Abstract
Imidacloprid (IMD) may affect proteolysis, aspartate aminotransferase (AST), alanine aminotransferase (ALT), alkaline phosphatase (ALP), and global DNA methylation in honeybees. Queens, drones, and workers aged 1 or 20 days were exposed (free-flying colonies) to IMD (5 ppb and 200 ppb) in their diet. As a result, the colony depopulation did not occurred. IMD disturbed hemolymph/cuticle proteolysis; deactivated most of the cuticle protease inhibitors, activated hemolymph thiol and metal proteases and cuticle thiol proteases; downregulated ALP, ALT, AST; and increased DNA methylation in a caste- and age-dependent manner. The response in queens and workers differed, possibly due to eusocial evolution. Higher IMD dose had greater effects. The responses of ALP, ALT, AST, and DNA may reflect acceleration of biochemical senescence and epigenetic adaptation to IMD. All these biochemical side effects may lead to colony depopulation during future biotic/abiotic stress.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
1 Introduction
Imidacloprid (IMD) is a widely used neonicotinoid insecticide in agriculture (Simon-Delso et al. 2015). Sublethal doses of neonicotinoids impaired larval development, gland and neurobiological functions (Blacquière et al. 2012; Henry et al. 2015; Mužinić and Želježić 2018), longevity, resistance to biotic and abiotic stress (Van der Sluijs et al. 2013; Dively et al. 2015), foraging, and orientation (Colin et al. 2019; Jacob et al. 2019) in honeybee workers. IMD also increased the replacement rate, impaired reproductive traits, decreased body weight in queens (Williams et al. 2015; Chaimanee et al. 2016), and handicapped the sperm quality of drones (Ciereszko et al. 2017). Chronic, sublethal effects were more prevalent than acute toxicity (Hladik et al. 2018).
Because of all these harmful phenotypic effects, the biochemical side effects of IMD have been examined in honeybees, focusing on detoxification, immunocompetence, and antioxidative barriers, three mutually dependent systems of biochemical defense. Detoxification (Zhu et al. 2017) was impaired by IMD, consequently making bees more vulnerable to this pesticide and to other xenobiotics. Immunocompetence (Sánchez-Bayo et al. 2016; O’Neal et al. 2018) was suppressed by IMD, which in turn facilitated pathogen invasions, including Varroa destructor and Nosema ceranae. Harmful xenobiotics, including IMD, may induce oxidative stress that disturbs many metabolic pathways, including the antioxidative barrier itself (Chaimanee et al. 2016; Balieira et al. 2018).
However, knowledge of the biochemical side effects of IMD in honeybees is limited since we expect that both hemolymph and cuticle proteolysis, which are involved in honeybee resistance to harmful xenobiotics (Davies 1986; Grzywnowicz et al. 2009; Strachecka et al. 2010, 2012a), might also be impaired by IMD. Consequently, we hypothesized that exposure to IMD may lead to the destabilization of the honeybee hemolymph proteolytic system (hypothesis 1) and also impair the cuticle proteolytic layer (hypothesis 2), which is a crucial component of body surface defense against environmental stress including pathogens (Strachecka et al. 2012a, 2012b).
Metabolic disturbances caused by harmful xenobiotics in honeybees decreased the activities of the following biomarkers: aspartate aminotransferase (AST), alanine aminotransferase (ALT), and alkaline phosphatase (ALP) (Bajda et al. 2014; Strachecka et al. 2016). Exposure of bees to IMD should also change the activities of these biomarkers (hypothesis 3), but to our knowledge, this has not been studied. To the best of our knowledge, IMD-induced methylation of DNA has not been studied in honeybees either, only in target pests (Dong et al. 2013). However, DNA methylation is important for regulating gene expression in bees (Lyko and Maleszka 2011; Glastad et al. 2014; Cardoso-Júnior et al. 2018). It can be altered quickly by environmental factors, which may lead to further adaptation (Flores et al. 2013). Therefore, we hypothesized that exposure to IMD affects global DNA methylation in honeybees (hypothesis 4).
To examine these four hypotheses, we fed bees food containing 5 ppb (trace) and 200 ppb (more adverse) field relevent concentrations of IMD (Tapparo et al. 2011; Pohorecka et al. 2012; Henry et al. 2015; Colin et al. 2019) and monitored their cuticle/hemolymph proteolytic compounds, hemolymph ALP, ALT, and AST, and DNA global methylation in the free-flying colonies.
2 Materials and methods
2.1 Rearing colonies and experimental design
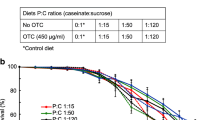
Fifteen rearing colonies of equal strength and structure, each headed by an Apis mellifera carnica sister queen and populating 2 hive supers (frame 360 × 260 mm), were divided into 3 groups of 5 colonies. The bees from two of these groups were fed sugar-water syrup (5:3 w/w) containing 5-ppb (IMD-5 group) or 200-ppb (IMD-200 group) IMD (Bayer Health Care AG, Leverkusen, Germany) diluted in water. The third, control group, was fed syrup only. Feeders (capacity 0.5 kg, restricted bee access) were refilled every 5 days after removing the unused syrup. Pastry made of the fresh pollen and bee food (API-Fortune HF 1575; ICKO, Bollène, France) containing 0-ppb, 5-ppb, and 200-ppb IMD was given to control, IMD-5, and IMD-200 colonies, respectively. Levels of IMD in the syrup and pastry amounted to 4.2 ppb and 196 ppb, respectively, when determined 3 months after preparation, using the method of Pohorecka et al. (2012). IMD levels were assayed (Pohorecka et al. 2012) in the bodies of about 100 workers at 1–10 days of age, sampled from each rearing colony fed IMD-200, and found to be 0.35 ± 0.24 ng/bee during rearing of the workers. IMD residuals were not found in bees feed IMD-5. The colonies were located in a place where the natural bee food was almost absent. In each group, 1-day-old (preimaginal IMD exposure) and 20-day-old (preimaginal + imaginal IMD exposure) workers/queens/drones were reared and kept. The experimental design was caste × age × group, i.e., 3 × 2 × 3 = 18 subclasses.
2.2 Rearing workers and drones exposed to IMD
Each queen was caged in a queen-excluder comb cage (1 drone + 2 worker combs) for 24 h to lay eggs and 4 weeks after beginning the IMD feeding in each of the rearing colonies. After releasing the queens, combs containing the eggs remained in the cages in their original colonies. Then, 24 h before the worker/drone emergence, the combs were artificially incubated (34.5 °C), the emerging bees were captured at 6-h intervals and either transported to the laboratory to assay 1-day-old workers/drones or marked with the colony-specific colors and returned into their original colonies for future assaying of the 20-day-old workers/drones. The drones were kept in lower hive supers isolated from the bottom and the top with queen excluders.
2.3 Rearing the queens exposed to IMD
Three queen-less nuclei were established in each of the three groups and fed the same group diet after the worker/drone examining was completed. Each of them contained a mix of workers from their original group. Next, 1-day-old larvae from all colonies in a given group were grafted into the nuclei belonging to the same group. Subsequently, the sealed queen cells were individually caged and incubated (34.5 °C), and the emerging queens were either transported to the laboratory (1 day old) or returned to their original nuclei to assay 20-day-old queens.
2.4 Sample processing
One pooled sample of fresh hemolymph was obtained (Łoś and Strachecka 2018) from each colony from the IMD-5, IMD-200, and control groups from both the 1-day-old and 20-day-old queens/drones/workers (90 pooled samples). Hemolymph from five queens/drones/workers constituted one pooled sample and was obtained using five capillaries (one capillary per bee). All five capillaries were placed into one sterile tube containing 150 μl of ice-cooled 0.6% NaCl and immediately frozen at – 80 °C for further analyses.
Ten workers/queens/drones were additionally captured from each colony according to the scheme described above, placed in a separate tube, and frozen at – 80 °C (90 pooled samples per 10 bees). Then, each sample was defrosted, shaken/rinsed in 10 ml of 1% (v/v) detergent (Triton X-100; Sigma-Aldrich) solution in distilled water, and centrifuged for 5 min at 775.44×g (3400 rpm). The solutions, which contained the cuticle proteins, were filtered through Miracloth and frozen again at – 40 °C for further biochemical analyses. Finally, five workers/queens/drones aged 1 and 20 days were acquired from each colony for each group and age and then frozen at – 40 °C for 1–2 months. After thawing, DNA was individually extracted from their heads and thoraces using the DNeasy Blood & Tissue Kit (Qiagen, Germany) following the manufacturer’s instructions. The DNA samples were stored at – 25 °C and later used to determine the global DNA methylation.
2.5 Biochemical analyses
The activities of acidic, neutral, and alkaline proteases were assayed (substrate – hemoglobin) in three respective buffers: 100-mM glycine-HCl at pH 2.4, 100-mM Tris-HCl at pH 7.0, and 100-mM glycine-NaOH at pH 11.2 using the methods described by Łoś and Strachecka (2018). The activities of the natural inhibitors of the acidic, neutral, and alkaline proteases were determined basing on the Lee and Lin (1995) method. In addition, the proteolytic activities after the addition of pepstatin A, phenylmethylsulfonyl fluoride, iodoacetamide, and o-phenanthroline were determined to detect the active enzyme centers (thiol, metal, asparagine, or serine proteases). The global DNA methylation was determined using the extracted DNA and the Imprint Methylated DNA Quantification Kit (MDQ1-96RXN, Sigma, USA) following the manufacturer’s instructions. The protocol for determining the activities of AST, ALT, and ALP and other details of all analytical procedures used was previously described by Łoś and Strachecka (2018).
2.6 Statistical analysis
One-way ANOVA and Duncan tests (p ≤ 0.05) were used to compare the averages of the control, IMD-5, and IMD-200 groups (treatment) nested within each caste (workers, drones, queens) and age (1 day old, 20 days old). The Bliss transformation was used in the case of DNA m5C. Principal component analysis (PCA, correlation matrix) was performed (Statistica version 12.0, StatSoft Inc., USA) for all biochemical characteristics taking into consideration treatment, caste, and age. The optimal number of principal components was determined based on the Cattell criterion. The data matrix for PCA had 11 columns and 90 rows for hemolymph and 7 columns and 90 rows for cuticle. Three-way ANOVA (treatment * caste * age) and PCA rotation procedure were also performed, and the results are shown in Online Resource 3, as they did not change the main findings.
3 Results
Amount of the consumed syrup, brood surface, and area of combs fully populated by bees was monitored during the IMD exposure in each of the 15 rearing colonies. We found no differences in these traits between the groups (Tukey test p ≥ 0.095). Therefore, IMD did not cause colony depopulation or differences in syrup consumption.
3.1 Effects of IMD on the proteolytic systems: hypotheses 1 and 2
IMD downregulated most acidic and alkaline hemolymph proteases in queens and all of them in workers and drones (Table I). Neutral proteases were upregulated in queens, independently of their age, whereas the response was age-dependent in drones and workers. IMD mostly downregulated all hemolymph protease inhibitors in queens but upregulated them in the other castes (in 100% in workers).
IMD downregulated the queen cuticle proteases, but they were mostly upregulated in drones and workers in IMD-200. Therefore, the responses were caste- and treatment-dependent (interactions). The neutral and alkaline cuticle protease inhibitors were completely suppressed by IMD in all castes. The acidic protease inhibitors were downregulated in queens. In the other castes, the inhibitors were either down- or upregulated depending on the treatment. The characteristics’ values for Tables I and II are given in Online Resource 1.
Generally, (1) higher IMD dose had greater effects, (2) protease downregulation was often not accompanied by upregulation of their inhibitors and vice versa, and (3) IMD influenced some proteases or their inhibitors depending on the enzyme type, bee age, and caste. The age-related changes of the proteolytic systems were analyzed in Online Resource 2.
Checking activities of protease centers shows that thiol and metal proteases were activated by IMD in hemolymph, but only the thiol proteases on cuticles in all castes. Asparagine and serine proteases were active, regardless of whether the bees were exposed to IMD.
3.2 Effects of IMD on ALT, AST, ALP and global DNA methylation: hypotheses 3 and 4
IMD downregulated ALT, AST, and ALP (Table II) and increased global DNA methylation. The higher the IMD dose, the greater the response. The response of all characteristics did not depend on caste and age. The values for all of these characteristics increased with age (Online Resource 2).
3.3 Effects of experimental factors and the trait correlation matrix
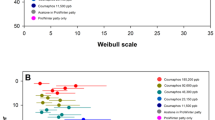
The first two PCA components (PC1 and PC2) define 83% of the variation on cuticles but only 58% of it in hemolymph; i.e., more factors influenced the hemolymph trait variation. In hemolymph, positive correlations (Figure 1) appeared between ALT, AST, and ALP and between all proteases, but negative correlations appeared between proteases and the global DNA methylation. Proteases were not correlated with their inhibitors. The inhibitors did not affect PC1 and PC2. ALT, AST, and ALP affected PC1. The remaining traits affected PC2. On cuticles, all proteases and the acid protease inhibitor were positively correlated with each other (the remaining inhibitors were not active); thus, the response to IMD was more clear. Bee age directly affected PC1 and PC2 only in hemolymph (Figure 2; Figs. 2, 3, and 4 should be analyzed together with the Figure SM5 included in Online Resource 3), so on cuticles, more inter-factor interactions might occur. Direct effects of the treatment were revealed both in hemolymph and on cuticles, but in hemolymph, the gradient reflecting the IMD dose was observed, whereas on cuticles, the only difference was between exposed and unexposed bees (Figure 3). No direct effects of caste on PC1 and PC2 were revealed, so caste has to affect the trait variation interacting with other factors (Figure 4). ANOVA confirmed all these findings (Online Resource 3).
4 Discussion
4.1 Proteolysis: hypotheses 1 and 2
Downregulation of most hemolymph proteases in workers exposed to IMD (Table I) is in line with the findings of Wilde et al. (2016), although they analyzed homogenates from the entire worker body. So, we have shown that efficient proteolysis of the toxic peptides formed during antioxidative activity (Davies 1986) can be indirectly impaired by IMD. This side effect might be particularly harmful to foragers facing toxic xenobiotics in ecosystems including contemporary, intensive agriculture, as we saw this impairment particularly in the 20-day-old workers. Proper proteolysis is also necessary for apoptosis (Eldeeb et al. 2018), phagocytosis, cellular adhesion, molecular signaling, melanization, zymogene activation, transport through the cell membranes, activation of antimicrobial peptides, etc. (see Bode et al. 1999; Evans et al. 2006; Strachecka et al. 2012a). Therefore, it is essential for worker bee’s metabolism and for expanding its life span. Upregulation of the hemolymph protease inhibitors by IMD occurred especially in workers, which may be the additional factor disturbing the balance between proteases and their inhibitors (Farady and Craik 2010) and, consequently, worker proteolytic activity. On the other hand, Bania and Polanowski (1999) suggested that insects may protect their cuticles against external threats by synthesizing protease inhibitors. Thus, our workers could attempt to overcome the harmful effects of IMD in this way.
The cuticle proteases and their inhibitors are essential for the honeybee cuticle defense (Grzywnowicz et al. 2009). Harmful xenobiotics impaired this system, which facilitated pathogen infections (Strachecka et al. 2012a, b; Frączek et al. 2013). The cuticle protease activities were disturbed by IMD in workers and drones dependent on the dose, caste, and age. In queens, the enzymes were downregulated. The cuticle protease inhibitors were almost all deactivated. Thus, IMD may handicap the cuticle defense, particularly in queens.
Balieira et al. (2018) showed that IMD decreases the concentrations of thiol proteins in honeybees. In this study, thiol and metal proteases were activated by IMD in hemolymph and only thiol proteases in the cuticle. This also occurred when bees were treated with formic acid (Strachecka et al. 2012a). However, Balieira et al. (2018) employed different protocols than we did (see conclusions of Henry et al. 2015). Therefore, more studies are needed to explain these discrepancies. Concluding, we have confirmed our hypotheses 1 and 2 about the harmful effects of IMD on the hemolymph and cuticle proteolysis. Simon-Delso et al. (2015), Sánchez-Bayo et al. (2016), and Mužinić and Želježić (2018) described harmful effects of neonicotinoids in honeybees, but they did not mentioned about impairment of the hemolymph and cuticle proteolysis. Thus, this study provides new information. Van der Sluijs et al. (2013) and Dively et al. (2015) demonstrated that pesticides, even when they do not cause direct bee mortality, make colonies potentially more sensitive to future pathogens or xenobiotic stresses. Therefore, we believe that, although colony depopulation was not observed during the exposure time in the present study, the metabolic side effects we observed can potentially make colonies more sensitive to future stressors, such as pathogens or xenobiotics.
The responses of the hemolymph and cuticle proteases and their inhibitors to IMD differed between the castes, particularly between queens and workers (Table I). This suggests that queens are biochemically different from workers, which may reflect different mechanisms of eusocial evolution (Münch et al. 2008; Amdam 2011) at the biochemical level. On the other hand, queens may have been exposed to less IMD than workers, as they were fed food processed by nurse bees. However, both 1-day-old queens and workers emerged from larvae were fed the processed food, but their responses to IMD were different.
This may be surprising, as our bees were fed with IMD, particularly in the bees fed during the preimaginal stages, though Colin et al. (2019) revealed that IMD delivered only during the larval stage strongly influences worker foraging. This requires further study.
4.2 ALT, AST, ALP, and the global DNA methylation: hypotheses 3 and 4
Similar to pathogens and other xenobiotics (Bajda et al. 2014; Strachecka et al. 2015, 2016), ALT, AST, and ALP were downregulated, independent of the caste and age, when bees were exposed to IMD (Table II). This confirmed our hypothesis 3 and pointed out the harmful, physiological side effects of IMD. Strachecka et al. (2015) revealed that the activities of these biomarkers increased from the 1st to the 30th day of the workers’ lives but then dropped when senescence became serious. A similar drop was observed in this study as a result of exposure to IMD (Online Resource 2), suggesting that IMD accelerates biochemical senescence.
The IMD-induced increase in the DNA global methylation (Table II) confirmed our hypothesis 4. This also suggested that honeybees can adapt to IMD through epigenetic mechanisms (Flores et al. 2013; Dong et al. 2013) or that at least the far-reaching IMD caused nonadaptive epigenetic changes occur. Similar epigenetic responses have already been observed in honeybees exposed to other harmful xenobiotics (Strachecka et al. 2012b, 2016). On the other hand, the global DNA methylation level increased with honeybee age in the present study (Online Resource 2), which corresponds to the results of Lyko and Maleszka (2011) and Cardoso-Júnior et al. (2018). Glastad et al. (2014) showed that demethylation of the honeybee genome by RG108 increases worker life span. Curcumin decreased global DNA methylation while increasing the worker life span (Strachecka et al. 2015). Thus, IMD seems to accelerate senescence at the epigenetic level.
In conclusion, considering that Jacob et al. (2019) estimated median LC50 values for IMD of 22.78 ng μL-1 and Tapparo et al. (2011) reported a lethal dose of 102 mg L-1, our results show that sublethal doses of neonicotinoids may cause honeybee colony failure not only because the exposed workers start foraging when younger, which unbalances the age-based division of labor (Jacob et al. 2019), but also because of faster biochemical/epigenetic bee senescence. The impairment in proteolysis we revealed here is also a new important fact that may help explain controversial mechanisms of colony depopulation when it is exposed to sublethal neonicotinoid doses in rural regions.
References
Amdam G.V. (2011) Social context, stress, and plasticity of ageing. Aging Cell 10, 18–27
Bajda M., Łoś A., Merska M. (2014) Effect of amphotericin B on the biochemical markers in the Hemolymph of honey bees. Med. Weter. 70, 766–769
Balieira K.V.B., Mazzo M., Bizerra P.F.V., de J. S. Guimarães A.R., Nicodemo D. et al (2018) Imidacloprid-induced oxidative stress in honey bees and the antioxidant action of caffeine. Apidologie 49, 572–562
Bania J., Polanowski A. (1999) Bioinsecticides and insect defense mechanisms. Postepy Biochem. 45, 143–150
Blacquière T., Smagghe G., van Gestel C.A., Mommaerts V. (2012) Neonicotinoids in bees, a review on concentrations, side-effects and risk assessment. Ecotoxicology 21, 973–992
Bode W., Fernandez-Catalan C., Nagase H., Maskos K. (1999) Endoproteinase – protein inhibitor interaction. APMIS 107, 3–10
Cardoso-Júnior C.A.M., Guidugli-Lazzarini K.R., Hartfelder K. (2018) DNA methylation affects the lifespan of honey bee (Apis mellifera L.) workers - Evidence for a regulatory module that involves vitellogenin expression but is independent of juvenile hormone function, Insect Biochem. Mol. Biol. 92, 21–29
Chaimanee V., Evans J.D., Chen Y., Jackson C., Pettis J.S. (2016) Sperm viability and gene expression in honey bee queens (Apis mellifera) following exposure to the neonicotinoid insecticide imidacloprid and the organophosphate acaricide coumaphos. J. Insect Physiol. 89, 1–8
Ciereszko A., Wilde J., Dietrich G.J., Siuda M., Bąk B. et al (2017) Sperm parameters of honeybee drone exposed to imidacloprid. Apidologie 48, 211–222
Colin T., Meikle W.G., Wu X., Barron A.B. (2019) Traces of a neonicotinoid induce precocious foraging and reduce foraging performance in honey bees. Environ. Sci. Technol. 53, 8252–8261
Davies K.J.A. (1986) Intracellular proteolytic systems may function as secondary antioxidant defenses, a hypothesis. J. Free Radicals Biol. Med. 2, 155–173
Dively G.P., Embrey M.S., Kamel A., Hawthorne D.J., Pettis J.S. (2015) Assessment of chronic sublethal effects of imidacloprid on honey bee colony health. PLoS One 10, 155–173
Dong D.-Z., Dong Y.-F., Ge Z.-M., Wang D.-M., Chen J. et al (2013) Effects of imidacloprid on DNA methylation polymorphism in the brown planthopper, Nilaparvata lugens (Hemiptera, Delphacidae). Acta Entomol. Sin. 56, 39–46
Eldeeb M.A., Fahlman R.P., Esmaili M., Ragheb M.A. (2018) Regulating apoptosis by degradation, the N-end rule-mediated regulation of apoptotic proteolytic fragments in mammalian cells. Int. J. Mol. Sci. 19, 3414
Evans J.D., Aronstein K., Chen Y.P., Hetru C., Imler J.L. et al (2006) Immune pathways and defence mechanisms in honey bee Apis mellifera. Insect Mol. Biol. 15, 645–656
Farady C.J., Craik C.S. (2010) Mechanisms of macromolecular protease inhibitors. Chembiochem. 11, 2341–2346
Flores K.B., Wolschin F., Amdam G.V. (2013) The role of methylation of DNA in environmental adaptation. Integr. Comp. Biol. 53, 359–372
Frączek R., Żółtowska K., Lipiński Z., Dmitryjuk M. (2013) The mutual influence of protein from Varroa destructor extracts and from honeybee hemolymph on their proteolytic activity - in vitro study. Acta Parasitol. 58, 317–323
Glastad K.M., Hunt B.G., Goodisman M.A.D. (2014) Evolutionary insights into DNA methylation in insects. Curr. Opin. Insect Sci. 1, 25–30
Grzywnowicz K., Ciołek A., Tabor A., Jaszek M. (2009) Profiles of the body-surface proteolytic system of honey bee queens, workers and drones, Ontogenetic and seasonal changes in proteases and their natural inhibitors. Apidologie 40, 4–19
Henry M., Cerrutti N., Aupinel P., Decourtye A., Gayrard M. et al (2015) Reconciling laboratory and field assessments of neonicotinoid toxicity to neonicotinoid toxicity to honeybees. Proc. Biol. Sci. 282, 1819
Hladik M.L., Main A.R., Goulson D. (2018) Environmental Risks and Challenges Associated with Neonicotinoid Insecticides. Environ. Sci.Technol. 52, 3329–3335
Jacob C.R.O., Malaquias J.B., Zanardi O.Z., Silva C.A.S., Jacob J.F.O. et al (2019) Oral acute toxicity and impact of neonicotinoids on Apis mellifera L. and Scaptotrigona postica Latreille (Hymenoptera: Apidae). Ecotoxicology 28, 744–753
Lee T., Lin Y. (1995) Trypsin inhibitor and trypsin – like protease activity in air – or submergence – grown rice (Oryza sativa L.) coleoptiles. Plant Sci. 106, 43–54
Łoś A., Strachecka A. (2018) Fast and cost-effective biochemical spectrophotometric analysis of solution of insect “blood” and body surface elution. Sensors 18, 1494
Lyko F., Maleszka R. (2011) Insects as innovative models for functional studies of DNA methylation. Trends Genet. 27, 127–164
Münch D., Amdam G., Wolschin F. (2008) Ageing in a eusocial insect, molecular and physiological characteristics of life span plasticity in the honey bee. Funct. Ecol. 22, 407–421
Mužinić V., Želježić D. (2018) Non-target toxicity of novel insecticides. Arh. Hig. Rada Toksikol. 69, 86–102
O’Neal S.T., Anderson T.D., Wu-Smart J.Y. (2018) Interactions between pesticides and pathogen susceptibility in honey bees. Curr. Opin. Insect Sci. 26, 57–62
Pohorecka K., Skubida P., Miszczak A., Semkiw P., Sikorski P. (2012) Residues of neonicotinoid insecticides in bee collected plant materials from oilseed rape crops and their effect on bee colonies. J. Apic. Sci. 56(2), 115–134
Sánchez-Bayo F., Goulson D., Pennacchio F., Nazzi F., Goka K. et al (2016) Are bee diseases linked to pesticides? — A brief review. Environ. Int. 89–90, 7–11
Simon-Delso N., Amaral-Rogers V., Belzunces L.P., Bonmatin J.M., Chagnon M., Downs C., Furlan L., Gibbons D.W., Giorio C., Girolami V., Goulson D., Kreutzweiser D.P., Krupke C.H., Liess M., Long E., McField M., Mineau P., Mitchell E.A., Morrissey C.A., Noome D.A., Pisa L., Settele J., Stark J.D., Tapparo A., van Dyck H., van Praagh J., van der Sluijs J., Whitehorn P.R., Wiemers M. (2015) Systemic insecticides (neonicotinoids and fipronil), trends, uses, mode of action and metabolites. Environ. Sci. Pollut. Res. Int. 22, 5–34
Strachecka A., Gryzińska M., Krauze M. (2010) The influence of environmental pollution on the protective proteolytic barrier of the honey bee Apis mellifera mellifera. Pol. J. Environ. Stud. 19, 855–859
Strachecka A., Paleolog J., Borsuk G., Olszewski K. (2012a) The influence of formic acid on the body surface proteolytic system at different developmental stages in Apis mellifera L. workers. J. Apic. Res. 51, 252–262
Strachecka A., Paleolog J., Olszewski K., Borsuk G. (2012b) Influence of amitraz and oxalic acid on the cuticle proteolytic system of Apis mellifera L. workers. Insects 3, 821–832
Strachecka A., Olszewski K., Paleolog J. (2015) Curcumin stimulates biochemical mechanisms of Apis mellifera resistance and extends the apian life-span. J. Apic. Sci. 59, 129–140
Strachecka A., Olszewski K., Paleolog J. (2016) Varroa treatment with bromfenvinphos markedly suppresses honeybee biochemical defence levels. Entomol. Exp. Appl. 160, 57–71
Tapparo A., Giorio C., Marzaro M., Marton D., Soldà L., Girolami V. (2011) Rapid analysis of neonicotinoid insecticides in guttation drops of corn seedlings obtained from coated seeds. J. Environ. Monit. 13, 1564–1568
van der Sluijs J.P., Simon-Delso N., Goulson D., Maxim L., Bonmatin J.-M. et al (2013) Neonicotinoids, bee disorders and the sustainability of pollinator services. Curr. Opin. Environ. Sustain. 5, 293–305
Wilde J., Frączek R.J., Siuda M., Bąk B., Hatjina F. et al (2016) The influence of sublethal doses of imidacloprid on protein content and proteolytic activity in honey bees (Apis mellifera L.). J. Apic. Res. 55, 212–220
Williams G.R., Troxler A., Retschnig G., Roth K., Yanez O. et al (2015) Neonicotinoid pesticides severely affect honey bee queens. Sci. Rep. 5, 14621
Zhu Y.C., Yao J., Adamczyk J., Luttrell R. (2017) Synergistic toxicity and physiological impact of imidacloprid alone and binary mixtures with seven representative pesticides on honey bee (Apis mellifera). PLoS One 12, 1–16
Funding
The lab part was financed from ZKB/MN-5 and ZKZ/DS-5 Ministry of Science and Higher Education, Poland.
Author information
Authors and Affiliations
Contributions
JW concept; JW, MS, BB designed/made field – AS, ŁW lab part. AS, ŁW, JP statistics. AS, JP, JW evaluated results; JP interpreted results and wrote the paper. All authors read and approved final manuscript.
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Additional information
Manuscript editor: Monique Gauthier
L'imidaclopride affecte de façon marquée la protéolyse de l'hémolymphe, les biomarqueurs, la méthylation globale de l'ADN et la couche protéolytique de la cuticule chez les abeilles domestiques occidentales.
abeille domestique / imidaclopride / protéolyse / méthylation de l'ADN / biomarqueur.
Imidacloprid beeinflusst die Hämolymphproteolyse, Biomarker, die globale DNA Methylierung und die proteolytische Kutikulaschicht de Westlichen Honigbiene.
Honigbiene / Imidacloprid / Proteolyse / DNA-Methylierung / Biomarker.
Publisher’s note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Paleolog, J., Wilde, J., Siuda, M. et al. Imidacloprid markedly affects hemolymph proteolysis, biomarkers, DNA global methylation, and the cuticle proteolytic layer in western honeybees. Apidologie 51, 620–630 (2020). https://doi.org/10.1007/s13592-020-00747-4
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s13592-020-00747-4