Abstract
The objective of this study was to compare how well the detection of Paenibacillus larvae in samples of live bees or in accumulated winter debris collected from the bottom of beehives relates to symptoms of American foulbrood in honey bee colonies. Fifty-eight colonies in one commercial beekeeping operation were inspected for disease symptoms and assayed for P. larvae using culture-based techniques and PCR. The results show that culture-based techniques are more accurate than recently published PCR methods for detecting the bacterium in clinically diseased colonies, and that the prognostic value of bacterial colony counts from bee samples is superior to colony counts from debris. However, if the objective is to monitor the prevalence of the bacterium irrespective of disease symptoms, the preferable method is PCR analysis of accumulated winter hive debris.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
1 Introduction
The honey bee larval disease American foulbrood (AFB) is one of the major pathogenic threats to beekeeping. The disease is highly contagious, capable of killing infected colonies and causing large economic losses worldwide. AFB is caused by the Gram-positive, spore-forming bacterium Paenibacillus larvae (Genersch et al. 2006). Spores are only infectious to young larvae that become infected by consuming spore-contaminated food (Hitchcock et al. 1979). As few as 10 spores are enough to infect and kill a larva (Brødsgaard et al. 1998; Woodrow 1942). The most conspicuous symptom in colonies suffering from clinical AFB disease is irregular brood capping, with capped and uncapped cells scattered irregularly across the brood frames. The caps are dark, sunken and often punctured, emitting a ‘foul’ odor. The brown, glue-like remains of the dead larvae in these cells form a highly characteristic ropey thread when pulled out with a wooden stick or a match, which is a reliable field diagnosis for AFB. The remains of the larvae finally form a hard scale in the bottom of the cell. Such scales contain millions of bacterial spores that remain viable for decades (Genersch 2010; Hasemann 1961).
Traditional diagnosis of AFB is based on the observation of clinical symptoms in the hive as described above (De Graaf et al. 2006) and microbial cultivation of material from infected colonies. Several diagnostic protocols have been published for the cultivation of P. larvae from extracted honey (De Graaf et al. 2006; Nordström and Fries 1995; Pernal and Melathopoulos 2006), beehive debris (Bzdil 2007; Titera and Haklova 2003) and adult bees (Gillard et al. 2008; Lindström and Fries 2005). A direct comparison between these showed that cultivating the bacteria from live bees relates better to colony-level disease symptoms than cultivating from honey samples (Lindström and Fries 2005; Nordström et al. 2002). Culture-based laboratory P. larvae assays are definitive for confirming the diagnosis of AFB suspected colonies, but may be less suitable for surveillance and epidemiological studies. Early detection of sub-clinical levels of P. larvae in an apiary or beekeeping practice is imperative for effective preventative disease control. One common objection to culture-based AFB diagnosis methods has been that many spores do not germinate in vitro. For instance, Dingman and Stahly (1983) concluded that only 6 % of P. larvae spores germinate under culture conditions. A more recent study (Forsgren et al. 2008) also shows that there is a significant variability in the germination rate on solid media between different P. larvae genotypes.
In recent years, a number of PCR-based protocols have been published as an alternative to traditional cultivation-based techniques for the diagnosis, characterization and quantification of P. larvae. Conventional PCR has been used to detect P. larvae in brood, foulbrood scales and in honey (Alippi et al. 2004; Bakonyi et al. 2003; Dobbelaere et al. 2001; Lauro et al. 2003) as well as in beehive debris (Ryba et al. 2009). More recently quantitative, real-time PCR-based methods have been developed for the quantification of P. larvae in honey and larvae (Han et al. 2008; Martinez et al. 2010).
Several of these molecular protocols were developed for the detection of P. larvae DNA in vegetative bacterial cells cultivated on agar medium or in total DNA isolated from larvae with an active infection (Dobbelaere et al. 2001; Lauro et al. 2003). However, for epidemiological and quantitative studies it is imperative to be able to detect not only the vegetative cells but also the bacterial spores from different materials such as honey, debris, adult bees and wax. There are specific protocols available for the extraction of genetic material from the endospores using commercially available extraction kits (Ryba et al. 2009).
Analysis of the debris collected from the bottom of the beehive is a simple and non-invasive method for assessing the health status of a bee colony. However, it is not known how well the published P. larvae hive debris assays, either PCR-based (Ryba et al. 2009) or culture-based (Titera and Haklova 2003), relate to colony-level disease symptoms, compared to similar assays of adult bees. Moreover, we still know relatively little about the consistency of culture- or PCR-based assays for the detection and quantification of P. larvae from other types of honey bee colony samples.
The main objectives of this study were therefore to: (1) evaluate how well two types of beehive samples (adult bees and hive debris) relate to actual disease symptoms in the colony (2) to compare two methods for assaying bee and debris samples, by real-time PCR and by microbial cultivation on semi-selective media. By sampling adult bees and debris from the same hives, assessing the samples with the two different methods and applying separate logistic regressions to the data, this study provides the first statistical evaluation of some of the current diagnostic methods of AFB.
2 Materials and methods
2.1 Sampling
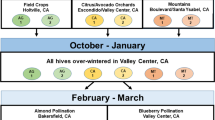
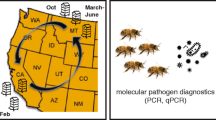
A total of 58 colonies in five honey bee (Apis mellifera L) apiaries in a commercial beekeeping operation in Uppsala County, Sweden were subjected to visual brood inspection for AFB symptoms in May 2012. Two types of samples were collected from these colonies; adult bees from the brood nest and hive debris. Two to three hundred bees were shaken into a plastic bag, killed by freezing, and kept frozen until processed. The debris was collected from the bottom board representing the accumulated hive debris from the winter period. Disease symptoms were observed in 11 of the inspected colonies, but in several cases only a few diseased larvae were found. In the rest of the colonies (n = 47) there were no visual symptoms of disease in the brood.
2.2 Cultivation of P. larvae from bees
The bee samples were cultivated on MYPGP-agar plates as described by Nordström and Fries (1995). In brief, 100 worker bees were placed into a filter grinding bag (Neoreba®), and 20 mL of sterile 0.9 % NaCl was added. The bees were crushed and the fluid produced centrifuged for 10 min at 27,000×g. The resulting pellet was resuspended in 2 mL sterile NaCl, heat shocked at 85 °C for 10 min and spread onto 3 MYPGP-agar plates (10 μL each). The plates were incubated at 36 °C in 5 % CO2 for 7 days and the numbers of P. larvae colonies were counted. The data was normalized and presented as colony forming units (CFU) per bee.
2.3 Cultivation of P. larvae from hive debris
The hive debris was processed using the Tween 80 method (Bzdil 2007). Briefly, 1 g of debris was mixed with 8.5 mL sterile water and 0.5 % Tween 80 (pre-warmed to 70 °C), carefully mixed and immersed for 30–60 min in a 70 °C water bath. The tubes were removed from the water bath and the samples settled for 2 h (room temperature) before 2 mL of the liquid was transferred to a new tube and mixed with an additional 2 mL of sterile water. The samples were mixed thoroughly, heated at 85 °C for 10 min and spread onto 3 MYPGP-agar plates (10 μL each). The plates were incubated at 36 °C in 5 % CO2 for 7 days and the numbers of P. larvae colonies were counted. The data is presented as CFU per gram hive debris.
2.4 Comparison of DNA extraction methods
A pilot experiment was performed to evaluate two previously published extraction protocols for isolating genomic bacterial DNA from P. larvae spores (Martinez et al. 2010; Ryba et al. 2009). A stock spore suspension of P. larvae (CCUG 48979 Forsgren et al. 2008) in sterile ddH2O was prepared and checked under a phase-contrast light microscope (×1,000) to confirm presence of endospores and absence of vegetative cells. The spores were counted in a Helber Bacteria Counting Chamber (Hawksley, UK) and tenfold dilutions (107, 106, 105 and 104 spores/mL) of the suspension were prepared. Bees and hive debris from healthy colonies were spiked with different concentrations of P. larvae spores (Table I). Briefly, one whole bee or 1 g debris was crushed in 1 mL of spore suspension, before being subjected to DNA extraction using two commercial extraction kits, QIAamp® DNA Mini Kit (Qiagen)(Ryba et al. 2009) and UltraClean Soil DNA isolation kit (Mo Bio Laboratories Inc., Martinez et al. 2010), following the manufacturer’s instructions.
2.5 Bacterial DNA extraction from bee samples
One hundred worker bees were placed into a filter grinding bag (Neoreba®), and 20 mL of sterile ddH2O was added. The bees were crushed and 1 mL of the fluid produced was immediately subjected to DNA extraction using the QIAamp® genomic DNA isolation mini kit for Gram-positive bacteria (Qiagen). The DNA was eluted with 100 μL elution buffer and stored at −20 °C until further use.
2.6 Bacterial DNA extraction from hive debris
One gram of debris was subjected to DNA extraction following the protocol of Ryba and co-workers (Ryba et al. 2009). Briefly, 1 g of debris was mixed with 5 mL of sterile, distilled water and shaken at room temperature for 1 h. One hundred microliters of the suspension was immediately subjected to DNA extraction using the QIAamp® genomic DNA isolation mini kit for Gram-positive bacteria (Qiagen). The DNA was eluted with 100 μL elution buffer and stored at −20 °C until further use.
2.7 Real-time PCR for P. larvae 16s RNA sequence
The quantitative real-time PCR assay used here was modified from Martinez et al. (2010). Briefly, the reaction contained 10 μL of SsoFast™ EvaGreen® Supermix (Bio-Rad), 0.4 μM each of primers PL2-Fw and PL2-Rev, 2 μL of template, and the final reaction volume was adjusted to 20 μL with nuclease-free water. A negative control containing water instead of DNA template was included in each run. All real-time qPCR reactions (standards, unknown samples and controls) were performed in duplicate in neighboring wells on the sample plate. The results reported are an average of the duplicates. The amplification and data acquisition was carried out using a CFX Connect® (Bio-Rad) real-time PCR machine under the following cycling condition: enzyme activation step, 98 °C for 2 min, PCR cycling (40 cycles of 98 °C for 5 s (denaturation), 58 °C for 5 s (annealing/extension) and data collection. The identity of the amplified product was confirmed using a melting curve analysis, by raising the temperature from 55 to 95 °C in 0.5 °C increments with a hold of 1 s at each increment. The assay specificity and the presence/absence of non-specific amplification products were determined through the melting temperature(s) (T m) of the amplified product(s). Each set of PCR assays included serial dilutions of DNA extracted from known concentrations of P. larvae spores (total microscopic count) as external standards, for relating the qPCR data to spore counts (Forsgren et al. 2008). Standard curves were prepared by using serial dilutions of target DNA ranging from 102 to 108 as quantification standards in every run. The quantification data was normalized to tissue or debris weight and converted to bacterial cells per bee or bacterial cells per gram debris.
2.8 Statistical analyses
To determine if the sample type (adult bees vs. debris) influenced the amount of bacteria detected with the two assessment methods (culture vs. PCR), we used two-sample t tests to test for difference in means, and F tests to test for differences in variances. To evaluate the effectiveness of the four combinations of sample types and detection methods in detecting AFB-symptomatic colonies, we fitted separate logistic regressions to each of the four combinations, using the statistical package R, version 3.0.0 (R Core Team 2013). We fitted the health status of the hive, healthy or diseased, as a binary response variable, and the log10-transformed amount of bacteria detected by each of the four combinations as the predictor. We evaluated the predictive power of the four combinations from the model output, through the Akaike’s Information Criterion (AIC) statistic. A lower AIC value for a particular combination of detection method and sample type indicates a better fit of the model to the data, and therefore a superior ability to predict clinical symptoms. However, the AIC score is only a relative measure of model performance and does not give the absolute goodness-of-fit of a model. Therefore, we present two additional criteria for evaluating the method–sample type combinations. First, the amount of variation explained by each model is presented by a pseudo-R 2 (Nakagawa and Schielzeth 2012), given by formula 1—(residual deviance/null deviance). R 2 takes values between 0 and 1, and the higher the values; the more variation in the data is explained by the model. Second, to get an estimate of the detection sensitivity for each combination of sample type and detection method, we calculated the predictor value at which 95 % (95-quantile; q 95) of the colonies were diagnosed with clinical symptoms, using the function
where a is the slope and b is the intercept of each logistic regression. A low q 95-value predicted from a specific model means that a low amount of bacteria is necessary for detecting clinically diseased colonies. Consequently, an assessment method with a lower q 95-value is more sensitive than a method with a high q 95-value.
3 Results
3.1 Cultivation of P. larvae from bees versus debris
Eleven of the inspected colonies showed clinical signs of AFB. All the adult bee samples (11/11) and all but one debris sample (10/11) from the diseased colonies were positive for P. larvae when cultured as described above (Figure 1b). Moreover, in all the sampled colonies, irrespective of disease symptoms, 34 out of 58 debris samples and 33 out of 58 bee samples were found positive for P. larvae when cultured (Figure 1a). A Welch two-sample t test showed that significantly more bacteria could be detected in the debris samples than in bee samples, using the culture method (mean 2.56 and 1.09, respectively, t = −3.74, DF = 103.69, P = 0.0003; Figure 2). The two sample types also differed in their variance, with the debris samples generally showing more variance than the bee samples (F test; F = 0.52, DF = 57, P = 0.015).
3.2 Real-time PCR detection of P. larvae from bees versus debris
A Welch two-sample t test showed that on average, there were significantly more bacteria detected in the debris samples than in bee samples, using qPCR (mean 3.46 and 1.35, respectively, t = −4.72, DF = 101.50, P = 7.4 × 10−6; Figure 2). The sample types also differed in their variance, with the debris samples generally showing more variance than the bee samples (F test; F = 0.48, DF = 57, P = 0.006).
3.3 Comparison of DNA extraction methods
There were clear differences between the two extraction methods regarding the efficiency in extracting DNA from P. larvae spores. The QIAamp® DNA Mini Kit proved to be more efficient for DNA isolation from pure P. larvae spore suspensions, with six out of six extracted aliquots of the 103 suspension producing a PCR product, whereas product was only detected from 1 out of 6 bacterial DNA templates of the same concentration extracted using the UltraClean Soil DNA isolation kit (Table I). Hence, the bee and debris samples in this study were extracted using QIAamp® DNA Mini Kit, according to the manufacturer’s instructions. However, when a healthy bee spiked with bacterial spore suspension was extracted, the UltraClean Soil DNA isolation kit was superior to the QIAamp® DNA Mini Kit (see Table I), indicating a better efficiency in removing inhibitory compounds from the bee extract.
3.4 Predicting health status of colonies from amount of bacteria
The accuracy of predicting colony health status using bacterial amounts was higher for bee samples than for debris samples, as indicated by the better model fit (lower AIC value and higher R 2 value), irrespective of the detection method used (Table II). Moreover, for each sample type, culturing bacteria is always superior to PCR. Plotting the fitted models (Figure 3) shows that the combination of bee samples and culture-based assays (steepest slope) is far better at predicting colony health status than the three other combinations (shallower slopes). The combination also had the lowest level of bacteria needed to predict 95 % of diseased colonies (Table II).
Raw data and fitted logistic regressions showing the probability of detecting American foulbrood disease) as a function of log10-transformed number of bacteria in sample, using two different assessment methods; culture (left panel) and PCR (right panel) and two different sample types; adult bees (crosses and dotted lines) and PCR (open circles and solid lines).
4 Discussion
Our results clearly demonstrate that if the objective is to find clinically diseased colonies, the prognostic value of bacterial colony counts from bee samples is superior to bacterial colony counts from debris. All the bee samples from visibly diseased colonies contained detectable P. larvae spores using cultivation, whereas one sample of debris gave a false negative result (i.e., disease symptom in the colony, but no colonies produced using microbial cultivation). However, using bacterial colony counts for the quantification of P. larvae in hive debris and adult bee samples revealed significantly higher spore loads in the debris (Figure 2). This is not surprising since bees collected from the brood chamber represent a “snapshot” of the actual disease status of the brood, whereas hive debris collected during the wintering period reflects an historical accumulation of bacterial spores from the colony during that time. However, in all the sampled colonies combined, irrespective of disease symptoms, the results for the two materials are comparable, with 34 out of 58 debris samples and 33 out of 58 bee samples found positive for P. larvae when cultured (Figure 1a).
Significantly more spores were detected in the hive debris than in the bee samples not only when colony counts were used, but also using the PCR assay. It is likely that the bacterial loads are higher in hive material accumulated for months than in live bees whose life-span is measured in weeks. Moreover, the probable presence of PCR-inhibitors in adult bee samples and the difficulty to extract bacterial DNA from the endospores may lead to a general underestimation of bacterial loads in all sample materials, using PCR-based methods.
Culture-based detection methods allow the quantification of microorganisms with considerable precision, but PCR- based assays are often said to be superior for many reasons. However, quantification of PCR products should not necessarily be equated with the number of organisms present in the original sample (Vaneechoutte and Van Eldere 1997). Amplification of targets from clinical samples may have rather limited sensitivity, in contrast to what is often reported from research studies (Vaneechoutte and Van Eldere 1997). This can clearly be seen by our data where we can detect P. larvae in some of the sampled bee colonies using bacterial colony counts but not with the PCR method. This may depend on a variety of factors such as inhibitors of the PCR reaction and/or failures in the extraction protocols (Wilson 1997). Rusenova et al. (2012) reported low sensitivity in their PCR assay when using standard DNA extraction protocols. Incubation of the samples in a rich nutrient medium for 1 h at 37 °C considerably improved the protocol, presumably because the spore envelope is altered and the bacterial DNA has become more accessible for extraction.
Martinez et al. (2010) reported an assay detection limit of 2 P. larvae spores g−1 honey contrasting to earlier reports of a detection limit of 105 spores g−1 of honey (Ryba et al. 2009) and 283 spores g−1 of honey (Alippi et al. 2004). These differences may very well be due to differences in extraction methods, highlighting the possibility that comparative evaluation of DNA extraction and purification may be more important than comparisons of different amplification methods. Our comparison of two commercial extraction kits revealed large differences in their efficiencies for isolating bacterial DNA from pure P. larvae spore suspensions. Much to our surprise, DNA isolated with the UltraClean Soil DNA isolation kit failed to consistently produce a PCR product when used on suspensions with less than 103 spores mL−1, whereas QIAamp® DNA Mini Kit showed to be much more effective for the isolation of pure bacterial DNA (Table I), but was less effective when bee or debris was added. Possibly, the UltraClean Soil DNA isolation kit is more effective for the removal of polymerase inhibitors, but more work needs to be done to further evaluate the DNA extraction and purification of P. larvae spores.
Bee samples were more accurate than debris samples for predicting colony health status using P. larvae quantification, irrespective of detection method used. The combination of using pooled samples of adult bees and bacterial colony counts on MYPGP-agar to assess the bacterial load is superior to any of the other combinations (Figure 2). This is probably due to: (1) samples of adult bees better reflects the actual health status of the bee colony, and (2) the superiority of culture-based assessment of the bacterial loads to the PCR-based protocols used here, due to the detection limitations in the nucleic acid amplification (Vaneechoutte and Van Eldere 1997). However, it should be noted that the predictions from the logistic regressions were based on data from only 11 colonies for which clinical symptoms were observed. These results should therefore be validated through follow-up evaluations of molecular vs. cultivation detection techniques involving a larger sample size of symptomatic colonies.
To conclude, analyzing adult bee samples comparing PCR assay and culture, CFU counts proved to be superior. P. larvae was detected in 100 % of the bee samples from diseased colonies using cultivation whereas the bacteria was only detected in 8 out of 11 colonies using PCR assay, presumably depending on polymerase inhibitors in the bees (Boncristiani et al. 2011) and/or failures the DNA extraction protocols (Wilson 1997). Assaying bee samples using PCR was the least sensitive method for detecting symptoms of AFB when looking at the total number of bee colonies assayed (Figure 1a). However, using the PCR method on debris samples revealed not only the highest bacterial levels but also the highest proportion of infected bee colonies (40 out of 58 investigated bee colonies). If the objective is to detect sub-clinical levels of P. larvae irrespective of disease symptoms, hive debris and PCR is clearly the material and method preferred. However, in order to apply and improve any PCR assay for the detection and quantification of P. larvae spores in various bee related materials, such as adult bees, wax, debris, and pollen, DNA extraction and purification techniques need to be compared and validated.
References
Alippi, A.M., Lopez, A.C., Aguilar, O.M. (2004) A PCR-based method that permits specific detection of Paenibacillus larvae subsp. larvae, the cause of American foulbrood of honey bees, at the subspecies level. Lett. Appl Microbiol 39(1), 25–33
Bakonyi, T., Derakhshifar, I., Grabensteiner, E., Nowotny, N. (2003) Development and evaluation of PCR assays for the detection of Paenibacillus larvae in honey samples: comparison with isolation and biochemical characterization. Appl Environ Microbiol 69(3), 1504–1510
Boncristiani, H., Li, J., Evans, J.D., Pettis, J., Chen, Y. (2011) Scientific note on PCR inhibitors in the compound eyes of honey bees, Apis mellifera. Apidologie 42(4), 457–460
Brødsgaard, C., Ritter, W., Hansen, H. (1998) Response of in vitro reared honey bee larvae to various doses of Paenibacillus larvae larvae spores. Apidologie 29, 569–578
Bzdil, J. (2007) Detection of Paenibacillus larvae spores in the debris and wax of honey bee by the Tween 80 method. Acta Veterinaria Brno 76(4), 643–648
De Graaf, D.C., Alippi, A.M., Brown, M., Evans, J.D., Feldlaufer, M., Gregorc, A., Hornitzky, M. (2006) Diagnosis of American foulbrood in honey bees: a synthesis and proposed analytical protocols. Lett Appl Microbiol 43(6), 583–590
Dingman, D.W., Stahly, D.P. (1983) Medium promoting sporulation of Bacillus larvae and metabolism of medium components. Appl Environ Microbiol 46(4), 860–869
Dobbelaere, W., De Graaf, D.C., Peeters, J.E., Jacobs, F.J. (2001) Development of a fast and reliable diagnostic method for American foulbrood disease (Paenibacillus larvae subsp. larvae) using a 16S rRNA gene based PCR. Apidologie 32, 363–370
Forsgren, E., Stevanovic, J., Fries, I. (2008) Variability in germination and in temperature and storage resistance among Paenibacillus larvae genotypes. Vet Microbiol 129, 342–349
Genersch, E. (2010) American Foulbrood in honeybees and its causative agent, Paenibacillus larvae. J. Inv. Pathol. 103(nul)
Genersch, E., Forsgren, E., Pentikäinen, J., Ashiralieva, A., Rauch, S., Kilwinski, J., Fries, I. (2006) Reclassification of Paenibacillus larvae subsp. pulvifaciens and Paenibacillus larvae subsp. larvae as Paenibacillus larvae without subspecies differentiation. Int J Syst Evol Microbiol 56, 501–511
Gillard, M., Charriere, J.D., Belloy, L. (2008) Distribution of Paenibacillus larvae spores inside honey bee colonies and its relevance for diagnosis. J. Inv. Pathol. 99(1), 92–95
Han, S.H., Lee, D.B., Lee, D.W., Kim, E.H., Yoon, B.S. (2008) Ultra-rapid real-time PCR for the detection of Paenibacillus larvae, the causative agent of American Foulbrood (AFB). J. Inv. Pathol. 99, 8–13
Hasemann, L. (1961) How long can spores of American foulbrood live? Am. Bee J. 101, 298–299
Hitchcock, J.D., Stoner, A., Wilson, W.T., Menapace, D.M. (1979) Pathogenicity of Bacillus pulvifaciens to honey bee larvae of various ages (Hymenoptera: Apidae). J. Kansas Entomol. Soc. 52(2), 238–246
Lauro, F.M., Favaretto, M., Covolo, L., Rassu, M., Bertoloni, G. (2003) Rapid detection of Paenibacillus larvae from honey and hive samples with a novel nested PCR protocol. Int. J. Food Microbiol. 81(3), 195–201
Lindström, A., Fries, I. (2005) Sampling of adult bees for detection of American foulbrood (Paenibacillus larvae subsp. larvae) spores in honey bee (Apis mellifera) colonies. J. Apic.Res. 44(2), 82–86
Martinez, J., Simon, V., Gonzalez, B., Conget, P. (2010) A real-time PCR-based strategy for the detection of Paenibacillus larvae vegetative cells and spores to improve the diagnosis and the screening of American foulbrood. Lett Appl Microbiol 50(6), 603–610
Nakagawa, S., Schielzeth, H. (2012) A general and simple method for obtaining R2 from generalized linear mixed-effects models. Meth. Ecol. Evol. 4, 133–142
Nordström, S., Fries, I. (1995) A comparison of media and cultural conditions for identification of Bacillus larvae in honey. J Apic Res 34(2), 97–103
Nordström, S., Forsgren, E., Fries, I. (2002) Comparative diagnosis of American foulbrood using samples of adult bees and honey. J. Apic. Sci. 46, 5–12
Pernal, S.F., Melathopoulos, A.P. (2006) Monitoring for American foulbrood spores from honey and bee samples in Canada. Apiacta 41, 99–109
R Core Team. (2013). R: A language and environment for statistical computing. Vienna, Austria: R Foundation for Statistical Computing,. Retrieved from ISBN 3-900051-07-0, URL http://www.R-project.org/
Rusenova, N. V., Parvanov, P., Stanilova, S. (2012) Development of multiplex PCR for fast detection of Paenibacillus larvae in putrid masses and in isolated bacterial colonies. Appl Biochem Microbiol 49(1), 79–84
Ryba, S., Titera, D., Haklova, M., Stopka, P. (2009) A PCR method of detecting American foulbrood (Paenibacillus larvae) in winter beehive wax debris. Vet Microbiol 139, 193–196
Titera, D., Haklova, M. (2003) Detection method of Paenibacillus larvae larvae from winter beehive debris. Apiacta 38(3), 131–133
Vaneechoutte, M., Van Eldere, J. (1997) The possibilities and limitations of nucleic acid amplification technology in diagnostic microbiology. J Med Microbiol 46(3), 188–194
Wilson, I.G. (1997) Inhibition and facilitation of nucleic acid amplification. Appl Environ Microbiol 63(10), 3741–3751
Woodrow, A.W. (1942) Susceptibility of honeybee larvae to individual inoculations with spores of Bacillus larvae. J. Econ. Entomol. 35, 892–895
Acknowledgements
This project had financial support from The Swedish Board of Agriculture through the EU-financed National Program for Production and Sale of Honey. We like to thank Emilia Semberg, Asmayet Kelemeworki and Emmy Sundström for excellent technical assistance.
Author information
Authors and Affiliations
Corresponding author
Additional information
Manuscript editor: Peter Rosenkranz
Valeur du pronostic établi à partir d’abeilles et de débris de la ruche dans la détection de la loque américaine dans les colonies d’abeilles
Paenibacillus larvae / AFB / culture / PCR / surveillance sanitaire / régression logistique
Die prognostische Aussagekraft bei der Verwendung von Bienen und Gemüll für den Nachweis der Amerikanischen Faulbrut in Bienenvölkern
Amerikanische Faulbrut / Paenibacillus larvae / Kultur / PCR / logistische Regression
Rights and permissions
Open Access This article is distributed under the terms of the Creative Commons Attribution License which permits any use, distribution, and reproduction in any medium, provided the original author(s) and the source are credited.
About this article
Cite this article
Forsgren, E., Laugen, A.T. Prognostic value of using bee and hive debris samples for the detection of American foulbrood disease in honey bee colonies. Apidologie 45, 10–20 (2014). https://doi.org/10.1007/s13592-013-0225-6
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s13592-013-0225-6