Abstract
Cannabidiol (CBD) is a pharmacologically active ingredient for use in medical, cosmetic, and food products. CBD forms from cannabidiolic acid (CBDA) with the process of decarboxylation by heating cannabis (Cannabis sativa L.) material. During the production of CBD-rich material, decarboxylation should be performed in precise conditions regarding temperature and time. The experiments were performed by heating hemp samples at 100, 110, 120, 130, and 140 °C for 180 min. Materials were sampled every 20 min and cannabinoid content was analyzed using HPLC, followed by calculation of kinetic parameters. Experimental results showed an exponential reduction of CBDA in the samples during heating. CBD simultaneously increased, and after a specific point, CBD started degrading. The optimal conditions were 140 °C for 30 min. At the level of molecular orbitals, reaction steps, and reaction coordinates, along with the corresponding changes in molecular energy, the molecular mechanism of CBDA decarboxylation and CBD formation was described. Computational analysis has confirmed that the mechanism of CBDA decarboxylation is a direct beta-keto acid pathway. The course of CBDA decarboxylation depends on the time, temperature, and chemical composition of the sample.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
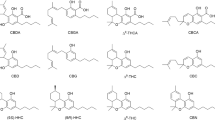
Cannabinoids are interesting organic compounds for present-day medicines, food, and cosmetic products. Cannabidiol (CBD) and ∆9-tetrahydrocannabinol (∆9-THC) are the most significant cannabinoids from cannabis plants because they are the most abundant and have the most pronounced pharmacological effects (Atakan 2012). Together with other cannabis compounds, such as terpenes, they interact with cannabinoid receptors and non-cannabinoid receptors, modulating pain, spasticity, sedation, and appetite (Russo 2011). They have neuroprotective, antioxidant, anti-inflammatory, and anticonvulsant properties and indirectly induce apoptosis of tumor cells as they stimulate ceramide secretion. Clinical trials are showing their effects in the treatment of cancer, epilepsy, Parkinson's disease, Alzheimer's disease, multiple sclerosis, anxiety, depression, and other conditions (Russo 2011; Atakan 2012; Ożarowski et al. 2021).
Cannabinoids are generated from cannabinoid acids synthesized by cannabis (Cannabis Sativa L.) in the glandular trichomes as resin components. Cannabinoid acids have lower bioavailability and less biological effects than decarboxylated (neutral) cannabinoids, because they are more hydrophilic and harder to cross lipophilic barriers, and have a lower affinity for binding with cannabinoid receptors (Moreno-Sanz et al. 2020; Ryu et al. 2021). Neutral cannabinoids are produced after a heat-assisted nonenzymatic decarboxylation of cannabinoid acids (Ryu et al. 2021). Decarboxylation of cannabinoid acids is for example achieved through smoking, vaping, cooking, or baking the cannabis material. At the production of pharmaceutical and food products with cannabinoids, decarboxylation is an important part of manufacturing that needs to be precisely controlled (Potter 2014; Moreno-Sanz et al. 2020; Olejar et al. 2021). Knowledge about the chemical mechanism, kinetics, and thermodynamics of the reactions is therefore of the utmost importance.
Previous studies have found that during CBDA decarboxylation, the bond between the carbon from the carboxyl group and the alpha carbon (C–C bond) cleaves, CO2 is released and CBD is formed (Veress et al. 1990; Perrotin-Brunel et al. 2011; Wang et al. 2016; Citti et al. 2018). For decarboxylation to occur, electrons from the cleaved C–C bond of carboxylic acid should be stabilized. CBDA is an aromatic carboxylic acid, containing a substituted aromatic ring, which allows stabilization of electrons from the cleavage of the C–C bond, shown in Fig. 1 (Brenneisen 2007; Kluger et al. 2013).
Results from studies by Verres et al. (1990), Perrotin-Brunel et al. (2011), Wang et al. (2016), Citti et al. (2018), and Moreno et al. (2020) have experimentally confirmed that decarboxylation of cannabinoid acids is a pseudo first order reaction, resulting in an exponentially decreasing amount of cannabinoid acids and increasing number of neutral analogs, cannabinoids. During thermal decarboxylation, at some point, neutral analogs start to degrade (Veress et al. 1990; Perrotin-Brunel et al. 2011; Wang et al. 2016; Citti et al. 2018; Moreno et al. 2020). Moreno et al. (2020) have performed a combined experimental and computational study of CBDA decarboxylation similar as Perrotin-Brunel et al. (2011) who did the same but with THCA. They found that cannabinoid acids’ decarboxylation is a direct keto-enol mechanism, catalyzed by short-chain organic acids with two transition states and keto intermediate (Perrotin-Brunel et al. 2011). It was found that a hydroxyl group on the 2nd carbon directly participates in the decarboxylation, while other bound groups have a smaller or indirect effect (Hu et al. 2016; Gao et al. 2016).
Published data of CBDA decarboxylation and CBD formation kinetics and mechanism are insufficient. In this study, we present experimental work of the CBDA decarboxylation and CBD formation kinetics with a computational approach.
Materials and methods
Materials
Kief samples were prepared from dried hemp [Cannabis sativa L. (Finola)], sieved through a 100 mm sieve. Cannabinoid content was analyzed with HPLC. Standard solutions for HPLC analysis were CBD in acetonitrile and CBDA in methanol (Sigma-Aldrich Corporation).
Decarboxylation experiments
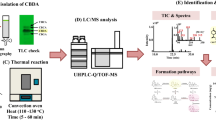
Decarboxylation experiments were performed in an oven at temperatures: 100, 110, 120, 130, and 140 °C for 180 min with air ventilation. The procedure for experiments was modified according to Chuchev and BelBruno (2007) and Atakan (2012). The experiments were performed separately at each temperature. Sample of 500 mg was weighted into 9 separate Petri dishes, 5 cm in diameter and 2 cm in height (one for each time point). Uncovered Petri dishes were placed in the reactor, which was preheated to a certain temperature for 1 h. Petri dishes were separately removed from the reactor at 20 min intervals for 180 min.
Sample preparation for HPLC analysis
From each vessel with decarboxylated content, 200 mg of sample was weighed (Mettler Toledo XS205 Dual range) in the 15 mL plastic tubes. Then, 10 mL of ethanol (CARLO ERBA Reagents, Val de Reuil, France) was added using an automatic pipette. Plastic tubes were transferred to the ultrasonic homogenizer (Bandelin Sonorex Digitec DT 100 H) at 35 kHz, 25 °C, and 15 min. After homogenization, 1.5 mL of the sample was filtered through a filter (0.22 μm Nylon pore size, Q-Max® RR Syringe Filters) into dark vials for HPLC analysis.
HPLC analysis
Samples were analyzed with HPLC at 40 °C with a flow rate of 2 mL/min and a photo-diode array detector (PDA) detection at 220 and 280 nm (Shimadzu Corporation, Japan, Shimadzu 20AD XR UFCL XR system, PDA SPD-M20A, RF-10A fluorescence detector, and Phenomenex Kinetex® C18 column 2.6, 100A, 100 × 4.6 mm, ID 1.7 µm particle size). Mobile phases A and B were used, prepared from acetonitrile, water, and 3-fluoro-acetic acid (J. T. Baker; Deventer, The Netherlands). Mobile phase A was water with 2% acetonitrile and 0.1% 3-fluoro-acetic acid. Mobile phase B was acetonitrile with 2% water and 0.1% 3-fluoro-acetic acid. Elution was performed by gradient method: 0.01–0.20 min 61% B, 0.20–0.21 min 61–43% B, 0.21–50 min 43–56% B. HPLC analysis was managed with the computer program LC Solution Shimadzu 1.24 SP1. The identities of CBDA and CBD were confirmed by comparing retention times and absorption spectra with reference standard compounds of CBDA and CBD. Mass fractions of CBDA and CBD in the samples was determined from calibration curves at following concentrations (mg/mL): 0.000, 0.001, 0.005, 0.010, 0.050, 0.100, 0.167, 0.250, 0.500, 0.750, 1.000; gaining R2 for CBDA 0.9988 and for CBD 0.9996.
Calculation of kinetic and thermodynamic parameters of CBDA decarboxylation
The approximation is that CBDA decarboxylation is a pseudo-first-order reaction resulting in the generation of CBD which can be further degraded under certain conditions, which are described previously. CBDA decarboxylates into CBD at rate constant k1 after which CBD is degraded to its decomposition products at rate constant k2. The rate of CBDA decarboxylation is described as a decrease of CBDA (− d[CBDA]) in time (dt) or a product of the rate constant (k1) and the mass fraction of CBDA in the sample (Eq. 1) (Pagni 2006) (pp. 385).
Equation 1 can be integrated and written as a linear relationship of the natural logarithm of the CBDA mass fraction (y-axis) and time (x-axis), with the slope value representing rate constant (k1) (2) (Pagni 2006) (pp. 385).
[CBDA] is a CBDA mass fraction at a certain time (t) of decarboxylation. [CBDA0] is CBDA mass fraction at the beginning of the reaction. CBD during CBDA decarboxylation can be calculated with Eq. (3), showing the relationship of CBDA and CBD amounts and their rate constants.
We approximate CBD formation and degradation as two separate first-order pseudo reactions, where in the first reaction CBD is formed, and in the second, CBD is degraded. The rate of CBD formation is described as an increase of CBD in time (d[CBD]/dt) or product of the rate constant (k2a) and the CBD mass fraction in the sample (Eq. 4) (Pagni 2006) (pp. 385). The rate of CBD degradation is described as a decrease of CBD in time (− d[CBD]/dt) or product of the rate constant (k2b) and the CBD mass fraction in the sample (Eq. 5) (Pagni 2006) (pp. 385).
After integrating Eqs. (4 and 5) for CBD formation and CBD degradation, we got equations showing a linear relationship of the natural logarithm of the CBD mass fraction and time (Eqs. 6, 7)
The half-life span (t1/2) for a first-order reaction is calculated with Eq. (8) (Pagni 2006) (pp. 385).
Linearizing and plotting the Arrhenius equation shows a relation between the natural logarithm of k and the reciprocal temperature in Kelvin, 1/T (Eq. 9). From the slope of the linear Arrhenius equation, EA/R, R is the universal gas constant [8.31441 J (Kmol)−1] and activation energy (EA) can be calculated. From the section on the y axis, ln(A), pre-exponential, or frequency factor (A) can be calculated.
The Eyring Eq. (10) describes the dependence of the reaction rate constant and the constant for the transition of reactants to the transition state (K‡). The transmission coefficient (κ) describes the probability that the transition state will result in the product and for the reactions resulting with its product, the approximation is κ = 1. kB is the Boltzmann constant (1.380662 × 10−23 JK−1) and h is the Planck constant (6.626176 × 10−34 Js).
Gibbs’s free energy of activation (ΔG‡) can be calculated from the relation with K‡. It can also be calculated with k, by incorporating K‡ from Eq. (10) to get 11. ΔG‡ can also be calculated with ΔH‡ and TΔS‡ (12).
Expressed k from Eq. (11) shows the relation between k and ΔG‡ (Eq. 13)
By incorporating ΔG‡ from Eq. (12) into (13), the relation of k with ΔH‡ and ΔS‡ give Eq. (14)
By linearizing Eq. (14), ln(k) is expressed with Eq. (15)
Dependence of ln(kh/kBT) versus 1/T is given by transforming Eqs. (15) to (16)
Equation (16) can be transformed as the dependence of ln(k/T) versus 1/T (17). From the slope of Eq. (17), (ΔH‡)/R, enthalpy of activation (ΔH‡) can be calculated. From the section on the y axis, ln(kB/h) + (ΔS‡)/R, the entropy of activation (ΔS‡) can be calculated.
Computational analysis of CBDA decarboxylation
GaussView 6.0.16. was used for drawing molecular structures, inserting inputs, and analyzing outputs of calculations. Calculations of decarboxylation mechanisms were performed by Gaussian 09W Revision D.01. First, structures of reactants, transition states, intermediates, and products were created. Then, geometries of the structures were optimized and the computation was performed using HF and B3LYP theories with a 6-31G* basis set. Computation of decarboxylation mechanism of 2-HBA was performed, following decarboxylation mechanism of CBDA. Using the QST2 module, transition state structures were calculated. Values of the potential energy were calculated using the SCAN model. For the 2-HBA molecule, the B3LYP/6-31G* was used, and for the CBDA molecule, the HF/6-31G*. The structures of transition states were determined from values of frequencies. Transition states have one negative, imaginary frequency, while the structures of reactants, intermediates, and products have real, positive frequencies.
From the computational analysis, ΔG‡ and ΔH‡ were determined under standard conditions (298.15 K and 1 atm). If we assume that ΔH‡ varies insignificantly with temperature, then ΔG‡1 at different temperatures is calculated from ΔG‡2 determined at 298.15 K by using Van't Hoff Eq. (18) and by incorporating ln(K‡) from Eq. 11 into (18) (Eq. 19).
ΔS‡ is calculated by expressing it from Eqs. (12, 20)
Results
Experimental analysis of CBDA decarboxylation to CBD formation and degradation
We found that the amounts of CBDA and CBD depend on the duration and temperature of the process. CBDA mass fraction decreased with time exponentially, as shown with a green curve in Fig. 2. CBDA decarboxylated completely after 140 min at 130 °C and after 60 min at 140 °C, while at 100–120 °C, it did not completely decarboxylate even after the end of the experiment at 180 min. Simultaneously, the amount of CBD increased to its maximum value and after that, depending on the temperature, CBD started decreasing, shown as the red curve in Fig. 2. Degradation is shown with the blue curve, showing the loss of total CBD content (which was calculated with SUM = 0.877 × CBDA + CBD).
Kinetic and thermodynamic parameters of CBDA decarboxylation
To observe the influence of temperature on the rate of CBDA decarboxylation, values of the natural logarithm of CBDA mass fractions were calculated and plotted in dependence on time. Slope values represent rate constants at each temperature. As the temperature increased, the rate constant of the reaction increased, which means the reaction was faster (Fig. 3).
Linearity of the lines in Fig. 3 with R2 values close to 1 (between 0.95 and 0.98) supports the assumption of the first-order kinetics of CBDA decarboxylation. Second-order kinetics was also evaluated by calculating R2 values of reciprocal data gaining R2 values between 0.61 and 0.92. We proceeded by plotting the Arrhenius Eq. (9) to calculate activation energy and frequency factor which is a "macroscopic" description of the rate constant that origin from the approximation of simple conversion of one compound to another and results in activation energy for the total reaction. We calculated activation energy, EA = 65.84 kJ/mol and frequency factor, A = 3.47 × 105 s−1. The Eyring equation provides a "microscopic" description of the reaction rate constant and a mechanistic description of reaction in multi-steps, which includes transition states with the determination of Gibbs energy of activation for each transition state. By plotting Eyring Eq. (17), we calculated enthalpy of activation, ΔH‡ = 62.42 kJ/mol, and entropy of activation, ΔS‡ = − 149.84 J/Kmol (Fig. 4). Gibbs’s free energies of activation (ΔG‡) at different temperatures were calculated with Eq. 12.
To make a possible comparison of experimentally determined CBDA decarboxylation results from our study with the results from previous studies, we calculated kinetic and thermodynamic parameters at temperatures, at which the parameters were determined in our study by extrapolation with Eyring Eq. (17) (Table 1).
Kinetic and thermodynamic parameters of CBD formation and degradation
To analyze the effects of temperature on the rate of CBD formation and degradation, we approximated the processes as two separate pseudo-first-order reactions, where in the first reaction CBD was formed, and in the second, CBD was degraded. The Endpoint of formation was the point after which CBD decreased. By plotting Eqs. (6 and 7), we calculated reaction constants for CBD formation and degradation. After a certain time at a certain temperature, CBD decreased, faster with higher temperatures (Fig. 5).
Values of R2 for CBD formation were distant from 1, which means the relationship between the natural logarithm of CBD concentration and time during CBD formation was not linear. Therefore, presented CBD formation was not just a simple pseudo-first-order reaction, which is why it should have been broken down into more reactions with smaller intervals. In contrast, R2 values for CBD degradation were close to 1, so there was a linear relationship between the natural logarithm of CBD mass fraction and time.
We proceeded by plotting Arrhenius Eq. (9) and Eyring Eq. (17) for CBD formation and degradation. For CBD formation, activation energy was EA = 90.03 kJ/mol and frequency factor, A = 2.38 × 108 s−1. Further, enthalpy of activation was determined to be ΔH‡ = 86.85 kJ/mol and entropy of activation, ΔS‡ = − 94.92 J/Kmol (Fig. 6a). For CBD degradation, activation energy was determined to be EA = 28.81 kJ/mol, frequency factor, A = 1.72 × 10–1 s−1, enthalpy of activation was determined to be ΔH‡ = 25.55 kJ/mol and entropy of activation, ΔS‡ = − 270.06 J/Kmol (Fig. b). Gibbs’s free energies of activation (ΔG‡) at different temperatures were calculated with Eq. (12).
To compare CBD formation and degradation with other studies, we calculated kinetic and thermodynamic parameters by extrapolating results at the same temperatures, using Eyring Eq. (17) (Table 2).
Decarboxylation model
CBDA decarboxylation kinetics model could be created with Eqs. (1 and 3). A simple model of CBDA decarboxylation at 140 °C, which is found to be an optimal temperature, if heating for 30 min, is shown in Fig. 7. It is possible to create a model which would take into consideration all points in cannabis processing that can lead to decarboxylation.
Computational analysis of CBDA decarboxylation
To compare and support experimental results with computational analysis, the mechanism of CBDA decarboxylation with Gaussian software was explored. CBDA has a hydroxyl group on beta carbon of the aromatic ring, so it could be studied as a derivative of 2-hydroxybenzoic acid (2-HBA) or salicylic and olivetolic acid (Fig. 1). As the hydroxyl group is the closest group to the site of decarboxylation, the hypothesis is that it participates in the process. Other bounded groups on the aromatic ring of CBDA (limonene at 3rd, hydroxyl at 4th, and pentyl at 6th position) have a smaller or indirect effect.
Computation was performed on 2-HBA with B3LYP/6-31G* and on CBDA with HF/6-31G*. The structure of 1st transition state was determined using the QST2 module, and the structure of 2nd transition state was determined using the SCAN module. For 2-HBA decarboxylation, structures with imaginary frequency − 2068.63 cm−1 for 1st transition state and − 902.30 cm−1 for 2nd transition state were determined. For CBDA decarboxylation, the 1st transition state has an imaginary frequency of − 2504.02 cm−1, and the 2nd transition state, − 393.25 cm−1 (Fig. 8).
The mechanism involves the transfer of hydrogen from the hydroxyl group to alpha carbon, after which the keto intermediate is formed. Then, the carboxyl group rotates toward the keto group and hydrogen moves from the carboxyl to the keto group, cleaving the C–C bond (Fig. 9d).
a–c Frontier molecular orbitals (FMO) for the reactant state of the reactive part of the molecule, and the corresponding electron density cloud (99%, BGR scale, blue positive, red negative). d Reaction pathway of CBDA decarboxylation as the dependence of relative energy on the reaction coordinate. The computation was performed in Gaussian with HF/6-31G*. Decarboxylation goes through a transition state in which H+ transfers from –OH to α–C (TS1) and a keto intermediate (M1) is formed. H+ is then transferred from –COOH to C=O at keto intermediate after which tautomerization into enol form (TS2) occurs, following C–C bond cleavage (P)
From the difference in energy of the initial reactant structure and transition states for 2-HBA and CBDA decarboxylation, it was found that 1st transition state determines the reaction rate. From ∆H‡ and ∆G‡ within 1st transition state (as a rate-determining step), ∆G‡ at different temperatures were calculated using Eq. (19), and ∆S‡ was calculated with Eq. (20). For the 2-HBA decarboxylation, 1st transition state has ∆G‡ = 317.179 kJ/mol and ∆H‡ = 319.639 kJ/mol. For CBDA decarboxylation, 1st transition state has ∆G‡ = 317.683 kJ/mol and ∆H‡ = 319.206 kJ/mol (Table 3).
Computational analysis was performed to examine the mechanism of decarboxylation of 2-HBA and CBDA with the thermodynamic parameters at different temperatures. The mechanism and two transition states for 2-HBA and CBDA decarboxylation were determined. Energy barriers were determined for decarboxylation at 25 °C. Energy barriers for the 1st transition state for 2-HBA 317.179 kJ/mol and CBDA, 317.683 kJ/mol determine the reaction rate, where hydrogen is transferred from the hydroxyl group to alpha carbon and the keto intermediate is formed. After that, hydrogen transfers from the carboxyl group to the keto group (tautomerization into enol form), C–C bond cleaves with the formation of CO2 and phenol (in the case of 2-HBA) or CBD (in the case of CBDA).
Comparing experimental and computational analysis
In the present study, computationally determined thermodynamic parameters are higher than experimentally determined parameters. In computational analysis, the entropy is positive, which is why its contribution is higher in determining the Gibbs energy of activation, compared to the experimental analysis, where entropy is negative, so enthalpy contributes more (Table 4).
Discussion
Experimental analysis of CBDA decarboxylation with CBD formation and degradation
Previous studies showed that CBDA decarboxylation at temperatures below 100 °C is a simple stoichiometric conversion to CBD with a CBD loss of up to 2% (Citti et al. 2018). At temperatures, 100 °C and higher, CBDA decarboxylation, CBD formation and a subsequent loss of CBD through degradation and evaporation runs faster (Veress et al. 1990; Wang et al. 2016; Citti et al. 2018; Moreno et al. 2020).
In this study, obtained rate constants were smaller than rate constants from other studies, except for a study by Moreno et al. (2007). Results suggests that in biomass material, flowers and kief, CBDA decarboxylation is slower than in extracts, which corresponds to the study of Wang et al. (2016). Activation energy and frequency factor for CBDA decarboxylation in kief and flowers are lower in contrast to the ones in extracts. Entropy has a negative value and relative to enthalpy, indicating that Gibbs energy solely depends on enthalpy, which is consistent with previous studies (Veress et al. 1990; Wang et al. 2016; Citti et al. 2018; Moreno et al. 2020). Gibbs energy has a positive value, which confirms CBDA decarboxylation is not spontaneous and endothermic reaction (Table 1). It can be assumed that the chemical composition of the sample could energetically "facilitate" decarboxylation. More energy is required for CBDA decarboxylation in non-extracted samples than in extracted samples. Compounds, such as cannabinoids, organic acids, terpenes, or waxes are more concentrated in the extracts than in non-extracted samples (Potter 2014), which could facilitate CBDA decarboxylation (Chuchev and BelBruno 2007; Hu et al. 2016). Another reason for the less energy required to decarboxylate CBDA in extracts is that preserved cell walls of trichomes on flowers act protectively, while those protective structures are not present in the extracts (Richins et al. 2018).
CBD loss occurs due to elevated temperatures that lead to its evaporation or degradation. The outcome also depends on the type and mass of the sample. The same was observed in other studies. Citti showed decarboxylation in closed reactors results in a minor loss of CBDA and CBD, probably because CBD does not evaporate from the reactor (Citti et al. 2018). Similar results were shown by Moreno et al. (2020). Cannabinoid decomposition to unknown components was smaller in a closed reactor, but decarboxylation occurred faster, probably because the temperature in a closed reactor is more consistent (Moreno et al. 2020). The greater amount of plant material placed in the reactor resulted in a slower process of CBDA conversion and a smaller loss of total molar concentration. They described that this effect could be attributed to the slower heat transfer rate through the bigger mass of the sample material, or due to the reduced surface area exposed to air circulation and evaporation (Citti 2018).
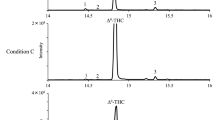
To understand CBD loss during decarboxylation, there is a need for analysis of its degradation product/s. Gaoni et al. (1971), Golombek et al. (2020) and Marzullo et al. (2020) described CBDA, when exposed to heat, undergoes intramolecular cyclization into Δ9-THC, which can be further degraded into CBN in the presence of oxygen, or to more thermodynamically stable Δ8-THC (Gaoni et al. 1971; Golombek et al. 2020; Marzullo et al. 2020). Elsohly and Slade (2006) showed that CBD can undergo photo-oxidation or pyrolysis to cannabielsoin (CBE) (Elsohly and Slade 2006). With the rapid development of new methods for chemical analysis, it is possible to more accurately distinguish among small molecular differences. Layton et al. (2016) showed different degradation products of CBD under basic and acidic conditions (Layton et al. 2016).
Cannabis samples are very complex in terms of their constituents, hence the whole picture of what influences CBD during the decarboxylation process with the determination of byproducts during decarboxylation remains unknown.
Computational analysis of CBDA decarboxylation
We used HF/6-31G*, and for 2-HBA, B3LYP/6-31G* protocols. The decarboxylation mechanism in this study started from an optimized syn-conformer that has different energy when compared to the previously described conformers (Chuchev and BelBruno 2007; Hu et al. 2016; Gao et al. 2016). We analyzed hydrogen transferred from the hydroxyl group to the alpha carbon as the first step. Earlier studies analyzed the rotation of the syn-conformer into the anti-conformer as the first step, which was followed by the transfer of hydrogen from the carboxyl group to the alpha carbon (Chuchev and BelBruno 2007). All studies of Δ9-THCA decarboxylation consistently show the formation of keto intermediates and direct beta-keto acid pathway (Perrotin-Brunel et al. 2011).
Different energy barriers can be calculated with different model mechanisms, that used different starting molecule structures, with different energy differences between reactants, intermediates, and products (Chuchev and BelBruno 2007; Perrotin-Brunel et al. 2011; Hu et al. 2016; Gao et al. 2016). The comparisons of different mechanisms show that acid and water can reduce the energy barriers because water molecules can facilitate the "transfer" of hydrogen to alpha carbon, which is followed by tautomerization that can facilitate the breaking of the C–C bond. Short substituents on benzoic acid further facilitate decarboxylation (Hu et al. 2016) (Table 5).
Since the rate of CBDA decarboxylation depends on the temperature applied, it would be necessary to include dependence on different temperatures in the computational analysis of quantum–mechanical programs such as Gaussian and GAMESS (The General Atomic and Molecular Electronic Structure). In the computational analysis, it is important to consider the structure of the initial reactant molecule from which the reaction proceeds and the angle at which the reaction molecule collides. If the molecules are not positioned correctly concerning the orientation of HOMO (highest occupied molecular orbital) and LUMO (lowest unoccupied molecular orbital), the reaction will not occur.
Mechanism of CBDA decarboxylation
Based on the literature data, experimental and computational results, the mechanism of CBDA decarboxylation can be assumed. Decarboxylation happens through rotation from the most stable syn-conformer into the anti-conformer. Hydrogen is then transferred from the hydroxyl group to the alpha carbon and tautomerization takes place from the keto intermediate. This is followed by the transfer of hydrogen from the carboxyl group to the keto group and with tautomerization to the enol form. Subsequently, the C–C bond cleaves and CO2 goes away, resulting in the formation of CBD (Fig. 10).
Presented results show that a further increase in accuracy in the determination of optimal conditions for decarboxylation reaction could be achieved by following decarboxylation reaction at time intervals shorter than 20 min. The reaction mechanism can be described separately in the open and closed reactors and with a larger sample mass suitable for industrial production. For more precise computational analysis, it is necessary to know the chemical composition of the sample to have an insight into which of the constituents would have the most impact on CBDA decarboxylation.
Conclusions
We confirmed that CBDA decarboxylation is a pseudo-first order reaction resulting in an exponential decrease of CBDA and an increase of CBD mass fractions. CBD starts to degrade after a specific time at a specific temperature. The course of CBDA decarboxylation depends on time, temperature, and the chemical composition of the sample. Experimentally, the optimal conditions to turn CBDA to CBD are heating at 140 °C for 30 min. Steps presented in our study could be used to create a model for different conditions for individual use. Computational analysis has shown that the mechanism of CBDA decarboxylation is a direct beta-keto acid pathway in which the reaction rate depends on the transfer of hydrogen from the hydroxyl group to the alpha ring carbon after which hydrogen from the carboxyl group is transferred to the keto group and the alpha carbon to a carboxyl group, resulting in CBD formation. The computational analysis of the reaction mechanism has suggested that in the future the rate of decarboxylation could be controlled by using different salts of the CBDA which can influence the structure of the intermediary state.
Abbreviations
- CBD:
-
Cannabidiol
- CBDA:
-
Cannabidiolic acid
- ∆9-THC:
-
∆9-Tetrahydrocannabinol
- 2-HBA:
-
2-Hydroxybenzoic acid
- HPLC:
-
High-performance liquid chromatography
- PDA:
-
Photo-diode array detector
- ΔH ‡ :
-
Enthalpy of activation
- ΔS ‡ :
-
Entropy of activation
- E A :
-
Activation energy
- ΔG ‡ :
-
Gibbs’s free energy of activation
- K ‡ :
-
Transition state
- κ :
-
Transmission coefficient
References
Atakan Z (2012) Cannabis, a complex plant: different compounds and different effects on individuals. Ther Adv Psychopharmacol 2:241–254. https://doi.org/10.1177/2045125312457586
Brenneisen R (2007) Chemistry and analysis of phytocannabinoids and other cannabis constituents. In: ElSohly MA (ed) Marijuana and the cannabinoids. Humana Press, Totowa, pp 17–49
Chuchev K, BelBruno JJ (2007) Mechanisms of decarboxylation of ortho-substituted benzoic acids. J Mol Struct Theochem 807:1–9. https://doi.org/10.1016/j.theochem.2006.12.004
Citti C, Pacchetti B, Vandelli MA, Forni F, Cannazza G (2018) Analysis of cannabinoids in commercial hemp seed oil and decarboxylation kinetics studies of cannabidiolic acid (CBDA). J Pharm Biomed Anal 149:532–540. https://doi.org/10.1016/j.jpba.2017.11.044
Elsohly M, Slade D (2006) Chemical constituents of marijuana: the complex mixture of natural cannabinoids. Life Sci 78:539–548. https://doi.org/10.1016/j.lfs.2005.09.011
Gao L, Hu Y, Zhang H, Liu Y, Song Z, Dai Y (2016) DFT computational study on decarboxylation mechanism of salicylic acid and its derivatives in the anionic state. J Mol Struct 1116:56–61. https://doi.org/10.1016/j.molstruc.2016.03.005
Gaoni Y, Mechoulam R (1971) Isolation and structure of.DELTA.+- tetrahydrocannabinol and other neutral cannabinoids from hashish. J Am Chem Soc 93:217–224. https://doi.org/10.1021/ja00730a036
Golombek P, Müller M, Barthlott I, Sproll C, Lachenmeier DW (2020) Conversion of cannabidiol (CBD) into psychotropic cannabinoids including tetrahydrocannabinol (THC): a controversy in the scientific literature. Toxics 8:41. https://doi.org/10.3390/toxics8020041
Hu Y, Gao L, Dai Z, Sun G, Zhang T, Jia S, Dai Y, Zhang X (2016) DFT investigation on the decarboxylation mechanism of ortho hydroxy benzoic acids with acid catalysis. J Mol Model 22:56. https://doi.org/10.1007/s00894-016-2923-2
Kluger R, Howe GW, Mundle SOC (2013) Chapter two - avoiding CO2 in catalysis of decarboxylation. In: Williams IH, Williams NH (eds) Advances in physical organic chemistry. Academic Press, pp 85–128
Layton C, Runco J, Aubin AJ (2016) Forced degradation of cannabidiol. Application note. Waters Corporation
Marzullo P, Foschi F, Coppini DA, Fanchini F, Magnani L, Rusconi S, Luzzani M, Passarella D (2020) Cannabidiol as the substrate in acid-catalyzed intramolecular cyclization. J Nat Prod 83:2894–2901. https://doi.org/10.1021/acs.jnatprod.0c00436
Moreno T, Dyer P, Tallon S (2020) Cannabinoid decarboxylation: a comparative kinetic study. Ind Eng Chem Res 59:20307–20315. https://doi.org/10.1021/acs.iecr.0c03791
Moreno-Sanz G, Ferreiro Vera C, Sánchez-Carnerero C, Nadal Roura X, de Medina S, Baena V (2020) Biological activity of Cannabis sativa L. extracts critically depends on solvent polarity and decarboxylation. Separations 7:56. https://doi.org/10.3390/separations7040056
Olejar KJ, Hatfield J, Arellano CJ, Gurau AT, Seifried D, Heuvel BV, Kinney CA (2021) Thermo-chemical conversion of cannabis biomass and extraction by pressurized liquid extraction for the isolation of cannabidiol. Ind Crops Prod 170:113771. https://doi.org/10.1016/j.indcrop.2021.113771
Ożarowski M, Karpiński TM, Zielińska A, Souto EB, Wielgus K (2021) Cannabidiol in neurological and neoplastic diseases: latest developments on the molecular mechanism of action. Int J Mol Sci 22:4294. https://doi.org/10.3390/ijms22094294
Pagni R (2006) Modern physical organic chemistry (Eric V. Anslyn and Dennis A. Dougherty). J Chem Educ 83:387. https://doi.org/10.1021/ed083p387
Perrotin-Brunel H, Buijs W, van Spronsen J, van Roosmalen MJE, Peters CJ, Verpoorte R, Witkamp G-J (2011) Decarboxylation of Δ9-tetrahydrocannabinol: kinetics and molecular modeling. J Mol Struct 987:67–73. https://doi.org/10.1016/j.molstruc.2010.11.061
Potter DJ (2014) A review of the cultivation and processing of cannabis (Cannabis sativa L.) for production of prescription medicines in the UK. Drug Test Anal 6:31–38. https://doi.org/10.1002/dta.1531
Richins RD, Rodriguez-Uribe L, Lowe K, Ferral R, O’Connell MA (2018) Accumulation of bioactive metabolites in cultivated medical Cannabis. PLoS ONE 13:e0201119. https://doi.org/10.1371/journal.pone.0201119
Russo EB (2011) Taming THC: potential cannabis synergy and phytocannabinoid-terpenoid entourage effects. Br J Pharmacol 163:1344–1364. https://doi.org/10.1111/j.1476-5381.2011.01238.x
Ryu BR, Islam MJ, Azad MOK, Go E-J, Rahman MH, Rana MS, Lim Y-S, Lim J-D (2021) Conversion characteristics of some major cannabinoids from hemp (Cannabis sativa L.) raw materials by new rapid simultaneous analysis method. Mol Basel Switz 26:4113. https://doi.org/10.3390/molecules26144113
Veress T, Szanto JI, Leisztner L (1990) Determination of cannabinoid acids by high-performance liquid chromatography of their neutral derivatives formed by thermal decarboxylation: I. Study of the decarboxylation process in open reactors. J Chromatogr A 520:339–347. https://doi.org/10.1016/0021-9673(90)85118-F
Wang M, Wang Y-H, Avula B, Radwan MM, Wanas AS, van Antwerp J, Parcher JF, ElSohly MA, Khan IA (2016) Decarboxylation study of acidic cannabinoids: a novel approach using ultra-high-performance supercritical fluid chromatography/photodiode array-mass spectrometry. Cannabis Cannabinoid Res 1:262–271. https://doi.org/10.1089/can.2016.0020
Acknowledgements
Authors would like to acknowledge coworkers from Freyherr, from Faculty for Pharmacy, Department for Pharmaceutical Biology, University of Ljubljana, and to Srećko Valić, Professor from Faculty of Medicine, the University of Rijeka for inter-collaboration, support, and critical suggestions in performing our study.
Funding
The APC was funded by the Slovenian Research Agency (ARRS), research program P1-0208. Part of the research was financially supported by the research program P4-0127. High-performance computing at the University of Rijeka is supported by European Fund for Regional Development (ERDF) and by the Ministry of Science, Education and Sports of the Republic of Croatia under the project number RC.2.2.06-0001.
Author information
Authors and Affiliations
Contributions
TF: investigation, data curation, writing—original draft preparation, validation; SK: resources, writing—review and editing; ŽMS: methodology, software, resources, writing—review and editing, supervision; ET: conceptualization, methodology, writing—review and editing, supervision, validation. All authors have read and agreed to the published version of the manuscript.
Corresponding author
Ethics declarations
Conflict of interest
The authors declare no conflict of interest.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Fućak, T., Kreft, S., Svedružić, Ž.M. et al. Mechanism and kinetics of CBDA decarboxylation into CBD in hemp. J. Plant Biochem. Biotechnol. 32, 608–621 (2023). https://doi.org/10.1007/s13562-023-00847-z
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s13562-023-00847-z