Abstract
Purpose
Immunotherapy using PD-L1 blockade is effective in only a small group of cancer patients, and resistance is common. This emphasizes the importance of understanding the mechanisms of cancer immune evasion and resistance.
Methods
A genome-scale CRISPR-Cas9 screen identified Bap1 as a regulator of PD-L1 expression. To measure tumor size and survival, tumor cells were subcutaneously injected into both syngeneic WT mice and immunocompromised mice. The phenotypic and transcriptional characteristics of Bap1-deleted tumors were examined using flow cytometry, RNA-seq, and CUT&Tag-seq analysis.
Results
We found that loss of histone deubiquitinase Bap1 in cancer cells activates a cDC1-CD8+ T cell-dependent anti-tumor immunity. The absence of Bap1 leads to an increase in genes associated with anti-tumor immune response and a decrease in genes related to immune evasion. As a result, the tumor microenvironment becomes inflamed, with more cDC1 cells and effector CD8+ T cells, but fewer neutrophils and regulatory T cells. We also found that the elimination of Bap1-deleted tumors depends on the tumor MHCI molecule and Fas-mediated CD8+ T cell cytotoxicity. Our analysis of TCGA data further supports these findings, showing a reverse correlation between BAP1 expression and mRNA signatures of activated DCs and T-cell cytotoxicity in various human cancers.
Conclusion
The histone deubiquitinase Bap1 could be used as a biomarker for tumor stratification and as a potential therapeutic target for cancer immunotherapies.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
1 Introduction
While immunotherapy targeting PD-L1-mediated tumor tolerance has achieved remarkable clinical success in various cancers, it has only exhibited durable responses in a subset of patients [1, 2]. This suggests that a deeper and broader understanding of cancer immunity is necessary. The low response rates and durations of PD-L1 blockade agents can be attributed to two reasons: firstly, the mechanisms regulating PD-L1 expression for immune evasion in different cancer contexts are not completely understood. Secondly, there are tumor immune evasion mechanisms that are independent of PD-L1.
Recent evidence supports the notion that modulators of histone acetylation and histone methylation play context-dependent roles in anti-tumor immune responses or tumor immune evasion [3, 4]. Certain inhibitors targeting these epigenetic regulators have shown promising anti-tumor immune responses in preclinical studies [5]. For instance, inhibition of lysine-specific demethylase 1 (LSD1), a well-known demethylase of H3K4 and H3K9, has been found to stimulate anti-tumor immunity by activating the expression of endogenous retroviruses in cancers [6,7,8]. However, to our knowledge, there have been no reports on the role of regulators of histone ubiquitination in anti-tumor immunity.
Here, our study revealed that the loss of a histone deubiquitinase Bcra1 associated protein 1 (Bap1) in tumors leads to an anti-tumor immune response that is dependent on CD8+ T cell cytotoxicity. This response is characterized by increased infiltration of cDC1 and CD8+ effector cells, while neutrophils and regulatory T cells (Tregs) show reduced infiltration. As a result, the tumors are eventually cleared in host mice. Furthermore, we demonstrated that the eradication of Bap1-deleted tumors in syngeneic mice relies on the expression of tumor MHCI molecules and Fas-mediated CD8+ T cell cytotoxicity. Supporting our findings, we observed that lower levels of BAP1 were associated with significant increases in mRNA signatures of activated DCs and T-cell cytotoxicity in various human cancer types. In Bap1-deleted tumors, which have an inflamed microenvironment, we observed significant upregulation of genes associated with anti-tumor immune responses, such as Ccl7, Ccl11, Thbs2, Mavs, and Ifitm1. Conversely, genes related to immune evasion, including Cxcl1, Cxcl2, Tgfb3, Smad3, and Epha2, were downregulated in Bap1-deleted tumors. These findings have important implications for the development of cancer immunotherapies by targeting chromatin modulators.
2 Materials and methods
2.1 Mice
NSG (NOD.Cg-PrkdcscidIl2rgtm1Wjl/SzJ) mice (005557), Rag2−/− mice (008449), Tcra−/− mice (002116), Batf3−/− mice (013755), Prf1−/− mice (002407), and OT-I (C57BL/6-Tg (TcraTcrb)1100Mjb/J) mice (003831) were purchased from The Jackson Laboratory. C57BL/6 and BALB/c wild-type mice were ordered from Shanghai SLAC Laboratory Animal Co., Ltd. Female mice at 6–8 weeks of age were used for the experiment. All mice used were maintained under a specific pathogen-free condition at the Laboratory Animal Research Center of Tongji University.
2.2 Cancer cell lines
Colorectal adenocarcinoma cells MC38 cell line (ENH204-FP) was purchased from Kerafast (Boston, MA). Breast cancer cells 4T1 (CRL-2539), colon carcinoma cells CT26 (CRL-2638), mammary carcinoma cells EMT6 (CRL-2755), and melanoma cells B16-F10 (CRL-6322) were obtained from ATCC. All the cells were cultured according to the supplier’s recommendations and were maintained in a 37 °C, 5% CO2 atmosphere in a Thermo Forma incubator (Thermo Fisher, Waltham, MA).
2.3 Antibodies
Fluorochrome-labeled monoclonal antibodies (clones denoted in parentheses) against.
Cd274-APC (10 F.9G2), CD45-APC-Cy7 (30-F11), CD3-APC (17A2), CD4-BV421 (RM4-5), CD8-FITC (53 − 6.7), γδT-PE (GL3), CD11b-APC (M1/70), Ly6C-PE (AL-21), MHCII-PerCPCy5.5 (M5/114.15.2), F4/80-Alexa-flour700 (BM8), CD24-BV421 (M1/69), CD11c-FITC (N418), CD103-PE-Cy7 (2E7) were purchased from Biolegend. CD95-PE (SA367H8), and H-2 K-MHCI (36-7-5) were purchased from BD Biosciences. Antibodies for cells deletion and pathways blockade including CD4 (GK1.5), CD19 (1D3), CD8a (YTS169.4), IFNγ (H22), CTLA4 (9H10), PD-1 (29 F. 1A12) were purchased from BioXcell. BAP1 (D1W9B), H2AK119 (D27C4), Myc-Tag (9B11), β-Tubulin (D3U1W), Anti-rabbit IgG-HRP, Anti-mouse IgG-HRP antibodies were ordered from Cell Signaling Technology. Anti-H3K4me3 and anti-H3K27ac polyclonal antibodies were purchased from Active Motif. PE-conjugated H-2Kb/SIINFEKL Dextramer was purchased from Immudex.
2.4 Lentivirus production and transduction
Lentivirus packaging was carried out when HEK293FT cells were at a density of 70-80% confluency. 5.4 µg lentiviral transfer vector, 5.4 µg psPAX2, 1.2 µg pMD2.G and 12 µL PLUS reagent (Invitrogen, 11514015) were mixed in 3 mL Opti-MEM (Gibco, 31985062) and incubate for 5 min at room temperature. 36µL Lipofectamine LTX (Invitrogen, 15338100) was added to the mixture and further incubated for 30 min. Then incubated mixture was added to HEK293FT cells with 5 mL Opti-MEM and persistently incubated for 4–8 h, after which the mixture was replaced with DMEM (Corning, 10-013-CV) medium with 10% FBS + 1% PS. Two days after transfection, viral supernatant was collected, filtered through a 0.45 μm filter, and stored at −80 °C.
1.0 × 106 cells were resuspended in 500 µl diluted viral supernatant with 4 µg/mL polybrene. The cell-lentivirus suspension was incubated at 37 °C for 30 min before being transferred to a DMEM medium. The medium was changed the next day. The amounts of reagents were scaled up or down depending on the number of cells to be transduced.
2.5 Generation of stable Cas9-expressing MC38 cell lines
MC38 cells were transduced with lentivirus supernatant containing EF1α-Cas9-T2A-tdTomato elements (Supplementary Fig. 1a). Transduced cells were cultured for 8 days, after which tdTomato+ cells were single-cell sorted into 96-well plates (MoFlo XDP). The Cas9 function was validated by transducing with a dual reporter construct (Supplementary Fig. 1d; addgene, 67980) and examined by flow cytometry (Supplementary Fig. 1e).
2.6 Generation of genome-wide mutant libraries and screening
3.0 × 107 MC38 cells were transduced with a pre-determined volume of lentiviral supernatant of the Mouse Improved Genome-wide Knockout CRISPR Library v2 [9] (addgene, 67988) that gave rises to 30% transduction efficiency measured by BFP expression. The library has 90,230 gRNAs against 18,424 mouse genes. Two independent infections were performed. After stable expression of BFP, the mutant cells (BFP+) were sorted and cultured in multi-layer flasks. At every passage, 5.0 × 107 cells were seeded in new flasks. The first-round sorting of PD-L1low/− cells was conducted on day 12 (Supplementary Fig. 1g).
2.7 Sequencing of gRNAs and statistical analysis
Genomic DNA extraction from sorted cells was performed as described previously [10]. gRNAs were amplified using the following primers:
-
Forward primer: ACACTCTTTCCCTACACGACGCTCTTCCGATCTCTTGTGGAAAGGACGAAACA;
-
Reverse primers: TCGGCATTCCTGCTGAACCGCTCTTCCGATCTCTAAAGCGCATGCTCCAGAC.
To cover the complexity of the gRNA library, approximately 72 µg genomic DNA was analyzed for the control unsorted mutant samples. All the genomic DNA was analyzed for sorted PD-L1 negative cells. The end PCR product was purified using SPRI beads and transferred for Illumina sequencing.
Enrichment gRNAs and their targeting genes were analyzed using the MAGeCK statistical algorithm [11] by comparing read counts from sorted mutant cells with counts from unsorted mutant cells.
2.8 Generation of stable knockout cell lines
Several stable knockout MC38, CT26, 4T1, and EMT6 cell lines were generated using Cas9-gRNAs to create a frame-shift mutation by indel generation. The gRNAs used to generate stable knockout cell lines were listed in Supplementary Table S3. The protein expression of indicated gene knockout was validated by western blot.
2.9 Tumor mouse models
The syngeneic mice were injected in the flank subcutaneously with 1.0 × 106 tumor cells The MC38 cells were injected into C57BL/6 mice. CT26, 4T1, and EMT6 cells were injected into BALB/c mice. Tumor lengths and widths were measured every 2–3 days with a caliper. Tumor volumes were determined using the formula: ½ × L×W2 where L is the major axis and W is the minor axis. Mice were sacrificed when tumors reached 1000 mm3 or upon ulceration.
Where indicated, the C57BL/6 mice were treated with anti-CD4 antibodies (500 µg/mouse/injection, day −3, 0, 3, 6, 9, 12,15, 18, 21 after tumor cell injection, i.p.), anti-CD8 (300 µg/mouse/injection, day 0, 1, 2 after tumor cell injection, i.p.), anti-CD19 (300 µg/mouse/injection, day −3, 0, 5, 10, 15, 20 after tumor injection, i.p) for CD4, CD8, and CD19 cells depletion respectively and were treated with anti-IFNγ (200 µg/mouse/injection, day −5, −1, 0, 1, 2, 3, 4, 5 after tumor injection, i.p) for IFNγ neutralization. In tumor fragment implantation experiment, the tumors initial growth (reached ~ 800 mm3) in Rag2−/− mice were dissected into 3–5 mm3 fragments and were implanted into C57BL/6 mice. When tumor volume reached 50 ~ 200 mm3 (around day16 after tumor fragments implantation), the mice were administrated with anti-PD-1 (250 µg/mouse/injection, day 16, 19, 22, 25 after tumor fragments implantation, i.p.) and anti-CTLA-4 (200 µg /mouse/injection, day 16, 19, 22 after tumor fragments implantation, i.p.) for PD-L1 and CTLA-4 signals blocking.
2.10 Flow cytometry analysis of tumor immune cells infiltration
Upon collection of WT and Bap1-KO tumors from the C57BL/6 mice 10 days after injection, tumors were dissociated by the Tumor Dissociation Kit (Miltenyi, 130-096-730). ACK lysis buffer (Gibco, A10492-01) was used to remove the red blood cells. A Dead Cell Removal kit (Miltenyi, 130-090-101) was used to remove dead cells. Cells were washed in FACS buffer (PBS + 2% fetal bovine serum) and blocked with anti-mouse CD16/32 antibody (2.4G2) before staining with indicated fluorescence-conjugated antibodies on ice for 20 min. The cells were washed in FACS buffer before analysis on LSRFortessa™X-20 Cell Analyzer (BD Biosciences).
2.11 scRNA-seq analysis of tumor immune cell infiltration
CD45+ cells isolated from WT and Bap1-KO tumors were resuspended with 1×PBS containing 0.04% BSA. Cell viability was detected with 0.4% trypan blue staining and counted by Countess II cell counter (Invitrogen). The cells with more than 90% viable were used for scRNA-seq. The cell suspension was loaded into Chromium microfluidic chips with v3 chemistry and barcoded with a 10× Chromium Controller (10X Genomics). mRNA from the barcoded cells was reverse-transcribed into cDNA, and the sequencing library was constructed according to the instructions of Chromium Single Cell 3ʹ Reagent kits v3 (10×Genomics). Sequencing was performed with NovaSeq 6000 according to Illumina instructions (Berry Genomics Corporation, Beijing, China).
2.12 Analysis scRNA-seq data
Cell Ranger (version 3.0.2) was employed to analyze the output of Chromium single-cell 3′ RNA-seq. For quality control, we filtered the cells with higher than 13% mitochondrial contents and cells of fewer than 500 UMIs or fewer than 200 features. Features expressed in fewer than 3 cells were removed in the analysis. Scrublet (version 0.2.1) was used to determine the doublets in the datasets.
SCANPY was used for data processing and visualization. The following steps were performed in order: data normalization, log-transformation, highly variable genes (HVG) selection and principal component analysis (PCA). Based on the Harmony-corrected latent space, T-distributed stochastic neighbor embedding (t-SNE) was performed using SCANPY. SCCAF [12] was used to determine the discriminative ability of the clusters.
2.13 Tumor cell bulk RNA-seq
1.0 × 106 WT-BFP and Bap1-KO-BFP MC38 cells were injected into Rag2−/− mice subcutaneously. On day 10, tumors were collected and dissociated as described above. BFP+ tumor cells were purified by FACSAria™ Fusion cell sorter (BD Biosciences). Total RNA was extracted by TRIzol reagent (Ambion, 15596-026). Sequencing libraries were generated by using the NEBNext UltraTM RNA Library Prep Kit (NEB, E7770). The library preparations were sequenced on an Illumina Novaseq 6000 platform.
2.14 Differential gene expression analysis of RNA-seq
Differentially expressed genes (DEGs) were identified using the FindMarkers function in Seurat with the parameter test.use = “Wilcox” by default. False detection rate (FDR) was estimated using the Benjamini–Hochberg procedure. The genes with a false detection rate (FDR) lower than 0.01 were used for the heatmap. The genes with a log2 fold change value > 1 are considered differential genes.
2.15 Tumor cell CUT&Tag-seq
WT-BFP and Bap1-KO-BFP MC38 cells derived from Rag2−/− mice as described above were subjected to CUT&Tag-seq. A Universal CUT&Tag Assay Kit for Illumina (Vazyme, TD903-01) was used to construct CUT&Tag libraries targeting H3K4me3, H2AK119, and H3K27ac in both WT and Bap1-KO MC38 cells. Library preparation involved 100,000 cells and included primary antibodies against H2AK119, H3K27ac, or H3K4me3, each at a 1:100 dilution. After Qubit and Bioanalyzer evaluations to verify library quality, samples were pooled for sequencing. The sequencing process was executed using an Illumina Novaseq 6000 platform, achieving a sequencing depth of 6G base pair raw data.
2.16 Tumor cell ATAC-seq
Bap1-KO and WT MC38 cells under the stimulation of IFNγ (20 ng/mL) for 48 h were subjected to ATAC-seq. A Hyperactive ATAC-Seq Library Prep Kit for Illumina (Vazyme, TD711) was used to generate libraries from cultured WT and Bap1-KO MC38 cells. Similar to the CUT&Tag procedure, approximately 100,000 cells were harvested for library preparation. After confirming library quality through Qubit and Bioanalyzer assessments, Illumina Novaseq 6000 sequencing was carried out, achieving a sequencing depth of 12G base pair raw data.
2.17 CUT&Tag and ATAC-seq data processing
Initial processing of CUT&Tag and ATAC-seq raw reads involved adaptor trimming using Trimmomatic (v.0.39). Quality assessment was performed using FastQC (https://www.bioinformatics.babraham.ac.uk/projects/fastqc/) before alignment, ensuring optimal library quality. Paired-end sequencing reads were aligned to the mouse genome (mm10) using Bowtie 2. PCR duplicates were removed using SAMtools. To generate bigwig files, DeepTools2 was employed. Peak calling was conducted using MACS2 (v.2.2.6). Subsequently, coverage plots for both CUT&Tag and ATAC-seq results were generated using DeepTools2. All correlation graphs were produced using R scripts within RStudio.
2.18 T-cell adoptive transfer
For OT-I T cell adoptive transfer, Rag2−/− mice were injected subcutaneously with 5.0 × 105 OVA-expressing control WT or Bap1-KO MC38 cells. OT-I CD8+T cells were isolated from the spleens of OT-I transgenic mice using the CD8+ T cell isolation mouse kit (Miltenyi, 130-104-075) as per the manufacturer’s instructions. and 2.5 × 106 activated OT-I T cells were injected into recipient mice via tail vein on day 3 after tumor cells injection. For CAR-T cell adoptive transfer, Rag2−/− mice were injected subcutaneously with 2.0 × 105 CD19-expressing MC38 cells. CD8+ T cells were isolated from the spleens of C57BL/6 mice as described above, anti-CD19 CAR-expressing vector retrovirus [13] was used to infect CD8+ T cells to generate CAR-T cells, and 1.7 × 106 activated CAR-T cells were injected via tail vein to mice on the day 3 after tumor cells injection.
2.19 In vitro killing assays
Purified OT-I CD8+ T cells were activated in 24-well plates with anti-mouse CD3ε antibody (5 µg/mL), anti-mouse CD28 antibody (1 µg/mL) in the presence of mIL-2 (20 ng/mL), and mIL-7 (2 ng/mL) cytokines. 5.0 × 104 OVA-expressing control WT or Bap1-KO MC38 tumor cells were seeded in 24-well plates a day before adding activated OT-I T cells to the wells at ratios of 1:1, 5:1, 10:1, 20:1 (Tumor cells versus T cells). After being co-cultured for 2 days, cells were stained with trypan blue and counted by Countess™ II automated cell counter. For the apoptosis experiment, tumor cells were treated with 100 ng/mL IFNγ for three days before analysis of cell apoptosis by APC Annexin V Apoptosis Detection Kit with 7-AAD (Biolegend, 640930).
2.20 Histochemistry
C57BL/6 mice were injected subcutaneously with 1.0 × 106Bap1-KO and WT tumor cells. Tumors were collected at day 10, and fixed in 4% paraformaldehyde for 24 h, dehydrated, and embedded in paraffin. And paraffin-soaked tumors were put into a paraffin slicer (Leica, RM2016) for slicing and tissue slices were stored at room temperature. 20 μm tissue slices were deparaffinized and rehydrated, after sealing by 3% BSA for 30 min, and the slices were stained with Anti-CD8α Rabbit antibody (Servicebio, GB114196-100) in the 4 °C for a whole night. PBS was used to wash the samples three times before adding FITC-conjugated goat anti-rabbit IgGs (servicebio, GB22303). The cell nucleus was stained for 10 min by the DAPI. CD8+T cells in tumor tissues were examined by Ortho-Fluorescent Microscopy (Nikon).
For TUNEL staining, after the tissue slice was deparaffinized and rehydrated, 20 µg/ml protease K working solution was added to cover the tissue, and incubated at 37 °C for 22 min. After that, the sections were rinsed twice with PBS. The permeabilised working solution is 0.1% triton. TMR (red) Tunel Cell Apoptosis Detection Kit (servicebio, G1502) was used for TUNEL staining. Images were acquired and photographed using Ortho-Fluorescent Microscopy and Imaging system (Nikon).
2.21 Luciferase reporter assay
Fragments of the DNAs containing the promoter region of Cd274 were amplified by PCR and cloned into the PGL4.10 luciferase reporter vector (Promega, E6651). Site-directed mutagenesis of the Elf4 and Irf1 binding sites in the fragment of the Cd274 promoter was performed using a site-directed mutagenesis kit (Takara, R401). Different MC38 stable cell lines were seeded in 24-well plates overnight and transfected in triplicate using Lipofectamine LTX with 0.5 mg of the vector constructed and 0.5 mg of the Renilla pRL-SV40 vector (Promega, E2231) as an internal control per well. After 36 h of transfection, relative luciferase units (RLUs) were measured using the Dual-Luciferase Reporter Assay System kit (Promega, E1910) according to the manufacturer’s instructions. Firefly RLUs/Renilla RLUs ratios were determined as relative luciferase activities.
2.22 Chromatin immunoprecipitation (ChIP)
Chromatin was prepared from different MC38 cells. SimpleChIP Plus Enzymatic Chromatin IP Kit (CST, 9005) was used for the ChIP experiment. For each IP, 5 ul of Myc-Tag antibody (CST, 2276) was used. After immunoprecipitation, qPCR was conducted to calculate the DNA enrichment in a CFX96 real-time PCR system (Bio-Rad) with TB Green Premix Ex Taq™ II (Tli RNaseH Plus) (Takara, RR820A). The primer pairs for the Cd274 promoters are listed in Supplementary Table S4.
2.23 Immunoprecipitation
Co-immunoprecipitation was carried out in HEK293T cells which were co-transfected with pcDNA3.1(+) and pcDNA3.1(+)-Bap1 or pcDNA3.1(+)-Elf4-Myc-DDK and pcDNA3.1(+)-Bap1. Cells were lysed in NP-40 lysis buffer (Beyotime, P0013F). For immunoprecipitation with Elf4, lysates were incubated with prewashed Pierce c-Myc magnetic beads (Thermo Fisher Scientific, 88842) for 30 min at room temperature, the beads were washed with 5× TBST buffer (125mM Tris, 0.75 M NaCl, 0.25% Tween-20 Detergent) 3 times and ultrapure water once and boiled with SDS sample buffer. For immunoprecipitation with Bap1, lysates were incubated overnight with anti-Bap1 antibody (CST, 13187), then Dynabeads Protein G (Invitrogen, 10003D) was added and incubated for 1 h, the beads were washed with PBS 3 times and boiled with SDS sample buffer. Immunoblotting was performed.
2.24 Analysis of immune cells infiltration in human cancers
The mRNA expression profiles of 33 cancers in the TCGA dataset were obtained from the website cBioPortal (http://www.cbioportal.org/). Abbreviations and the full name of the cancer were provided (Supplementary Table S5). To investigate the effect of BAP1 expression on immune infiltration in tumors, we used the computational method xCell (https://github.com/dviraran/xCell) to calculate the immune infiltration scores of 36 immune cells in patients. For each tumor, the patients were grouped according to quartiles of BAP1 expression, and those in the upper quartile (highly expressed ones) and in the lower quartile (lowly expressed ones) were used for the analysis. The Wilcoxon test was applied to compare the distribution of immune scores between the two groups. The average mRNA expression levels of the genes GZMB and IFNγwere used to measure the cytotoxicity of T cells. The Spearman’s correlation between BAP1 expression and the cytotoxicity of T cells was calculated. All analyses and graphing were done using R software.
2.25 Statistics
All the experiments were not blinded to allocation in our study. Number of mice and sample size were chosen to be suitable for clarifying the statistical differences between the experimental groups. For individual experiments, sample size was based on preliminary experiments or at least two independent repeated experiments were done. For the treatment of the mice and next-generation sequencing samples, all experimental groups are randomly assigned. Unpaired Student’s t-test and one-way ANOVA were used for calculating P values (including qRT-PCR, tumor volume statistics etc.). Statistical tests used were accessible in Figure legends.
3 Results
3.1 CRISPR-Cas9 screening identifies regulators of PD-L1 in mouse cancer cell lines
To identify genes associated with cancer-intrinsic immune evasion mediated by PD-L1 that might be necessary for immunotherapies, we performed a genome-scale CRISPR-Cas9-knockout (KO) screen in a mouse colorectal adenocarcinoma cells MC38. MC38 cells are known to have strong PD-L1 expression, which helps these cells evade immune-mediated killing [14]. Using empirically defined rules [15, 16], we first generated MC38-Cas9 clone3 that had a constitutive expression of functional Cas9 and PD-L1 (Supplementary Fig. 1a–e); we then mutagenized these MC38-Cas9 cells using a previously described mouse genome-wide guide RNA (gRNA) library containing 90,230 gRNAs that target 18,424 protein-coding genes [9] (Supplementary Fig. 1f). CRISPR-Cas9-mutagenized cells were sampled at different time points and were subjected to deep DNA sequencing to identify gRNAs that were enriched in sorted PD-L1− cells relative to unsorted mutant cells: specifically, the unsorted mutant cells (denoted as “S1”) and a mixture of sorted PD-L1low and PD-L1− (5% total mutant cells) cells (“S2”) were sampled after first round sorting on day 12; we also sampled the PD-L1− cells that were isolated after a couple of rounds of sorting (“S3”) (Fig. 1a and Supplementary Fig. 1g).
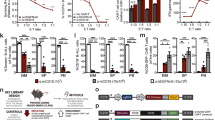
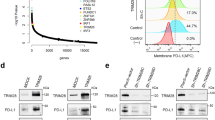
CRISPR-Cas9 screening identifies Bap1 and Elf4 as regulators of PD-L1 in mouse cancer cells lines. (a) Workflow for a genome-scale, pooled CRISPR-Cas9 screening to identify PD-L1 regulators in cancer cells (refer to Supplementary Fig. 1G, with additional details in the Methods). S1 indicates unsorted MC38 mutant cells. S2 indicates the sorted MC38 PD-L1low and PD-L1− mutant cells (5% of total mutant cells). S3 indicates the PD-L1− mutant cells (after 4 rounds PD-L1 negative sorting). Each sample has 2 replicates. (b–d) Scatter plots showing the enriched gRNA targeting genes in S2 (b, S2 vs. S1), in S3 (c, S3 vs. S1), and in S3 (d, S3 vs. S2). The gRNA abundance was analyzed using the algorithm MAGeCK. (e) FACS plots showing the surface expression of PD-L1 in the indicated MC38 cells. Bap1-WT: Bap1 wild-type cells (black line); Bap1-KO1: Bap1 knock-out clone 1 (red line); Cd274-KO: Cd274 knock-out (dashed line). (f) Western blot image showing the protein levels of Bap1, H2AK119Ub, and PD-L1 in MC38 cells. β-tubulin was used as an internal control. (g) qRT-PCR analysis of mRNA expression for Bap1 and Cd274 in the indicated genotypes of MC38 cells. (h) FACS plots showing the surface expression of PD-L1 in the indicated MC38 cells. Elf4-WT: Elf4 wild-type cells (black line); Elf4-KO1: Elf4 knock-out clone 1 (red line). (i) qRT-PCR showing Elf4 and Cd274 mRNA levels in indicated cell types. (j) FACS plots showing the surface expression of PD-L1 in Bap1-KO1 MC38 cells transgenically complemented with retroviral MSCV vectors for expression of human BAP1-C91A and BAP1-WT. MSCV-empty vector was used as a control. BAP1-C91A: a catalytically inactive mutant variant of BAP1. (k) qRT-PCR analysis of mRNA expression for BAP1 and Cd274. Data are representative of two (b–d) or three (e–k) independent experiments. ***P < 0.001, **P < 0.01, *P < 0.05 by unpaired student’s t-test (g and k), or one-way ANOVA (I). ns not significant. Error bars indicate SEMs
MAGeCK [11] analysis of gRNA abundances in the S1, S2, and S3 samples revealed as expected that the most highly enriched gRNAs in the S2 and S3 PD-L1low/− cell population were those targeting PD-L1 (encoded by Cd274) itself (Fig. 1b–d and Supplementary Fig. 1h). We also detected that the S3 PD-L1− cell population had significant enrichment for gRNAs that target the Bap1 and Elf4 genes (Fig. 1c, d and Supplementary Fig. 1h), which have no previously reported functional connection to the regulation of PD-L1 expression. We conducted experiments that confirmed deletion of either Bap1 or Elf4 using specific single gRNAs led to a marked reduction expression of both proteins and mRNAs of PD-L1 in MC38-Cas9 cells (Fig. 1e–i and Supplementary Fig. 1i, j). The decreased expression of PD-L1 in Bap1-KO cells was further validated in various mouse cancer cell lines (colon carcinoma cells CT26, mammary carcinoma cells EMT6, and breast cancer cells 4T1) with different gRNAs targeting Bap1 (Supplementary Fig. 1k–m).
The Bap1 protein is a deubiquitinase that can remove histone monoubiquitin modifications from Lys119 (H2AK119Ub) sites [17]. We found an obvious increase in the total extent of H2AK119Ub in Bap1-KO MC38 cells compared to WT cells (Fig. 1f) and re-expression of WT human BAP1 but not a catalytical inactive BAP1 Cys91-Ala (C91A) variant [18] in Bap1-KO cells rescued both cell-surface PD-L1 accumulation levels and Cd274 mRNA expression (Fig. 1j, k), indicating that Bap1-mediated regulation of PD-L1 expression is dependent on its enzymatic activity. We also found that the re-expression of WT transcription factor Elf4 in Elf4-KO MC38 cells resulted in the restoration of PD-L1 expression (Fig. 1i).
3.2 Bap1 and Elf4 independently regulate basal PD-L1 expression in mouse cancer cell lines
Elf4 is a member of the ETS transcription factor family [19]. To confirm Elf4-mediated Cd274 expression, we conducted bioluminescence firefly luciferase and chromatin immunoprecipitation (ChIP) PCR assays. Based on JASPAR [20] analysis, a putative Elf4 binding site and two known IRF1 binding sites [21] were identified within the Cd274 promoter (Supplementary Fig. 2). To examine the role of the predicted ETS motif in Elf4-mediated Cd274 expression, Cd274 promoter constructs with and without ETS motif were cloned into pGL4.10 firefly luciferase plasmids (Fig. 2a). These constructs were individually transfected into Elf4-KO or Elf4 ectopically restored Elf4-KO MC38 cells for bioluminescence assay. The results revealed that Cd274 promoter activity decreased significantly after Elf4 deletion, but could be restored by Elf4 re-expression (Fig. 2b). Moreover, in the presence of Elf4, the Cd274 promoter activity was also significantly reduced when the ETS motif was deleted (Fig. 2b), indicating that Elf4 regulates Cd274 expression by binding to this ETS motif. The ChIP-qPCR assay confirms that Elf4 only binds the region containing the ETS motif within the Cd274 promoter (Fig. 2c, d). Thus, these assays demonstrate that Elf4 directly binds an ETS motif around the Cd274 transcription start site and regulates its expression in MC38 cells.
Bap1 and Elf4 independently regulate the transcription of Cd274. (a) Schematic representation of firefly luciferase construct with Cd274 promoter. Cd274 promoter: wild type Cd274 promoter sequence; Cd274-▵ETS promoter: Cd274 promoter sequence with ETS motif deleted. (b) and (e) Histograms showing the relative activity of Cd274 promoter in indicated MC38 cells. Relative promoter activities were examined by measuring the relative activity of luciferase. The cells were co-transfected with Renilla pRL-SV40 constructs and PGL4.10 constructs with and without Cd274 promoter. Luciferase activity was measured 36 h post-transfection. Relative luciferase activities were determined by the ratio of firefly relative light units (RLUs) to Renilla RLUs. Elf4-KO1 + Elf4-Myc-DDK: Elf4 knock-out clone 1 with Elf4 cDNA fused with Myc-DDK tags overexpression. Bap1-KO2 + Myc-DDK-Bap1: Bap1 knock-out clone 2 with Bap1 cDNA fused with Myc-DDK tags overexpression. (c) Schematic diagram showing the design of the primers for ChIP-qPCR assay on Elf4 binding activity within Cd274 promoter. (d) and (h) ChIP-qPCR showing the binding activities of Elf4 and Bap1 in different regions of Cd274 promoter in indicated cells. Results were represented as percent enrichment relative to input. Bap1-Elf4 DKO + Elf4-Myc-DDK (Myc-DKK-Bap1): Bap1 and Elf4 double knock-out clone with Elf4 (Bap1) cDNA fused with Myc-DDK tags overexpression. (f) and (g) Co-IP examining the interaction between Elf4 and Bap1 proteins. Bap1-cDNA and Elf4-Myc-DKK cDNA were cloned into pcDNA3.1 constructs and were transfected into HEK293FT cells. The anti-Myc tag antibody or anti-Bap1 antibody was used for immunoprecipitation. Immunoglobulin (Ig) G was used as a negative control in (g). Precipitated proteins were detected by western blot using the antibody as indicated. Results were represented as percent enrichment relative to input. Data are representative of two (b and d–h) independent experiments. ***P < 0.001, **P < 0.01, *P < 0.05 by one-way ANOVA (b, d–e, and h). ns not significant. Error bars indicate SEMs
Consistent with the reduction in Cd274 mRNA expression after Bap1 deletion, the Cd274 promoter activity was significantly decreased in Bap1-KO MC38 cells (Fig. 2e). However, when Bap1 expression was reintroduced, the Cd274 promoter activity rebounded (Fig. 2e). As Bap1 and Elf4 are the two prominent novel regulators of PD-L1 in our screening (Fig. 1d), we investigated the potential coordination between Bap1 and Elf4 in regulating Cd274 expression. Co-immunoprecipitation (Co-IP) experiments revealed no interaction between Bap1 and Elf4 proteins (Fig. 2f, g). Additionally, ChIP-qPCR analysis demonstrated that the absence of Bap1 did not affect the binding activity of Elf4 to the Cd274 promoter (Fig. 2d). In contrast, while Elf4 exhibited restricted binding activity within the Cd274 promoter (Fig. 2d), Bap1 displayed broad binding sites across the promoter, with its binding activity unaffected by Elf4 deletion (Fig. 2h). These findings suggest that Bap1 and Elf4 independently regulate the constitutive basal expression of Cd274 in mouse cancer cell lines.
3.3 Bap1-deleted cancer cells are sensitive to CD8+ T cell-mediated eradication
Their regulation of PD-L1 expression motivated us to investigate the impacts of Bap1 and Elf4 deletion on anti-tumor immunity. Bap1-KO, Elf4-KO, and control WT cells were subcutaneously injected into syngeneic WT mice or immunocompromised mice for tumor growth examination (Fig. 3a). Similar to WT tumors, the Elf4-KO MC38 tumors grew rapidly and the animals had to be euthanized after 28 days (Supplementary Fig. 3a). By contrast, Bap1-KO MC38 cells exhibited poor growth under the same conditions and were eventually eliminated (Fig. 3b, c), with the maximum observed tumor volume occurring around day 12 (Supplementary Fig. 3b). Similar results on Bap1 deletion were obtained with other tumor models in syngeneic BALB/c mice using various cancer cell lines (CT26, 4T1, and EMT6) (Fig. 3b, c and Supplementary Fig. 3b).
The effects of Bap1-KO cancer cells in syngeneic mice. (a) Mouse syngeneic model and experimental design. (b) Volumes of the indicated tumors in syngeneic mice. MC38 cells were subcutaneously injected into WT C57BL/6 mice (n = 5 for WT; n = 6 for Bap1-KO). CT26 (n = 10 for WT; n = 5 for Bap1-KO), 4T1 (n = 6 for WT; n = 6 for Bap1-KO), and EMT6 (n = 4 for WT; n = 4 for Bap1-KO) cells were subcutaneously injected into WT BALB/c mice. Western blot images showing the protein expression levels of Bap1 in the indicated cancer cell lines. β-tubulin was used as an internal control. (c) Survival analysis of mice in (b). (d) The volumes of indicated tumors in NSG (n = 6 for WT; n = 6 for Bap1-KO), Rag2−/− (n = 9 for WT; n = 8 for Bap1-KO), Tcra−/− (n = 6 for WT; n = 6 for Bap1-KO), or WT C57BL/6 mice that were treated with anti-CD8α (n = 6 for WT; n = 8 for Bap1-KO), anti-CD4 (n = 5 for WT; n = 6 for Bap1-KO), or anti-CD19 (n = 6 for WT; n = 6 for Bap1-KO) depletion antibodies. The antibody amounts and treatment timings were described in the methods. (e) The volumes of WT B16.F10 or WT MC38 tumors in WT C57BL/6 mice that were injected with Bap1-KO MC38 cells (n = 5 for WT B16.F10; n = 5 for WT MC38). 40 days after Bap1-KO MC38 tumor clearance, the mice were injected with WT B16.F10 cells or WT MC38 cells on the contralateral side. (f) The volumes of WT B16.F10 or WT MC38 tumors in WT C57BL/6 mice with anti-CD8α treatment (n = 3 for WT B16.F10; n = 3 for WT MC38). The mice were injected with Bap1-KO MC38 cells, 40 days after Bap1-KO MC38 tumor clearance, and then the mice were injected with anti-CD8α depletion antibodies before injections of WT B16.F10 cells or WT MC38 cells on the contralateral side on the same day. Data are representative of three (b–c) or two (d–f) independent experiments. ***P < 0.001 by unpaired student’s t-test (b–f). ns not significant. Error bars indicate SEMs
Although both Elf4 and Bap1 are essential regulators for PD-L1 expression in cultured cells (Fig. 1e, h), their deletions led to contrasting outcomes after implantation. While Elf4 deletion in MC38 cells does not alter the rate of tumor growth compared to control WT cells, Bap1 deletion in MC38 cells led to efficient clearance of the cancer cells. To investigate this further, we conducted additional experiments to assess the expression of PD-L1 on Elf4-KO and Bap1-KO MC38 cells cells isolated from the syngeneic mice a few days after injection (before Bap1-KO cells were eliminated). Intriguingly, both Elf4-KO and Bap1-KO MC38 cells showed a restored PD-L1 expression level to that of WT control cells in the WT C57BL/6 mouse, not restored in the Rag2−/− mouse (Supplementary Fig. 3c), suggesting that there are in vivo factors that induce the PD-L1 expression independent of Elf4 and Bap1, likely derived from lymphocytes. It’s well known that the cytokine IFNγ, mainly produced by active T cells, can upregulate PD-L1 through JAK-STAT-IRF1 axis [21]. Therefore, we evaluated the PD-L1 expression on Elf4-KO and Bap1-KO MC38 cells under IFNγ stimulation. Both Elf4-KO and Bap1-KO cells exhibited similar levels of PD-L1 as the WT control cells (Supplementary Fig. 3d), indicating IFNγ induced PD-L1 expression is not dependent of Elf4 and Bap1. These findings suggest that elimination of Bap1-deleted MC38 cells by the host immune system may be independent of PD-L1.
To confirm that the lower basal level of PD-L1 expression on Bap1-KO MC38 cells (Fig. 1e) does not trigger anti-tumor immunity upon injection into mice, we treated Bap1-KO MC38 cells with IFNγ to restore their PD-L1 expression before implantation. However, even with restored PD-L1 expression, these pre-treated Bap1-KO MC38 cells were still efficiently cleared in C57BL/6 mice (Supplementary Fig. 3e). In addition, Bap1-KO MC38 cells overexpressing Cd274 were also effectively eliminated after injection (Supplementary Fig. 3f). These results further confirm that the eradication of MC38 cells triggered by Bap1 deletion is independent of PD-L1 expression.
Moreover, the overexpression of human WT BAP1 in Bap1-KO MC38 cells promoted their growth in host mice (Supplementary Fig. 3g–j). This indicates that Bap1 has a distinctive role in anti-tumor immunity, independent of its association with PD-L1. As a result, our focus will be on investigating the mechanisms underlying the eradication of Bap1-deleted MC38 cells by the host immune system in the subsequent part of our study.
To identify which effector immune cells are involved in the observed eradication of Bap1-deleted tumors, WT (MC38-Cas9) and Bap1-KO MC38 cells were injected into various immunocompromised mice, including NSG (NOD.Cg-PrkdcscidIl2rgtm1Wjl/SzJ) (lack of B, T and innate lymphoid cells), Rag2−/− (lack of B and T cells), Tcra−/− (T cells deficiency), CD8+ T-cell-depleted (anti-CD8α treated), CD4+ T-cell-depleted (anti-CD4 treated), and CD19+ B-cell-depleted (anti-CD19 treated) C57BL/6 mice (Fig. 3a and Supplementary Fig. 3k). Upon injection of WT cells, tumors developed rapidly in all of the immunocompromised mice (Fig. 3d and Supplementary Fig. 3l). However, Bap1-KO MC38 cells formed tumors rapidly in certain immunocompromised mice, namely NSG, Rag2−/−, Tcra−/−, and CD8+ T-cell-depleted mice, but did not form tumors in mice depleted of CD4+ T-cells or CD19+ B-cells (Fig. 3d and Supplementary Fig. 3l). These findings strongly suggest that the eradication of Bap1-KO MC38 tumors is reliant on CD8+ T cells.
Notably, we also detected an apparent protective immune memory response. Upon re-challenging C57BL/6 mice that had been cured from their primary Bap1-KO MC38 tumors with WT MC38 and WT B16.F10 cells, we observed WT MC38 tumors didn‘t grow in these mice (Fig. 3e and Supplementary Fig. 3m) and we found that functionally reactive CD8+ T cells developed during the initial immune response later supported the rejection of the MC38 cells (Fig. 3f).
3.4 Loss of Bap1 increases tumor inflammation
To understand how the loss of Bap1 activates anti-tumor immunity, we conducted a comparison of the immune subsets present in the tumor microenvironment of Bap1-KO and control WT (MC38-Cas9) tumors. The tumors were collected approximately 10 days after the injection of MC38 cells in C57BL/6 mice (before obvious shrinkage of Bap1-KO MC38 tumors occurred). These tumors were then subjected to flow cytometry and single-cell RNA-sequencing (scRNA-seq) analyses (Fig. 4a).
Bap1 deletion leads to an inflamed tumor microenvironment. (a) Schematic of experimental design. Tumors were isolated from the C57BL/6 mice ~ 10 days after injection. (b) Comparison of the ratios of CD45+ T cells to BFP+ tumor cells in WT and Bap1-KO MC38 tumors (n = 16 for WT; n = 14 for Bap1-KO). Left FACS plots representing the gating strategies of CD45+ cells in the tumors. Right histogram showing the rations of CD45+ cells to BFP+ tumor cells. (c) Comparison of the ratios of T subsets to tumor cells in indicated MC38 tumors (n = 16 for WT; n = 14 for Bap1-KO). (d) Comparison of the ratios of CD103+dendritic cells (cDC1) to tumor cells in indicated MC38 tumors (n = 15 for WT; n = 13 for Bap1-KO). (e) Immunofluorescence assay showing CD8α+ T cell infiltration into tumors (n = 3 for WT; n = 3 for Bap1-KO). Green represents CD8α+ cells, and blue (DAPI) represents the nucleus. (f) A t-SNE plot of scRNA-seq of CD45+ cells from WT and Bap1-KO tumors. Each point represents an individual cell. Colors indicate different cell types. Treg, regulatory T cells; cDC, conventional dendritic cells; pDC, plasmacytoid dendritic cells; NK, natural killer cells. (g) A t-SNE plot showing the sample distribution. WT indicates the CD45+ cells derived from WT MC38 tumors. KO represents the CD45+ cells derived from Bap1-KO MC38 tumors. (h) Bar plots showing the percentages of immune cells from the WT and Bap1-KO tumors in the indicated cell types. (i) The t-SNE plot showing the re-clustering of CD8+ T cells from the tumors. 9 sub-clusters in CD8+ T cells were identified. (j) A t-SNE plot showing the sample distribution in CD8+ T cells. WT indicates the CD8+ T cells derived from WT tumors; KO represents the CD8+ T cells derived from Bap1-KO tumors. (k) and (l) Projection of the naïve CD8 T cell (k) and effector CD8 T cell signatures (l) in t-SNE plots. The signature genes of naïve CD8 T cells and effector CD8 T cells were obtained from a previous publication [56]. The color key indicates the average of normalized signature gene expressions of naïve CD8 T cell or effector CD8 T cell over the randomly sampled gene expressions. (m) The ratios of effector T cells (Teff) to regulatory T cells (Treg) in WT and Bap1-KO tumors by scRNA-seq analysis. Data are representative of three (b–d) or two (e) independent experiments. ***P < 0.001, **P < 0.01, *P < 0.05 by unpaired student’s t-test (b–e). ns not significant
Flow cytometry analysis showed that Bap1-KO MC38 tumors had significantly higher populations of CD45+ leukocytes, CD3+, CD4+, CD8+, and γδ T cells, CD103+DCs (cDC1 cells), and M1 macrophages compared to WT tumors, (Fig. 4b–d and Supplementary Fig. 4a–c). Immunostaining of tumor sections confirmed a notable increase in the number of CD8α+ cells, which diffusely infiltrated the Bap1-KO tumors (Fig. 4e), accompanied by a substantial increase in apoptotic cells, as determined by TUNEL staining (Supplementary Fig. 4d).
The results of the scRNA-seq analysis of CD45+ cells in the MC38 tumor microenvironment once again demonstrated the increased proportions of CD8+ T cells and cDC1 cells, while revealing a decrease in neutrophils and Tregs in Bap1-KO tumors (Fig. 4f–h and Supplementary Fig. 5a, b). Specifically focusing on the analysis of CD8+ T cells, we observed an enrichment of cells with CD8+ T effector, CD8+ memory precursor, and IFNγ signatures in Bap1-KO tumors compared to control WT tumors (Fig. 4i–l and Supplementary Fig. 5c–g). Additionally, Bap1-KO tumors exhibited a higher ratio of effector CD8+ T cells to Tregs compared to control WT tumors (Fig. 4m). Overall, these findings suggest that the deletion of Bap1 leads to a more favorable anti-tumor immune microenvironment.
3.5 Bap1-deleted cancer cells evoke a cDC1-dependent and Fas-mediated anti-tumor response
The finding of the increase in the proportion of cDC1 cells in Bap1-KO tumors prompted us to examine whether cDC1 cells are required for the observed Bap1-KO tumor eradication. We injected Bap1-KO or WT (MC38-Cas9) cells into Batf3−/− mice (Fig. 5a), which can not develop cDC1 cells [22]. In contrast to the successful elimination observed for the Bap1-KO tumors in WT mice (Fig. 3b), the Batf3−/− mice were unable to clear Bap1-KO tumors (Fig. 5b), indicating that cDC1 cells indeed are essential for eradication of Bap1- KO tumors.
Bap1 deletion stimulates cDC1-dependent and Fas-mediated anti-tumor response. (a) Schematic of experimental design. (b) The volumes of indicated WT (black) or Bap1-KO (red) MC38 tumors in Batf3−/− mice (n = 6 for WT; n = 6 for Bap1-KO). (c) FACS plots showing the percentages of antigen-specific CD8+ T cells by tetramer SIINFEKL/H2Kb staining in indicated tumors in WT C57BL/6 mice and Batf3−/− mice. Tumors were isolated 10 days after injections. (d) Statistical graph showing the percentages of antigen-specific CD8+ T cells by tetramer SIINFEKL/H2Kb staining in indicated tumors in WT C57BL/6 mice (n = 3 for WT; n = 4 for Bap1-KO). (e) Tumor volumes of Bap1-B2m DKO MC38 clones in WT C57BL/6 mice. Each line represents a tumor. (f) Tumor volumes of Bap1-Tap1 DKO and Bap1-Tap2 DKO MC38 clones in WT C57BL/6 mice. Each line represents a tumor. (g) Tumor volumes of WT or Bap1-KO MC38 tumors in Prf1−/− mice (n = 6 for WT; n = 6 for Bap1-KO). (h) Tumor volumes of Bap1-Fas DKO MC38 clones in WT C57BL/6 mice. Each line represents a tumor. (i) Tumor volumes of Bap1-Fas DKO MC38 clones in Prf1−/− mice. Each line represents a tumor. (j) Tumor volumes of control WT or Bap1-KO tumors in mice treated with anti-IFNγ antibody (n = 4 for WT; n = 4 for Bap1-KO). Mice were intraperitoneally given IFNγ at day −5, −1, 0, 1, 2, 3, 4, and 5 after tumor cells injection (200 µg/mouse/injection). Data are representative of two (b–j) independent experiments. ***P < 0.001, *P < 0.05 by unpaired student’s t-test (b, d, g and j). ns not significant. Error bars indicate SEMs
To determine the role of cDC1-mediated antigen presentation in the increase of CD8+ T cells in Bap1-KO tumors, we conducted an experiment involving the expression of full-length ovalbumin (OVA) in Bap1-KO or WT MC38 cells (Supplementary Fig. 6a). We then injected these cells into Batf3−/− or WT C57BL/6 mice (Fig. 5a). Our findings demonstrated that in WT mice, there was a significantly higher proportion of the antigen-specific CD8+ T cells recognizing the SIINFEKL epitope from OVA (in the context of H2k(b)) in Bap1-KO tumors compared to WT tumors (Fig. 5c, d). However, in Batf3−/− mice, no antigen-specific CD8+ T cells were detected in either Bap1-KO or WT tumors (Fig. 5c). These results suggest that the infiltration of tumor antigen-specific CD8+ T cells promoted by Bap1 deletion is dependent on cDC1 cells. Consequently, our findings indicate that Bap1 deletion in cancer cells stimulates an anti-tumor response that relies on cDC1-CD8+ T cell activation.
Having observed a significant increase in antigen-specific CD8+ T cells in Bap1-KO tumors, we sought to investigate the role of tumor antigen presentation in Bap1-KO tumor clearance. We suppressed the expression of major histocompatibility complex class 1 (MHCI) molecules in Bap1-KO MC38 cells by deleting B2m, Tap1, or Tap2 (Supplementary Fig. 6b, c), each of which is essential for MHCI molecule generation [23]. Following the injection of these double knockout (DKO) cancer cells into WT C57BL/6 mice, all DKO cancer cell types (including Bap1-B2m DKO, Bap1-Tap1 DKO, and Bap1-Tap2 DKO) formed rapidly growing tumors (Fig. 5e, f). These findings suggest that the presence of MHCI molecules on the surface of tumor cells is necessary for the antigen presentation and subsequent eradication of Bap1-KO tumors.
The perforin-granzyme and Fas pathways are two major mechanisms through which CD8+ T cell exert cytotoxicity [24]. To investigate the involvement of these pathways in Bap1-KO MC38 tumor clearance, we injected Bap1-KO cells into Prf1−/− mice, which lack perforin and exhibit impaired granule exocytosis [25]. Interestingly, Bap1-KO tumors were efficiently eliminated in the absence of perforin (Fig. 5g). Additionally, we generated Bap1-Fas DKO MC38 cells (Supplementary Fig. 6d) and injected them into both WT and Prf1−/− mice. The DKO cells formed tumors in both types of mice (Fig. 5h, i), supporting that the eradication of Bap1-KO tumors is dependent on the Fas-mediated CD8+ T cell cytotoxicity pathway.
We hypothesized that IFNγ could potentially contribute to the eradication of Bap1-KO MC38 tumors, considering an increased expression of the IFNγ signature in CD8+ T cells from Bap1-KO tumors (Supplementary Fig. 5e). To investigate this, WT C57BL/6 mice were administered with an anti-IFNγ neutralizing antibody, after which they were implanted with Bap1-KO or WT MC38 cells. In mice treated with anti-IFNγ antibodies, both Bap1-KO and WT MC38 cells formed tumors (Fig. 5j), suggesting the clearance of Bap1-KO tumors requires IFNγ. Moreover, Bap1-KO MC38 cells exhibited greater sensitivity to IFNγ induced cell apoptosis in vitro (Supplementary Fig. 6e).
In summary, these findings highlight the importance of tumor MHCI molecules and Fas-mediated CD8+ T cell cytotoxicity in the elimination of Bap1-deleted tumors. Additionally, the presence of IFNγ appears to play a critical role in this process.
3.6 Bap1 orchestrates tumor resistance to immunity
Bap1, an H2A histone deubiquitinase, has been linked to chromatin remodelling and gene transcription [17, 26, 27]. To understand how Bap1 affects anti-tumor immunity, we investigated the changes in transcriptome and epigenome in tumor cells after Bap1 deletion. Specifically, we analyzed MC38 tumor cells isolated from the Rag2−/− mice 10 days post-implantation using RNA-seq, quantitative RT-PCR (qRT-PCR) and cleavage under targets and tagmentation sequencing (CUT&Tag-seq). We also collected in vitro cultured MC38 cells and performed transposase-accessible chromatin with high-throughput sequencing (ATAC-seq) analysis under stimulation with IFNγ (Fig. 6a).
The gene expressions and their associated epigenomes change after Bap1 deletion. (a) Schematic of experimental design. MC38 tumors isolated from the Rag2−/− mice on day 10 after injection were subjected to bulk RNA-seq, qRT-PCR, and CUT&Tag-seq. MC38 cells stimulated under IFNγ (20 ng/mL) for 48 h in vitro were subjected to ATAC-seq. (b) Heatmap showing the indicated gene expressions analyzed by RNA-seq in WT and Bap1-KO cells. Colour scales show gene expressions. Log2FoldChange > 1, FDR < 0.01. (c) qRT-PCR analysis of the expressions of the indicated genes in WT and Bap1-KO cells. All values are normalized to the expression of β-actin. (d) UCSC genome browser tracks showing the chromatin accessibilities and three relevant histone marks (H2AK119Ub, H3K27ac, and H3K4me3) around the Bap1, Slc7a11, Cxcl1, Epha2, Thbs2 genes loci. Differential accessibility or histone modification regions between WT and Bap1-KO cell types were highlighted. ***P < 0.001, **P < 0.01 by unpaired student’s t-test (c). ns not significant. Error bars indicate SEMs
Our analysis reveals that Bap1-KO MC38 tumor cells exhibit 488 upregulated genes and 949 downregulated genes compared to the WT control group (FDR < 0.01 and log2foldchange > 1) (Supplementary Table S1). Interestingly, we observed that Bap1-KO MC38 tumor cells exhibited higher expression of a known Bap1 regulation gene Slc7a11 [28]. Furthermore, the accessibility of the Slc7a11 locus around its promoter region was increased, accompanied by enhanced H3K27ac and H3K4me3 modifications associated with active gene regulation, compared to the WT control group (Fig. 6b–d), These findings not only demonstrate the reliability of our multi-omics data but also validate the functional implications of Bap1 in regulating gene expression (Supplementary Fig. 7b–d and Supplementary Tables S1, 2).
The RNA-seq analysis revealed differential expression of several chemokines between Bap1-KO and control WT tumor cells (Fig. 6b and Supplementary Table S1). Notably, consistent with our previous observations (Fig. 4c, d and h), we observed a significant increase in chemokines that promote the infiltration of tumor cDC1 and CD8+ T cells, such as Ccl7 [29], and Ccl11 [30], upon Bap1 deletion. Conversely, chemokines associated with the recruitment of tumor-associated neutrophils including Cxcl1 [31] and Cxcl2 [32], were found to be significantly reduced (Fig. 6b, c). Additionally, CUT&Tag-seq analysis revealed a decrease in the extent of H3K27ac modification around the Cxcl1 promoter (Fig. 6d). These findings suggest that multiple chemokines regulated by Bap1 contribute to shaping the tumor immune microenvironment.
The gene expression analysis also identified several well-known immune-evasion genes that exhibited significant decreases in MC38 tumor cells following Bap1 deletion (Supplementary Table S1), for example, Tgfb3, Smad3, and Epha2 (Fig. 6b, c). Consistently, the regions around their promoters in Bap1-KO cells showed reduced levels of H3K27ac and H3K4me3 modifications compared to WT controls (Fig. 6d and Supplementary Fig. 7a). Additionally, Epha2 locus displayed decreased accessibility (Fig. 6d). Tgfb3 is a ligand and Smad3 is a downstream effector of the TGFβ pathway, which plays roles in shaping tumor microenvironment by inhibiting T-cell infiltration and restraining anti-tumor immunity [33]. Epha2, on the other hand, is a tyrosine kinase, and its deletion in pancreatic ductal adenocarcinoma has been shown to increase T-cell infiltration, rendering the tumor more sensitive to immunotherapy [34].
Furthermore, Epha2 has been shown to promote tumor angiogenesis [35]. Another gene, Runx1 [36], which is known to stimulate angiogenesis, exhibited significantly reduced expression in Bap1-KO tumor cells. Additionally, the expression levels of anti-angiogenesis genes such as Thbs2 [37,38,39] and Ndrg1 [40] were found to be significantly increased in Bap1-KO tumor cells (Fig. 6b, c). Thbs2, in particular, encodes thrombospondin-2, which is a potent inhibitor of tumor growth and angiogenesis [39]. It is worth noting that Thbs2 displayed the largest fold-change (~ 100 times) among the differentially expressed genes shown in Fig. 6. The regulatory effect of Bap1 on Thbs2 transcription was confirmed by ATAC-seq and CUT/Tag-seq analyses, which demonstrated that the deletion of Bap1 resulted in increased accessibility of the Thbs2 locus and higher H3K27ac and H3K4me3 levels in MC38 tumor cells (Fig. 6d). Taken together, these findings suggest that Bap1 plays a role in modulating the tumor microenvironment by promoting tumor angiogenesis.
In addition, Bap1 also appear to influence the innate immune response. We observed that the expression of innate immune response genes, such as Mavs [41] and Ifitm1 [42], was significantly increased in Bap1-KO MC38 tumor cells after Bap1 deletion (Fig. 6b and Supplementary Table S1). This suggests that Bap1 may contribute to shaping the tumor immune microenvironment through the regulation of the innate immune response.
Taken together, the multi-omics analysis reveals that Bap1 deletion leads to multiple changes within tumor cells, resulting in a cDC1-CD8+ T cell-mediated cytotoxicity in an immune-stimulatory tumor microenvironment. These findings highlight the role of Bap1 as an epigenetic orchestrator that modulates the susceptibility of tumors to immune system attack.
3.7 Clinical relevance of BAP1 to tumor immunity
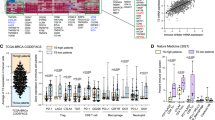
To examine how our findings in murine cancer may be clinically relevant, we analyzed transcriptome data from 33 different human cancer types in The Cancer Genome Atlas (TCGA) dataset. We specifically looked at the correlation between BAP1 expression levels and T-cell cytotoxicity, as well as various immune cell scores using xCell [43]. Our analysis revealed that a negative correlation between BAP1 mRNA levels and the extent of T-cell cytotoxicity in a variety of human cancer types (Fig. 7a). For instance, in lung squamous cell cancer (LUSC) and breast cancer (BRCA), BAP1-low tumors (bottom 25% of BAP1 expression) showed significant increases in signatures of T-cell cytotoxicity, CD8+ T cell and activated DCs compared to BAP1-high tumors (top 25% of BAP1 expression) (Fig. 7b and Supplementary Fig. 8a–c). These findings indicate that BAP1 expression is inversely associated with the infiltration of various immune cells across different human cancers.
Effects of Bap1 deletion on the efficacies of immune checkpoints blockade and CAR-T cell immunotherapies. (a) Volcano plot showing Spearman’s correlation and estimated significance of BAP1 with T cell cytotoxicity mRNA levels from RNA-seq data across 33 TCGA cancer types. Red dots indicate significant correlations (P < 0.01). T-cell cytotoxicity is derived from the average mRNA expression levels of GZMB and IFNG. (b) Comparison of the scores for T cell cytotoxicity, CD8+ T cells, and activated myeloid DCs between BAP1-high (upper quartile of BAP1 expression) and BAP1-low (lower quartile of BAP1 expression) groups in lung squamous cell cancer (LUSC). CD8+ T cells and activated myeloid DC scores were computed by xCell [43]. (c) Experimental design for the tumor fragment implantation experiment. (d) Tumor growth after treatment with anti-PD-1 and anti-CTLA-4 antibodies in C57BL/6 mice that were implanted with tumor fragments. Tumors were treated with three doses of anti-PD-1 (250 µg/mouse/dose) and anti-CTLA-4 (200 µg/mouse/dose) at day 16, 19, and 22 after tumor fragment implantation (right panel) or treated with PBS as a control (left panel) (n = 3 for WT; n = 3 for Bap1-KO). (e) Experimental design for the CAR-T cell (anti-CD19) therapy. CD8+ T cells were isolated from C57BL/6 mice and were transduced with anti-CD19 CAR [13] to generate CAR-T cells. CAR-T cells were injected via tail vein into Rag2−/− mice on day 3 after tumor cells injection. (f) The volumes of CD19-expressing Bap1-KO or CD19-expressing WT tumors in Rag2−/− mice under the treatment without (left panel, n = 3 for WT; n = 3 for Bap1-KO) and with (right panel, n = 4 for WT; n = 6 for Bap1-KO) anti-CD19 CAR-T cells. (g) The volumes of OVA-expressing Bap1-KO or WT tumors in Rag2−/− mice (n = 5 for WT; n = 6 for Bap1-KO) after the injection of activated OT-I T cells. Data are representative of two (d, f and g) independent experiments. ***P < 0.001, **P < 0.01, *P < 0.05 by a two-side Wilcoxon ran-sum test (b), or unpaired student’s t-test (d, f and g). ns not significant. Error bars indicate SEMs
To investigate the implications of these findings on immunotherapies, we conducted an experiment simulating the clinical setting of patients with fully established tumors treated with immune checkpoint inhibitors. Initially, we inoculated WT or Bap1-KO MC38 cells in immune-deficient Rag2−/− mice to induce tumor formation. Once the tumors grew to a size of 800 mm3, we performed surgery and implanted tumor fragments into syngeneic WT C57BL/6 mice (Fig. 7c). Surprisingly, the growth of both the Bap1-KO and WT tumors was comparable in WT C57BL/6 mice (Fig. 7d). However, when we treated them with monoclonal antibodies targeting PD-1 and CTLA-4, we observed a significant suppression of Bap1-KO tumors compared to WT tumors (Fig. 7d). This suggests that Bap1-deleted tumors are more susceptible to the immunotherapeutic effects of immune checkpoint inhibitors.
We further investigated whether the deletion of Bap1 could enhance the effectiveness of CAR-T cell therapy. We introduced CD19 expression in both Bap1-KO MC38 and WT control cells (Supplementary Fig. 8d) and implanted them in Rag2−/− mice. Once the tumors became visible, we treated the mice with CD8+ CAR-T cells (anti-CD19) derived from C57BL/6 mice (Fig. 7e). Interestingly, the administrated CAR-T cells displayed greater efficacy in inhibiting the growth of Bap1-KO tumors compared to WT tumors (Fig. 7f). Additionally, we observed that Bap1-KO (OVA+) tumors were more susceptible to OT-I T cell-mediated killing, both in vivo and in vitro (Fig. 7g and Supplementary Fig. 8e). These findings suggest that Bap1 deletion can sensitize tumor cells to CD8+ T cell-mediated killing, thereby indicating that modulating Bap1 expression by inhibiting Bap1 catalytic activity can potentially enhance the efficacy of immunotherapies.
4 Discussion
A better understanding of the mechanisms that regulate PD-L1 expression in cancer cells could improve the efficacy of immune checkpoint blockade therapy [1]. In this study, an unbiased genetic screening approach was used to identify two new regulators of PD-L1: the transcription factor Elf4 and the histone deubiquitinase Bap1. These two regulators were found to independently control the constitutive basal expression of PD-L1 in various cultured cancer cells, separating from the regulation of PD-L1 expression by IFNγ.
We fortuitously found that deletion of Bap1 in cancer cells can trigger an anti-tumor immunity, regardless of the expression of PD-L1. BAP1 is a well-known tumor suppressor gene [44] that was initially identified in metastasizing uveal melanoma [45]. Since then, it has been found to be commonly mutated or lost in various types of cancer, including mesothelioma [46,47,48]. Interestingly, studies have shown that patients with BAP1 germline mutations have a better prognosis in mesothelioma [49, 50]. This improved survival may be linked to the tumor immune microenvironment affected by BAP1 mutations [51], as the reduction in BAP1 is correlated with tumor inflammation [52]. Consistent with these findings, our experiments using MC38 mouse models revealed that deletion of Bap1 in cancer cells led to significant increases in cDC1 cells and CD8+ T cells, with markers indicating their effector and memory precursor functions, as well as increased levels of IFNγ.
We have discovered that the genes affected by Bap1 deletion span various categories, including chemokines, immune evasion, angiogenesis, and innate immune response. This suggests that Bap1, as a chromatin modifier, plays a crucial role in regulating gene expression [27] and orchestrating tumor immunity. Interestingly, Bap1-deleted tumors exhibit enrichment for anti-angiogenesis-related genes, which may contribute to their increased vulnerability to immune checkpoint inhibitors [53, 54].
Our results showed that when Bap1 was deleted, tumors became more susceptible to combined anti-PD-L1 and anti-CTLA-4 monoclonal antibodies therapy, as well as CAR-T cell therapy. These findings were consistent with a recent report that analyzed data from the JAVELIN Renal 101 and CheckMate-009/010/025 trials [55]. The report identified a BAP1-related signature that can predict the efficacy of immune checkpoint inhibitors-based immunotherapies. The inclusion of this additional evidence strengthens the argument for testing CAR-T cell therapy in BAP1-mutated tumors.
We successfully identified two new regulators of PD-L1 expression using the CRISPR/Cas9 screen. However, a limitation of our screening method arose from the stringent PD-L1 negative sorting across several rounds. As a result, the cells we selected were predominantly enriched with gRNAs targeting Cd274 directly and its most potent regulators. This might have led to the oversight of some regulators that modulate Cd274 expression at less optimal levels. Our findings clearly show that Bap1 deletion induces multiple alterations in tumor cells, such as the upregulation of genes linked to anti-tumor immune responses. This upregulation is paired with a stimulating tumor microenvironment, leading to the elimination of Bap1-deleted tumors. Yet, the precise mechanism by which Bap1 deletion in tumor cells activates anti-tumor responses remains a subject of ongoing research.
In conclusion, our study provides compelling evidence for the involvement of Bap1 in tumor immunity and its significance in the context of immunotherapies. These findings offer new possibilities for the development of targeted therapies and personalized treatment approaches for BAP1-mutated cancers.
Data availability
The bulk RNA-seq, scRNA-seq, ATAC-seq, and CUT&Tag data described in this study have been deposited in the BioStudies database under accession numbers E-MTAB-10587, E-MTAB-10610, E-MTAB-10593, E-MTAB-10585 respectively. Additional information about DEGs in the bulk RNA-Seq data is included in Supplementary Table S1. The ATAC-seq results are in Supplementary Table S2. All oligos used in the study are listed in Supplementary Tables S3 and 4. All other relevant data are available from the corresponding author on request.
References
H. Yamaguchi, J.M. Hsu, W.H. Yang, M.C. Hung, Mechanisms regulating PD-L1 expression in cancers and associated opportunities for novel small-molecule therapeutics. Nat. Rev. Clin. Oncol. 19, 287–305 (2022). https://doi.org/10.1038/s41571-022-00601-9
P. Sharma, S. Hu-Lieskovan, J.A. Wargo, A. Ribas, Primary, adaptive, and Acquired Resistance to Cancer Immunotherapy. Cell. 168, 707–723 (2017). https://doi.org/10.1016/j.cell.2017.01.017
J. Dunn, S. Rao, Epigenetics and immunotherapy: the current state of play. Mol. Immunol. 87, 227–239 (2017). https://doi.org/10.1016/j.molimm.2017.04.012
S.J. Gallagher, E. Shklovskaya, P. Hersey, Epigenetic modulation in cancer immunotherapy. Curr. Opin. Pharmacol. 35, 48–56 (2017). https://doi.org/10.1016/j.coph.2017.05.006
S.J. Hogg, P.A. Beavis, M.A. Dawson, R.W. Johnstone, Targeting the epigenetic regulation of antitumour immunity. Nat. Rev. Drug Discov. 19, 776–800 (2020). https://doi.org/10.1038/s41573-020-0077-5
W. Sheng, M.W. LaFleur, T.H. Nguyen, S. Chen, A. Chakravarthy, J.R. Conway, Y. Li, H. Chen, H. Yang, P.H. Hsu, E.M. Van Allen, G.J. Freeman, D.D. De Carvalho, H.H. He, A.H. Sharpe, Y. Shi, LSD1 ablation stimulates anti-tumor immunity and enables checkpoint blockade. Cell. 174, 549–563e519 (2018). https://doi.org/10.1016/j.cell.2018.05.052
Y. Qin, S.N. Vasilatos, L. Chen, H. Wu, Z. Cao, Y. Fu, M. Huang, A.M. Vlad, B. Lu, S. Oesterreich, N.E. Davidson, Y. Huang, Inhibition of histone lysine-specific demethylase 1 elicits breast tumor immunity and enhances antitumor efficacy of immune checkpoint blockade. Oncogene. 38, 390–405 (2019). https://doi.org/10.1038/s41388-018-0451-5
Y. Liu, B. Debo, M. Li, Z. Shi, W. Sheng, Y. Shi, LSD1 inhibition sustains T cell invigoration with a durable response to PD-1 blockade. Nat. Commun. 12, 6831 (2021). https://doi.org/10.1038/s41467-021-27179-7
K. Tzelepis, H. Koike-Yusa, E. De Braekeleer, Y. Li, E. Metzakopian, O.M. Dovey, A. Mupo, V. Grinkevich, M. Li, M. Mazan, M. Gozdecka, S. Ohnishi, J. Cooper, M. Patel, T. McKerrell, B. Chen, A.F. Domingues, P. Gallipoli, S. Teichmann, H. Ponstingl, U. McDermott, J. Saez-Rodriguez, B.J.P. Huntly, F. Iorio, C. Pina, G.S. Vassiliou, K. Yusa, A CRISPR dropout screen identifies genetic vulnerabilities and therapeutic targets in Acute myeloid leukemia. Cell. Rep. 17, 1193–1205 (2016). https://doi.org/10.1016/j.celrep.2016.09.079
H. Koike-Yusa, Y. Li, E.P. Tan, C. Velasco-Herrera Mdel, K. Yusa, Genome-wide recessive genetic screening in mammalian cells with a lentiviral CRISPR-guide RNA library. Nat. Biotechnol. 32, 267–273 (2014). https://doi.org/10.1038/nbt.2800
W. Li, H. Xu, T. Xiao, L. Cong, M.I. Love, F. Zhang, R.A. Irizarry, J.S. Liu, M. Brown, X.S. Liu, MAGeCK enables robust identification of essential genes from genome-scale CRISPR/Cas9 knockout screens. Genome Biol. 15, 554 (2014). https://doi.org/10.1186/s13059-014-0554-4
Z. Miao, P. Moreno, N. Huang, I. Papatheodorou, A. Brazma, S.A. Teichmann, Putative cell type discovery from single-cell gene expression data. Nat. Methods. 17, 621–628 (2020). https://doi.org/10.1038/s41592-020-0825-9
J.N. Kochenderfer, Z. Yu, D. Frasheri, N.P. Restifo, S.A. Rosenberg, Adoptive transfer of syngeneic T cells transduced with a chimeric antigen receptor that recognizes murine CD19 can eradicate lymphoma and normal B cells. Blood. 116, 3875–3886 (2010). https://doi.org/10.1182/blood-2010-01-265041
V.R. Juneja, K.A. McGuire, R.T. Manguso, M.W. LaFleur, N. Collins, W.N. Haining, G.J. Freeman, A.H. Sharpe, PD-L1 on tumor cells is sufficient for immune evasion in immunogenic tumors and inhibits CD8 T cell cytotoxicity. J. Exp. Med. 214, 895–904 (2017). https://doi.org/10.1084/jem.20160801
M. Li, J.S.L. Yu, K. Tilgner, S.H. Ong, H. Koike-Yusa, K. Yusa, Genome-wide CRISPR-KO screen uncovers mTORC1-Mediated Gsk3 regulation in naive pluripotency maintenance and dissolution. Cell. Rep. 24, 489–502 (2018). https://doi.org/10.1016/j.celrep.2018.06.027
J.S.L. Yu, K. Yusa, Genome-wide CRISPR-Cas9 screening in mammalian cells. Methods. 164–165, 29–35 (2019). https://doi.org/10.1016/j.ymeth.2019.04.015
J.C. Scheuermann, A.G. de Ayala Alonso, K. Oktaba, N. Ly-Hartig, R.K. McGinty, S. Fraterman, M. Wilm, T.W. Muir, J. Muller, Histone H2A deubiquitinase activity of the polycomb repressive complex PR-DUB. Nature. 465, 243–247 (2010). https://doi.org/10.1038/nature08966
K.H. Ventii, N.S. Devi, K.L. Friedrich, T.A. Chernova, M. Tighiouart, E.G. Van Meir, K.D. Wilkinson, BRCA1-associated protein-1 is a tumor suppressor that requires deubiquitinating activity and nuclear localization. Cancer Res. 68, 6953–6962 (2008). https://doi.org/10.1158/0008-5472.CAN-08-0365
M.A. Suico, T. Shuto, H. Kai, Roles and regulations of the ETS transcription factor ELF4/MEF. J. Mol. Cell. Biol. 9, 168–177 (2017). https://doi.org/10.1093/jmcb/mjw051
J.A. Castro-Mondragon, R. Riudavets-Puig, I. Rauluseviciute, R.B. Lemma, L. Turchi, R. Blanc-Mathieu, J. Lucas, P. Boddie, A. Khan, N. Manosalva Perez, O. Fornes, T.Y. Leung, A. Aguirre, F. Hammal, D. Schmelter, D. Baranasic, B. Ballester, A. Sandelin, B. Lenhard, K. Vandepoele, W.W. Wasserman, F. Parcy, A. Mathelier, JASPAR 2022: the 9th release of the open-access database of transcription factor binding profiles. Nucleic Acids Res. 50, D165–D173 (2022). https://doi.org/10.1093/nar/gkab1113
A. Garcia-Diaz, D.S. Shin, B.H. Moreno, J. Saco, H. Escuin-Ordinas, G.A. Rodriguez, J.M. Zaretsky, L. Sun, W. Hugo, X. Wang, G. Parisi, C.P. Saus, D.Y. Torrejon, T.G. Graeber, B. Comin-Anduix, S. Hu-Lieskovan, R. Damoiseaux, R.S. Lo, Ribas, Interferon receptor signaling pathways regulating PD-L1 and PD-L2 expression. Cell. Rep. 19, 1189–1201 (2017). https://doi.org/10.1016/j.celrep.2017.04.031
K. Hildner, B.T. Edelson, W.E. Purtha, M. Diamond, H. Matsushita, M. Kohyama, B. Calderon, B.U. Schraml, E.R. Unanue, M.S. Diamond, R.D. Schreiber, T.L. Murphy, K.M. Murphy, Batf3 deficiency reveals a critical role for CD8alpha + dendritic cells in cytotoxic T cell immunity. Science. 322, 1097–1100 (2008). https://doi.org/10.1126/science.1164206
D.R. Peaper, P. Cresswell, Regulation of MHC class I assembly and peptide binding. Annu. Rev. Cell. Dev. Biol. 24, 343–368 (2008). https://doi.org/10.1146/annurev.cellbio.24.110707.175347
P. Golstein, G.M. Griffiths, An early history of T cell-mediated cytotoxicity. Nat. Rev. Immunol. 18, 527–535 (2018). https://doi.org/10.1038/s41577-018-0009-3
D. Kägi, B. Ledermann, K. Bürki, P. Seiler, B. Odermatt, K.J. Olsen, E.R. Podack, R.M. Zinkernagel, H. Hengartner, Cytotoxicity mediated by T cells and natural killer cells is greatly impaired in perforin-deficient mice. Nature. 369, 31–37 (1994). https://doi.org/10.1038/369031a0
A. Campagne, M.K. Lee, D. Zielinski, A. Michaud, S. Le Corre, F. Dingli, H. Chen, L.Z. Shahidian, I. Vassilev, N. Servant, D. Loew, E. Pasmant, S. Postel-Vinay, M. Wassef, R. Margueron, BAP1 complex promotes transcription by opposing PRC1-mediated H2A ubiquitylation. Nat. Commun. 10, 348 (2019). https://doi.org/10.1038/s41467-018-08255-x
N.A. Fursova, A.H. Turberfield, N.P. Blackledge, E.L. Findlater, A. Lastuvkova, M.K. Huseyin, P. Dobrinic, R.J. Klose, BAP1 constrains pervasive H2AK119ub1 to control the transcriptional potential of the genome. Genes Dev. 35, 749–770 (2021). https://doi.org/10.1101/gad.347005.120
Y. Zhang, J. Shi, X. Liu, L. Feng, Z. Gong, P. Koppula, K. Sirohi, X. Li, Y. Wei, H. Lee, L. Zhuang, G. Chen, Z.D. Xiao, M.C. Hung, J. Chen, P. Huang, W. Li, B. Gan, BAP1 links metabolic regulation of ferroptosis to tumour suppression. Nat. Cell. Biol. 20, 1181–1192 (2018). https://doi.org/10.1038/s41556-018-0178-0
M. Zhang, W. Yang, P. Wang, Y. Deng, Y.T. Dong, F.F. Liu, R. Huang, P. Zhang, Y.Q. Duan, X.D. Liu, D. Lin, Q. Chu, B. Zhong, CCL7 recruits cDC1 to promote antitumor immunity and facilitate checkpoint immunotherapy to non-small cell lung cancer. Nat. Commun. 11, 6119 (2020). https://doi.org/10.1038/s41467-020-19973-6
Q. Gao, S. Wang, X. Chen, S. Cheng, Z. Zhang, F. Li, L. Huang, Y. Yang, B. Zhou, D. Yue, D. Wang, L. Cao, N.R. Maimela, B. Zhang, J. Yu, L. Wang, Y. Zhang, Cancer-cell-secreted CXCL11 promoted CD8(+) T cells infiltration through docetaxel-induced-release of HMGB1 in NSCLC. J. Immunother Cancer. 7, 42 (2019). https://doi.org/10.1186/s40425-019-0511-6
T. Chao, E.E. Furth, R.H. Vonderheide, CXCR2-Dependent Accumulation of Tumor-Associated neutrophils regulates T-cell immunity in pancreatic ductal adenocarcinoma. Cancer Immunol. Res. 4, 968–982 (2016). https://doi.org/10.1158/2326-6066.CIR-16-0188
T. Jamieson, M. Clarke, C.W. Steele, M.S. Samuel, J. Neumann, A. Jung, D. Huels, M.F. Olson, S. Das, R.J. Nibbs, O.J. Sansom, Inhibition of CXCR2 profoundly suppresses inflammation-driven and spontaneous tumorigenesis. J. Clin. Invest. 122, 3127–3144 (2012). https://doi.org/10.1172/JCI61067
S. Mariathasan, S.J. Turley, D. Nickles, A. Castiglioni, K. Yuen, Y. Wang, E.E. III Kadel, H. Koeppen, J.L. Astarita, R. Cubas, S. Jhunjhunwala, R. Banchereau, Y. Yang, Y. Guan, C. Chalouni, J. Ziai, Y. Senbabaoglu, S. Santoro, D. Sheinson, J. Hung, J.M. Giltnane, A.A. Pierce, K. Mesh, S. Lianoglou, J. Riegler, R.A.D. Carano, P. Eriksson, M. Hoglund, L. Somarriba, D.L. Halligan, M.S. van der Heijden, Y. Loriot, J.E. Rosenberg, L. Fong, I. Mellman, D.S. Chen, M. Green, C. Derleth, G.D. Fine, P.S. Hegde, R. Bourgon, T. Powles, TGFbeta attenuates tumour response to PD-L1 blockade by contributing to exclusion of T cells. Nature. 554, 544–548 (2018). https://doi.org/10.1038/nature25501
N. Markosyan, J. Li, Y.H. Sun, L.P. Richman, J.H. Lin, F. Yan, L. Quinones, Y. Sela, T. Yamazoe, N. Gordon, J.W. Tobias, K.T. Byrne, A.J. Rech, G.A. FitzGerald, B.Z. Stanger, R.H. Vonderheide, Tumor cell-intrinsic EPHA2 suppresses anti-tumor immunity by regulating PTGS2 (COX-2). J. Clin. Invest. 129, 3594–3609 (2019). https://doi.org/10.1172/jci127755
D.M. Brantley-Sieders, W.B. Fang, D.J. Hicks, G. Zhuang, Y. Shyr, J. Chen, Impaired tumor microenvironment in EphA2-deficient mice inhibits tumor angiogenesis and metastatic progression. Faseb j. 19, 1884–1886 (2005). https://doi.org/10.1096/fj.05-4038fje
K. Iwatsuki, K. Tanaka, T. Kaneko, R. Kazama, S. Okamoto, Y. Nakayama, Y. Ito, M. Satake, S. Takahashi, A. Miyajima, T. Watanabe, T. Hara, Runx1 promotes angiogenesis by downregulation of insulin-like growth factor-binding protein-3. Oncogene. 24, 1129–1137 (2005). https://doi.org/10.1038/sj.onc.1208287
T.R. Kyriakides, Y.H. Zhu, L.T. Smith, S.D. Bain, Z. Yang, M.T. Lin, K.G. Danielson, R.V. Iozzo, M. LaMarca, C.E. McKinney, E.I. Ginns, P. Bornstein, Mice that lack thrombospondin 2 display connective tissue abnormalities that are associated with disordered collagen fibrillogenesis, an increased vascular density, and a bleeding diathesis. J. Cell. Biol. 140, 419–430 (1998). https://doi.org/10.1083/jcb.140.2.419
T.R. Kyriakides, K.J. Leach, A.S. Hoffman, B.D. Ratner, P. Bornstein, Mice that lack the angiogenesis inhibitor, thrombospondin 2, mount an altered foreign body reaction characterized by increased vascularity. Proc. Natl. Acad. Sci. U S A 96, 4449–4454 (1999). https://doi.org/10.1073/pnas.96.8.4449
M. Streit, L. Riccardi, P. Velasco, L.F. Brown, T. Hawighorst, P. Bornstein, M. Detmar, Thrombospondin-2: a potent endogenous inhibitor of tumor growth and angiogenesis. Proc. Natl. Acad. Sci. U S A 96, 14888–14893 (1999). https://doi.org/10.1073/pnas.96.26.14888
X. Dai, Y. Fu, Y. Ye, Increased NDRG1 expression suppresses angiogenesis via PI3K/AKT pathway in human placental cells. Pregnancy Hypertens. 21, 106–110 (2020). https://doi.org/10.1016/j.preghy.2020.05.009
B.J. Hwang, L.C. Tsao, C.R. Acharya, T. Trotter, P. Agarwal, J. Wei, T. Wang, X.Y. Yang, G. Lei, T. Osada, H.K. Lyerly, M.A. Morse, Z.C. Hartman, Sensitizing immune unresponsive colorectal cancers to immune checkpoint inhibitors through MAVS overexpression. J. Immunother Cancer. 10 (2022). https://doi.org/10.1136/jitc-2021-003721
X. She, S. Shen, G. Chen, Y. Gao, J. Ma, Y. Gao, Y. Liu, G. Gao, Y. Zhao, C. Wang, C. Jiang, P. Wang, H. Qin, H. Gao, Immune surveillance of brain metastatic cancer cells is mediated by IFITM1. EMBO J. 42, e111112 (2023). https://doi.org/10.15252/embj.2022111112
D. Aran, Cell-type Enrichment Analysis of Bulk transcriptomes using xCell. Methods Mol. Biol. 2120, 263–276 (2020). https://doi.org/10.1007/978-1-0716-0327-7_19
M. Carbone, J.W. Harbour, J. Brugarolas, A. Bononi, I. Pagano, A. Dey, T. Krausz, H.I. Pass, H. Yang, G. Gaudino, Biological mechanisms and clinical significance of BAP1 mutations in Human Cancer. Cancer Discov. 10, 1103–1120 (2020). https://doi.org/10.1158/2159-8290.CD-19-1220
J.W. Harbour, M.D. Onken, E.D. Roberson, S. Duan, L. Cao, L.A. Worley, M.L. Council, K.A. Matatall, C. Helms, A.M. Bowcock, Frequent mutation of BAP1 in metastasizing uveal melanomas. Science. 330, 1410–1413 (2010). https://doi.org/10.1126/science.1194472
M. Bott, M. Brevet, B.S. Taylor, S. Shimizu, T. Ito, L. Wang, J. Creaney, R.A. Lake, M.F. Zakowski, B. Reva, C. Sander, R. Delsite, S. Powell, Q. Zhou, R. Shen, A. Olshen, V. Rusch, M. Ladanyi, The nuclear deubiquitinase BAP1 is commonly inactivated by somatic mutations and 3p21.1 losses in malignant pleural mesothelioma. Nat. Genet. 43, 668–672 (2011). https://doi.org/10.1038/ng.855
J.R. Testa, M. Cheung, J. Pei, J.E. Below, Y. Tan, E. Sementino, N.J. Cox, A.U. Dogan, H.I. Pass, S. Trusa, M. Hesdorffer, M. Nasu, A. Powers, Z. Rivera, S. Comertpay, M. Tanji, G. Gaudino, H. Yang, M. Carbone, Germline BAP1 mutations predispose to malignant mesothelioma. Nat. Genet. 43, 1022–1025 (2011). https://doi.org/10.1038/ng.912
M. Nasu, M. Emi, S. Pastorino, M. Tanji, A. Powers, H. Luk, F. Baumann, Y.A. Zhang, A. Gazdar, S. Kanodia, M. Tiirikainen, E. Flores, G. Gaudino, M.J. Becich, H.I. Pass, H. Yang, M. Carbone, High incidence of somatic BAP1 alterations in sporadic malignant mesothelioma. J. Thorac. Oncol. 10, 565–576 (2015). https://doi.org/10.1097/jto.0000000000000471
S. Pastorino, Y. Yoshikawa, H.I. Pass, M. Emi, M. Nasu, I. Pagano, Y. Takinishi, R. Yamamoto, M. Minaai, T. Hashimoto-Tamaoki, M. Ohmuraya, K. Goto, C. Goparaju, K.Y. Sarin, M. Tanji, A. Bononi, A. Napolitano, G. Gaudino, M. Hesdorffer, H. Yang, M. Carbone, A subset of Mesotheliomas with Improved Survival Occurring in carriers of BAP1 and other germline mutations. J. Clin. Oncol. 36, JCO2018790352 (2018). https://doi.org/10.1200/JCO.2018.79.0352
F. Baumann, E. Flores, A. Napolitano, S. Kanodia, E. Taioli, H. Pass, H. Yang, M. Carbone, Mesothelioma patients with germline BAP1 mutations have 7-fold improved long-term survival. Carcinogenesis. 36, 76–81 (2015). https://doi.org/10.1093/carcin/bgu227
M. Ladanyi, F. Sanchez Vega, M. Zauderer, Loss of BAP1 as a candidate predictive biomarker for immunotherapy of mesothelioma. Genome Med. 11, 18 (2019). https://doi.org/10.1186/s13073-019-0631-0
R. Shrestha, N. Nabavi, Y.Y. Lin, F. Mo, S. Anderson, S. Volik, H.H. Adomat, D. Lin, H. Xue, X. Dong, R. Shukin, R.H. Bell, B. McConeghy, A. Haegert, S. Brahmbhatt, E. Li, H.Z. Oo, A. Hurtado-Coll, L. Fazli, J. Zhou, Y. McConnell, A. McCart, A. Lowy, G.B. Morin, T. Chen, M. Daugaard, S.C. Sahinalp, F. Hach, S. Le Bihan, M.E. Gleave, Y. Wang, A. Churg, C.C. Collins, BAP1 haploinsufficiency predicts a distinct immunogenic class of malignant peritoneal mesothelioma. Genome Med. 11, 8 (2019). https://doi.org/10.1186/s13073-019-0620-3
D. Fukumura, J. Kloepper, Z. Amoozgar, D.G. Duda, R.K. Jain, Enhancing cancer immunotherapy using antiangiogenics: opportunities and challenges. Nat. Rev. Clin. Oncol. 15, 325–340 (2018). https://doi.org/10.1038/nrclinonc.2018.29
A.W. Griffioen, Anti-angiogenesis: making the tumor vulnerable to the immune system. Cancer Immunol. Immunother. 57, 1553–1558 (2008). https://doi.org/10.1007/s00262-008-0524-3
K. Liu, Y. Huang, Y. Xu, G. Wang, S. Cai, X. Zhang, T. Shi, BAP1-related signature predicts benefits from immunotherapy over VEGFR/mTOR inhibitors in ccRCC: a retrospective analysis of JAVELIN renal 101 and checkmate-009/010/025 trials. Cancer Immunol. Immunother. 72, 2557–2572 (2023). https://doi.org/10.1007/s00262-023-03424-4
S. Kurtulus, A. Madi, G. Escobar, M. Klapholz, J. Nyman, E. Christian, M. Pawlak, D. Dionne, J. Xia, O. Rozenblatt-Rosen, V.K. Kuchroo, A. Regev, A.C. Anderson, Checkpoint blockade immunotherapy induces dynamic changes in PD-1(-)CD8(+) tumor-infiltrating T cells. Immunity. 50, 181–194e186 (2019). https://doi.org/10.1016/j.immuni.2018.11.014
Acknowledgements
We would like to acknowledge the flow cytometry core facility of Frontier Science Center for Stem Cell Research and Laboratory Animal Research Center of Tongji Laboratory. The authors thank Kosuke Yusa for providing a mouse genome-wide CRISPR v2 library.
Funding
This work was supported by the National Key R&D Program of China (2021YFA1100903, 2021YFA1100900) and the Natural Science Foundation of China (82371736).
Author information
Authors and Affiliations
Contributions
H.C. and M.L. carried out experiments and analyzed the data. Y.Y. performed the CRISPR-Cas9 screening experiment with the help of M.L., S.H.O. performed the CRISPR-Cas9 screening data analysis. L.Z., Z.Z., X.X., Y.Z., J.W., X.L., K.L. and Y.L. performed the experiments. H.W. provided intellectual inputs for the manuscript. J.Z. analyzed the TCGA human cancer samples. Z.M. performed the transcriptome data analysis. X.C. performed ATAC-seq and CUT&Tag data analysis. J.Z. conceived the study and wrote the manuscript. Y.Y. conceived the study, designed the research, analyzed the data and wrote the manuscript. All authors provided critical feedback on the manuscript and reviewed the manuscript.
Corresponding authors
Ethics declarations
Ethics approval and consent to participate
All the mice used in this study were accommodated in the Tongji University animal care facility and were maintained under pathogen-free conditions. All the experiments were approved and conducted following the guidelines of the Animal Care Committee of Tongji University under approval number TJAB03420101.
Dual publication
The results/data/figures in this manuscript have not been published elsewhere, nor are they under consideration by another publisher.
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Chang, H., Li, M., Zhang, L. et al. Loss of histone deubiquitinase Bap1 triggers anti-tumor immunity. Cell Oncol. (2024). https://doi.org/10.1007/s13402-024-00978-y
Accepted:
Published:
DOI: https://doi.org/10.1007/s13402-024-00978-y