Abstract
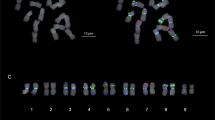
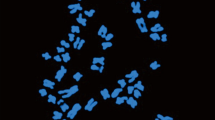
Solanum is the most representative and largest genus of the family Solanaceae with numerous important crop species including potato, tomato and eggplant, etc. Wild species are crucial resources for elucidating interspecific relationships and breeding in this genus. Nightshades are various wild plant species belonging to the Solanum, consisting of many morphological variants collectively known as the Solanum nigrum complex. They are exploited for food, medicines, animal feed, and spiritual uses. They have especially proven useful in Solanum crop breeding. However, only a limited number of molecular cytogenetic studies have been reported, which has impeded comparative analyses that would allow a better understanding of Solanum genome evolution and facilitate breeding improvements. In this study, chromosomal level genome constitution and genome size were analyzed in three wild species of Solanum through triple-color fluorescence in situ hybridization (FISH) and flow cytometry. The chromosome number was 2n = 2x = 24 in S. americanum, 2n = 4x = 48 in S. villosum, and 2n = 6x = 72 in S. nigrum. Each pair of 5S and 45S rDNA signals was observed in diploid species; however, they proportionally increased with the ploidy level with two and three pairs in tetraploid and hexaploid species, respectively. Telomeric signals were detected in all the chromosome terminal regions of the three species. The diploid genotypes have an average genome size of 2.65 pg/2C, whereas the tetraploid and hexaploid species have approximately two- and three-fold larger genome sizes with 5.06 and 7.70 pg/2C, respectively. Our results suggest that the diploid species S. americanum and the tetraploid species S. villosum may be potential parents of the hexaploid species S. nigrum. This data will be helpful for future taxonomic, cytogenetic, phylogenetic, and evolutionary studies in Solanum as well as breeding improvement of this genus.




Similar content being viewed by others
References
Bennetzen JL, Ma J, Devos KM (2005) Mechanisms of recent genome size variation in flowering plants. Ann Botany 95(1):127–132. https://doi.org/10.1093/AOB/MCI008
Bourge M, Brown SC, Siljak-Yakovlev S (2018) Flow cytometry as tool in plant sciences, with emphasis on genome size and ploidy level assessment. Genet Appl 2(2):1–12. https://doi.org/10.31383/GA.VOL2ISS2PP1-12
Chen J, Liu X, Zhu L, Wang Y (2013) Nuclear genome size estimation and karyotype analysis of Lycium species (Solanaceae). Sci Hort 151:46–50. https://doi.org/10.1016/J.SCIENTA.2012.12.004
Doležel J, Greilhuber J, Suda J (2007) Estimation of nuclear DNA content in plants using flow cytometry. Nat Protoc 2(9):2233–2244. https://doi.org/10.1038/nprot.2007.310
Hembree WG, Ranney TG, Lynch NP, Jackson BE (2020) Identification, genome sizes, and ploidy of Deutzia. J Am Soc Hortic Sci 145(2):88–94. https://doi.org/10.21273/JASHS04779-19
Jacoby A, Labuschagne MT (2006) Hybridization studies of five species of the Solanum nigrum complex found in South Africa and two cocktail tomato cultivars. Euphytica 149:3(3):303–307. https://doi.org/10.1007/S10681-005-9078-Z
Jiang J (2019) Fluorescence in situ hybridization in plants: recent developments and future applications. Chromosome Res 27(3):153–165. https://doi.org/10.1007/S10577-019-09607-Z
Levan A, Frega K, Sandberg A (2009) Nomenclature for centromeric position on chromosomes. Hereditas 52:201–220. https://doi.org/10.1111/j.1601-5223.1964.tb01953.x
Li J, Zhu K, Wang Q, Chen X (2021) Genome size variation and karyotype diversity in eight taxa of Sorbus sensu stricto (Rosaceae) from China. Comp Cytogenet 15(2):137–148. https://doi.org/10.3897/COMPCYTOGEN.V15I2.58278
Lobdell MS, Shearer K (2022) Genome sizes, ploidy levels, and base compositions of Styrax species and cultivars. HortScience 57(3):478–484. https://doi.org/10.21273/HORTSCI16294-21
Manoko MLK, van den Berg RG, Feron RMC, van der Weerden GM, Mariani C (2007) AFLP markers support separation of Solanum nodiflorum from Solanum americanum sensu stricto (Solanaceae). Plant Syst Evol 267(1):1–11. https://doi.org/10.1007/S00606-007-0531-4
Melo CAF, Martins MIG, Oliveira MBM, Benko-Iseppon AM, Carvalho R (2011) Karyotype analysis for diploid and polyploid species of the Solanum L. Plant Syst Evol 293(1–4):227–235. https://doi.org/10.1007/s00606-011-0434-2
Mohyuddin A, Kurniawan TA, Khan Z, Nadeem S, Javed M, Dera AA, Iqbal S, Awwad NS, Ibrahium HA, Abourehab MAS, Rabea S, Elkaeed EB, Asghar MN, Saeed S (2022) Comparative insights into the antimicrobial, antioxidant, and nutritional potential of the Solanum nigrum complex. Processes 10(8):1455. https://doi.org/10.3390/PR10081455
Nguyen TH, Waminal NE, Lee DS, Pellerin RJ, Ta TD, Campomayor NB, Kang BY, Kim HH (2021) Comparative triple-color FISH mapping in eleven Senna species using rDNA and telomeric repeat probes. Hortic Environ Biotechnol 62(6):927–935. https://doi.org/10.1007/S13580-021-00364-9/FIGURES/3
Ojiewo CO, Murakami K, Masinde PW, Agong SG (2007) Polyploidy breeding of african nightshade (Solanum section Solanum). Int J Plant Breed 1(1):10–21
Ojiewo CO, Agong SG, Murakami K, Masuda M (2015) Chromosome duplication and ploidy level determination in african nightshade Solanum villosum Miller. 81(2):183–188. https://doi.org/10.1080/14620316.2006.11512048
Park TH (2016) The complete chloroplast genome sequence of potato wild relative species, Solanum nigrum. Mitochondrial DNA B Resour 1(1):858–859. https://doi.org/10.1080/23802359.2016.1250133
Pellerin RJ, Waminal NE, Kim HH (2018) Triple-color FISH karyotype analysis of four Korean wild Cucurbitaceae species. Hortic Sci Technol 36(1):98–107. https://doi.org/10.12972/KJHST.20180011
Peniton E, Waminal N, Kim T-H, Kim H (2019) FISH karyotype comparison between wild and cultivated Perilla species using 5S and 45S rDNA probes. Plant Breed Biotechnol 7:237–244. https://doi.org/10.9787/PBB.2019.7.3.237
Poczai P, Hyvönen J (2011) On the origin of Solanum nigrum: can networks help? Mol Biol Rep 38(2):1171–1185. https://doi.org/10.1007/S11033-010-0215-Y/FIGURES/3
Powell AF, Feder A, Li J, Schmidt MH, Courtney L, Alseekh S, Jobson EM, Vogel A, Xu Y, Lyon D, Dumschott K, McHale M, Sulpice R, Bao K, Lal R, Duhan A, Hallab A, Denton AK, Bolger ME, Fernie AR, …, Usadel B (2022) A Solanum lycopersicoides reference genome facilitates insights into tomato specialized metabolism and immunity. Plant J 110(6):1791–1810. https://doi.org/10.1111/tpj.15770
Robert ML, Lim Y, Hanson K, Sanchez-teyer L, Bennett F, Leitch MD, A. R., Leitch IJ (2008) Wild and agronomically important Agave species (Asparagaceae) show proportional increases in chromosome number, genome size, and genetic markers with increasing ploidy. Bot J Linn Soc 158(2):215–222. https://doi.org/10.1111/j.1095-8339.2008.00831.x
Särkinen T, Poczai P, Barboza GE, van der Weerden GM, Baden M, Knapp S (2018) A revision of the old world black nightshades (morelloid clade of Solanum L. Solanaceae). PhytoKeys 106:1–223. https://doi.org/10.3897/PHYTOKEYS.106.21991
Schneider CA, Rasband WS, Eliceiri KW (2012) NIH Image to ImageJ: 25 years of image analysis. Nat Methods 9(7):671–675. https://doi.org/10.1038/nmeth.2089
Simonin KA, Roddy AB (2018) Genome downsizing, physiological novelty, and the global dominance of flowering plants. PLoS Biol 16(1):e2003706. https://doi.org/10.1371/JOURNAL.PBIO.2003706
van Biljon A, Labuschagne M, Koen E (2010) Microsatellite-based assessment of five Solanum nigrum complex species and their progeny. Acta Agriculturae Scandinavica - Section B Soil and Plant Science 60(6):494–499. https://doi.org/10.1080/09064710903233839
Vásquez-Espinal A, Yañez O, Osorio E, Areche C, García-Beltrán O, Ruiz LM, Cassels BK, Tiznado W (2019) Theoretical study of the antioxidant activity of quercetin oxidation products. Front Chem 7:818. https://doi.org/10.3389/FCHEM.2019.00818/BIBTEX
Vrána J, Simková H, Kubaláková M, Cíhalíková J, Doležel J (2012) Flow cytometric chromosome sorting in plants: the next generation. Methods (San Diego Calif) 57(3):331–337. https://doi.org/10.1016/j.ymeth.2012.03.006
Waminal NE, Kim HH (2012) Dual-color FISH karyotype and rDNA distribution analyses on four Cucurbitaceae species. Hortic Environ Biotechnol 53(1):49–56. https://doi.org/10.1007/s13580-012-0105-4
Waminal NE, Pellerin RJ, Kim N, Jayakodi M, Park JY, Yang T, Kim HH (2018) Rapid and efficient FISH using pre-labeled oligomer probes. Sci Rep 8(8224):1–10. https://doi.org/10.1038/s41598-018-26667-z
Younas A, Yousaf Z, Rashid M, Riaz N, Aftab A, Bushra H, And S, Yasin H (2019) Karyological studies of pharmaceuticaly important species belonging to genus Solanum for evaluation of interspecific phylogenetic relationships. J Plant 1(1):24–31
Zhang R, Zhang L, Wang W, Zhang Z, Du H, Qu Z, Li XQ, Xiang H (2018) Differences in codon usage bias between photosynthesis-related genes and genetic system-related genes of chloroplast genomes in cultivated and wild Solanum species. Int J Mol Sci 19(10):3142. https://doi.org/10.3390/IJMS19103142
Acknowledgements
This study was funded by grant number 2020R1A2C1101915 of the National Research Foundation of Korea.
Author information
Authors and Affiliations
Contributions
HHK is the supervisor and project administrator. THN carried out the experiments, analyzed the data, and wrote the original draft, reviewed, and edited the manuscript. SJP and BYK reviewed and edited the manuscript. All authors read and approved the final version of the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare that they have no conflicts of interest.
Additional information
Communicated by Seon-In Yeom.
Publisher’s Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Nguyen, H.T., Park, S.J., Kang, B.Y. et al. Comparative triple-color FISH mapping and genome size advances understanding of the cytogenetic diversity in wild Solanum species. Hortic. Environ. Biotechnol. 64, 811–817 (2023). https://doi.org/10.1007/s13580-023-00522-1
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s13580-023-00522-1