Abstract
Genome-wide overwintering gene expression studies in bees are of critical importance to understand their survival, life cycles, and fitness. In this study, we use RNA sequencing to characterize the genes and gene functions associated with overwintering adult females compared with active season females for the small carpenter bee, Ceratina calcarata. We found extensive changes in gene expression associated with overwintering, including an underrepresentation of genes related to muscle fibers and an overrepresentation of genes related to lipid metabolic processes. These data suggest inactive, overwintering bees invest heavily in the production of proteins related to fat storage and divest transcriptional activity away from cellular and muscular structural processes. This study provides the first characterization of overwintering gene expression as important baseline data of a major life history event relevant to survival and health for a wild bee pollinator.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
1 Introduction
Wild bees are ubiquitous pollinators of agricultural crops (Klein et al. 2007; Stubbs et al. 1997), and thus, understanding their physiology and overwintering survival is paramount to protecting and conserving this invaluable resource. Many bees are well-adapted to temperate climates, but phenologies are changing, and the sustainability of wild bee populations is a major concern (Bartomeus et al. 2011, 2013; Ollerton et al. 2014). To date, there are few studies on wild bee overwintering physiology (Skandalis et al. 2011; Sakagami et al. 1981; Yocum et al. 2005, 2006; Wasielewski et al. 2013, 2014; Alford 1969) and even fewer that focus on the molecular underpinning of the overwintering process (Yocum et al. 2005, 2006; Wasielewski et al. 2014; Torson et al. 2015). Most studies focus on single life stages in lab-reared environments. No studies to date have characterized the genes and gene functions associated with overwintering individuals compared with active season adults in natural environments.
Carpenter bees of the tribes Ceratinini and Xylocopini are bees with truly cosmopolitan distributions (Michener 1979). They are found on all continents and ecoregions except the Antarctic (Michener 1979; Rehan and Schwarz 2015). These species are known for their cold hardiness which is tightly linked to their adult overwintering strategy (Michener 1979; Skandalis et al. 2011). Carpenter bees remain in their natal nests, above ground during the winter months, forcing them to cope with freezing temperatures as opposed to avoiding them in the way honey bees or bumble bees do (Alford 1969; Southwick 1991; Skandalis et al. 2011). Their flexible habitat requirements, large geographical range, and ability to thrive in areas that experience subfreezing temperatures makes them ideal for studies aimed at characterizing overwintering physiology. Moreover, carpenter bees are an emerging model system for studying the molecular genetic basis of wild bee physiology and behavior. Recently, the first small carpenter bee, Ceratina calcarata, transcriptome was published providing a valuable resource for genome-wide characterization of physiological processes underlying bee health (Rehan et al. 2014). This bee is economically important because of its generalist pollination services (Kennedy et al. 2013). Transcriptional profiling of wild bees is the first step to better understanding their adaptations to overwintering and molecular responses to climate change.
In this study, we focused on C. calcarata, a small carpenter bee found in eastern North America. This species ranges from Georgia north to Ontario and east to Nova Scotia (Rehan and Sheffield 2011; Shell and Rehan 2015). Unlike many temperate insects (Danks 1991; Denlinger 2002), C. calcarata overwinters as an adult (Rehan and Richards 2010a). C. calcarata are a subsocial species, meaning females build their nests solitarily but remain on the nest and continue to care for their offspring until they reach adulthood. Adult male and female offspring spend the winter months in their natal nest and disperse to establish new nests in spring (Rehan and Richards 2010a). Here, we present the first genome-wide examination of overwintering gene expression patterns in a small carpenter bee. We characterize gene expression profiles of field-collected overwintering C. calcarata and contrast these against previously published active season gene expression profiles in the same species (Rehan et al. 2014). Based on existing overwintering gene expression literature (Colinet et al. 2007, 2012; Dunning et al. 2013, 2014; Lee et al. 1998; Purac et al. 2008; Ragland et al. 2010; Rinehart et al. 2007, 2010; Robich et al. 2007; Torson et al. 2015; Qin et al. 2005; Yocum et al. 2005, 2015), we predict that differential expression would be found in genes relating to heat shock proteins, membrane lipids, cuticular proteins, and cytoskeletal and muscular alterations. We also provide comparisons of our results to previous studies on overwintering gene expression and cold tolerance in other insects, in order to assess whether similar or different genes and pathways are associated with overwintering across taxa.
2 Methods
2.1 Bee sampling and field collections
Twenty overwintering females were collected from independent nests during mid-winter (January) and were flash frozen with liquid nitrogen in the field. C. calcarata overwinter as adults in their natal nests (Rehan and Richards 2010a, b). Internal and external nest temperatures were taken using a digital, noncontact, infrared thermometer (Micro Temp MT100). Three temperature readings were taken on the exterior surface of each stem followed by three temperature readings of the interior of the nest. Ambient temperatures were −0.7 °C (±4.71 °C) and internal nest temperature was 0.1 °C (±3.81 °C) at the time of nest collection (Figure S1). Weather data for Durham, NH was taken from the Global Historical Climate Network (http://www.ncdc.noaa.gov/climate-information).
The non overwintering groups included five distinct groups of females representing all three active seasons, including mothers from spring, summer, and autumn, as well as two distinct categories of daughters from autumn (dwarf and regular-sized; Rehan et al. 2014). The overwintering females collected were also regular-sized daughters, as mothers and dwarf daughters do not survive past the active season (Rehan and Richards 2010a).
2.2 RNA extraction, sequencing, and quality filtering
Brain tissue was dissected for RNA extraction in RNAlater from flash frozen samples. The methods used for RNA extraction, RNA-Seq library preparation, and sequencing were the same as those reported in Rehan et al. (2014) and are as follows: total RNA was extracted from the brain tissue of 20 overwintering adult females using the RNeasy Kit (Qiagen) followed by quality control of extractions using spectrophotometry (NanoDrop) and BioAnalyzer (Aligent). Brain tissue was used so that overwintering gene expression profiles could be directly compared to previous active season, female brain gene expression profiles of C. calcarata examined by Rehan et al. (2014). An RNA-Seq library for the sample was prepared from these 20 pooled brains using Illumina’s TruSeq RNA-Seq sample preparation kit, and sequencing was run on an Illumina HiSeq 2000. Library preparation and 100 bp paired end sequencing were conducted at the Iowa State University DNA Facility. After sequencing, the quality of the read data was checked using FastQC (Andrews 2010), and the data were overall high quality. There were some lower quality bases present, mostly near the ends of reads; however, this is a typical observation with Illumina data and does not indicate an issue. The adapter sequences (added during the library preparation) were removed using fastx_clipper (Fastx Toolkit Version 0.0.13) (Gordon and Hannon 2010). The final stage of quality control was implemented using the Trim Perl script (Nikhil Joshi, unpublished; code available from http://wiki.bioinformatics.ucdavis.edu/index.php/Trim.pl) with a quality threshold of greater than or equal to 20 and a length threshold of 50 bases. Quality filtering and adapter removal resulted in removal of approximately 20 % of reads from the library (Table S1). Raw data have been submitted to the NCBI Sequence Read Archive (SRA) with accession number SRX541305.
2.3 Read mapping, read counts, and differential expression analyses
The C. calcarata transcriptome was previously assembled and described by Rehan et al. (2014). The transcriptome consists of 358,709 transcripts. Homologs for 152,809 of the transcripts were identified using Blastx of NCBI’s nonredundant (NR) database with an E value threshold of 1e−4. Of the annotated transcripts, 88 % had best blast hits to other bee species, and 9 % had hits to other hymenopterans (ants and wasps) (Rehan et al. 2014). We used this transcriptome to generate read counts for genes expressed in overwintering bees.
The paired end reads from the library were mapped to the C. calcarata transcriptome using Bowtie2 (version 2.1.0) (Langmead et al. 2009). Quantification of the abundance of aligned transcripts was conducted using eXpress (version 1.3.1) (Roberts and Pachter 2012). Differential expression between the active period (spring, summer, and autumn) and the inactive period (winter) of the C. calcarata life cycle were calculated using DESeq (version 1.12.0) (Anders and Huber 2010) found in the Bioconductor Repository using the statistical software R (version 3.0.1).
2.4 Annotation and gene ontology analyses
The Blast2GO (Conesa et al. 2005) suite was used to identify homologs of the differentially expressed transcripts by running a tBLASTx of NCBI’s nonredundant database (E value threshold ≤1e−4). Functional annotation of the C. calcarata transcripts were based on the best BLAST hits to Drosophila melanogaster sequences in the NCBI Entrez protein database (E value threshold ≤1e−4). DAVID (Huang et al. 2009) was used to test for significant (P value <0.05) enrichment of the gene ontology (GO) terms associated with the differentially expressed transcripts.
2.5 Comparative transcriptomics
Lists of differentially expressed transcripts, uniquely expressed transcripts (expressed in only one female group), and lists of enriched GO terms from overwintering C. calcarata were compared to transcriptomic studies of cold-related gene expression on fruit fly (D. melanogaster; Zhang et al. 2011; Telonis-Scott et al. 2009), flesh fly (Sarcophaga crassipalpis; Rinehart et al. 2010), Asian tiger mosquito (Aedes albopictus; Poelchau et al. 2013), and the leafcutter bee (Megachile rotundata; Torson et al. 2015; Yocum et al. 2015). We identified putatively homologous sequences between C. calcarata and D. melanogaster using tBLASTx (E value ≤1e−4). Homologous sequences from transcriptomic studies on other species were also determined using best BLAST comparisons to D. melanogaster. With these putatively homologous sequences, we tested for significant overlap in differentially expressed genes, unique genes, and GO terms between pairs of species using a two-tailed hypergeometric test.
3 Results
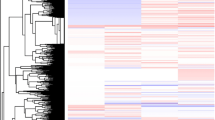
Differential expression (DE) analysis identified 2703 DE transcripts in the overwintering C. calcarata adult females compared to previously published brain gene expression profiles in active season females of this species (Rehan et al. 2014; Figure 1). Of the DE transcripts from overwintering females, 658 were upregulated (24 %), while the remaining 2045 (76 %) were downregulated (Figure 1). Comparing DE transcripts revealed an extensive difference in the number of DE transcripts between overwintering (1428 DE transcripts on average) and active season (283 DE transcripts on average) life history stages (Figure 2).
Relative expression profiles of differentially expressed transcripts from the six focal time points assayed scaled by library size and mean transcript expression (values between −2 and 2). Overwintering Ceratina calcarata daughters had dramatically different transcript expression profiles with 2703 DE transcripts and 4996 uniquely expressed transcripts compared to previously published brain gene expression profiles in active season (spring, summer, autumn) females of this species combined (Rehan et al. 2014).
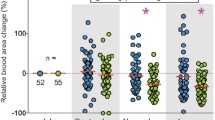
The number of differentially expressed transcripts (DETs) in all pairwise comparisons among overwintering and active season females in Ceratina calcarata. Overwintering females had a five times greater number of differentially expressed transcripts (on average 1428 DETs; red and orange shading) versus comparisons among females across the active season (on average 283 transcripts; yellow shading).
3.1 Functional annotation and GO enrichment
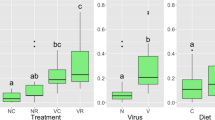
Among the 2703 overwintering DE transcripts, 2688 (99 %) had best blast hits (E-value ≤1e−4) to D. melanogaster, of which 182 transcripts had functional annotation in DAVID (Table S2). GO analysis revealed 33 significantly enriched GO terms (Table S3, Figure 3). Among the 658 upregulated genes, we found enrichment for several GO terms associated with lipid metabolism, including five biological process terms [lipid catabolic process (GO:0016042), phospholipid metabolic process (GO:0006644), cellular lipid metabolic process (GO:0044255), cellular lipid catabolic process (GO:0044242), lipid metabolic process (GO:0006629)], one cellular component term [lipid particle (GO:0005811)], and three molecular function terms [phospholipase C activity (GO:0004629), phospholipase activity (GO:0004620), lipase activity (GO:0016298] (Table S3). Significant downregulated genes contributed to the negative enrichment of four cellular component GO terms related to muscle development: [sarcomere (GO:0030017), myofibril (GO:0030016), contractile fiber part (GO:0044449), contractile fiber (GO:0043292)].
Analysis of gene ontology (GO) levels, revealed over- (green) or under- (red) represented categories in Ceratina calcarata females between overwintering (this study) and active season (Rehan et al. 2014). The three separate GO categories (biological process, cellular component, and molecular function) provide overlapping information, as a single gene can have multiple annotations from the GO categories. P values represent uncorrected results for each hypergeometric test. A subset of 13/33 overwintering GO terms is shown here; for the full list, see Table S3.
3.2 Transcripts unique to overwintering C. calcarata
Standard analyses of DE transcripts do not allow for the identification of transcripts that are uniquely expressed to a particular group (i.e., present in one group but absent in others), even though these transcripts provide valuable information about extreme gene expression differences. Therefore, we examined presence/absence of transcripts from each sequencing library to identify transcripts unique to each of the female groups. There were relatively few unique transcripts (expressed in only one female group) for each of the active season females (range 1103–2258, average 1819, Table S1) compared to overwintering females (14,996 transcripts). Of these unique overwintering transcripts, 14,671 (97.8 %) had best blast hits (E value <1e−4) to D. melanogaster, of which 1249 transcripts had functional annotation in DAVID, including cuticular protein genes, heat shock protein genes, as well as isoforms of both lipid and Actin genes (Table S4). The GO analysis revealed unique overwintering transcripts had significant positive enrichment (FDR ≤0.05) for actin filament-based process (GO:0030029), actin cytoskeleton organization (GO:0030036), and muscle cell differentiation (GO:0042692).
3.3 Comparative transcriptomics
We compared our lists of overwintering differentially expressed transcripts, uniquely expressed transcripts, and our list of enriched GO terms to transcriptomic studies examining gene expression responses to cold in fruit fly (D. melanogaster; Zhang et al. 2011; Telonis-Scott et al. 2009), flesh fly (S. crassipalpis; Rinehart et al. 2010), Asian tiger mosquito (A. albopictus; Poelchau et al. 2013), and leafcutter bee (M. rotundata; Torson et al. 2015; Yocum et al. 2015; Table S5). Although none of these studies is directly comparable to our study due to differences in life stages, tissues examined, and technologies (see Table S5), we used these comparisons to begin to address whether there are any common patterns of gene activity related to overwintering between flies and bees.
Overall, we found no significant overlap between our list of overwintering DE transcripts with those reported in diapause and cold exposure gene expression studies in Diptera and leafcutter bees (hypergeometric tests, P > 0.05; Table S5). In addition, we found no significant overlap between our list of enriched overwintering GO terms with those reported from DE transcripts in Diptera and leafcutter bees (hypergeometric tests, P > 0.05; Table S5).
We also looked at unique overwintering transcripts from C. calcarata and although the overlap was not significant (hypergeometric tests, P > 0.05), we found some individual transcripts of interest that overlapped between bees and flies. Three genes overlapped between previous studies on cold exposure in Drosophila and unique overwintering transcripts in C. calcarata. Upheld, a gene associated with muscle cell components (GO:0044449, GO:0030017, GO:0030016), was uniquely expressed in overwintering C. calcarata and upregulated in D. melanogaster subjected to repeated cold exposure (Zhang et al. 2011). We also found two transcripts uniquely expressed in overwintering C. calcarata (CG11674, mRNA splicing; CG1529, zinc and nucleic acid binding) that were also significantly upregulated in artificially selected lines of cold resistant D. melanogaster (Telonis-Scott et al. 2009).
4 Discussion
Here, we present the first genome-wide overwintering gene expression study in a small carpenter bee. These data are the first comparison of gene expression across the entire colony cycle of a wild bee and have provided novel gene lists and gene ontology categories involved with overwintering physiology and cold hardiness in C. calcarata. We found substantial changes in overwintering gene expression with 2703 DE transcripts in comparison to active season females of this species. The change in gene expression is extensive, suggesting large-scale shifts in transcriptional activity during overwintering. Interestingly, the majority (76 %) of differential expression was the result of transcript downregulation during overwintering and includes members of the Actin gene family, an indicator of overall transcriptional activity level. Far fewer genes (24 %) were upregulated, and these were significantly positively enriched for lipid metabolic processes. These results suggest that overwintering bees have a large shut down in the production of proteins related to cellular and muscular structural maintenance, in favor of a large, concomitant shift to the production of proteins related to lipid storage and metabolism. Greatly increased lipid stores have been proposed to be an essential adaptation for overwintering, especially in adult Hymenoptera (Strassmann et al. 1984; Toth et al. 2009); thus, our transcriptional data agree well with physiological studies on increased fat body size in overwintering adult bees and wasps (Wasielewski et al. 2014). However, our transcriptome data are from brains, not fat bodies, and therefore suggest that metabolic shifts for overwintering are not only occurring in fat bodies but are also evident in neural tissues.
Many of the identified genes and molecular processes related to overwintering in C. calcarata were similar to those previously reported for other insect species, such as increased lipid metabolism, and alterations in proteins related to the structure of the cuticle and muscle tissue. In addition, there was some overlap between C. calcarata unique overwintering transcripts and genes related to cold response in D. melanogaster (Zhang et al. 2011; Telonis-Scott et al. 2009). Despite these gross similarities, our quantitative bee-fly comparisons of cold-related gene expression patterns (on both the transcript and GO levels) did not uncover significant overlaps. This may be attributable to species-specific molecular responses to cold, as suggested by a recent comparative study that also reported a lack of conserved gene expression related to dormancy between C. elegans, D. melanogaster, and S. crassipalpis (Ragland et al. 2010; Colinet et al. 2012). Alternatively, the lack of conservation may be explained by the fact that these studies examined different life stages and tissues (Table S5). In addition, these results are complicated by the fact that for all the studies we compared, we used D. melanogaster genes as a common reference for functional annotation. This could be problematic because many genes related to cold response many not be well-characterized in D. melanogaster, a naturally tropical species (Sinclair et al. 2007). At this point, it remains unclear whether differences between gene expression studies are due to differing overwintering strategies, technical differences between studies, or whether there is an overall lack of genetic similarity of nonmodel species to D. melanogaster. Further studies in nonmodel insects with diverse overwintering strategies are needed to fully understand the molecular underpinnings of these differing responses to extreme temperature changes.
Overall, our results do suggest some similarities between overwintering gene expression in C. calcarata with known molecular mechanisms associated with temperature response in other insects. Below, we synthesize some of our specific findings for C. calcarata with previous studies on other insects in order to better understand the patterns we report and pave the way for future studies on the most informative genes and/or pathways.
4.1 Heat shock proteins
Previous studies have found that numerous genes belonging to the heat shock protein (HSP) gene family respond to exposures to low temperatures (Colinet et al. 2007, 2012; Ragland et al. 2010; Rinehart et al. 2007, 2010; Robich et al. 2007; Qin et al. 2005; Yocum et al. 2005). We observed three heat shock proteins (heat shock proteins 60, cognate 2, and cognate 3) uniquely expressed in overwintering females, which lends some support for the importance of HSPs during diapause in the small carpenter bee (Table S4). This is in accordance with the concept that HSP genes are generally upregulated during diapause or cold exposure compared to active season insects.
Although there is a general consensus that HSP genes are upregulated during diapause, some studies have also found variation in the magnitude of upregulation and the time points at which these genes are activated. Colinet et al. (2010) found that in Drosophila HSP gene, upregulation occurs during a recovery period (directly following cold exposure when temperature is increased) rather than during the cold exposure itself. This may explain the lack of upregulation we found in HSP genes, as our samples were collected mid-winter when the bees were still enduring cold temperature exposure and did not experience a recovery period.
4.2 Cytoskeletal and muscular alterations
In this study, we found downregulation of two actin genes, Actin 87E and 57B and underrepresentation of sarcomere myofibril and contractile fiber cellular components (Figure 3). In addition, we found unique expression of the myofilin gene and underrepresentation of several GO terms relating to muscular components including actin filament-based process and actin cytoskeleton organization (Table S3). Downregulation of actin genes during diapause has also been found in the leafcutter bee, M. rotundata (MRactin; Yocum et al. 2005) and the gypsy moth, L. dispar (130 and 45 kDa actin proteins; Lee et al. 1998). Conversely, diapause studies in two Diptera, C. pipiens and S. crassipalpis, showed upregulation of putative actin genes (DQ322244 and FL483261, respectively) (Yocum et al. 2005; Lee et al. 1998; Robich et al. 2007; Rinehart et al. 2010). A recent transcriptional response study on the leafcutter bee, M. rotundata found upregulation of actin-related protein 2,3 complex subunit 3-like in fluctuating temperature environments (Torson et al. 2015). These opposing results may not be unexpected due to the fact that actin genes are a very large family of genes, and it is likely that different members of the family are inversely regulated at a given time (Rinehart et al. 2010).
4.3 Cuticular proteins
Cuticle differentiation and reorganization is a common response to cold temperatures found in many insects (Dunning et al. 2013, 2014; Colinet et al. 2012; Purac et al. 2008). Although we did not find differential gene expression in this gene family in our brain transcription assays, we did identify two cuticular protein-coding genes (cuticular protein 49AC and 60D) that were uniquely expressed in overwintering females. Upregulation of cuticular genes and proteins during overwintering was also found in the Antarctic springtail, C. antarcticus, and parasitic wasp A. colemani, but downregulation was found in another parasitic wasp, P. volucre (Colinet et al. 2007, 2010; Purac et al. 2008). Qin et al. (2005) found upregulation of transcripts that are potentially responsible for encoding membrane proteins in D. melanogaster; however, the exact function of those transcripts were unknown, and there was no overlap with our list of differentially regulated transcripts. Comparative studies between populations and species of New Zealand alpine stick insects in the genus Micrarchus suggest that overwintering-related gene expression patterns are highly dependent on genotype and environmental conditions (Dunning et al. 2013, 2014). If duration of cold exposure is critical, then it is possible that the females collected in our study may have not experienced the necessary conditions required to induce a change in cuticular gene regulation.
4.4 Lipid metabolism
In this study, we found GO enrichment for lipid metabolism (Table S3; Figure 3). Many ecological and physiological studies have identified lipid storage and metabolism as a necessity for insect survival during overwintering periods and exposure to cold temperatures (Toth et al. 2009; Strassmann et al. 1984; Wasielewski et al. 2014). However, to our knowledge, there are few studies that have found significant overwintering-related differential expression associated with lipid metabolic processes. Colinet et al. (2012) found that a lipid carrier was downregulated during diapause in the parasitic wasp, P. volucre, possibly signifying that the lipids were being stored.
5 Conclusions
This study provides an important first genome-wide study of overwintering gene expression in small carpenter bees to which future bee species that naturally experience a range of environmental conditions and overwintering temperatures can be compared. Previous studies have noted the importance of wild bees to natural and agroecosystems (Klein et al. 2007; Stubbs et al. 1997); therefore, understanding overwintering survival is the key in furthering necessary wild bee conservation. C. calcarata is an ideal species to examine the molecular genetic mechanisms of cold tolerance due to their cold hardiness and large geographical range.
Future studies including additional comparisons of gene expression as individuals enter and exit diapause would provide additional insights into the time course and specificity of overwintering gene regulation in the small carpenter bee, C. calcarata. Additional information could be gleaned from lab-based studies where temperature exposure and duration can be controlled. Comparative studies between this and other Ceratina species across their eastern North American range would be of interest to determine the relative roles of local adaptation and lineage-specific gene regulation in response to climate change.
References
Alford, D. (1969) A study of the hibernation of bumblebees (Hymenoptera: Bombidae) in Southern England. J. Anim. Ecol. 38, 149
Anders, S., Huber, W. (2010) Differential expression analysis for sequence count data. Genome Biol. 11, R106
Andrews, S. (2010) FastQC: A Quality Control Tool for High Throughput Sequence Data. Available at: http://www.bioinformatics.bbsrc.ac.uk/projects/fastqc/
Bartomeus, I., Ascher, J.S., Wagner, D., Danforth, B.N., Colla, S., Kornbluth, S., Winfree R. (2011) Climate-associated phenological advances in bee pollinators and bee-pollinated plants. Proc. Nat. Acad. Sci. 108, 20645–20649
Bartomeus, I., Ascher, J.S., Gibbs, J., Danforth, B.N., Wagner, D.L., Hedtke, S.M., Winfree, R. (2013) Historical changes in northeastern US bee pollinators related to shared ecological traits. Proc. Nat. Acad. Sci. 110, 4656–4660
Colinet, H., Nguyen, T., Cloutier, C., Michaud, D., Hance, T. (2007) Proteomic profiling of a parasitic wasp exposed to constant and fluctuating cold exposure. Insect Biochem. Molec. Biol. 37, 1177–1188
Colinet, H., Lee, S., Hoffmann, A. (2010) Temporal expression of heat shock genes during cold stress and recovery from chill coma in adult Drosophila melanogaster. FEBS J. 277, 174–185
Colinet, H., Renault, D., Charoy-Guevel, B., Com, E. (2012) Metabolic and proteomic profiling of diapause in the aphid parasitoid Praon volucre. PLOS One 7, e32606
Conesa, A., Gotz, S., Garcia-Gomez, J., Terol, J., Talon, M., Robles, M. (2005) Blast2GO: a universal tool for annotation, visualization and analysis in functional genomics research. Bioinformatics 21, 3674–3676
Danks, H. (1991) Winter Habitats and Ecological Adaptations for Winter Survival. In: Lee, R., Denlinger, D. (eds.) Insects at Low Temperatures, pp. 231–259. Chapman and Hall, New York
Denlinger, D. (2002) Regulation of diapause. Annu. Rev. Entomol. 47, 93–122
Dunning, L., Dennis, A., Park, D., Sinclair, B., Newcomb, R., Buckley, T. (2013) Identification of cold-responsive genes in a New Zealand alpine stick insect using RNA-Seq. Comp. Biochem. Physiol. Part D Genom. Proteomic. 8, 24–31
Dunning, L., Dennis, A., Sinclair, B., Newcomb, R., Buckley, T. (2014) Divergent transcriptional responses to low temperature among populations of alpine and lowland species of New Zealand stick insects (Micrarchus). Mol. Ecol. 23, 2712–2726
Gordon, A., Hannon, G. (2010) FASTQ/A Short-Reads Pre-Processing Tools. Available at: http://hannonlab.cshl.edu/fastx_toolkit
Huang, D., Sherman, B., Lempicki, R. (2009) Systematic and integrative analysis of large gene lists using DAVID bioinformatics resources. Nat. Protoc. 4, 44–57
Kennedy, C.M., Lonsdorf, E., Neel, M.C., Williams, N.M., Ricketts, T.H., et al. (2013) A global quantitative synthesis of local and landscape effects on wild bee pollinators in agroecosystems. Ecol. Lett. 16, 584–599
Klein, A., Vaissiere, B., Cane, J., Steffan-Dewenter, I., Cunningham, S., Kremen, C., Tscharntke, T. (2007) Importance of pollinators in changing landscapes for world crops. Proc. R. Soc. B 274, 303–313
Langmead, B., Trapnell, C., Pop, M., Salzberg, S. (2009) Ultrafast and memory-efficient alignment of short DNA sequences to the human genome. Genome Biol. 10, R25
Lee, K., Hiremath, S., Denlinger, D. (1998) Expression of actin in the central nervous system is switched off during diapause in the gypsy moth, Lymantria dispar. J. Insect Physiol. 44, 221–226
Michener, C. (1979) Biogeography of the bees. Ann. Missour. Bot. Gard. 66, 277
Ollerton, J., Erenler, H., Edwards, M., Crockett, R. (2014) Extinctions of aculeate pollinators in Britain and the role of large-scale agricultural changes. Science 346, 1360–1362
Poelchau, M., Reynolds, J., Elsik, C., Denlinger, D., Armbruster, P. (2013) RNA-Seq reveals early distinctions and late convergence of gene expression between diapause and quiescence in the Asian tiger mosquito, Aedes albopictus. J. Exp. Biol. 216, 4082–4090
Purac, J., Burns, G., Thorne, M., Grubor-Lajsic, G., Worland, M., Clark, M. (2008) Cold hardening processes in the antarctic springtail, Cryptopygus antarcticus: clues from a microarray. J. Insect Physiol. 54, 1356–1362
Qin, W., Neal, S., Robertson, R., Westwood, J., Walker, V. (2005) Cold hardening and transcriptional change in Drosophila melanogaster. Insect Mol. Biol. 14, 607–613
Ragland, G., Denlinger, D., Hahn, D. (2010) Mechanisms of suspended animation are revealed by transcript profiling of diapause in the flesh fly. Proc. Nat. Acad. Sci. 107, 14909–14914
Rehan, S., Richards, M. (2010a) Nesting biology and subsociality in Ceratina calcarata (Hymenoptera: Apidae). Can. Ent. 142, 65–74
Rehan, S., Richards, M. (2010b) The influence of maternal quality on brood sex allocation in the small carpenter bee, Ceratina calcarata. Ethology 116, 876–887
Rehan, S., Schwarz, M. (2015) A few steps forward and no steps back: long-distance dispersal patterns in small carpenter bees suggest major barriers to back-dispersal. J. Biogeogr. 42, 485–494
Rehan, S., Sheffield, C. (2011) Morphological and molecular delineation of a new species in the Ceratina dupla species-group (Hymenoptera: Apidae: Xylocopinae) of eastern North America. Zootaxa 2873, 35–50
Rehan, S., Berens, A., Toth, A. (2014) At the brink of eusociality: transcriptomic correlates of worker behaviour in a subsocial bee. BMC Evol. Biol. 14, 260
Rinehart, J., Li, A., Yocum, G., Robich, R., Hayward, S., Denlinger, D. (2007) Up-regulation of heat shock proteins is essential for cold survival during insect diapause. Proc. Nat. Acad. Sci. 104, 11130–11137
Rinehart, J., Robich, R., Denlinger, D. (2010) Isolation of diapause-regulated genes from the flesh fly, Sarcophaga crassipalpis, by suppressive subtractive hybridization. J. Insect Physiol. 56, 603–609
Roberts, A., Pachter, L. (2012) Streaming fragment assignment for real-time analysis of sequencing experiments. Nat. Methods 10, 71–73
Robich, R., Rinehart, J., Kitchen, L., Denlinger, D. (2007) Diapause-specific gene expression in the northern house mosquito, Culex pipiens L. identified by suppressive subtractive hybridization. J. Insect Physiol. 53, 235–245
Sakagami, S., Tanno, K., Enomoto, O. (1981) Cold resistance of the small carpenter bee Ceratina flavipes restudied. Low Temp. Sci. 39, 1–7
Shell, W, Rehan, S. (2015) Recent and rapid diversification of the small carpenter bees in eastern North America. Biol. J. Linn. Soc. doi:10.1111/bij.12692
Sinclair, B., Gibbs, A., Roberts, S. (2007) Gene transcription during exposure to, and recovery from, cold and desiccation stress in Drosophila melanogaster. Insect Molec. Biol. 16, 435–443
Skandalis, D., Richards, M., Sformo, T., Tattersall, G. (2011) Climate limitations on the distribution and phenology of a large carpenter bee, Xylocopa virginica (Hymenoptera: Apidae). Can. J. Zool. 89, 785–795
Southwick, E. (1991) Overwintering in Honey Bees: Implications for Apiculture. In: Lee, R., Denlinger, D. (eds.) Insects at Low Temperature, pp. 446–460. Chapman and Hall, New York
Strassmann, J.E., Lee, R.R., Rojas, R.R., Baust, J.G. (1984) Caste and sex-differences in cold-hardiness in the social wasps, Polistes annularis and Polistes exclamans (Hymenoptera, Vespidae). Insectes Soc. 31, 291–301
Stubbs, C., Drummond, F., Allard, S. (1997) Bee conservation and increasing Osmia spp. in Maine lowbush blueberry fields. Northeast. Nat. 4, 133
Telonis-Scott, M., Hallas, R., McKechnie, S.W., Wee, C.W., Hoffmann, A.A. (2009) Selection for cold resistance alters gene transcript levels in Drosophila melanogaster. J. Insect Physiol. 55, 549–55
Torson, A.S., Yocum, G.D., Rinehart, J.P., Kemp, W.P., Bowsher, J.H. (2015) Transcriptional responses to fluctuating thermal regimes underpinning differences in survival in the solitary bee Megachile rotundata. J. Exp. Biol. 218, 1060–1068
Toth, A.L., Bilof, K.B.J., Henshaw, M.T., Hunt, J.H., Robinson, G.E. (2009) Lipid stores, ovary development, and brain gene expression in Polistes metricus females. Insectes Soc. 56, 77–84
Wasielewski, O., Wojciechowicz, T., Giejdasz, K., Krishnan, N. (2013) Overwintering strategies in the red mason solitary bee physiological correlates of midgut metabolic activity and turnover of nutrient reserves in females of Osmia bicornis. Apidologie 44, 642–656
Wasielewski, O., Szczepankiewicz, D., Giejdasz, K., Wojciechowicz, T., Bednarova, A., Krishnan, N. (2014) The potential role of adiponectin- and resistin-like peptides in the regulation of lipid levels in the hemolymph of over-wintering adult females of Osmia bicornis. Apidologie 45, 491–503
Yocum, G., Kemp, W., Bosch, J., Knoblett, J. (2005) Temporal variation in overwintering gene expression and respiration in the solitary bee Megachile rotundata. J. Insect Physiol. 51, 621–629
Yocum, G., Kemp, W., Bosch, J., Knoblett, J. (2006) Thermal history influences diapause development in the solitary bee Megachile rotundata. J. Insect Physiol. 52, 1113–1120
Yocum, G., Rinehart, J., Horvarth, D., Kemp, W., Bosch, J., Alroobi, R., Salem, S. (2015) Key molecular processes of the diapause to post-diapause quiescence transition in the alfalfa leafcutting bee Megachile rotundata identified by comparative transcriptome analysis. Physiol. Entomol. 40, 103–112
Zhang, J., Marshall, K.E., Westwood, J.T., Clark, M.S., Sinclair, B.J. (2011) Divergent transcriptomic responses to repeated and single cold exposures in Drosophila melanogaster. J. Exp. Biol. 214, 4021–4029
Acknowledgments
We thank Amy Geffre for assistance with RNA extraction and quality control and members of the Rehan and Toth labs for reviewing the manuscript. Funding from the University of New Hampshire to SMR and Iowa State University to ALT supported this work.
Author information
Authors and Affiliations
Corresponding author
Additional information
Manuscript editor: Klaus Hartfelder
Profilage transcriptionnel de l’expression du gène associé à l’hivernage chez l’abeille charpentière Ceratina calcarata
Ceratini / Xylocopinae / diapause / résistance au froid / survie / abeille
Erfassung des Genexpressionsprofils der Überwinterungsphase der Keulhornbiene Ceratina calcarata
Ceratini / Xylocopini / Diapause / Kälteresistenz / Überleben von Bienen
Rights and permissions
Open Access This article is distributed under the terms of the Creative Commons Attribution 4.0 International License (http://creativecommons.org/licenses/by/4.0/), which permits unrestricted use, distribution, and reproduction in any medium, provided you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made.
About this article
Cite this article
Durant, D.R., Berens, A.J., Toth, A.L. et al. Transcriptional profiling of overwintering gene expression in the small carpenter bee, Ceratina calcarata . Apidologie 47, 572–582 (2016). https://doi.org/10.1007/s13592-015-0402-x
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s13592-015-0402-x