\mathbf {M}_2\) are transition probability matrices. Then we may show that both \(\mathbf {x}\) and \(\mathbf {y}\) satisfying (3) and (4) are unique:
Lemma 2
For \(\alpha \in (0,1)\) , there exist unique \(\mathbf {x}\) and \(\mathbf {y}\) satisfying (3) and (4).
Proof
From Lemma 1, we get that
therefore
denote the radius of \(\mathbf {M_1}\mathbf {M_2}\) as \(\rho (\mathbf {M_1}\mathbf {M_2})\) and it is not larger than its norm, we get \(\rho (\mathbf {M_1}\mathbf {M_2})\le 1\).
then
where \(\mathbf {I}\) is identity matrix because \(\rho ((1-\alpha )\mathbf {M_1}\mathbf {M_2})=(1-\alpha )\rho (\mathbf {M_1}\mathbf {M_2})\le (1-\alpha )<1\) and \(\rho ((1-\alpha )\mathbf {M_2}\mathbf {M_1})\le 1-\alpha <1\). Therefore, \(\mathbf {I}-(1-\alpha )\mathbf {M_1}\mathbf {M_2}\) and \(\mathbf {I}-(1-\alpha )\mathbf {M_2}\mathbf {M_1}\) are nonsingular,
\(\square\)
Now consider the iteration in (5) and (6), we may give the following Lemma:
Lemma 3
The iteration guarantees the sequences \(\{\mathbf {x_k}\}\) and \(\{\mathbf {y_k}\}\) satisfying \(\Vert \mathbf {x}_k\Vert _1=1\) and \(\Vert \mathbf {y}_k\Vert _1=1\).
Proof
The iteration can be written as the following
let \(e=[1,1,\ldots ,1]^{\rm T}\), then
If \(\Vert \mathbf {x_k}\Vert _1=1\), then \(\Vert \mathbf {x_{k+1}}\Vert _1=1\). The same consequence can be obtained for sequence \(\{\mathbf {y_k}\}\). \(\square\)
Based on previous two lemmas, we may give the following remarks.
Remark 1
-
1.
If we set \(\Vert \mathbf {x_0}\Vert _1=\Vert \mathbf {y_0}\Vert _1=1\), then the \(\Vert \cdot \Vert _1=1\) can keep during the iteration.
-
2.
\(\Vert \mathbf {M_1}\Vert _1=\Vert \mathbf {M_2}\Vert _1=1\) is established under the condition \(\Vert \mathbf {x}\Vert _1=1\) which the iteration promised.
Then we are ready to present the following Theorem:
Theorem 1
The iterations (5) and (6) are convergent when \(\alpha \in (2/3,1)\).
Theorem 1 can be proved easily via the similar proof of Lemma 2. This theorem demonstrates the convergence of the iterative scheme in the second stage of Algorithm 1. Then according to Algorithm 1, it is easy to get:
then
It is not difficult to see that the convergence of \(\{\mathbf {y}(t)\}\) is guaranteed if \(\{\mathbf {x}(t)\}\) converges. Therefore, it is enough that we only consider the convergence of \(\{\mathbf {x}(t)\}\) here. For simplicity, let
where \(F(\mathbf {x}(t))=(\mathbf {I}-(1-\alpha )\mathbf {M_1}(t)\mathbf {M_2}(t))^{-1}\alpha \mathbf {p}\).
Let us consider the mapping
where \(\mathbf {F}(\mathbf {x})=(\mathbf {I}-(1-\alpha )\mathbf {M_1}(\mathbf {x})\mathbf {M_2}(\mathbf {x}))^{-1}\alpha \mathbf {p}\). \(\mathbf {M}_1(\mathbf {x})= \mathcal {A}^{(1)}\times _{2}\mathbf {x}\) and \(\mathbf {M}_2(\mathbf {x})=(\mathcal {A}^{(2)}\times _{2}\mathbf {x})^{\rm T}\).
Denote \(\mathbf {Q}(\mathbf {x})=(\mathbf {I}-(1-\alpha )\mathbf {M_1}(\mathbf {x})\mathbf {M_2}(\mathbf {x}))^{-1}\), \(\mathbf {G}(\mathbf {x})=(\mathbf {Q}(\mathbf {\mathbf {x}}))^{-1}=\mathbf {I}-(1-\alpha )\mathbf {M_1}(\mathbf {x})\mathbf {M_2}(\mathbf {x})\) and let \(\mathbf {L}(\mathbf {x})=\mathbf {M}_1(\mathbf {x})\mathbf {M}_2(\mathbf {x})\), we have:
In addition, if rewrite
and
where \(\mathcal {A}^{(1)}(1)\) is unfolding of tensor \(\mathcal {A}^{(1)}\) from 1st direction, \(\mathcal {A}^{(2)}(3)\) is the unfolding of tensor \(\mathcal {A}^{(2)}\) from 3rd direction, we may get
Then
In addition, we may also get
Therefore, we have
In conclusion, when \(\alpha >\frac{2}{3}\), \(\Vert \frac{{\rm d}F(\mathbf {x})}{{\rm d}\mathbf {x}}\Vert _1<1\), it is easy to see that the mapping F is a contract mapping. From the Banach fixed point theorem, the Eq. (7) converges to a unique fixed point \(\mathbf {x}_*\) which means the algorithm 1 converges.
Rights and permissions
About this article
Cite this article
Shen, C., Pan, J., Zhang, S. et al. Multiple networks modules identification by a multi-dimensional Markov chain method. Netw Model Anal Health Inform Bioinforma 4, 32 (2015). https://doi.org/10.1007/s13721-015-0106-1
Received:
Revised:
Accepted:
Published:
DOI: https://doi.org/10.1007/s13721-015-0106-1