Abstract
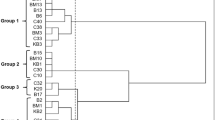
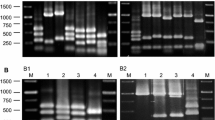
Traditional cheeses made without starter cultures can be characterised by the attribute of instability. The addition of autochthonous starter cultures can ensure stability without compromising the characteristics of the final product. This study aimed to characterise the autochthonous lactic acid bacteria (LAB) population in “Vastedda della valle del Belìce” cheeses, which have a protected designation of origin (PDO) status, in order to develop an ad hoc starter culture to be used in its future production. Winter and spring productions were analysed to ensure isolation of specific LAB that had adapted to perform fermentation at low temperatures. Plate counts revealed total microbial numbers nearing 109 CFU.g−1. All of the cheese samples were dominated by coccus-shaped LAB. When enterobacteria were present, their concentrations were at similar levels (3.3–5.6 Log CFU.g−1) in both seasons. All of the colonies that differed in morphological appearance were isolated and differentiated on the basis of phenotypic characteristics and genetic polymorphisms, as analysed by random amplification of polymorphic DNA-polymerase chain reaction. A total of 74 strains were identified and further genotyped by sequencing the 16S rRNA gene, resulting in the identification of 16 LAB species belonging to five genera (Enterococcus, Lactobacillus, Lactococcus, Leuconostoc and Streptococcus). The species most frequently present were Streptococcus gallolyticus subsp. macedonicus, Streptococcus thermophilus, Lactococcus lactis and Leuconostoc mesenteroides. The 74 strains were also investigated in vitro for general dairy parameters such as acidification capacity, diacetyl generation and antibacterial activity. Several strains of the most frequently represented species displayed traits relevant to the production of PDO “Vastedda della valle del Belìce”.


Similar content being viewed by others
References
Abd-el-Malek Y, Gibson T (1948) Studies in the bacteriology of milk. I. The streptococci of milk. J Dairy Res 15:233–240
Aponte M, Fusco V, Andolfi R, Coppola S (2008) Lactic acid bacteria occurring during manufacture and ripening of Provolone del Monaco cheese: detection by different analytical approaches. Int Dairy J 18:403–413
Antonsson M, Molin G, Ardo Y (2003) Lactobacillus strains isolated from Danbo cheese as adjunct cultures in a cheese model system. Int J Food Microbiol 85:159–169
Baruzzi F, Maturante A, Morea M, Cocconcelli PS (2002) Microbial community dynamics during the Scamorza Altamurana cheese natural fermentation. J Dairy Sci 85:1390–1397
Beresford TP, Fitzsimons NA, Brennan NL, Cogan TM (2001) Recent advances in cheese microbiology. Int Dairy J 11:259–274
Chamba JF, Irlinger F (2004) Secondary and adjunct cultures. In: Fox PF, McSweeney PLH, Cogan TM, Guinee TP (eds) Cheese: chemistry, physics and microbiology. Elsevier, London
Corsetti A, Settanni L, Braga TM, De Fatima Silva Lopes M, Suzzi G (2008) An investigation on the bacteriocinogenic potential of lactic acid bacteria associated with wheat (Triticum durum) kernels and non-conventional flours. LWT—Food Part Sci Technol 41:1173–1182
De Angelis M, Corsetti A, Tosti N, Rossi J, Corbo MR, Gobbetti M (2001) Characterization of non-starter lactic acid bacteria from Italian ewe cheeses based on phenotypic, genotypic, and cell wall protein analyses. Appl Environ Microbiol 67:2011–2020
Didienne R, Defargues C, Callon C, Meylheuc T, Hulin S, Montel MC (2012) Characteristics of microbial biofilm on wooden vats (‘gerles’) in PDO Salers cheese. Int J Food Microbiol 156:91–101
Donnelly CW (2004) In: Fox PF, McSweeney PLH, Cogan TM, Guinee TP (eds) Cheese: chemistry, physics and microbiology. London, Chapman & Hall
Fitzsimons NA, Cogan TM, Condon S, Bereford T (1999) Phenotypic and genotypic characterization of nonstarter lactic acid bacteria in mature cheddar cheese. Appl Environ Microbiol 65:3418–3426
Franciosi E, Settanni L, Carlin S, Cavazza A, Poznanski E (2008) A factory-scale application of secondary adjunct cultures selected from lactic acid bacteria during “Puzzone di Moena” cheese ripening. J Dairy Sci 91:2981–2991
Franciosi E, Settanni L, Cavazza A, Poznanski E (2009) Biodiversity and technological potential of wild lactic acid bacteria from raw cows’ milk. Int Dairy J 19:3–11
Harrigan WF, McCance ME (1976) Laboratory methods in food and dairy microbiology. Academic Press, New York
Jack RW, Tagg JR, Ray B (1995) Bacteriocins of gram-positive bacteria. Microbiol Rev 59:171–200
Jackson CR, Fedorka-Cray PJ, Barrett JB (2004) Use of a genus- and species-specific multiplex PCR for identification of enterococci. J Clin Microbiol 42:3558–3565
Jans C, Lacroix C, Meile L (2012) A novel multiplex PCR/RFLP assay for the identification of Streptococcus bovis/Streptococcus equinus complex members from dairy microbial communities based on the 16S rRNA gene. FEMS Microbiol Lett 326:144–150
King N (1948) Modification of Voges–Proskauer test for rapid colorimetric determination of acetyl methyl carbimol plus diacetyl in butter. Dairy Industries 13:860–866
Lortal S, Di Blasi A, Madec M-N, Pediliggieri C, Tuminello L, Tanguy G, Fauquant J, Lecuona Y, Campo P, Carpino S, Licitra G (2009) Tina wooden vat biofilm. A safe and highly efficient lactic acid bacteria delivering system in PDO Ragusano cheese making. Int J Food Microbiol 132:1–8
Micari P, Sarullo V, Sidari R, Caridi A (2007) Physico-chemical and hygienic characteristics of the Calabrian raw milk cheese, Caprino d’Aspromonte. Turk J Vet Anim Sci 31:55–60
Moatsou G, Kandarakis I, Moschopoulou E, Anifantakis E, Alichanidis E (2001) Effect of technological parameters on the characteristics of Kasseri cheese made from raw or pasteurized ewes’ milk. Int J Dairy Technol 54:69–77
Mucchetti G, Bonvini B, Remagni MC, Ghiglietti R, Locci F, Barzaghi S, Francolino S, Perrone A, Rubiloni A, Campo P, Gatti M, Carminati D (2008) Influence of cheese-making technology on composition and microbiological characteristics of Vastedda cheese. Food Control 19:119–125
Parente E, Cogan TM (2004) In: Fox PF, McSweeney PLH, Cogan TM, Guinee TP (eds) Cheese: chemistry, physics and microbiology. London, Chapman & Hall
Parente E, Moschetti G, Coppola S (1998) Starter cultures for Mozzarella cheese. Ann Microbiol Enzim 48:89–109
Poyart C, Quesne G, Trieu-Cuot P (2002) Taxonomic dissection of the Streptococcus bovis group by analysis of manganese-dependent superoxide dismutase gene (sodA) sequences: reclassification of ‘Streptococcus infantarius subsp. coli’ as Streptococcus lutetiensis sp. nov. and of Streptococcus bovis biotype 11.2 as Streptococcus pasteurianus sp. nov. Int J Syst Evol Microbiol 52:1247–1255
Qadri SMH, Desilva MI, Zubairi S (1980) Rapid test for determination of esculin hydrolysis. J Clin Microbiol 12:472–474
Salvadori del Prato O (1998) Tecnologia Lattiero-Casearia. Edagricole, Bologna
Schillinger U, Lücke FK (1989) Antibacterial activity of Lactobacillus sake isolated from meat. Appl Environ Microbiol 55:1901–1906
Schlegel L, Grimont F, Ageron E, Grimont PAD, Bouvet A (2003) Reappraisal of the taxonomy of the Streptococcus bovis/Streptococcus equinus complex and related species: description of Streptococcus gallolyticus subsp. gallolyticus subsp. nov., S. gallolyticus subsp. macedonicus subsp. nov. and S. gallolyticus subsp. pasteurianus subsp. nov. Int J Syst Evol Microbiol 53:631–645
Settanni L, Di Grigoli A, Tornambé G, Bellina V, Francesca N, Moschetti G, Bonanno A (2012) Persistence of wild Streptococcus thermophilus strains on wooden vat and during the manufacture of a Caciocavallo type cheese. Int J Food Microbiol 155:73–81
Settanni L, Franciosi E, Cavazza A, Cocconcelli PS, Poznanski E (2011) Extension of Tosèla cheese shelf-life using non-starter lactic acid bacteria. Food Microbiol 28:883–890
Settanni L, Massitti O, Van Sinderen D, Corsetti A (2005) In situ activity of a bacteriocin-producing Lactococcus lactis strain. Influence on the interactions between lactic acid bacteria during sourdough fermentation. J Appl Microbiol 99:670–681
Settanni L, Moschetti G (2010) Non-starter lactic acid bacteria used to improve cheese quality and provide health benefits. Food Microbiol 27:691–697
Sherman JM (1937) The streptococci. Bacteriol Rev 1:3–97
Stackebrandt E, Goebel BM (1994) Taxonomic note: a place for DNA-DNA reassociation and 16S rRNA sequence analysis in the present species definition in bacteriology. Int J Syst Bacteriol 44:846–849
Stackebrandt E, Ebers J (2006) Taxonomic parameters revisited: tarnished gold standards. Microbiol Today 33:152–155
Todaro M, Francesca N, Reale S, Moschetti G, Vitale F, Settanni L (2011) Effect of different salting technologies on the chemical and microbiological characteristics of PDO Pecorino Siciliano cheese. Eur Food Res Technol 233:931–940
Tsakalidou E, Manolopoulou E, Kabaraki E, Zoidou E, Pot B, Kersters K, Kalantzopoulos G (1994) The combined use of whole-cell protein extracts for the identification (SDS-PAGE) and enzyme activity screening of lactic acid bacteria isolated from traditional Greek dairy products. Syst Appl Microbiol 17:444–458
Tsakalidou E, Zoidou E, Pot B, Wassill L, Ludwig W, Devriese LA, Kalantzopoulos G, Schleifer KH, Kersters K (1998) Identification of streptococci from Greek Kasseri cheese and description of Streptococcus macedonicus sp. nov. Int J Syst Bacteriol 48:519–527
Vaska VL, Faoagali JL (2009) Streptococcus bovis bacteraemia: identification within organism complex and association with endocarditis and colonic malignancy. Pathology 41:183–186
Weisburg W, Barns SM, Pelletier DA, Lane DJ (1991) 16S ribosomal DNA amplification for phylogenetic study. J Bacteriol 173:697–703
Williams AG, Choi SC, Banks JM (2002) Variability of the species and strain phenotype composition of the non-starter lactic acid bacterial population of Cheddar cheese manufactured in a commercial creamery. Food Rev Int 35:483–493
Acknowledgments
This work was financially supported by the project for industrial research and training PON01_02249 “Application of molecular biotechnologies and pro-technological microorganisms for the characterisation and valorisation of dairy and bakery chains of typical products” of the Italian Ministry of Education, University and Research (CUP: B11C11000430005). Mr. Simon Gill is also thanked for proof-reading the final version in English.
Author information
Authors and Affiliations
Corresponding author
About this article
Cite this article
Gaglio, R., Francesca, N., Di Gerlando, R. et al. Identification, typing and investigation of the dairy characteristics of lactic acid bacteria isolated from “Vastedda della valle del Belìce” cheeses. Dairy Sci. & Technol. 94, 157–180 (2014). https://doi.org/10.1007/s13594-013-0150-5
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s13594-013-0150-5