Abstract
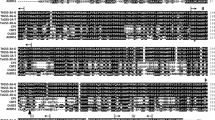
Cotton fibre quality is a multigenic trait. Genetic modification of different genes to achieve high quality fibre is difficult without knowing the mechanism lying behind genes interaction. Based on background knowledge an attempt to explore the potential structural interactions between Gossypium hirsutum Wlim5 domain1 and Gossypium hirsutum ACTIN-1 proteins was done in current study. Sequence features of the LIM domain1 of GhWlim5 protein were identified through multiple sequence alignment analysis, and a phylogenetic tree was built to identify evolutionary relationships between sequences. Conservation indicated the evolutionary importance of side chain residues and the presence of several aliphatic and/or bulky residues, which stabilize the protein core and facilitate packing of zinc fingers. The structures of GhWlim5 domain1 and GhACTIN-1 proteins were modelled and validated through computational methods. Validation of GhACTIN-1 and GhWlim5 domain1 structures indicated good structural quality with 99.7% and 100% of the favoured number of residues in allowed regions and Z-score, within the ranges of − 9.87 and − 4.17, respectively. Docking analysis indicated various possible modes of interaction between these two proteins with favourable binding affinities. Based on our strong binding interaction results between GhWlim5 domain1 and GhACTIN-1 proteins, we further investigated the role of over-expression of GhWlim5 by transformation in cotton plants under fibre specific promoter and transgenic plants displayed significant increases in fibre strength.












Similar content being viewed by others
Abbreviations
- ABPs:
-
Actin binding proteins
- LIM:
-
LIN-11, ISL-1 and MEC-3
- HMM:
-
Hidden Markov model
- PDB:
-
Protein data bank
- IDT:
-
Integrated DNA technologies
- GUS:
-
β-glucuronidase
- EDTA:
-
Ethylenediaminetetraacetic acid
- RT-qPCR:
-
Quantitative reverse transcription polymerase chain reaction
- GAPDH:
-
Glyceraldehyde 3-phosphate dehydrogenase
References
Ahad A, Ahmad A, Din S, Rao AQ, Shahid AA, Husnain TJF (2015) In silico study for diversing the molecular pathway of pigment formation: an alternative to manual coloring in cotton fibers. Front Plant Sci 6:751
Ahmad A et al (2015) In-silico determination of insecticidal potential of Vip3Aa-Cry1Ac fusion protein against lepidopteran targets using molecular docking. Front Plant Sci 6:1081
Arnaud D et al (2012) Expression analysis of LIM gene family in poplar, toward an updated phylogenetic classification. BMC Res Notes 5(1):102. https://doi.org/10.1186/1756-0500-5-102
Bach I (2000) The LIM domain: regulation by association. Mech Dev 91(1–2):5–17
Chen CY et al (2002) The regulation of actin organization by actin-depolymerizing factor in elongating pollen tubes. Plant Cell 14(9):2175–2190
Cheung AY et al (2002) Rab2 GTPase regulates vesicle trafficking between the endoplasmic reticulum and the Golgi bodies and is important to pollen tube growth. Plant Cell 14(4):945–962
Dong C-H, Xia G-X, Hong Y, Ramachandran S, Kost B, Chua N-H (2001) ADF proteins are involved in the control of flowering and regulate F-actin organization, cell expansion, and organ growth in Arabidopsis. Plant Cell 13(6):1333–1346
Franklin-Tong VE (1999) Signaling and the modulation of pollen tube growth. Plant Cell 11(4):727–738
Fu Y, Li H, Yang Z (2002) The ROP2 GTPase controls the formation of cortical fine F-actin and the early phase of directional cell expansion during Arabidopsis organogenesis. Plant Cell 14(4):777–794
Gutierrez R, Lindeboom JJ, Paredez AR, Emons AM, Ehrhardt DW (2009) Arabidopsis cortical microtubules position cellulose synthase delivery to the plasma membrane and interact with cellulose synthase trafficking compartments. Nat Cell Biol 11(7):797–806. https://doi.org/10.1038/ncb1886
Han L-B et al (2013) The dual functions of WLIM1a in cell elongation and secondary wall formation in developing cotton fibers. Plant Cell 113:116970
Han L, Li Y, Sun Y, Wang H, Kong Z, Xia G (2016) The two domains of cotton WLIM1a protein are functionally divergent. Sci China Life Sci 59(2):206–212
Hooft RW, Sander C, Vriend G (1997) Objectively judging the quality of a protein structure from a Ramachandran plot. Bioinformatics 13(4):425–430
Huang S, An YQ, McDowell JM, McKinney EC, Meagher RBJTPJ (1996) The Arabidopsis thaliana ACT4/ACT12 actin gene subclass is strongly expressed throughout pollen development. Plant J 10(2):189–202
Jaakola L, Pirttila AM, Halonen M, Hohtola A (2001) Isolation of high quality RNA from bilberry (Vaccinium myrtillus L.) fruit. Mol Biotechnol 19(2):201–203. https://doi.org/10.1385/mb:19:2:201
Kadrmas JL, Beckerle MC (2004) The LIM domain: from the cytoskeleton to the nucleus. Nat Rev Mol Cell Biol 5(11):920
Kandasamy MK, McKinney EC, Meagher RB (2002) Functional nonequivalency of actin isovariants in Arabidopsis. Mol Biol Cell 13(1):251–261
Kelley LA, Mezulis S, Yates CM, Wass MN, Sternberg MJ (2015) The Phyre2 web portal for protein modeling, prediction and analysis. Nat Protoc 10(6):845–858
Komis G, Luptovciak I, Doskocilova A, Samaj J (2015) Biotechnological aspects of cytoskeletal regulation in plants. Biotechnol Adv 33(6):1043–1062
Kost B, Mathur J, Chua N-H (1999) Cytoskeleton in plant development. Curr Opin Plant Biol 2(6):462–470
Kuroda S et al (1996) Protein–protein interaction of zinc finger LIM domains with protein kinase C. J Biol Chem 271(49):31029–31032
Latif A et al (2015) Herbicide-resistant cotton (Gossypium hirsutum) plants: an alternative way of manual weed removal. BMC Res Notes 8(1):453. https://doi.org/10.1186/s13104-015-1397-0
Li H, Lin Y, Heath RM, Zhu MX, Yang Z (1999) Control of pollen tube tip growth by a Rop GTPase-dependent pathway that leads to tip-localized calcium influx. Plant Cell 11(9):1731–1742
Li X-B, Fan X-P, Wang X-L, Cai L, Yang W-C (2005) The cotton ACTIN1 gene is functionally expressed in fibers and participates in fiber elongation. Plant Cell 17(3):859–875
Li F et al (2009) Modified fiber qualities of the transgenic cotton expressing a silkworm fibroin gene. Chin Sci Bull 54(7):1210–1216
Li Y et al (2013) A cotton LIM domain-containing protein (GhWLIM5) is involved in bundling actin filaments. Plant Physiol Biochem 66:34–40
Li L, Li Y, Wang NN, Li Y, Lu R, Li XB (2015) Cotton LIM domain-containing protein Gh PLIM 1 is specifically expressed in anthers and participates in modulating F-actin. Plant Biol 17(2):528–534
McDowell JM, Huang S, McKinney EC, An Y-Q, Meagher RB (1996) Structure and evolution of the actin gene family in Arabidopsis thaliana. Genetics 142(2):587–602
Pierce BG, Wiehe K, Hwang H, Kim B-H, Vreven T, Weng Z (2014) ZDOCK server: interactive docking prediction of protein–protein complexes and symmetric multimers. Bioinformatics 30(12):1771–1773
Rao AQ, Irfan M, Saleem Z, Nasir IA, Riazuddin S, Husnain T (2011) Overexpression of the phytochrome B gene from Arabidopsis thaliana increases plant growth and yield of cotton (Gossypium hirsutum). J Zhejiang Univ Sci B 12(4):326–334. https://doi.org/10.1631/jzus.B1000168
Sahu M et al (2013) In silico prediction and characterization of three-dimensional structure of Actin-1 of Arabidopsis thaliana. BioTechnol J Biotechnol Comput Biol Bionanotechnol 94:4
Satyavathi V, Prasad V, Lakshmi BG, Sita GLPS (2002) High efficiency transformation protocol for three Indian cotton varieties via Agrobacterium tumefaciens. Plant Sci 162(2):215–223
Seagull RW (1992) A quantitative electron microscopic study of changes in microtubule arrays and wall microfibril orientation during in vitro cotton fiber development. J Cell Sci 101(3):561–577
Thomas C et al (2007) The LIM domains of WLIM1 define a new class of actin bundling modules. J Biol Chem 282(46):33599–33608
Velyvis A, Qin J (2005) LIM domain and its binding to target proteins zinc finger proteins. Springer, New York, pp 99–105
Wiederstein M, Sippl MJ (2007) ProSA-web: interactive web service for the recognition of errors in three-dimensional structures of proteins. Nucleic Acids Res 35(suppl_2):W407–W410
Winder SJ, Ayscough KR (2005) Actin-binding proteins. J Cell Sci 118(4):651–654
Yang Z (1998) Signaling tip growth in plants. Curr Opin Plant Biol 1(6):525–530
Zhang J, Stewart JJJCS (2000) Economical and rapid method for extracting cotton genomic DNA. J Cotton Sci 4(3):193–201
Zhang M et al (2017) Overexpression of GhFIM2 propels cotton fiber development by enhancing actin bundle formation. J Integr Plant Biol 59(8):531–534
Zhao L et al (2016) Phylogenomic analyses of large-scale nuclear genes provide new insights into the evolutionary relationships within the rosids. Mol Phylogenet Evol 105:166–176
Zheng Q, Zhao Y (2007) The diverse biofunctions of LIM domain proteins: determined by subcellular localization and protein–protein interaction. Biol Cell 99(9):489–502
Acknowledgements
This work was supported by the National Centre of Excellence in Molecular Biology University of the Punjab and the School of Plant Sciences, University of Arizona, USA. We extend our gratitude to the Center for Electron Microscopy Zhejiang University in Hangzhou China, for SEM microscopy and to the Central Cotton Research Institute, Multan, Pakistan for fibre analysis.
Author information
Authors and Affiliations
Contributions
The concept and the experimental design were initiated by Adnan Iqbal and David W. Galbraith. Bio-informatics analyses was performed by Adnan Iqbal, Basit Jabbar and Muhammad Azam Ali. Cotton plant transformation was done by Adnan Iqbal and Ayesha Latif. Molecular characterization of transgenic cotton plants was done by Adnan Iqbal and Mukhtar Ahmad. Review of literature; Adnan Iqbal and Abdul Qayyum Rao. Data collection and analysis; Adnan Iqbal and Basit Jabbar. Preparation of main manuscript; Adnan Iqbal and Ahmad Ali Shahid. Critical review of manuscript; David W. Galbraith, Abdul Qayyum Rao and Tayyab Husnain; Approval of the final version of manuscript; Adnan Iqbal, Ambreen Gul, David W. Galbraith, Abdul Qayyum Rao and Tayyab Husnain. The manuscript was read by all authors and approved for publication.
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that there are no conflicts of interest.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
About this article
Cite this article
Iqbal, A., Latif, A., Galbraith, D.W. et al. Structure-based prediction of protein–protein interactions between GhWlim5 Domain1 and GhACTIN-1 proteins: a practical evidence with improved fibre strength. J. Plant Biochem. Biotechnol. 30, 373–386 (2021). https://doi.org/10.1007/s13562-020-00603-7
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s13562-020-00603-7